JOURNAL OF VIROLOGY, Oct. 1976, p.222-233 Copyright ©D1976 AmericanSocietyforMicrobiology
Vol.20,No. 1 Printed in U.S.A.
Physical Maps for
Herpes
Simplex
Virus
Type
1
DNA for
Restriction
Endonucleases Hind III, Hpa-1, and
X.
bad
NEIL M. WILKIE
ColdSpring Harbor Laboratory, Cold Spring Harbor, New York 11724, andInstituteofVirology,
Glasgow,Gll 5JR Scotland*
Received forpublication10May 1976
Ithas beenproposed that thegenomeofherpes simplexvirustype 1 (HSV-1)
consists oftwo internal unique sequences, S and L, bounded by two sets of
redundant sequences (P. Sheldrick and N. Berthelot, 1974). In this
arrange-ment, terminal sequences (TRS and TRL) arerepeated in an internal inverted
form
(IRS
and IRL) and delimitS and L. Furthermore, abody of evidencehasaccumulated that suggests that S and L themselvesareinverted, givingriseto
four related forms of the HSVgenome. Inthisstudytheorderingofrestriction
endonuclease fragments of HSV-1 DNA for physical maps has been studied usingmolecularhybridizationtechniquesandthecleavageofisolatedrestriction
endonuclease fragmentswithfurtherrestriction endonucleases. Physicalmaps
for the fragments produced by Hind III, Hpa-1, and X. bad have been
con-structedfor the four related forms of the HSV-1genome.
TRs
andIRS
werefoundtobe between 3.5x 106and 4.5 x 106daltons,
TR,
andIRLabout6x 106daltons, Sabout 8 x 106 to 9 x 106 daltons, andL about 6.8 x 106daltons.
The genome ofherpes simplex virus type 1
(HSV-1) consists ofalinearduplex DNA
mole-cule of about100 x 10"daltons.A true terminal
redundancy of about0.5 x 106 daltons (2, 3)is
contained insomewhat largerblocks of
termi-nal sequences which Sheldrick and Berthelot
(10)found to be repeatedin
internal,
inverted forms. Two unique sequences(S
= 107daltons;
L =75 x 106daltons) were foundtobebounded
by these inverted repetitions. It was
proposed
that Swasboundedatthe terminusby
TRs
andinternally by thesamesequence
inverted,
IRS.
Similarly, L was bounded byTR,
and IRL. Sheldrick and Berthelot (10) alsoproposed
thatthe unique sequences S and L might become relatively inverted by recombination events
(seeFig. 10).
Subsequent
workby
variouslaboratoriespro-videdclearevidence that thismodel, including
the relativeinversionsinS andL,was
substan-tially correct. Restriction endonuclease cleav-ageanalysis showed that submolar bands
pres-ent in
digests
ofboth HSV-1 and HSV-2 DNAcould be accounted for by the Sheldrick and Berthelot model
(1,
4, 5, 8, 11, 14). In digestswithendonucleaseswhich cleaveonlyinS and
L,two 0.5 Mterminalfragmentsaregenerated
from eachend. Four 0.25 M
fragments,
whichspan the internal inverted redundant
se-quences, also arise. If it isassumedthat
TRL/IRL
andTRS/IRS
are not identical, then enzymes that cleaveinSandL,butinonlytheTRL/IRL
paironthe
TRs/IRs
pair, generate amolarendfrom that terminus in which the redundant
sequence iscleaved. Two 0.5 Mfragmentsfrom
the otherterminusandtwointernal 0.5 M
frag-ments are also present (see Fig. 10). Direct
analysis of the terminalfragments found in
re-striction endonuclease digests of HSV-1 DNA, combinedwithsequence homology studies,
con-firmed thesehypotheses (15)and provided
fur-ther evidence that
TRL/IRL
andTRs/IRs
werenotidentical. Finally, the relative inversionsof
S and L in thefourgenome arrangements for
HSV-1were visualized directly inthe electron
microscope via both incomplete (5) and
com-plete partial denaturation maps (Delius and
Clements, in press). In the latter study,
TRL/
IRLand
TRs/IRs
haddifferentpartialdenatura-tion profiles.
Thisreport givesthe orderofHind III,
Hpa-1, and X. bad fragments in the four genome
arrangements for HSV-1DNA.
MATERIALS AND METHODS
Enzymes. Restriction endonucleases Hind III and Hpa-1 were prepared as described previously (9, 14). X. bad, anenzyme fromXanthamonus badrii, was
the kind gift of S. Zain. DNA polymerase I was purchased from the Boehringer Mannheim Corp., andDNaseIwas fromWorthington.
Nick translation. HSV-1 DNA was labeled with
a-32P-labeled deoxyribonucleotide triphosphates
(specific activity approximately 100 Ci/mmol; New
England Nuclear Corp.) essentially as described by 222
on November 10, 2019 by guest
http://jvi.asm.org/
PHYSICAL MAPS FOR HSV-1 DNA 223
Maniatis etal. (7). Reaction mixtures contained, in avolume of 100 ul: 1
Ag
ofpurified HSV-1 DNA; 50mMTris-hydrochloride, pH 7.8; 5 ,MMgCl2; 1mM
dithiothreitol; 50 ,ug of bovine serum albumin per
ml; 180 pmol of[a-32P]dTTP, [a-32P]dGTP,
[a-32P]-dCTP, and [a-32P]dATP; 2 units of DNA
polymer-ase I; 10-10 g of DNase I per ml. Incubation was at 16°C for 1 h. The DNA was then extracted with phenol and purified by gel filtration on Sephadex G-50.
The growth ofvi*usand DNA,end-labeling
stud-ies onHSV-1 DNA, restrictionendonuclease diges-tion, analysis and purification of DNAfragments,
theSouthern DNA-transfer technique (12), and hy-bridization methods were carried out as described in the previous paper (15).
RESULTS AND DISCUSSION The cleavage patterns of HSV-1 strain 17
DNA for Hind III, Hpa-1, X. bad, and an X. bad-Hpa-1 double digest are shown in Fig. 1. The molarityofeach submolarband isshown
for thesingle
digests.
The molecularweights
ofthe Hind III and Hpa-l fragments have been published previously (15), and theX.baddigest
is discussed below. The over-molar group of
fragments previously
designated
Hpa-1 o (15)areshown belowtobe derived fromtwo separa-ble maplocations, and this group hastherefore
been subdividedintoHpa-1o and Hpa-1o'.
Highly labeled HSV-1 DNA was needed for the studies discussed below, and 32P-labeled DNA ofhigh specific activity was obtained by nick translation as described under Materials and Methods. At the end of the reaction, DNA
withaspecific
radioactivity
of3 x 107cpm perjig
appeared to be intact (in the sense of double-strandmolecularweight)
and could bedigested with restriction endonucleases to give normal profiles. Figure lb showsanautoradiograph
ofnick-translated
HSV-1DNA, cleaved withHpa-1 and separated
by
electrophoresis
on anaga-roseslabgel. The profile obtained appears iden-tical tothatobtained with the unlabeled DNA
showninFig. la. The
autoradiogram
wasused as a template to enable the DNAbands tobeexcised
directly
forsubsequent purification.
Figure 2 shows that when the purified frag-ments were analyzed by gelelectrophoresis
theirmobility
hadnotaltered.Analysis of Hind
III/Hpa-1
double digests. To aid the construction of linkage groups for differentrestrictionendonucleasefragments, acomprehensivedescriptionof the Hind
III/Hpa-1doubledigest was obtained.
32P-labeled
Hpa-1 fragments were isolated as described above andfurther cleaved with Hind III. The reciprocal
experiment was also carried out. The analysis
of the Hind IIIcleavage productsof someHpa-1
fragments is shown in Fig. 2. The molecular
weights
oftheproductswereestimatedbyref-erence to the standard cleavage products of HSV-1 DNArun ontheoutside slots of the slab
gel.
The results of several experiments are
sum-marizedinFig.3.Each fragmentpresentinthe HindIII/Hpa-1 doubledigestwasshownto de-rive from the indicated Hind III and
Hpa-1
fragments by
further cleavage. Hind III orHpa-1 fragmentspresent aslimitfragments in the double digest (fragments not further cleaved by the alternative enzyme) are
indi-cated. The terminal fragments in the double
digestwerelocated by end-labelingstudies
sim-ilar to those described in the previous paper
(15), and their positions in single and double
digestsare indicated.
Sequence homology between fragments.
Theassignmentofdifferentrestriction
endonu-cleasefragmentstolinkage groups canalso be deduced from sequence homology studies. 32p_ labeledHpa-1fragmentswerehybridizedto
un-labeledHind IIIfragments (and vice versa) by usingtheDNAtransferorblottechnique
devel-oped by Southern (12) and described in our
previous publication (15). Some
cross-hybridi-zation results are presented in Fig. 4. It is
evidentfromthisfigure thatfragments which
appearedtobefairlypure on the basis oftheir relative mobilities upon gel electrophoresis (Fig. 2)are, infact, contaminated bysequences
from many different regions of the genome.
This contamination can also be seen in the
further cleavage analysis of some isolated
bands shown in Fig. 2. This problem of
se-quence contamination was alsoobservedwhen
DNA labeled in vivo was used, and it seems
morelikelytobeanartifactdue-tothe
purifica-tion and separation methods used rather than
to the method oflabeling. The contaminating
sequences in the 32P-labeledHpa-1 fragments
causehybridization to most Hind IIIfragments
at levels above background. Nevertheless, the
hybridization ofthe main Hpa-1 fragment in
any reaction can readily be distinguished as
moreintensebands of
radioactivity
ontheauto-radiographs. The data presentedinFig.5and6
summarize the data obtained in several such
experiments in which 32P-labeled Hind III
fragments were annealed to unlabeled Hpa-1
fragmentsand vice versa.
Construction ofa partial map. In the
pre-vious publication (15) the four terminal
frag-ments in Hind IIIdigestsof HSV-1 DNA were
identified asHind IIId, g, andmandeitherh or i (notseparated by thegel electrophoresis).
Hind III iisarbitrarilydesignatedas the
termi-nal 0.5 M band. In Hpa-1 digests, the three
terminalfragments wereidentifiedas a molar
VOL. 20, 1976
on November 10, 2019 by guest
http://jvi.asm.org/
224 WILKIE
a
.}~~
a
F
~~~~~~is
--- k
- p
a
b
-_ c
0 -
ci
e
-
f
g o
--
----,
Ik
-~
a1
n
S
aa
I)
cl
f
ic0
0
Hindc
IIIXbaci
Hpa
Hpa
0.5M
cl
g mIII0Q5M
accig(Hpa I)
0.5M
abcdq0. 25M
bceff
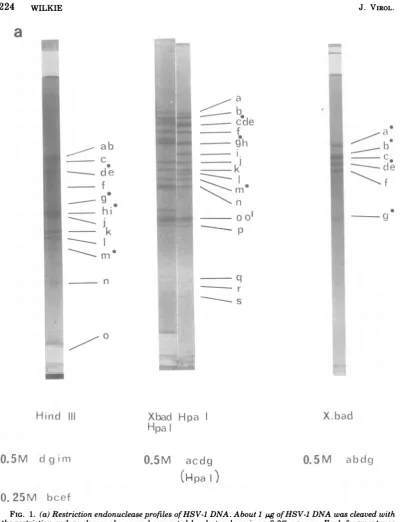
FIG. 1. (a)Restriction endonucleaseprofiles ofHSV-1 DNA. About1 pgofHSV-1DNAwascleaved with
the restriction endonucleases shown andseparatedbyelectrophoresison0.3%agarose.Eachfragmentwas
assignedaletterofthealphabet. Hpa-1 o',anover-molargroupoffragments,has beensubdivided into two
classes,o ando', for reasons explained in the text. Minor bands ofmolarities 0.25 or0.5 areindicated.
Terminalfragmentsareindicatedbythesolid circle(@). (b)Theprofile ofnick-translated HSV-1 DNA.32p_
labelednick-translatedDNA wascleaved withHpa-1andseparated byelectrophoresison aslabgel of0.7%
agarose.
J. VIROL.
.w ..,.',U..,7
..-r
[image:3.508.65.467.62.585.2],;^. iR..
.|..tr.2SX .||.
.,w: ., ;jE
iilllit
1a
.!L'
|,iF
:!Ei *.,]-';''
'.!,
r
-1111ruS
i >;
:ZE
F
Bt.
VblE
i44
q
r S
X.
b
ad
on November 10, 2019 by guest
http://jvi.asm.org/
PHYSICAL MAPS FOR HSV-1 DNA
225
VOL. 20, 1976
b
R
.
b!__.
_...
,
be.,
s4.
...
___;s.
__.5S|*
s
_^S
_we M .. '0, '.i:.
... ..
+:--;.,<. j: -w','s, 27r.; ,: - :*:
:- :@ 'r+.
<,.; +'s ',.
a. :u;^s9;s,2.- ,; { 0" w:; s
*;-+ ,,@ _ , }t ...
_L
w_|-
f-
on November 10, 2019 by guest
http://jvi.asm.org/
226 WILKIE
a b cde
_4
'
r I g i_m
m
Om
k
m7
FI 4h n
I. ft.f*
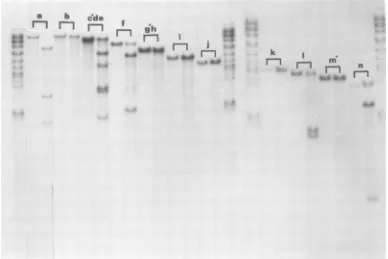
FIG. 2. Furthercleavage products of Hpa-1 fragments with HindIII. 32P-labeledHpa-1 fragments were
isolated as described. 2 x 103 to 2 x 104 cpm of each fragment (less than 1p) was mixed with 0.5 igof
unlabeled adenovirus-2 DNA and cleaved with Hind III. The cleaved fragments wereseparated by electropho-resis onslab gels of0.7% agarose. Undigested fragments were run inparallelslots.Ineachcasetheunlabeled adenovirus-2 DNA wasdigestedto limitproductsasdetermined byDNA-ethidium bromidefluorography.
Labeled DNA was determined by autoradiography. Standard Hpa-1 cleavageproducts of total HSV-1 DNA
areshownonthe outside slots. Hpa-1 fragments from thetermini are indicatedbythesolid circle(0).
fragment Hpa-1m andtwo 0.5 Mfragmentsin
Hpa-1 cde and hg. By a similar arbitrary deci-sion
Hpa-1
dandg aredesignatedas thetermi-nal fragments. Evidencewas alsopresentedin
the previous publication (15) to suggest that
Hind III d and i andHpa-1 m were from one
terminusof thegenome and Hind III gandm
andHpa-1candg from the other. Itcanbeseen
from Fig. 3that the terminal fragmentsin the
Hind
III/Hpa-1
double digests allarisefrom theDNAbands of the single digests which are, or contain, ends. The data of Fig. 3 are in
com-plete agreement withtheprevious assignment (15) of end fragments to different termini of the genome.
By combining this information and the data
summarized inFig. 3, 5, and 6, apartial map
fortheHind IIIandHpa-1fragments has been
constructed. Forexample, Fig. 5 and6
demon-stratethattheuniquefragments HindIIIj and
Hpa-1
j and n are sequence related. From theresultspresented in Fig. 2 and 3, it can be seen
that there is a Hpa- 1 site in Hind
IJIj
and a HindIII site inHpa- 1 n, but no Hind III site in Hpa- 1
j. Furthermore,
Hpa-1
nand Hind IIIj give rise to the same double-digest fragment "o." Themolecular weights of the further
cleavage
prod-ucts (notshown)suggestthatat oneend of this linkagegrouptheremustbe Hind III and
Hpa-1 sites very close together. The only other
se-quence homology for
Hpa-1
n inHind IIIdi-gests is inHindIIIab (Fig. 6). The
homologous
sequences must lie in the unique
fragment
HindIII a, sincebis a 0.25 Mfragment, andits
sequences must be present in other minor
bands because of the relativeinversions ofthe uniquesequencesSandL inHSV DNA.
Hpa-1
n must therefore span the Hind IIIa-HindIIIj
junction. Analogous reductive
analysis
of theresults shown in Fig. 3, 5, and 6 indicates
thatHpa-1 b, h, i, and q must be contained
withinHindIII a.Thefollowing large linkage
group is thusconstructed:
Hind III
a
1i1
A
A~n
(b,h, i, q) Hpa-1
J. VIROL.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:5.508.69.457.82.341.2]PHYSICAL MAPS FOR HSV-1 DNA 227
Hpa I fragments
bX cdxe a
'xe
f ijX cdxe;j
kx ma de c0dxe a;edxe0 J
(>I)p
q r
-ab;c;de;I ab;c;cle;g
hi-ox
f;ool a;c dXe;px
I
FIG. 3. Summaryoffurther cleavage experiments.
On the basisof further cleavage experiments, bands
ofDNA inHpa-1IHind III doubledigestswereshown
experimentally to have arisen from the cleavage of
HindIIIfragments with Hpa-1 andof Hpa-1
frag-mentswithHindIII. Solid circles(a) indicate frag-ments arising from termini. Crosses (X) indicate
fragments of single digestswhicharenotcleavedby
the secondenzymeand which existaslimitfragments
in the double digest. For example, double-digest
band k arisesfromtheHpa-1 cleavage ofHind III hi. SinceHpa-1 k isnotfurthercleaved with HindIII, double-digestband k andHpa-1kmustbe identical. Hind III i is a terminal fragment. Hpa-1 k must therefore be contained within either Hind III h or
Hind III i.
Space does not permita full analysisof the
construction of all the linkage groups here.
However, the data presentedinFig. 4, 5, and6
contain all the information necessary to
con-struct the partial map shown in Fig. 7. The
linkagegroup containingHind IIIa andj and
that containing Hind III h could not be
as-signedmappositionsonthe basisof the
experi-mentswithHindIII and Hpa-1describedsofar.
The limits of the map terminate at positions
wherethe Hind IlI and Hpa-1 sites are so close
together that no more overlaps could be ob-tained.
The tendency for Hind III c and de toanneal
to Hpa-1 b is notaccounted for in this partial
map. Since further mapping data below puts
Hpa-1 b firmly within Hind III a, it is thought
that this aberrant annealing data is due to
Hind IIIa sequencescontaminating Hind III c and de.
The cleavage pattern for Hpa-1 arises from
Hpa-1 sites in SandLandin either
TR,/IRs
orin
TRL/IRL
(1, 15).Therefore, one terminalfrag-ment mustbe present in molar amounts.Hpa-1
m hasbeen identifiedas a terminal fragment (15), but was originally quantified as 0.77 M
andwasthoughttobea 0.5Mband (Hpa-11 in
reference1). Fromthe resultspresentedhere it is clear that Hpa-1 m arises from a cleavage
within TRL and must be the molar end. The
reason for the apparently low amount of this
terminalfragment indigests is notcompletely understood. We havenoted thatinmany cases
terminal fragments from bothtermini are more
spread after gel electrophoresis than other
bands of DNA (N. M. Wilkie, J. B. Clements, and R. Cortini,
unpublished
observations) asthough there was some heterogeneity intheir molecular weights. Such heterogeneity and spreading tends to reduce the peak height of the bands and in some cases may lead to a
reduction in the observed amount ofDNA in
thebands. Thereasons fortheheterogeneityin
terminal fragmentsremains a matter for
specu-lation.
The hybridization and further cleavage
ex-periments also confirm the
general
model for HSV DNAfirstproposed bySheldrick and Ber-thelot (10).Bands ofDNA containingsubmolar fragments from one enzyme digest cross-hy-bridizetotheexpected submolar fragmentspro-duced by the second enzyme. For example, Hpa-1a (0.5M) annealsto all ofthe submolar fragmentsintheHindIIIdigest (Fig.4,6). The terminal Hpa-1 fragment m hybridizes to the
0.25 MHindIIIfragments andtothetwoHind
IIIterminal fragmentsd andi, but itdoesnot
anneal significantly to the Hind III terminal fragmentsg and m. This confirms the
assign-ment ofHpa-1 m to the same terminus from
which Hind III d and i arise. It might have
been expectedthat the actual terminal
redun-dancy of about 0.5 x 106 daltons previously proposed byother groups (2, 3, 10)mighthave
caused some cross-hybridization between
ter-minal fragments fromdifferent endsof the
ge-nome. However, thenearby inverted repeat of
the terminal sequences found in HSV-1 DNA
by Hyman et al. (6) and subsequently
con-shownto bederived from
Hind III fragments
a ab
b-- ab;de
c- . c
de __ ab;g
f- dAe
g- k
h ab
___. j;eX
k hi
I - de;hi
m I n.~ nx OC0 0
Im
a6 I)VOL. 20, 1976
on November 10, 2019 by guest
http://jvi.asm.org/
[image:6.508.50.242.69.420.2]228 WILKIE
32p
Hpa
fragments
a
cof
I
m no0
d
oab
Cd-ire
f
-9k.- *
i-k
-=W
a)I(
U.,.-c
-5
U0
Ca
b
i
k
ab-
+
c-de
-f _
h
*.
h
-k
-I
_m-n-
S
n-
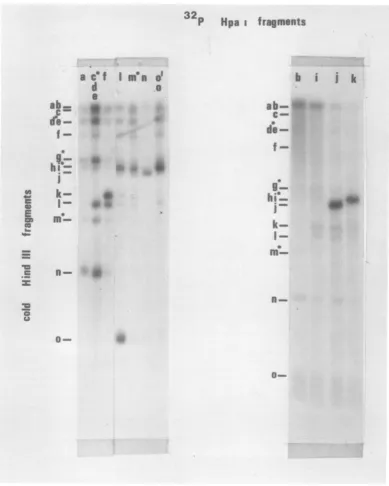
0-FIG. 4. Hybridizationof isolated 32P-labeledHpa-1 fragmentstounlabeled HindIIIfragments of HSV-1
DNA.Thetechnique was that described previously (12,15) and annealing of labeledDNAto unlabeled DNA onnitrocellulose membrane strips was determined byautoradiography. The positionsofalloftheunlabeled
DNAbands on nitrocellulose membrane strips were located by annealing with fragmented, unfractionated 32P-labeled HSV-1 DNA. Terminal fragments are indicated by the solid circle.
firmed by Wadsworth et al. (13) may cause
internal reassociation (in the unlabeled DNA)
which might interfere with the reassociation
reaction.
AnalysisofX. baddigests. X. bad fragments
of HSV-1 DNAcanbeseparatedinto sixbands
by electrophoresis on 0.3% agarose (Fig. 1). Analysisof themassdistribution of the DNAin J.
VIROL.-,^- Ai 0
on November 10, 2019 by guest
http://jvi.asm.org/
[image:7.508.75.465.73.560.2]PHYSICAL MAPS FOR HSV-1 DNA 229
32P-LABELED HIND III FRAGMENTS
ab c de f g
hi'
j k+ + + + + + ++ ++ ++ + + + + + +
1 me n 0
[image:8.508.67.430.71.201.2]+ ++ ++ ++ ++ ++ ++ (+) + ++ + + + + ++ ++
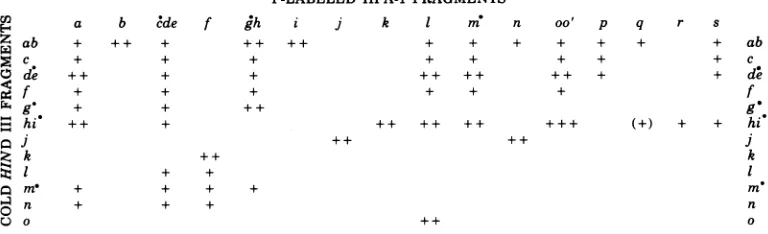
FIG. 5. Thehybridizationof 32P-labeledHind IIIfragmentsto unlabeledHpa-1 fragments.Solidcircles
(a)indicate terminalfragments.Theamountoflabel in each band(abovethat level causedby contaminating
sequences) wasassessedbyeyeforeachstripand allocated +, ++,or+ + +inorderofincreasing intensity.
E-l a b cde
Z ab + + + +
c + +
de ++ +
¢4
f + +x g + +
,_ hi + + +
Zk
SI +
om. + +
O n + +
0v
32P-LABELED HPA-1FRAGMENTS
e f gh i j k 1 m
++ ++ + + ++
n oo' p
+ + + + + + + + + + ++ ++ + + ++ ++ ++ ++ + + + ++ + ++ ++
q r s
+ ab
+ c
+ de f
FIG. 6. Thehybridization of32P-labeledHpa-1 fragmentstounlabeledHind IIIfragments.
those bands by the techniques described in a
previous publication (1) give themass
percent-ageof thesix bandsa, b,c,de,f,gas 19, 11.2,
26, 27, 13, 3.2. The molecular weights of these
fragments were estimated from slab gels and
from the further analysis and mapping ofX.
badfragmentstobe 34x 106,22 x 106, 21 x 106,
18 x 106, 15 x 106, and 6.5 x 10-6, respectively.
From thesefigures, and assumingamolecular
weight of about 98 x 206for HSV-1 DNA, the
molarities for a, b, c, de, f, and g could be
calculated to be 0.55, 0.51, 1.2, 1.46, 0.85, and
0.48.Thissuggeststhata, b, d,ore,andgare
0.5 M fragments and thatc, d ore, andfare
molar fragments. As before, d wasdesignated
asthe 0.5Mfragment, andewasdesignatedas
the molarfragment.
Analysis of X. bad/Hind III, and X. bad!
Hpa-1 double digests (e.g., Fig. 1) clearly
indi-catethat thereare only fourcleavage sites for
X. bad inHSV-1 DNA. One site lies in HindIII
I andHpa-1 e, two in Hind IIIa (one each in
Hpa-1 b and i),and the remaining site in Hind
IIIi (and its inversion Hind III
f)
andinHpa-11. These X. bad sites are shown inthe partial
mapinFig. 7. Thetwo cleavage sites inHind
IIIawereconfirmedby the furtherdigestion of
isolated Hind III ab with X. bad. Three further
cleavage productswereidentified with
molecu-larweights of15 x 106, 8 x 106, and 2.8 x 106.
The15 x 106fragments had thesamemobility asX. badf.
The fourcleavagesites all lie in the L region
of the genome and would give rise to seven
fragments because of the inversion of the L
region: anenzyme which cleaves only in the L
regionshould producetwo 0.5Mterminal
frag-ments which contain the Ssequences and two
0.5 Mfragmentsfrom the L terminus. All other
fragments should be molar. This fitsverywell
with the cleavage profile and the estimates of
molecular weights and molarities obtained for
the X. bad fragments. X. bada and b arethe
0.5 M terminifromthe S region; X. bad d andg
arethe 0.5 Mtermini fromL; andX. badc, e,
and fare the remaining internal molar
frag-ments from L. This analysis of the X. bad
cleavage patternoffers further confirmation of
the partial mapshown inFig. 7.
Completion of themap. The position of the
X. bad sites onthe HSV-1 DNA mapprovides
thenecessaryoverlaps that allow completion of
themap. 32P-labeled Hind IIIfragments k ando
andHpa-l-labeled fragments b, j, and k were
cna
E- b : f
° gh
t4
-,J k I m
rn
O oo' p b cde f gh j +++ k I m n oo' p+++ (+) + + hL k
m n VOL. 20, 1976
on November 10, 2019 by guest
http://jvi.asm.org/
[image:8.508.61.445.248.365.2]230 WILKIE
49 k
m
InI
0)
c A d
AoA
0pA
e f AA 0 i
&I
OA
mm ,|n > e o
c2 ^-
--A d
A60A
IAg In>
1 | k ,
A a
Ao
e A Ag a a o~~~~~pf
n2 f d
g a a A0A f A e A o'p
AoA
mti
t4
AA
n
a
a
bhiq A krA
HindI1I1|
Hpa a
X. bad t
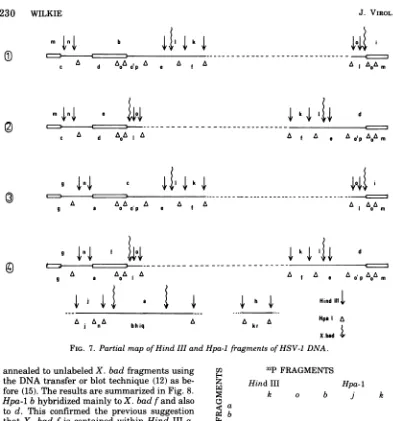
FIG. 7. Partial mapofHind III andHpa-1 fragments ofHSV-1 DNA.
annealed to unlabeledX. bad fragments using the DNAtransfer orblottechnique (12)as
be-fore(15). The resultsaresummarized inFig.8.
Hpa-1 bhybridized mainlyto X. badfand also
to d. This confirmed the previous suggestion
that X. bad f is contained within Hind IIIa.
Hpa-1j andk and HindIIIohybridized mainly
to X. bad c, whereas Hind III k hybridized
mainlytoX. bad de. SinceX. bad disa
termi-nalfragmentfromL,itssequencesmust also be
contained inonelarger0.5 Mfragment(a orb)
from the Sregion.ThehybridizationofHind III
k mainlytoX. bad de must thereforebe to the
unique fragment e. From the partial map
shown in Fig. 7 it canbe seen thatX. bad e
must lie adjacent to the Hind III b/l and c/l
junction. Since Hind III o and Hpa-1 k both
annealtoX. badc,Hind III h must lieonthe
Hind III o side of the gap in the partial map shown inFig. 7.Thus,X. badc must lieinthe
regionof the Hind IIIh/ojunction.Thisleads to
thefollowing unequivocalorderof X. bad
frag-mentsinthe four arrangements of thegenome shownin Fig. 7 and 10:
C,)
z
Ca
9: b
c
¢ de
mq
f
ZC g
04
u
32p FRAGMENTS Hind III
k b
++
++
Hpa-1
k
++ ++
+
FIG. 8. Thehybridization of 32P-labeledHindIII andHpa-1fragmentstounlabeled X. badfragments.
(1)aefcg (2)bc fed
(3)aefcg (4)bcfed
Sinceonlytwoarrangementsarepossiblefor L,there areonlytwopossibleX. bad maps.
SinceHpa-1 j andk bothannealed to X. bad
c,both must beadjacenttothe HindIIIoregion
inthepartialmapofFig.7.Theonlywaythese
linkagegroups canbearrangedintheLregion
k 1 d
Af A g
Ae
AAp&o
m
AI Am
I
J.
VIROL.-
on November 10, 2019 by guest
http://jvi.asm.org/
[image:9.508.65.458.51.472.2] [image:9.508.277.458.418.533.2]PHYSICAL MAPS FOR HSV-1 DNA 231
istherefore (in the order for the Hind III frag-ments): 1 kaj ho.
Hind IIIacontains twocleavage sites for X. bad, whichgiverise tothe threefurther
cleav-ageproductsat 15 x
106,
8 x106,
and2.8 x 106 daltons. Hpa-1 bandieachcontain oneof theseX. bad sites. Theonlyinteriorarrangement of Hpa-1b, h, andi thatallows the further
cleav-age productsactually observed is:
Hind III
a
1| h b
A A
Hpa-1 where t=X.bad; I =HindIII, and A=Hpa-1.
The results donotallow thelocation of
Hpa-1
q withinHind IIIa.Since
Hpa-1
q issmall (1.7x 106 daltons) this doesnot affect the relative order of
Hpa-1
b, h,
and i within Hind IIIa.(The hybridization of 32P-labeled Hpa-1 q to
unlabeledX. bad fragments should allow the unequivocal assignment of q.) The final
orien-tationofthemaps wasobtainedby
hybridizing
32P-labeled Hpa-1 fragments to unlabeled X.
bad/HindIII
double-digest
fragments using the DNA transfer or blottechnique
(14). There-sultsareshownin
Fig.
9.Hpa-1
ihybridizes
to twofragments,
asexpected
from thesmallsec-tion of the map shown
immediately
above. Hpa-1n hybridizes tothe smaller of thesetwofragments and also to a
higher-molecular-weight fragment.Hpa-1
j
annealsexclusively
to this larger
fragment,
which has amobility
thesame asthat of HindIIIj (whichis alimit
fragmentinthe X.bad/HindIIIdouble
digest).
This confirms the Hpa-1 maporder inthis re-gion as:h b i nj (kr)
Lqi
Taking all these results together, the
com-plete cleavage fragment maps for Hind III,
Hpa-1,
and X. bad shown in Fig. 10 can beconstructed. Theresultsdo not allow the final
orderofHpa-1o
andp,
ofHpa -1 k and r, or thefinal positionofHpa-1qwithin Hind III a to be
assigned. Thehybridizationresults with Hpa-1
s did not give a unique location on the map.
However, it ispossible thatHpa-1 s is in fact two fragments. In this case, the hybridization data shown inFig. 5 can beexplainedifone of
thefragments,
Hpa-1
s,liesintheo'p
region of themap, andthe otherfragment,
Hpa-1 s', liesin the linkage group kr within Hind III h. *-.
E
Cu
I
x0
=
=
Cu
,. 4,v
a
u
TIj"
P
Hpa
I
fragments
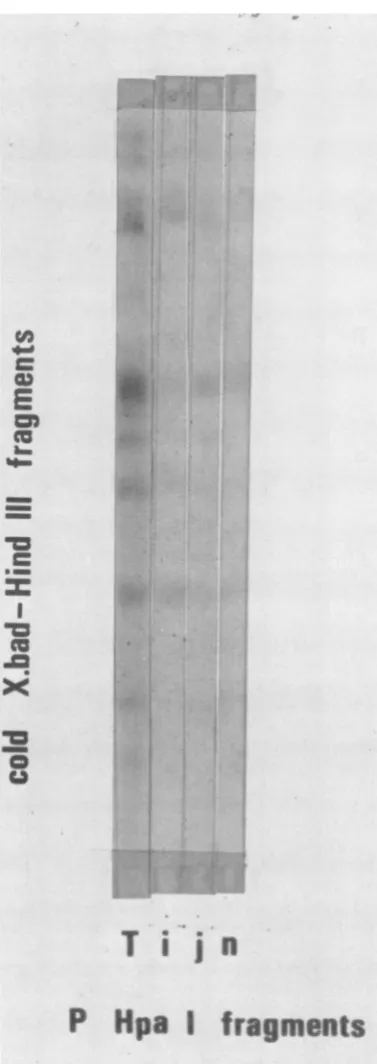
FIG. 9. Hybridization of32P-labeled Hpa-1
frag-mentsof HSV-1 DNA to unlabeled X. bad-Hind III double-digest fragments. T means the hybridization of fragmented, unfractionated 32P-labeled HSV-1 DNA to cold fragments.
VOL. 20, 1976
-v
on November 10, 2019 by guest
http://jvi.asm.org/
[image:10.508.259.448.69.602.2]232 WILKIE
TR5 S IIR IR L TRL
m n b I k a h I
[ G 1 Jk
A AAd A ^ daA
c d o ops e f h b i n j I9omskr
L q
msn e 0 h I a k I d
, A A A A A A ,
c d oI skr j n b h If e ops o m
b c f (- d
g4,n,
c , I k4 a j h oA AA, A A A A A AA A
g a o ops e f h b i n j skr
a e f c 9g
g
n,
f o h l a k I d4_
4,4 4,4.
4,4,4,a A-A AA A A A
i
A AA. [image:11.508.68.464.54.341.2]9%~ a kr eglJ n b h _ opsom
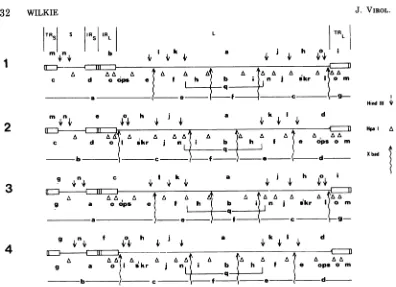
FIG. 10. Completephysical mapsforthe HindIII, Hpa-1, and X. badfragments of HSV-1 DNA. The
redundantsequences TRL, IRL, TRs, andIRsand the unique sequencesL andS, originallyproposed by
Sheldrick and Berthelot(10), areshown.
Accordingly, these map positions have been
tentativelyassignedtoHpa-1sand Hpa-1 s'.
The molecular weights of the fragments in
the maps shown in Fig. 10 give a genome
molecularweighttotalof 98 x 106.Sappearsto beabout 8-9x 106daltons, and L about 68 x 106
daltons. These figuresaresomewhatless than
those originally published by Sheldrick and
Berthelot (3), butareveryclosetothosefound
by Delius and Clements frompartial
denatura-tion data (in press). Themapping information
also allowslimitstobeplacedonthelengths of
the redundant sequences
TRs/IRs,
IRL, andTRL.TheX. bad endg is about 6.5x 106daltons
andarises fromacleavagepointnearthe TRL/L
junction (Fig. 10).Thus, 6.5 x 106daltons isan
upperlimitfor TRLand IRL. Datanotpresented
inthisreportshows that there isanEcoRI site
withinTR5 togiveaterminal fragment of 3.5 x
106 daltons. Since the Hind IIIterminal
frag-mentm,arising fromacleavagesiteinS,is 4.5
x 106 daltons, these give the upperand lower
limits for
TRs
andIRS.
In the map shown inFig. 10, TRL/IRLand
TRs/IRs
havebeen drawnas 6 x 106daltonsand4 x 106daltons,
respec-tively. These estimates are verysimilartothe
figures of 6 and 4% for TRL/IRL and
TRs/IRs
derived byClementsandDelius(in press) from
their complete
paltial-denaturation
maps ofHSV-1 DNA. Since there issomeindication (15)
that thetrueterminalredundancy of 0.5 x 106
daltons (2, 3, 10)mayberepeatedin the
inter-nal inversions of
TRs
and TRL, this sequencehas beenindicated in themapsshown in Fig. 10
as a separate division within each redundant
region.
Therestrictionenzyme mapsof HSV-1 DNA
for Hind III,Hpa-1, andX. bad shown inFig.
10shouldform the basis for furtherrestriction
endonuclease maps. Additional mapping work
should also eliminatetheremainingminor
un-certaintiesintheHpa-1maps.
ACKNOWLEDGMENTS
Thisworkwas carriedout at theCold Spring Harbor Laboratory duringtemporary leave of absence from the Institute ofVirologyinGlasgow. IamgratefultoJ.Watson formaking availablethefacilities at the ColdSpring
Har-borLaboratory andtoJ.Sambrook formanyhelpful discus-sions. JanetArrandand RitaCortiniprovidedinvaluable assistance.
LITERATURE CITED
1. Clements, J. B., R. Cortini, and N. M. Wilkie. 1976.
AnalysisofherpesvirusDNAsubstructurebymeans
ofrestriction endonucleases. J. Gen. Virol. 30:243-256.
2. Grafstrom,R.H.,J. C.Alwine,W. L.Steinhart,and C. W. Hill.1974.Terminalrepetitionsinherpessimplex J. VIROL.
2
3
HindIII *
Hpa A
Xbad
4
on November 10, 2019 by guest
http://jvi.asm.org/
PHYSICAL MAPS FOR HSV-1 DNA
233
virustype 1DNA. ColdSpring Harbor Symp. Quant. Biol. 39:679-681.
3. Grafstrom,R.W.,J.C.Alwine,W. L.Steinhart,C. W. Hill,and R. W. Hyman. 1975. The terminal repeti-tion of herpessimplex virus DNA. Virology 67:144-157.
4. Hayward, G. S., N. Frenkel, and B. Roizman. 1975. Anatomy of herpes simplex virus DNA: strain differ-encesandheterogeneityinthelocation of restriction endonuclease cleavage sites. Proc. Natl. Acad. Sci. U.S.A. 72:1768-1772.
5. Hayward, G. S., R. J. Jacob, S. C.Wadsworth,and B. Roizman. 1975. Anatomy of herpes simplex virus DNA:evidence for four populations ofmoleculesthat differ in the relativeorientationsof theirlongand short components. Proc. Natl. Acad. Sci. U.S.A. 72:4243-4247.
6. Hyman, R. W., S. Burke, and L. Kudler. 1975. A nearby inverted repeat of the terminal sequence of herpessimplex virus DNA. Biochem. Biophys. Res. Commun. 68:609-615.
7. Maniatas, T., A. Jeffrey, and D.G.Kleid.1975. Nucleo-tide sequences of the rightward operator ofphage lambda. Proc. Natl. Acad. Sci. U.S.A. 72:1184-1188. 8. Roizman, B., M. Kozak, R. W. Honess, and G. Hay-ward.1974. Regulation ofherpesvirus macromolecu-lar synthesis:evidence for multilevel regulation of herpes simplex 1 RNAand protein synthesis. Cold Spring HarborSymp. Quant.Biol. 39:687-701.
9. Sharpe,P.A., B.Sugden,and J.Sambrook.1975. De-tectionoftwo restrictionendonuclease activities in Haemophilus parainfluenzae using analytical aga-rose-ethidium bromide electrophoresis. Biochemistry 12:3055.
10. Sheldrick, P.and N. Berthelot. 1974. Inverted repeti-tions in thechromosome of herpes simplex virus. Cold Spring Harbor Symp. Quant. Biol. 39:667-678. 11. Skare,J., W. P. Summers, and W. C. Summers. 1975.
Structure and function of herpesvirus genomes. I. Comparisonoffive HSV-1 and two HSV-2 strains by cleavage of their DNA with EcoRI restriction endonu-clease. J. Virol. 15:726-732.
12. Southern,E. M. 1975. Detection of specific sequences among DNAfragments separated by gel electropho-resis. J. Mol.Biol. 98:503-518.
13. Wadsworth,S., G. S. Hayward, and B. Roizman. 1976. Anatomy of herpes simplex DNA. V. Terminally re-petitive sequences. J.Virol. 17:503-512.
14. Wilkie, N. M., J. B. Clements, J. C. M. Macnab, and J. H.Subak-Sharpe. 1974. The structure and biological properties of herpes simplex virus DNA. Cold Spring Harbor Symp. Quant. Biol. 39:657-666.
15. Wilkie,N.M.,and R. Cortini. 1976. Sequence arrange-ment inherpes simplex type 1DNA: identification of terminal fragments in restriction endonuclease di-gestsand evidence for inversions inredundant and unique sequences. J. Virol. 20:211-221.
VOL. 20, 1976