Copyright©1977 AmericanSocietyforMicrobiology Printed in U.S.A.
Control of Protein
Synthesis
in
Semliki Forest Virus-Infected
Cells
BAT-EL LACHMIl* AND LEEVI KAARIAINEN
DepartmentofVirology,Universityof Helsinki, SF-00290Helsinki29, Finland Receivedfor publication 3 August 1976
Protein synthesis in Semliki forestvirus-infected chicken embryo cells was
studied by labeling them with [35S]methionine for short periods at different
timesafter infection,with orwithout synchronization ofproteinsynthesisbythe
hypertonic block technique. The rate ofhost-cell protein synthesis declined
almost linearly in inverse correlation to the increase in the amount of
virus-specific RNA. At5.5hpostinfection, the host-cell proteinsynthesiswasreduced
byabout 70%. The viralstructuralproteins weredetectablewithcertaintyat3.5
h postinfection, and their rate of synthesis increased linearly parallel to the
amount of their messenger, the 26S RNA. This suggests that the rate of
synthesisof the structuralproteins is controlledatthe level oftranscription.The
rate of synthesis of the virus-specific nonstructural proteinsattained its
maxi-mum between 3 and 4 h postinfection and declined thereafter, whereas the
amountof theirmessenger, the42SRNA, continuedtoincreaselinearlyinthe
cells.Thus,the messengeractivityofthe42S RNA is reducedinthelatephaseof
infectioncompared with its activity inthe earlyphase.
The mode of expression ofthe genetic
infor-mation ofSemliki forest virus 42S RNA
(molec-ularweight, 4.0 x 106to 4.5 x 106) (19, 26) has
recently been elucidated to alarge extent.The
fourstructuralproteins(capsidprotein and the
glycoproteins El, E2, and E3) are translated as
apolyprotein with a molecularweightofabout
130,000 (16, 17). The messenger for the
struc-turalpolyproteinis theintracellular 26S RNA
(molecular weight, 1.6 x 106) (26), which is a
replicaof the 3' third of the 42S RNA (38). The
translation startsfromoneinitiation site ashas
beenshown by several investigators (5, 7, 9).
Four nonstructural proteins with molecualr
weights of70,000 (ns7O), 86,000 (ns86), 72,000
(ns72),and60,000(ns6O)wererecentlyshownto
be synthesized sequentially when a
tempera-ture-sensitive mutant (ts-1) of Semliki forest
virus was studied (18) by the hypertonicblock
technique (6, 24, 25). The four nonstructural
proteins were apparently synthesized as a giant
polyprotein with a molecular weight of about
290,000, which is presumably cleaved to give
two short-lived intermediates with molecular
weights of 155,000 (nsl55) and 135,000 (ns135)
(18). The messenger for the nonstructural
pro-teins is probably the viral 42S RNA genome,
whichinvitrodirects thesynthesisofproducts
identical to the nonstructural proteins isolated
' Present address:Department ofVirology,Israel Insti-tuteforBiologicalResearch, Tel-AvivUniversityMedical School, P.O.B. 19,Ness-Ziona,Israel.
from ts-1-infected cells (N. Glanville,B.Lachmi,
A. E. Smith, and L.Kaariainen, submitted for
publication). Support for the idea that the
non-structural proteins are made as a polyprotein
has also been obtained from in vitro studies, in
which the 42S RNA-directed product was
shown toyield only oneformyl-[35S]methionine
tryptic and pronase peptide, suggesting that there is only one active initiation site for the nonstructural proteins in this RNA (9).
Here we have studied the synthesis of the
structural and nonstructural proteins in
Sem-liki forest virus wild-type-infected cells. The
rate of synthesis ofthe nonstructural proteins
was maximal early in infection and declined
thereafter, despite the continuous increase in
the amount of the 42S RNA. Bycontrast, the
rate of synthesis of the structural proteins in
thecells increasedparalleltothe amountof 26S
RNA showing that the synthesis of structural
andnonstructuralproteins is controlledby
dif-ferent mechanisms.
MATERIALS AND METHODS
Virus and cells. Semliki forest virus prototype strain (13)and a ts-1 mutant (15, 16) were used in these studies. BHK-21 cells and secondary special
pathogen-freechicken embryo fibroblasts were cul-tivatedasbefore (16). All experiments werecarried outat39°C,therestrictive temperature,in the pres-enceof1 ugofactinomycin D per ml.
Labeling of virus-specific proteins. Confluent monolayers ofsecondary chickenembryofibroblasts 142
on November 10, 2019 by guest
http://jvi.asm.org/
ALPHAVIRUS-INFECTED
wereinfectedat amultiplicityof 50 or 500 PFU per cell andincubated at390C. At the times indicated,
the cellswerepulsedwith[35S]methionine,50 or200
jCi
per plate (210 to 250 Ci/mmol, Amersham/ Searle Corp.) in a methionine-free medium. The pulses were followed by chases in the presence of 10-fold the normal concentration of unlabeled methio-nine. The whole cells were taken into hot 2% sodium dodecyl sulfate (SDS) as described before (16, 17). In some experiments the cells were incubated in the presence of 335 mM NaCl for 30 or 40 min before the pulse was given in isotonic medium as described(18). Protein determinations were carried out
ac-cording to Lowry et al. (20) using bovine serum albumin as standard.
Polyacrylamide gel electrophoresis. Polyacryl-amide gel electrophoresis was carried out by the
discontinuoussystemdescribedby Neville (23) and modified as before (17). Electrophoresiswas carried out inslabgelsat 10 mAfor 6 to 7 h. Thegelswere dried and autoradiographed. In some experiments after electrophoresis the gels were impregnated
withPPO (2,5-diphenyloxazole) andthen dried and
fluorographed (2). Forquantitation, theindividual lanes, taken from the slab, were cut into 1-mm slices, solubilized in NCS (Nuclear Chicago Solubi-lizer) and counted in a toluene-based scintillation fluid. The quantitations of the gels were done
ac-cordingtothe methoddescribedbyMcAllister and
Wagner (21). This method enabled us to determine the percentage ofreductioninhost-cellprotein accu-mulationand the accumulation of virus-specific pro-teins.
The region of each gel that does not contain knownvirus-specific proteins (fractions 5 to 15) was chosen to normalize activity. The normalization ra-tio (N), which indicates thedegree ofreduction of host-cellproteinsynthesis,wasdeterminedby divid-ing the sum ofradioactivityinfractions 5 to 15 from virus-infectedcells(I)bythe sum ofradioactivityin the same region from mock-infected cells (U), i.e., N
=I/U.
The virus-specific protein accumulation in each infected cell gel fraction (V) was determinedbythe equationi-NU,where iwastheradioactivityinan infectedgelfraction and U was theradioactivityin the corresponding mock-infected gel fraction, and where N is thereductioninhost-cell protein
synthe-sisand NU is the host-cellbackgroundestimated for theinfectedcellgelfraction.
Labeling of viral RNA. Confluent monolayers
wereinfected at amultiplicityof 500 PFUand incu-bated at39°C.Atthe end of theadsorptionperiod,10
,ACi
perplateof[3H]uridine(28Ci/mmol)wasadded toeachplate, and the incubation was continued. At 1-h intervals, from 1.5 to 5.5 h postinfection, one plate was taken into 2% SDS, and the extracts wereanalyzed in15to30% (wt/wt) sucrose gradients as described before (15).
RESULTS
Identification of Semliki forest
virus-in-duced proteins. The structural and
nonstruc-tural proteins synthesized in chicken cells
in-fected withour ts-1 mutant at39°C are shown
inFig. 1. As shown, the viral proteinsynthesis
recovered more quickly from hypertonic block
ofinitiation than did the host-cell protein
syn-thesis (6, 18, 24). Wetriedtoreduce the
resid-ualhost-cellprotein synthesistotheminimum
in infected cellsby incubating them in
hyper-tonicmedium (335 mMNaCl) for 40 min. When
the hypertonic medium is replaced by an
iso-tonic one, synchronous initiation follows (6).
Therefore, infected cellswerelabeled
immedi-155
135
86
72
U.701
p62
mw
C
-El
-C
A
B
FIG. 1. Autoradiograph of7.5% polyacrylamide
slabgel of cellular extractsfromts-1-infected chicken
embryo cellsfollowing synchronizationof initiation byahigh-saltblock. Infected cells were incubated at 39°C for4h 20min, treated with high salt (335 mM NaCl) for 40 min, and then pulsed for 15 min with [35S]methionine (100 uCi per plate) upon restoration
ofsalt concentrations andchased for 15 min (A) or 60 min(B)inthepresence of cold methionine. The slab gelwas impregnated with PPO and then dried and
fluorographedfor1 week. C, El, and p62 are
struc-tural proteins and 155 (ns155), 135 (ns135), 86 (ns86), and 70(ns7O) are nonstructural proteins.
VOL. 22, 1977
on November 10, 2019 by guest
http://jvi.asm.org/
[image:2.508.284.427.200.539.2]ately with [35S]methionine for 15 min and
chasedinthepresence of excess unlabeled
me-thionine for 15 min (lane A) and 60 min (lane
B). Thefastestmigrating band in the
autoradi-ogramofadiscontinuous SDS slab gel was the
viral capsid protein (C) followed by the
enve-lope protein (El) and the precursor of enveenve-lope
proteins E2and E3 (p62). Thebands migrating
more slowly than p62 were the nonstructural
proteins with apparent molecular weights of
70,000, 72,000 (previously determined 78,000
from itsmigrationin11%gels[16]), and 86,000.
These proteins weredesignated ns7O, ns72, and
ns86 according to the recommendation of an
international group of virologists (1).Inlane A,
which represents the 15-min chase, two
addi-tional bands with apparent molecular weights
of 155,000 (nsl55) and 135,000 (nsl35) can be
seen. These proteins were scarcely detectable
after the 60-min chase suggesting that they
were precursors forthe nonstructural proteins.
The fourth nonstructural protein, ns6O, could
notbe separated from p62 using the usual
poly-acrylamide gel electrophoresis system. It has
only been demonstratedby C-terminal labeling
aftertreatmentwithpactamycin (18).
We have recently shown by tryptic peptide
analysis that ns155 and ns135aredifferent
pro-teins, the former being the precursor for ns7O
and ns86 and the latter for ns72 and probably
ns6O (Glanville et al., submitted for
publica-tion).
Nonstructural proteins in Semliki forest
vi-rus wild-type-infected cells. The function of
the more stable nonstructural proteins is not
yet known. SinceSindbis virus RNA-
tempera-ture-sensitive mutants fall into four
comple-mentation groups (3, 33), one would expect at
least some of the nonstructural proteins to be
involved in the replication of the viral RNA.
Since RNA synthesis is an earlyevent inthe
infectious cycle, the presence of the
nonstruc-tural proteins in wild-type-infected cells was
studied mainly during the early phase of the
growthcycle.
Wild-type-infected cells were labeled at
dif-ferenttimes postinfection both with and
with-out hypertonic salt treatment. The
autoradio-.
6
-,. ... S-*
86
86
L.-.r
I \
{-1
PfnE,#
; it _W
;_I0,o
*rC_3/1 >
__awL
"Q.",
..wj
4^EI
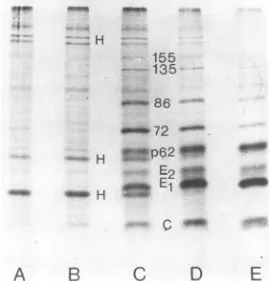
FIG. 2. Autoradiographof 7.5%discontinuousSDS-polyacrylamide gel of cellular extracts from Semliki forestviruswild-type-infectedandmock-infected cells. Infected chicken embryo cellswereincubatedat39°C
andat2.5h(A),3.5h(C), 4.5 h (D),and5.5h(E); one platewas treatedwithhigh salt (335mMNaCl) for 30min.The cellswerepulsedimmediately with[35S]methionineinisotonicmediumfor20minandchasedfor
5 min. Mock-infected cells (B) were not treated with high salt but were pulsed and chased similarlyat 5 h postinfection. Each sample applied to thegel contained about 180,000 cpm. C and El are the structural proteins. p62 is aprecursorofenvelope proteins E2 andE3. 72, 86, 135, and155 are the nonstructural proteins.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:3.508.81.436.332.585.2]graph ofa polyacrylamide slab gel in Fig. 2
shows the [35S]methionine-labeled proteins of
Semliki forest virus at 2.5, 3.5, 4.5, and 5.5h
postinfection. The infected cells were treated
with the hypertonicmedium for30 minprior to
the 20-minpulse given inisotonicmedium
fol-lowed bya5-minchase. Inadditionto theabove
describedstructuralproteins,theenvelope
pro-tein E2 is seen between p62 and El (Fig. 3).
The nonstructuralproteinsns155 and ns135are
clearly identifiable,as are ns86, Ms72,and ns7O.
Similar labeling carried out with cells not
ex-posed to the high salttreatment is shown in
Fig.3.The presenceofthe abovedescribed
non-structural proteins among host-cell proteins
canbeseenclearly at3.5hpostinfection (lane
C) with the exception of ns7O, which is not
properlyresolved from ns72.
Individual lanes of the dried slab gels were
cut into 1-mm slices and then solubilized and
counted in liquid scintillator (Fig. 4). To
esti-mate the proportions of viral structural and
nonstructuralproteins, the difference analysis
method of McAllister and Wagner was used
(21).Aregionofthe gel fromvirus-infected and
mock-infected cells (fractions5 to 15 in Fig. 4)
was chosen for evaluation of the reduction in
host-cell-specific protein synthesis. The total
radioactivityinthis region ofgels frominfected
cells was divided by the radioactivity in the
same region ofmock-infected cells. This
nor-malizationratioexpresses thedegreeof
inhibi-tionofhost-cellproteinsynthesis(Fig. 5A).The
multiplicationof theradioactivityin eachof the
mock-infected gel fractions by this ratio gave
the value ofresidual host-cell protein
synthe-sis. This value was then subtracted from the
radioactivity in the corresponding fractions of
thevirus-infected gelsto givethevirus-specific
radioactivity. The amount ofvirus-specific
ra-dioactivity atthe positions ofcapsid, envelope
proteins El, E2, and p62 were summed up to
yield the total radioactivity of structural
pro-teins synthesizedatdifferent timesafter
infec-155
^-
135
_72
l#*-At
p
Q2
sa
E2
El
El
C-,l
A
B
C
D
E
FIG. 3. Autoradiograph of7.5%discontinuousSDS-polyacrylamide gel ofcellularextractsfromSemliki
forestvirus-infectedandmock-infectedcells.Infectedchickenembryocellswereincubatedat39°Candat2h 10 min(A),3 h10min(C),4h 10 min(D), and5h 10 min(E)postinfection; theywerepulsed for20min with 50,Ci of[35S]methionineperplateandthen chasedforanadditional5min.Mock-infectedcells(B)were
pulsedandchasedsimilarlyat5h10minpostinfection. The slabgel wasdried andexposed for2 days.
Symbolsand amountsof radioactivityare asfor Fig.2. The mainhostproteinsaremarked withH. 40maw
H
on November 10, 2019 by guest
http://jvi.asm.org/
[image:4.508.121.392.310.597.2]94-x
U3-I
2
10 20 30 40 50 6 70 80 90 100
FRACTION NUMBER
8 B 155135 66 7n 62 E2 El C
7-
6-
5-o-
AU3
2-10 20 30 40 50 60 70 80 90 100
FRACTION NUMBER
FIG. 4. Quantitationofpolyacrylamide gels representingwild-typevirus-infectedcellsat3.5h(A)and5.5
h(B)postinfection, representinglanesC and EinFig. 3, respectively.Theprofile ofthequantitation of
mock-infectedcells (lane B inFig.3) isdrawnfor reference. Thearrowsshow thepositionsofnonstructural155
(nsl155),135(ns135),86(ns86), 72(ns72),and structuralproteins El,E2and C.Symbols: 0,virusinfected;
mock-infected.
tion. The result isexpressed asthe percentage
of totalradioactivity recovered from thegelsin
Fig. 5C.
Similarly, thevirus-specificradioactivitiesat
the positions of ns155, ns135, ns86, and ns72
plus ns7O were summed upto yield the total
amount of nonstructural proteins synthesized
duringthe 20-min pulse. In this quantitation
the fourth nonstructural protein ns6O, which cannot be distinguished from p62 by the gel
electrophoresis method used, was included in
the structural proteins. Thus the amount of
on November 10, 2019 by guest
http://jvi.asm.org/
[image:5.508.116.401.68.546.2]z
rj60-20
2 4 2 4 2 4
HOURS
FIG. 5. Synthesis rate ofhost-cell proteins (A), virus-specific nonstructuralproteins (B), and struc-turalproteins (C) compared with theamountof42S RNA (B) and 26S RNA (C)atdifferent timesafter infection. Lanes of the slabgelshowninFig.3were
cut into 1-mm slices and then solubilized and
counted in a liquid scintillator. The radioactivity
profile from mock-infected cells, showninFig. 4,was
usedtocalculate the degree of inhibition of host-cell protein synthesis and for the estimation of virus-specific nonstructural and structural proteins
ac-cording to the difference analysis method of Mc-Allister and Wagner(21).The radioactivities ofhost-cellproteins and viral nonstructural and structural proteinsarepresented aspercentageof the total
ra-dioactivity recovered from the gels. The cumulative amountof 42S RNA (B) and 26S RNA (C)at differ-enttimesofinfectionareshown for comparison.
La-beling of cells with[3Hiuridine was carriedoutas
describedinthelegendtoFig.6.Symbols:0,rateof
proteinsynthesis by the difference analysis method;
A, residual host protein synthesis determined by
dividingthesumofradioactivityinfractions5 to 15 (see Fig. 4) of infectedcells byrespectivesumfrom
uninfected cells x 100; 0, percent radioactivity in
42SRNA and in 26S RNA oftotal at 5.5 h post-infection.
radioactivity inthenonstructural proteins
rep-resentsanunderestimate(Fig. 5B). Therestof
theradioactivity inthe gelswastaken ashost
proteins givinganother estimate for the
resid-ual hostprotein synthesis (Fig. 5A).
There isaprogressive decreaseinthe rate of
host-cellprotein synthesis and asimultaneous
increase in therateofsynthesisofthe
nonstruc-turalproteins. Themaximum rate ofsynthesis
of the nonstructuralproteinsis reached at 3.5 h
postinfection, ata timewhen host-cell protein
synthesis is inhibited only by about 20%. At
thistime the synthesisrates ofstructural and
nonstructuralproteinsarealmostequal. At 5.5
h postinfection, the synthesis rate of the
non-structural proteins is about half of the
maxi-mum. The difference analysis revealed that
1.4%of the totalradioactivitywasin
nonstruc-tural proteins at 2.5 h postinfection, whereas
nostructural proteinscould be detected atthis
time.
Amount ofvirus-specificRNAs. Cumulative
labeling of wild-type-infected cells with
[3H]-uridine starting from 1 h postinfection was
carried out in the presence of actinomycin D.
Part of the cultures were removed at hourly
intervals, and the RNAs were analyzed, after
treatment with SDS, by sucrose gradient
cen-trifugation as shown in Fig. 6. The
radioactiv-ity in the viral RNAs was taken to represent
the total amount of these RNAs in the cell (35).
The main RNA species are the 42S RNA
ge-nomeand26S RNA, whichweresynthesizedin
a molar ratio of close to 1 at all times after
infection. The increase in the radioactivity in both RNA species is almost linear beginning from 2.5 h postinfection (Fig. 5). The increase in
the rate ofsynthesis of the structural proteins
correlates well with the increase in the amount
of the 26S RNA; however, there does not seem
tobe asimple correlation between the rate of
synthesis of the nonstructural proteins andthe
amountof the42S RNA, which is regarded as
themessengerofthese proteins (5, 8, 9).
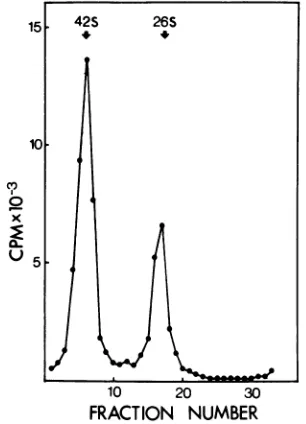
15 42S 26S
10
C,,
x
10 20 30
FRACTION NUMBER
FIG. 6. Sucrosegradient analysis ofRNAsfrom
Semlikiforestvirus-infectedcells.Secondarychicken
embryo fibroblasts were infected with 500PFU per cell and maintained in minimal essential medium
supplemented with 0.2% bovineserum albumin in the presence of1 Mgof actinomycin Dper ml. The cellswerelabeled with[3H]uridine(101iCiperplate) from1.0 to5.5hpostinfection. Thewhole cellswere dissolvedin 2%SDSin a buffer containing 0.15 M
NaCl, 0.001 MEDTA, and 0.01 MTris(TSE; pH 7.4). Theextract wasanalyzedin 15 to 30%(wtlwt) sucrosemadeinTSEcontaining0.1%SDS.
Centrif-ugation was for12 h at 24,000 rpm in an SW27
Spincorotor at230C.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:6.508.57.247.52.194.2] [image:6.508.281.434.320.532.2]148
LACHMI AND KAARIAINENDISCUSSION
The translation of Semliki forest virus-coded
proteins consists of two different processes. (i)
The structural proteins are translated concep-tually as a polyprotein. Due to the nascent
polysomal cleavage ofcapsid protein and the
rapid cleavage between envelope proteins El
and p62 (the precursor of E2 and E3), these
three proteins are the earliest detectable
prod-ucts of translation (6, 30). (ii) The
nonstruc-tural proteins areprobably also translated as a
polyprotein, which should have a molecular
weight to 300,000 (9, 18). Such a protein has not been found, which may indicate that a nascent
cleavage, similar tothatof the capsidprotein,
takes place, giving rise to the short-lived
pre-cursorsnsl55 and nsl35. The former gives rise
to two morestablepolypeptides, ns7Oandns86,
and the latter gives rise to polypeptides ns72
and ns6O (18; Glanville et al., submitted for
publication).
These nonstructural proteins have thus far
been demonstrated only in cells infected with
our temperature-sensitive mutant ts-1 (15-17)
where they accumulate in easily detectable
quantities (11, 12). Inthisstudy we have
dem-onstrated the presence of the nonstructural
pro-teins also inthe wild-type-infectedcells. Their
synthesis was followed during the early hours
of infection and reached its maximum rate at
3.5 h postinfection. At this time, they were
synthesized at a rate similar to that of the
structural proteins (Fig. 5). Estimates at 2.5 h
postinfection, although less reliable because of
the ongoing host-cell protein synthesis, would
suggest that more nonstructural than
struc-tural proteins were made at this time. All our
attempts todemonstratethesynthesis of either
nonstructural or structural proteins earlier
than 2.5 h postinfection have failed. It seems
that at least 10% of the amount of viral RNA
foundat 5.5 hpostinfection mustbepresented
before the proteins become detectable by the
methods used here. This already represents
several thousands ofnewly synthesized viral
RNAmolecules (35).
The fact that a given RNA can direct the
synthesisofaknownproductinacell-free
pro-tein-synthesizing system, together with the
finding that the same RNA is associated with
polysomes,has beenregardedassufficient
evi-dence ofitsroleas amessengerinthecell. The
alphavirus 26SRNAfulfills both thesecriteria
and is regarded as the cellular messenger for
the structural proteins (4, 6, 7, 8, 14, 22, 27, 31, 32, 37). According to the same criteria, the 42S
RNAshould be regarded asthe messenger for
the nonstructural proteins; it has been found
associated with the polysomesininfected cells
(22, 27, 31), and it stimulates the synthesis of
nonstructural proteinsinvitro (5, 8, 9,. 28). It
wastherefore interesting to correlate the rate
ofsynthesis of the structural and nonstructural
proteins with the amount of their respective
messenger RNAs. The rate of synthesis of the
structuralproteins and the amount of 26S RNA
rose with almostidentical kineticsthroughout
the infection (Fig. 5C), suggesting that the
amount of structural proteins synthesized is
dependent on the amount of their messenger
RNA.This situation is similar to thatfound for
the synthesis ofa-and ,3-globinchains in the
reticulocyte lysate (34).
The rate of synthesis of the nonstructural
proteins correlated poorly with the amount of
their 42S RNA messenger.The maximum rate
ofsynthesis of these proteins was observed at
3.5 hpostinfection when only about half of the
42S RNAfoundat 5.5 hpostinfection had been
synthesized (Fig. 5B). Aroughestimateof the
efficiency of42S RNAat 3.5and5.5h
postinfec-tion canbeobtainediftheamountof
radioactiv-ityincorporated into the nonstructural proteins
during the 20-min pulse is divided by the
amount of radioactivity accumulated in 42S
RNA from 1 hpostinfection. There is about a
sixfolddrop in the "messengerefficiency" of this
RNAbetween 3.5 and 5.5 h postinfection.
One apparent possibility to explain the
re-duced messenger activity of the 42S RNA is the
consumption of this RNA in the formation of
viral nucleocapsids (11, 29, 32), which would
makethe 42S RNA inactive as messenger. The
maximum rate ofsynthesisofthe nonstructural
proteins taking place at 3.5 h postinfection
wouldthen be the result of two processes: the
synthesis of enough 42S RNA and the lack of
structural proteins, i.e., capsid protein to
en-capsidate this RNA. Later, when the rate of
synthesis of the structural proteins increased,
all or almost all the newly synthesized 42S
RNA would become encapsidated, and only
smallamounts ofnonstructuralproteins could
betranslated dueto"leakiness" of the
encapsi-dation process. This would explain why the
bulkof42SRNAsynthesizedbetween4and5 h
postinfection isfoundinthe 140Snucleocapsid
(27, 29, 32, 36). The process of association of the
capsid protein with the 42S RNAisseemingly
fastand, at 5 to6 h, postinfection takes place
within 5 to 8 min after the protein has been
synthesized (29).
The mechanism of nucleocapsid assembly is
poorly understoodatpresent. Ithas been
dem-onstrated recently that the SFVcapsid protein
bindsrapidly the60Sribosomalsubunit inboth
infected cells (36) andinvitro(10). The
signifi-cance of thecapsid protein-60S complexinthe
nucleocapsid assembly (11) andinthe control of
on November 10, 2019 by guest
http://jvi.asm.org/
149
the translation of the nonstructural proteins is
under investigation.
ACKNOWLEDGMENTS
Wethank RitvaRajalaandMizjaSalonen for excellent technical assistance. Actinomycin D wasagiftfromMerck, Sharp and Dohme.
This work was supportedby grants fromtheSigrid Juse-lius Foundation and the Finnish Academy. B.L. is a recipi-entofascholarshipfrom the Finnish Ministry of Education.
LITERATURE CITED
1. Baltimore, D., D. C. Burke, M. C. Horzinek, A. S. Huang, L. Kiiriainen, E. M. Pfefferkorn, M. J. Schlesinger,S. Schlesinger, W. R. Schlesinger, and C. Scholtissek. 1976. Proposed nomenclature for al-phavirus polypeptides. J. Gen. Virol. 30:273. 2. Bonner, M. W., and R. Laskey. 1974. A film detection
method for tritium-labeled proteins and nucleic acids inpolyacrylamide gels. Eur. J. Biochem. 46:83-88. 3. Burge, B. W., and E. M. Pfefferkorn. 1966.
Comple-mentationbetweentemperature-sensitivemutants of Sindbis virus. Virology 30:214-223.
4. Cancedda,R., and M. J. Schlesinger. 1974. Formation ofSindbis virus capsid protein in mammalian cell-free extracts programmed with viral messenger RNA. Proc.Natl. Acad. Sci.U.S.A. 71:1843-1847. 5. Cancedda, R., L. Villa-Komaroff, H. Lodish, and M.
Schlesinger. 1975. Initiation sites for translation of Sindbis virus 42S and 26S messenger RNAs. Cell 6:215-222.
6. Clegg, J. C. S. 1975. Sequential translation of capsid and membrane proteins in alphaviruses. Nature (London) 254:454-455.
7. Clegg,J.C. S., and S. I. T. Kennedy. 1975. Initiation of the synthesis of the structural proteins ofSemliki forest virus. J. Mol. Biol. 97:401-411.
8. Glanville, N., J. Morser, P. Uomala, and L. Kkiri-ainen. 1976. Simultaneous translation of structural andnonstructural proteins from Semliki forest virus RNA in two eukaryotic systems in vitro. Eur. J. Biochem. 64:167-175.
9. Glanville,N., M. Ranki, J. Morser,L.Kaariainen, and A. E. Smith. 1976. Initiation of translation directed by 42S and 26S RNAs from Semliki forest virus in vitro. Proc. Natl. Acad. Sci. U.S.A. 73:3059-3063. 10. Glanville, N., and I. Ulmanen. 1976. Biological activity
of in vitro synthesized protein: binding of Semliki forest virus capsid protein to the large ribosomal subunit. Biochem. Biophys. Res. Commun. 71:393-399.
11. Kaariainen,L., S.Keranen, B. Lachmi, H.Soderlund, K. Tuomi, and I. Ulmanen. 1975. Replication of Sem-liki forest virus. Med.Biol. 53:342-352.
12. Kaariainen, L., B. Lachmi, and N. Glanville. 1976. Translational control in Semliki forest virus infected cells. Ann.Microbiol. (Paris) 127A:197-203. 13. Kaariainen, L., K. Simons, and C.-H. von Bonsdorff.
1969. Studies on subviralcomponents of Semliki for-estvirus.Ann. Med. Exp. Biol. Fenn. 47:235-248. 14. Kennedy, S. I. T. 1972.Isolation and identification of
thevirus-specific RNA species found on membrane-bound polyribosomes of chick embryo cells infected with Semliki forest virus. Biochem. Biophys. Res. Commun. 48:1254-1258.
15. Keranen, S., and L. Kairiainen. 1974. Isolation and basic characterization oftemperature-sensitive mu-tants fromSemliki forest virus. ActaPathol. Micro-biol.Scand. Sect. B82:810-820.
16. Keranen,S., and L.Kaariminen. 1975. Proteins synthe-sizedby Semliki forest virus and its 16 temperature-sensitivemutants. J. Virol. 16:388-396.
17. Lachmi, B., N. Glanville, S. Keranen, and L.
Kasri-ainen.1975.Tryptic peptideanalysisofnonstructural and structuralprecursorproteinsfrom Semliki forest virus mutant-infectedcells. J. Virol. 16:1615-1629. 18. Lachmi, B., and L.Kaariinen.1976.Sequential
trans-lation ofnonstructural proteins incells infected with Semliki forest virus mutant. Proc. Natl. Acad. Sci. U.S.A. 73:1936-1940.
19. Levin, J. G., and R. M. Friedman. 1971. Analysis of arbovirus ribonucleic acid forms by polyacrylamide gelelectrophoresis. J.Virol. 7:504-514.
20. Lowry, 0. H., N.J. Rosebrough, A. L. Farr, and R. J. Randall. 1951. Protein measurement with the Folin phenolreagent. J. Biol. Chem. 193:265-275. 21. McAllister, P. E., and R. R. Wagner. 1976. Differential
inhibition of host protein synthesis in L cells infected withRNA-temperature-sensitive mutants of vesicu-larstomatitis virus. J. Virol. 18:550-558.
22. Mowshowitz, D. 1973. Identification of polysomal RNA in BHK cells infected by Sindbis virus. J. Virol. 11:535-543.
23. Neville, D. M. 1971. Molecular weight determination of proteindodecyl sulfate complexes bygel electrophore-sis in adiscontinuousbuffer. J. Biol. Chem. 246:6328-6334.
24. Nuss,D. L., H. Oppermann, and G. Koch. 1975. Selec-tiveblockage of initiation of host cell protein synthe-sis in RNA-virus-infected cells. Proc. Natl. Acad. Sci. U.S.A. 72:1258-1262.
25. Saborio, J. L., S. S. Pong, and G. Koch. 1974. Selective andreversible inhibition of initiation of protein syn-thesis in mammalian cells. J. Mol. Biol. 85:195-211. 26. Simmons, D. T., and J. H. Strauss. 1972. Replicationof
Sindbis virus. I. Relative size and genetic content of 26S and 49S RNA. J. Mol. Biol. 71:599-613. 27. Simmons, D. T., and J. H. Strauss. 1974. Replication
of Sindbis virus. V. Polyribosomes and mRNA in infected cells. J. Virol. 14:552-559.
28. Simmons,D. T., and J. H. Strauss. 1974. Translation of Sindbis virus 26S RNA and 49S RNA in lysates of rabbit reticulocytes. J. Mol. Biol. 86:397-409. 29. Sdderlund,H. 1973. Kinetics of formation of the
Sem-liki forest virus nucleocapsid. Intervirology 1:354-361.
30. Soderlund,H. 1976. The post-translational processing of Semliki forest virus structural proteins in puromy-cintreatedcells. FEBS Lett. 63:56-58.
31. Soderlund, H., N. Glanville, and L.Kiiridinen.1973/ 74. Polysomal RNAs in Semliki forest virus-infected cells. Intervirology 2:100-113.
32. Soderlund, H., and L.Kiiriainen.1974.Association of capsid protein with Semliki forest virus messenger RNAs. Acta Pathol. Microbiol. Scand. Sect. B 82:33-40.
33. Strauss, E. G., M. Lenches, and J. H. Strauss. 1976. Mutants of Sindbis virus. I. Isolation and partial characterization of 89 new temperature-sensitive mu-tants.Virology 74:154-168.
34. Temple,G., and H. F. Lodish. 1975. Competititon be-tween alpha-globin and beta-globin messenger-RNA. Biochem. Biophys. Res. Commun. 63:971-979. 35. Tuomi, K., L. K11riainen, and H. Sdderlund. 1975.
Quantitation of Semliki forestvirusRNAs in infected cells using32P equilibrium labeling. Nucleic Acids Research2:555-565.
36. Ulmanen, I., H. S6derlund,and L. Kaariainen. 1976. Semliki forest virus capsid protein associates with the608ribosomal subunit in infected cells. J. Virol. 20:203-210.
37. Wengler, G., M.Beato, and B. A. Hackemack. 1974. Translation of 26Svirus-specificRNA from Semliki forest virus-infected cells in vitro. Virology 61:120-128.
38. Wengler, G., and G. Wengler. 1976. Localization of the 26S RNA sequence on the viral genome type 42S RNA isolated from SFV infected cells. Virology73:190-199. VOL. 22, 1977
on November 10, 2019 by guest
http://jvi.asm.org/
![FIG.1.NaCl)gel39°Cfluorographedofslabembryo[35S]methionine(ns86),minturalbysalt a Autoradiograph of 7.5% polyacrylamide gel ofcellular extracts from ts-1-infected chicken cells following synchronization of initiation high-salt block](https://thumb-us.123doks.com/thumbv2/123dok_us/1546762.107204/2.508.284.427.200.539/cfluorographedofslabembryo-methionine-minturalbysalt-autoradiograph-polyacrylamide-ofcellular-synchronization-initiation.webp)