0022-538X/81/120772-18$02.00/0
Sizes of Bacteriophage T4 Early mRNA's Separated by
Preparative
Polyacrylamide Gel Electrophoresis and Identified
by In Vitro Translation and by
Hybridization
to
Recombinant
T4
Plasmids
ELTON T. YOUNG* AND ROSEMARY CRONE MENARD
Department of Biochemistry, University ofWashington, Seattle, Washington98195
Received 13February 1981/Accepted1July 1981
We determined thesizes ofspecific T4prereplicative mRNA's by preparative
polyacrylamide gel electrophoresis, andweused thefollowingtwotechniques to
identify specific gene transcripts: cell-free protein synthesis accompanied by
sodium dodecyl sulfate-polyacrylamide gelelectrophoresistodistinguish T4 poly-peptidesandhybridizationtorecombinant plasmids containing T4 DNA of known genetic composition. In our first analysis, the use of nonsense and in-phase
deletion mutants allowed unambiguous identifications of the functional
tran-scripts that encoded genes 32, rIIB, and rIIA. In addition, we identified the
functional transcripts that encoded genes 43, 45, 30, 39, and 52, the ,B-glucosyl
transferase gene, and the deletion293 region. A single peak of mRNA activity
that coded for gp43, gp39, gprIIA, /3-glucosyl transferase, and the polypeptide
encoded in the deletion 293 region was present; the other polypeptides were
encoded inmultiplemRNAspecies. gp46andgp32wereencodedbytwomRNA's,
andgp52andgprIIBwereencodedby three mRNA's. By hybridizing fractionated,
pulse-labeled earlyRNA to cloned restrictionfragmentsof T4DNA,weidentified
the same specific transcripts for genes 43, 52, and nIB. In addition, a
lower-molecular-weight RNA (presumably degraded mRNA)waspresentevenin
pulse-labeled RNApreparations. The distribution ofpulse-labeled RNAs that hybrid-izedto gene39,gene30, generIIA,gene40plusgene41,andgene42plus the
,B-glucosyl transferase gene indicated extensive degradation. We detected
cotran-scriptionofgenesrIlA and rIIB by rehybridization of RNA first annealed toan
rIIB plasmid and then eluted and annealed to anrIlA plasmid. The size
distri-butions of normalandchloramphenicol-treated RNAsthathybridizedtoplasmids
containing T4immediate earlygene30, gene39, gene40plusgene41, andgene
42plusthe f3-glucosyl transferase gene werenotsignificantly different. An understanding of the arrangement of T4
prereplicative transcriptionunits wouldincrease
our knowledge of the regulation of T4 early genes significantly. The current model of this arrangement (28) proposes that
chlorampheni-col (CAM)-resistant immediate early genes (transcriptsaresynthesizedafter infectionby T4 in the presenceofCAM)arepromoterproximal
to CAM-sensitive delayed early genes (tran-scriptsare notsynthesizedafter infection in the presenceofCAM)andthatquasi-late or middle-mode genes are regulated differently than im-mediateearly and delayed early genes. The
fac-torsthatinfluence the transition from
immedi-ate early transcription to delayed early tran-scription andsubsequentlytomiddle-mode
tran-scription are poorly understood. The Esche-richia coli termination factor rho is thought to be involved in the shift from immediate early
transcription to delayed early transcription (3,
5), and the T4 motgene productisinvolved in the expression of at least some middle-mode genes (18,20,27).
Various aspects of the model summarized
above have beentested. Themostdirect
predic-tionsarethat the sizes ofsomeimmediateearly transcriptswouldincreaseifdelayedearly
tran-scription were allowed and that
delayed early
transcripts would be the distalportionsof
poly-cistronic transcripts whose
promoter-proximal
portions would be immediate early transcripts.
This proposition was tested for the D2 region
(33) andfor the internalproteinsIPIIandIPIII,
buttheresultswereequivocal;the D2 RNA size
wasinvariant, whereasthe IP RNA seemedto
besomewhatlargerinthe absence of CAMthan
thepresenceof CAM.
Ingeneral, thephysicalsizes ofT4transcripts
77'2
on November 10, 2019 by guest
http://jvi.asm.org/
T4 EARLY mRNA SIZES 773
have beendifficulttostudybecauseof the labil-ity of the RNAs. The T4 tRNA'sare an
excep-tion to this. The presumed primary transcript synthesized in vitro has beenidentified,and the processing of this transcriptto mature tRNAs' has been studied (10). A similar study of T4 mRNA's that code for early proteins has not
been performed, although such a study would clearlybe relevanttoanymodel of regulation of
prereplicative transcription. In a few instances
the size of the functionaltranscript that codes
for an early protein has been identified by its ability to code foran enzymatic activity or an immunologically active polypeptide (1, 29). For the nonessential rII genes and the adjacent D
region, the RNA has been identified by deletion hybridization analyses (31, 32, 39). Inthis case
there is a discrepancy concerning the physical
sizes of the rIIA and B transcripts (33, 39). Wedetermined the sizes ofspecific T4 early transcripts bypolyacrylamidegel electrophore-sis. The RNAswereidentified after
electropho-resis by the following two techniques: in vitro translation and separation of the labeled poly-peptides by sodium dodecyl
sulfate-polyacryl-amide gel electrophoresis and hybridization of radioactive RNAtoclonedrestrictionfragments of T4 DNA. Analyses of functional early mRNA's revealed that a number oftranscripts
occurred as multiple species, which indicated
that thereweremultiplepromoters or
termina-tionsitesorthatpost-transcriptional processing
occurred. Some of these same multiple
tran-scriptscouldbeidentifiedbyhybridization, but the radioactive RNA species corresponding to
many genes were present as heterogeneous ar-rays ofsmallfragments,whichrepresented
de-gradedfragments of the active mRNA's.
MATERIALS AND METHODS
Biochemicals and media. Most reagents were purchased from standardsources.[3S]methioninewas preparedasdescribedbyCrawford andGesteland (6). Mouse RNAwas a gift from Ursula Storb, and 3H-labeled RNA from the midi-variant ofbacteriophage
Q8 (MDVRNA; 22)was agiftfrom FrankRyan.The
acrylamide used for gel electrophoresis was passed
through a membrane filter (Millipore Corp.) before
use.M9SandM9S.1media have beendescribed
pre-viously byBolleetal.(2).
Bacteria, phage, and plasmids. E. coli BE, a
nonpermissive host for ambermutants,wasusedas a
hostforallinfections. Plasmidsweremaintainedin E.
coli strain 802(suII+rk-mk+).
Thebacteriophagestrains withmultiplemutations
wereconstructedby standard phagecrosses(Table1).
TherIIA deletionmutationEM66andthe rIIB
dele-tionmutation 196have beencharacterized byA.Bolle
(personal communication). These mutations contain
wholly internal in-phasedeletions which correspond
to losses ofapproximately 300 and 100 amino acids
fromrIIA andrIIB, respectively.
The plasmids containing T4 restriction fragments
werefrom thecollections of Mattson et al. (19) and
Selzeretal.(34).Theproperties of these plasmidsare
shown in Table2.The T4restrictionfragments in the
original collection of Mattson et al. were cloned into
pCR1andweredesignated bytheprefix pVH. Mattson
and Van Houwe transferred these restriction
frag-ments to pBR322 and gave them new designations (the600 series), whichweused. Thedesignations of
the T4 recombinantplasmids that were derived from
the 600 series by cleavage with different restriction
enzymes than the enzymes used in the originalcloning
procedure contain asuffix letter (A, B, etc). Restriction
maps of the rII regions showing the origins of the
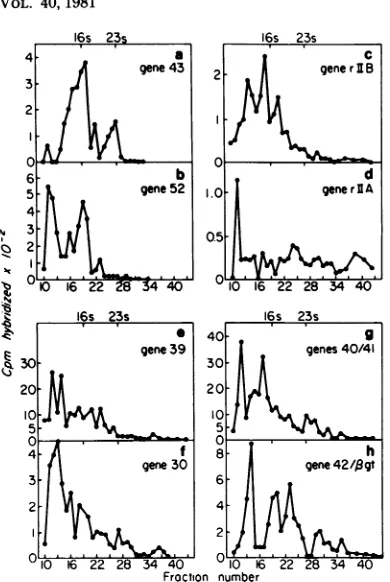
DNAfragments used in this work are shown in Fig. 1
(26a, 34).
TABLE 1. BacteriophageT4mutants
Mutant Description
32amA453 44amN82 Amino-terminal ambermutation ingene32 (no identifiable poly-peptide);nogene 44 polypep-tidedetected
32amA453 44amN82 rIIdell241coversall knownpoint rIIdell241 mutations inrIIAandrIIBand deletespartof theD region;no polypeptides from genes 32, 44, rIIA, and rIIB
32amH18 44amN82 18,000-dalton polypeptide from gene32;nogene 44polypeptide rIIBdell96 Internal, in-phase deletion ingene
rnIBmakinga22,000-dalton rIIB polypeptide
rIAdelEM66 Internal, in-phasedeletion in gene rIlA makinga55,000-dalton polypeptide
32amH18 44amN82 18,000-dalton gp32; 22,000-dalton rIIBdell96 gprIIB;nogp44 identified 32amH18 44amN82 18,000-dalton gp32;22,000-dalton
rIIBdell96 gprIIB;55,000-daltongprIIA;no rIIAdelEM66 gp44
TABLE 2. T4 recombinant plasmids
Plas- T4 Restriction
mid gene(s)a site(s)used Reference(s) mid
~~~for
cloning621 52 RI 19
622 43 RI 19
624 42,,fgt RI 19
625 30 RI 19
626A 39 HindIII 19;Young, unpub-lisheddata
627 40,41 RI 19
pTB35 rIIA HindIII 34
pABI rIIA HindIII, RI* 34;Selzer, unpublished data
pTB1O rIIAB HindIII 34
pABIV rIIB RI*,HindIII 34;Selzer, unpublished data
aNoothergenes havebeen identified by marker rescue tests.Plasmid621(gene 52)containsadditional DNA which hasnot beenidentified.Theplasmidscontaining genes 43, 30, 39, rIlA (pTB35 andpABI),and rIIB (pABIV) have been shownbymarker rescuetests tobewholly internal to the gene indicated.The T4rIIA and rIIBrestrictionenzymefragments usedin this work areshowninthe mapsinFig.1.Thelength of theT4fragmentinplasmid pTB1Ois 873 basepairs (26a).
VOL. 40,1981
on November 10, 2019 by guest
http://jvi.asm.org/
rIEA H
3B
H RI i RI RI- H RI W
GENE52 w RI
pTB I
[image:3.500.82.416.58.157.2]pTB10
FIG. 1. Mapsof T4 rIIA and rIIB restriction enzymefragments. The maps are drawn approximately to
scale. H, HindIII.
Nucleic acidpreparation.T4 DNA was prepared
frompurifiedT4 phage, and plasmid DNA was pre-pared by minor modifications of the cleared lysate procedure of Katz et al. (16), as described elsewhere
(42). Theethidium bromide and residual protein that
remained after centrifugation in CsCl were removed
by twophenolextractions.
Both radioactive RNA and nonradioactive RNA
were isolated from T4-infected E. coli BE cells, as
describedelsewhere (42).
Preparative electrophoresis of RNA on
poly-acrylamide gels. We used the preparative electro-phoresis apparatus of Hagen and Young (14), with modifications (13, 26). Denaturation of RNA samples with dimethyl sulfoxide before electrophoresis has
beendescribedpreviously(13, 26).Briefly,each RNA
sample dissolved in Tns-hydrochloride (pH
7.5)-EDTA wasminxedwith spectroscopy grade dimethyl
sulfoxidesothat the final concentrations in the sample
were90%dimethylsulfoxide, 10 mM Tris-hydrochlo-ride, and 3 mM EDTA. Then the sample was
incu-bated for5min at370C,cooled to room temperature,
andloadedonto agel. The percent recovery of RNA
activityfromthegelswasusually30to100%.
Cell-freeproteinsynthesis, electrophoresisof
polypeptides, anddensitometricanalysisof
au-toradiograms.AnS30extractwaspreparedfrom E.
coli MRE600 and used asdescribedpreviously (26).
Radioactive polypeptideswereanalyzedasdescribed
by Pachl andYoung (26).
Hybridizationtofilter-bound DNA. Filters
con-tainingimmobilizedT4phageorplasmidDNAwere
preparedand usedasdescribed elsewhere (42).Ifthe
annealed RNAwas tobe eluted foruseinasecond
experiment,the firstannealingwasperformedin 0.03
M sodiumcitrate-0.3 M sodium chloride(pH 7.0)-50%
formamide. The RNA was eluted by heating the
washedfilter in a solutioncontaining10mMTris(pH
8.8) and0.2mM EDTA for1minat1000Cand then
rapidly coolingthe solution. The filterwasremoved,
50
Ig
of tRNAwasadded,the solutionwasincubatedwith5ug ofRNase-free DNase for15minat300Cand
phenol extracted, and the RNA was recovered by
ethanolprecipitation (42).Elution of the RNA after
the filtersweretreated with RNasewasperformedas
describedelsewhere(42).
RESULTS
Sizes
offunctionalearly
transcripts.
RNAwas extracted 12 minafter infection ofE. coli
BE(su-)
with T4 32amA453 44amN82, asde-scribed above.(Ambermutants weredesignated bygenenumber, followed by alleledesignation; for example, the ambermutation A453isingene 32,and themutantcontaining thismutationwas
designated 32amA453.)About 400 ,ugofpurified RNA was denatured by treatment with 90% dimethyl sulfoxide before continuous-elution preparativeelectrophoresison apolyacrylamide gel (13) (Fig. 2). The radioactive
polypeptides
synthesized in vitro by the fractionated RNA
were analyzedby sodium dodecyl
sulfate-poly-acrylamide
gel electrophoresis. Theidentifica-tion of rIIB mRNA activity wasfacilitated by
using phage containing nonsense mutations in genes 32and44andinfectingansu-host. This allowedgprIIBtobeidentified andquantitated in the absence of comigrating gp32 and gp44
(23).(gprIIB,gp32,andgp44 arethe polypeptide products ofgenesrIIB,32,and44,respectively.)
Using
the double amber mutant had anaddi-tional advantage. Because bothgp32 and gp44 wererequired forDNAsynthesis and hence late
T4 RNA synthesis, this mutant did not make
any late RNA which could interfere with the identification ofearly mRNA species. Figure 2
showsanautoradiogram of the dried gel. Without
considering
theidentity
of the indi-vidual proteins synthesized by the fractionated RNA,weobservedtwodifferenttypesof behav-iorfor the mRNAspecies identified by in vitro translation. Some mRNA'smigrated
at one ormore
unique
rates,asshownby
thepresenceofmRNA activities for their cognate proteins in
one or more unique sets ofadjacent fractions collectedfromthegel. Other mRNA'sappeared
to bepolydisperse; their activities were spread rather continuously across the
gel.
For someRNA
species
this"tailing" appeared
to resultfromamajor
peak
whosemRNAactivity trailed throughout theremaining
fractions.However,in other casesthemRNA activitywasdistributed fairlyuniformly
over abroadrange ofmolecular weights. The polydispersity could have arisenfor one ormore reasons, including
incomplete
washing from the elution chamber,
aggregation
of theRNA,
physical
sizeheterogeneity
of the mRNA, and comigration of differentpolypep-- pTS35 b Asi
.0.
pABI
on November 10, 2019 by guest
http://jvi.asm.org/
T4 EARLY mRNA SIZES 775
V . a 9 - - _1-_. -.
A~~~~~~~~~~f
FIG. 2. Autoradiogramof T4 early polypeptides synthesized byfractionated32-44-RNA. A400-pgsample
of RNA isolated12minafterinfection of E. coli BE with T4 32amA453 44amN82 was denatured with 90%
dimethylsulfoxide for3minat37°C and thenfractionatedon apreparative gel (diameter, 5 cm)containing
2.25% acrylamide.TheRNAwaselectrophoresed at 15 mA for11 h, and the eluted RNA was concentrated by
ethanolprecipitation and then translated inacell-freeprotein-synthesizing system containing
[3S]methio-nine. The radioactivepolypeptides wereseparated by electrophoresis on a discontinuous sodium dodecyl
sulfate gel system containing equal volumes of 12.5, 10, and 7.5%acrylamidelayered sequentially over one
another. Thestacking gelcontained 3%acrylamide. After electrophoresis for3hat25mA, thegelwastreated
withdimethylsulfoxideandsubjectedtofluorographyasdescribed in thetext. Thepolypeptides synthesized
in vitrofromunfractionatedRNAs derivedfromcellsinfectedwithphagesofthe genotypesindicated above
theslotsareshownatthesidestoallowidentificationof specific T4polypeptides. The molecular weight scale
atthe topwasderivedfrom the 16S and 23S rRNA's present on this gel and from these and additional
molecularweightstandardsseparatedonanidentical gel (seeFig. 8). The two gelsusedin this analysis did
notseparatethe T4polypeptides identically. gp43 was the upper band in both gels. wt, Wild type.
tidesencodedby mRNA's of different but
over-lapping sizes. For those mRNA's which hadan
intensepeak ofactivity followed by trailing,we
assumed that the apparent
heterogeneity
wasprimarilyanartifact caused
by incomplete
elu-tion from the elution chamber and efficient translation of thelate-eluting
fractions in the cell-freesystem, as hasbeen observed with T7 mRNA(25, 26).Several proteins could be identified
readily
aftertranslation of the fractionated
RNA,
basedontheirrelative
electrophoretic
mobilities,
theirknownmolecular
weights,
andanalyses
ofambermutants (24, 40). The
autoradiogram
inFig.
2shows thegenes thatcoded for these
proteins;
they includedgene43,gene46,gene 30
plus
gene 39, gene 52,gene32, generlIB,
andgene IPIII.gp3O
and gp39comigrated
onthegel
shown inFig. 2;in otheranalyses (see Fig. 6) these gene
products were separated from each other, and
their individual mRNA activities could be iden-tified. The RNA forrIIA couldnotbeidentified
unambiguously.
Therewere tworeasonsforthis.gprIIA wassynthesized poorlyinvitro,and two different mRNA fractions coded for a protein
that wasapproximatelythesamesize asgprIIA.
These two mRNA species were identified in
fractions34 to 36 and39 to 60 of the
autoradi-ogramshowninFig.2.Theprotein encoded by themRNA that eluted in fractions34 to36had
the same
mobility
as thepolypeptide
that wasencoded
by
the deletion 293 (del 293) region (24).gprIIB appearedtobeidentified unambig-uouslyonthegel sincenogp32wasmade when unfractionated RNAwasused(Fig.
2).However,even
partial suppression
of the amino-terminalgene 32 amber mutation (A453) could cause
synthesis
ofenough
gp32 toallow ittobecon-fused with
gprIIB.
To overcome thisproblem
and to
identify
the mRNA forrILA,
weper-formedtwoexperiments.Inthe firstexperiment RNA from the
triple
mutant T4 32amA453 44amN82rIIdell241wasanalyzed
after fraction-ation on apreparative
polyacrylamide gel.
In this mutant both rIIA and rlIB were deleted completely.The translationproductswereana-lyzedasdescribedabove (Fig. 3).NorIIB poly-peptide wasfound on the gel, whereas mostof
the otherpolypeptides with molecular weights
less than 60,000 whichwerepresentontheFig.
2 gel were identified, and their mRNA's
mi-grated at
approxiimately
the same rates as onthegelshown inFig.2.Anexceptiontothis was the gene 52 message activity (see below). This
VOL. 40,1981
on November 10, 2019 by guest
http://jvi.asm.org/
[image:4.500.50.448.66.246.2]FIG. 3. Identification of rIIB message activity by analysis of RNA from anrIIdeletionmutant. RNA was extracted 12 minafterinfection of E. coliBEby T4 32amA453 44amN82rIIdell241andanalyzed as described
in thelegendtoFig.2andinthe text. The polypeptidessynthesized in vitroby unfractionated RNA isolated
12minafter infection by wild-typeT4 (T4 wt) T4 32amA453 44amN82, and T4 32amA453 44amN82rIIdell241
areshown in the threeslots on theleft. The elution positions of the 16S and 23S rRNA markers are indicated
onthe top ofthe autoradiogram. The open circle on the figure indicates the expected position of therIIB polypeptide.
was additional evidence that the polypeptide
labeledgprIIB in Fig. 2 wasthe product of the rIIB gene. Unfortunately, nopolypeptides with molecular weights of more than 60,000 were made in thesecond experiment. Thus,the
pre-sumptive rIIA message in Fig. 2 could not be
identified because neither of the two potential
rIIApolypeptideswassynthesized.
Only the middle-sized (Mr, 1.2 x
106)
gp52 message activity was present in the RNAiso-lated after infection with the deletion mutant
rII1241, whereas three peaks of gp52 message
activity were detected when RNA from wild-type T4 was analyzed. The right endpoint of del1241 was estimated to be 0.7 kilobase from the right or 3' end of rIIB (7, 8). Since it has beenestimated that rIlB and gene 52 are sepa-rated by about 3.5 kilobases (7, 40), the right endpoint ofdel1241 is about2.8kilobasesfrom gene52.None of the gene52messagesobserved waslarge enoughto startwithin thisregionand continuethrough gene52.
A morepositive identificationof
rnIB
mRNA andidentificationofrIlAmRNAwereachieved byusingin-phasedeletionmutants. de196 isaninternal, in-phase deletion within rlIB that pro-ducesashortenedpolypeptidewithamolecular weight of 22,000 (A. Bolle, personal communi-cation). delEM66 lies within rIIA, and coding also remains in phase in this case, so that an
rIIA polypeptide with a molecular weight of about 55,000 issynthesized (Bolle, personal
com-munication). These mutations wereintroduced
into the double amber mutant 32amH18
44amN82 to produce triple and quadruple
mu-tantswith thefollowinggenotypes:T432amH18
44amN82 delEM66, T4 32amH18 44amN82 del196, and T4 32amH18 44amN82 del196 delEM66. The gene 32 amber allele used for these experiments (H18) had the advantage that it produced a recognizable amber fragment with
a molecular weight of 18,000, so the gene 32 messagecould also beidentified. Figure4shows the identification of these polypeptides on a
sodiumdodecylsulfate-po;yacrylamide gel.The truncated gene 32, rIIA, and rnIB polypeptides
wereidentifiedeasilyandmigratedinregionsof the gel thatwere free of other radioactive
poly-peptides,asshownbythesamplIsisolated from thewild-typeandmutant-infected cells.
RNA from the triple mutant 32- 44- rIIB del196 was fractionated by preparative
poly-acrylamidegel electrophoresisessentiallyas de-scribed in the
legenid
toFig.2. Theeluted RNAwastranslated inacell-freeprotein-synthesizing
system, and the radioactive polypeptides were
separated bysodium dodecyl
sulfate-polyacryl-amide gel electrophoresis. Figure 5 shows an
autoradiogram of the driedgel.TherIIBdell96
polypeptide and the gene 32 amber fragment
H18 could be identified easily. The rIIBdell96
messageactivitymigratedmorerapidlythanthe wild-type rIlB message activity, as expected. Thiswasevidentby the relativepositions of the first rIIB message peaksand the positionofthe 16SrRNA(Fig.2and5). The molecularweights
and the differencesbetweenwild type anddell96
arediscussed below.gprIIAappearedtobe
pres-entinfractions68 to 74,whichalso containeda
small peak of rlIB message activity. Gene 43
message activity migrated slightly faster than theputativepolycistronicrlIA-B message.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:5.500.63.457.75.229.2]T4 EARLY mRNA SIZES 777
s:3 ti' L W
t \- (. \ \
rzJ
tTC\ -n C\ (\J (\J C %
\\\ 1, 1 1 1/
Gene
43
-r
lA-qo.
0,1X
_
*
rIIA
delEM66-
a
.*" a ..;.:i,.~ "
32?
_*
_.11--.2
r M/
~~~~~~ ...l 4- 4 S Wa
:e,h-' :§PA
-':.
FIG. 4. T4polypeptides synthesizedinvivoby
us-ingwild-type (wt)andmutantphage. CulturesofE. coli BE infected with T4phage were labeled with
[3S]methionineateither8to12 minor20to24min
(infectionwithwild-type phage only [slotlabeled T4
wt-16-20']). The radioactive polypeptides were
ex-tracted andseparatedon a 10%polyacrylamide by
electrophoresis for5hat25mA. Therelevant
geno-typesareshownatthetop;thegenescorresponding tothemarkedpolypeptidesareshownonthe left. ever, therewasaconsiderable amountof back-ground due to translation of endogenous mRNA's in the S30 extract, which interfered with theidentification of minor T4 polypeptides,
such as rIIA. Most of the other T4 mRNA's identified in Fig.2could also beidentified in Fig.
5,and the migrationratesof these mRNA'swere
approximately thesame onthe twogels. Some of thesemessages areidentifiedbygenenumber (Fig. 5).
Ashortened
rILA
polypeptide should besyn-thesized more
readily
in the cell-free system,therebymaking the rIIAmessageeasierto iden-tify. To do this, RNA extracted from cells
in-fected with the
quadruple
mutant32-44-del196delEM66
was fractionated, and the mRNAac-tivitywas
analyzed
asdescribed above. The rIIB del196, rlIAdelEM66,
and 32amH18 polypep-tideswere identifiedeasily. The mRNA activi-ties forrIlA
delEM66 and rIIB del196polypep-tides bothcomigratedat a ratethatrepresented
an
RNA
with a molecularweight
ofapproxi-mately1.1 x 106. The rIIB del196message activ-ity was alsopresent in lower-molecular-weight fractions,as observed inFig. 2and5. Noother
message
activity
forrIIAdelEM66wasdetected.An
analysis
of thedistribution ofgene32,rIIA,andrIIB messageactivities is
presented
below.Quantitation
offunctional
early
mes-sages.Tocomparetheamountsof
protein-syn-thesizing
activity
in the different mRNA's cod-ing for thesamepolypeptide andtoexamine theapparentcomigration of rIIA and rIIBmessage
activitiesmore
carefully, autoradiograms
ofgels such as the one shown in Fig. 2 were scannedwith amicrodensitometer. The results of these
measurements areshowninFig.6forgenes 43,
de1293,
52,nIB,
46, 30,and39 (from Fig. 2andotherdata not
shown)
and in Fig. 7 for genesrIIBdell96 and rIIAdelEM66(from the analysis of RNAs extracted from cells infected with the
mutant 32- 44- rIIBdell96 rIIAdelEM66). Of these nine genes,
only
genes43,del293,
39, andrIIA were represented bysingle messageswith uniquesizes. There weretwo
peaks
ofmessageactivity
for genes 32and46and threepeaks
ofmessage activity for genes rIIBdel196 and 52.
The
largest
RNA from therIIBregion
was notdetected in the RNA isolated from rIIB+-in-fected cells
(Fig.
6). Gene 30 messageactivity
migrated heterogeneously, with no apparent
well-definedpeak of
activity.
gp3O
andgp39were notseparatedonthegel shown inFig.
2;thus, itappeared that the gene 39 messageactivity was also heterogeneous. Numerous other message
activitieshavingoneof these types of behavior were observed (Fig. 2, 3, and 5). The genetic origins of most of these were not identified. A
conspicuousprotein havingamolecular
weight
of 20,000 to 25,000 was translated from four different messengers having molecular weights of0.25x 106,0.50x 106,0.8 x
106,
and1.0x106.
We foundavery similar distribution ofmessen-ger activity fora
protein
that had an identicalrle-
;196
-Vol- 40, 1981
on November 10, 2019 by guest
http://jvi.asm.org/
[image:6.500.47.238.76.529.2]OI.
~4 . -. * .~-. s- -. ...s - S
FIG. 5. AutoradiogramofT4earlypolypeptides synthesized by fractionated32t 44 rIIBdell96 RNA. RNA isolated 12minafter infection ofE.coli BE byT4 32amH1844amN82 rIIBdell96wasfractionatedona5.0-cm
2.0%,' acrylamide-0a5%agarosegel. The RNA wasdenatured before electrophoresis by incubatingit in 80%0 dimethyl sulfoxide for10minat37C The eluted RNA wasconcentratedbyethanolprecipitationandwas
translatedin a cell-freeprotein-synthesizingsystem as described in the text. The radioactivepolypeptides
wereseparatedona10%sodium dodecylsulfate-polyacrylamideget Thegelwasdried,andanautoradiogram
wasmade. The slotsatthe sides contained the radioactiveproteins programmed by unfractionatedRNAs isolatedafter infection ofE.coliBEbythephagestrainsshown above the slots. The genescorrespondingto
themarkedpolypeptidesareshown onthe sides. The elutionpositions of168 and 238 rRNA'sareindicated
atthetop. wt, Wildtype.
Molecular weight x10-6
20. ypu 0
I0 ~~~~80
0 ~~~~~~60gp39
50 d
40
gpd
4030
20 20
0
23034 842465054 22 2630 343842469 Frochon number
FIG. 6. Densitometricanalyses ofT4mRNA
activ-ities.Autoradiogramssuchasthose shown inFig.2, 3, and5 were scanned with a Joyce-Loebi double-beam recording microdensitometerequippedwith a
model JL-20013integrator.Theexposureschosenfor
analysis provided a linear response between film
darkeningasmeasuredonthe recorder and
radio-activity in thesample. Thegeneproducts analyzed
areindicatedonthefigure.
molecular weight when T4 late RNA was
ana-lyzed (42). This protein had the appropriate
molecular weight to be IPIII, but this identifi-cationwas notverified.
Sizes of functional early messages. RNA molecularweightcanbeestimatedfrom the time it takes an mRNAto be eluted from a gel be-cause there isalinearrelationship betweenlog
molecular weight and thereciprocal of the elu-tion time (13, 26). Thepreparative gelsused in
thisstudywerecalibratedwith RNAmarkersof
knownmolecular weights,ranging fromatRNA with a molecular weight of 2.5 x 104 to 28S
rRNAfrommouseliver cells (molecularweight
1.7 x
106).
These markers definedacurvefrom which the molecular weights of RNAs ofun-knownsize could be determined.Figure8shows
a curve for marker RNAs fractionated on a
preparative gel containing 2.25%
polyacryl-amide. Except fortRNA, all of the markerslay
on astraight line. Mostof the RNAs identified migrated in the linear region. The molecular weights of theT4mRNA's identified aboveare
shown in Table 3,together with the molecular weights ofthecorrespondingpolypeptides.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:7.500.67.460.64.313.2] [image:7.500.67.258.424.583.2]T4 EARLY mRNA SIZES 779
180
t 60
I6
~40
- ' 420 iv
^ C .o
l50
~40-
30-
,,10-2
Froction number
FIG. 7. DensitometricanalysesofrIImRNA
activ-ities. RNA isolatedafterinfection ofE.coliBEby T4
32amH18 44amN82 rIIBdell96 rIIAdelEM66 was
separatedon a2.25%polyacrylamide gel. TheRNA
was not denatured with dimethyl sulfoxide. After
electrophoresis and elution, the RNA wasanalyzed
asdescribedinthelegendtoFig.2and in thetext.
The radioactive polypeptides were separated on a
10%polyacrylamide gel, from which an
autoradi-ogramwasmade.TherIIBdel196, rIIAdelEM66,and
32amHl8polypeptideswereidentifiedby comparison
with the corresponding polypeptides synthesized by
unfractionated RNA,whichwerepresentonthesame
gel. Densitometric analyses ofthese three
polypep-tideswereperformedas described in the legendto
Fig. 6. The RNAmolecularweightscale atthe top
wasderivedfromanidenticalgelonwhichamixture
ofyeast and E. coli RNAswasfractionated.
ures2, 3,and5containmolecular
weight scales,
from which themolecular
weights
of other RNA species could be determined. Table3also shows the molecularweight of the mRNA that codedforT4
,B-glucosyl
transferase(designated
mfigt);
thisinformationwas obtained by measuring
f8-glucosyl
transferase enzymeactivity
after RNAfractionated by preparative
gel
electrophoresis
was
translated (data
notshown).
This RNAwasextracted from cells infected with a T4 mutant
containing adefectivea-glucosyl transferase to
avoid having to distinguish between the two enzymeactivities.
Table3shows theexcess codingcapacitiesof the genes and thus provides an indication of
whether the mRNA's were monocistronic or
polycistronic.The excesscoding capacityof the
polycistronic
rIIAB
transcriptwasnotrII-coding capacity. The size of this polycistronic RNA10 Z Q5
St 03
i
0.1
.05
th .03 .02
.01 .02 .03 .04 .05
/elution position 1D6 .07 .08
FIG. 8. Plot of logarithm ofmolecularweight
ver-susthereciprocalsoftheelutionpositions ofthe RNA
standards. Amixturecontaining
double-stranded3H-labeled Q/8MDV-1 RNA and nonradioactivemouse,
yeast,and E. coli RNAswasdenatured withdimethyl
sulfoxideandseparatedby electrophoresisat15mA
ona2.25%polyacrylamidegel. The elutionpositions
of the radioactive RNA speciesweredeterminedby
countingaportion of each sample inaliquid
scintil-lationcounter. Theelutionpositions ofthe
nonradio-active rRNA'sweredeterminedwithachartrecorder
monitoringthe elution buffer.Molecularweights of
1.7x 106, 1.3x 106, 1.07x 106, 0.7x 1(1,0.56x 106,
0.148x106, and0.074x106wereassumedformouse
28S,yeast26S, E. coli 23S,mouse18S,E. coli16S (17),
Q83MDV-1 double-stranded(ds)andQ/8 MDV-1 sin-gle-stranded(ss)RNAs(22), respectively. The
double-stranded QI) MDV-1 RNA, which waspresent in smallquantities, presumably represented
nondena-tured molecules.
decreased with each rII deletion introduced into the phage, and the excess coding capacity
re-mained approximately constant, indicating a
roughly quantitativeagreementbetween the size oftherII deletionandthedecrease in sizeofthe polycistronic RNA.
Sizes ofpulse-labeled early transcripts. The isolation and characterization of cloned fragments of T4 DNA (19, 34, 37, 38) provided
a convenient source of specific hybridization
probes for T4 messages (42). By hybridizing
pulse-labeled, fractionated RNA to filters
con-taining early genes, the sizes of both the func-tional and the nonfunctional transcripts could be measured. By annealing specific gene frag-mentswithpulse-labeledRNA rather than with
continuously labeled or unlabeled RNA, we
hopedtodetectfull-lengthtranscripts.
Radioactive T4 early RNA was prepared as
IF
4s\
VOL. 40,1981
on November 10, 2019 by guest
http://jvi.asm.org/
[image:8.500.248.443.53.287.2] [image:8.500.48.236.62.294.2]TABLE 3. RNAand polypeptide molecular weights of T4 mRNA activities
Excess coding
miRNA designa- M,of RNA opoyeida capacityof
Gene tion
(x106)
Ml ofpolypeptide mRNA (no.ofaminoacid resi-dues)
43 m43 1.2 112,000 200
del293 mdel293 1.0 103,000 70
46 m46a 0.95 71,000 300
m46b 1.5 71,000 850
39 m39 0.82 64,000 240
52 m52a 0.65 51,000 200
m52b 1.0 51,000 550
m52c 1.3 51,000 850
32 m32a 0.52 36,000 200
m32b 0.75 36,000 430
,Bgt mnfgt 0.67 46,000 250
rII transcripts
rIIABpolycistronicb rIIA+ rIIB 1.5 86,000+33,000 420
rIIA+rIIBdell96 1.3 86,000 +22,000c 320
rIIAdelEM66rIIBdell96 1.1 55,000 +22,000c 400
rIIBa. 0.55 33,000 250
rIIBb+ 0.88 33,000 580
rIIBdell96a 0.45 22,000 250
rIIBdell96b 0.78 22,000 580
aSee references24and 39.
bThe excesscodingcapacity is non-(rIIA+rlIB).
Bolle,personal
communication.
The Mr of gprIIA wassmaller
than thevalue of 95,000 reported byO'Farrell
etal., (24), but it agreed with the data reported here.
described elsewhere
(42)
and was fractionatedby
electrophoresis
oncontinuous-elutionprepar-ative
polyacrylamide
gels
as described above. Thefractionated RNAwashybridized
to nitro-cellulose filterscontaining
immobilizedplasmid
DNA. Each filter contained DNAfrom a
plas-mid
containing
asingle
T4restrictionfragment
ofknown
genetic
origin.
Insome cases, restric-tion fragmentswholly
internal to a gene wereused sothat the
transcript
could be associatedunambiguously
with one gene. In other cases,therestriction
fragment
usedasahybridization
probecontainedpartsoftwo
adjacent
genes. Inthesecases,the
transcripts
couldcomefromonegene orfrom bothgenes.
Figure 9 shows the results obtained
by
per-forming
hybridizations
toplasmids
containing
gene 43, gene52, gene
rIIB,
generIIA,
gene39,gene 40plusgene41, gene 42
plus
the/-gluco-syltransferase gene
(designated ,Bgt),
and gene30.
Significant
fractions of the RNase-resistantradioactivityrepresenting the RNAs from gene 43, gene 52, gene 42
plus
gene18gt,
and generIIB (Fig.8athroughcandg) hadthesamemolecular weights as the functional mRNA's described above. Forexample,
there was apeak
of RNAactivitythat
hybridized
to aninternalfragment
ofgene 43 which hada molecular
weight
of1.1x
106.
However, most of the RNAcomplemen-tarytogene43 wasmuchsmallerand
migrated
heterogeneously at molecular weights ranging from 0.1 x 106 to 0.8x 106. Aportion of the gene 52messagemigrated in discrete peaks with
mo-lecular weights of0.60x
106
and0.85 x106.
The sizes of these two species were similar to the sizes of two of the three functional gene 52 messages(Table 3). rIIB RNAs that migratedasdiscrete
peaks
withmolecular weights of0.50 x106
and0.80 x10c
alsocorrespondedclosely
in sizetothetwolower-molecular-weightrIIB mes-sagesdetectedbyin vitrotranslation.Morethan50% of the gene 52 and rIIB messages were
smallerthan the discretepeaks observed,as was
thecaseforgene 43.
A
polycistronic
rIIAB message was detectedbyinvitro translation (Fig. 6).Wethought that this would beamajor species in RNA thatwas
pulse-labeledat anearlytimesince it isbelieved
to be the initial transcript for rIIB (31). How-ever, norIIB RNAthatcorrespondedin molec-ular weight to the rIIA-B polycistronic RNA detected by in vitro translation was resolved
(Fig.9c). Hybridizationto aplasmid containing
aninternal HindIIIrestrictionfragmentor
rIlA
also failed to detectalargerIIAtranscript (Fig. 9d). Infact, we detected very
little
RNAcom-plementarytorIIA,probablybecause there was
less rIlA RNA than RNAs for the other early
genes studied (41). RNAs complementary to
gene 39, gene40plusgene 41, gene42plusgene
on November 10, 2019 by guest
http://jvi.asm.org/
[image:9.500.65.461.81.322.2]T4 EARLY mRNA SIZES 781
16S 23S 30540 ___165 __23S
20 20
10 16 22 26 22 28 34
FIG.9.racionaionofuls-laee T4eal
4N an 8eetofsecfcT Nsb hybii
gene30 genes40/41t
2 4
4fE-oiB y idtpf 4a ecie intheet
2- 4-~~~~~
010 1622283440 010 6 222834 40 Fraction number
FIG. 9. Fractionation ofpulse-labeled T4 early
RNA and detectionofspecific T4 RNAs by
hybridi-zation to recombinantplasmids. RNA waslabeled
with[3H]uracilbetween3and6minafter infection
ofE.coliBEbywild-typeT4asdescribed in thetext.
Thelabelingwasterminatedat 6min by adding cold
uracilandKCN,and the cellswererapidly disrupted
by hot sodiumdodecylsulfatelysis(14). About 50
jig
of RNA containing 5 x 106 cpm was subjected to
electrophoresis on a 2.25% polyacrylamide gel as
described in thelegendtoFig.2and in the text. The
RNAwas nottreated withdimethyl sulfoxide.
Frac-tions(1 ml;20minof elution time at a flow rate of
0.05ml/min)werecollected. Asmall portion of each
samplewasprecipitated with trichloroacetic acid to
determine the distribution of total radioactive RNA,
and0.50-ml portionsoffractions10through 45
(rep-resenting RNA species in the molecular weight range from0.025 x106to 2 x106)were incubated for 18 h at
67'C in siliconized scintillation vials containing
plasmid DNA immobilized on nitrocellulose filters
(see text). After 18 h, the scintillation vials were
chilledrapidly, and thefilterswereremoved, washed
several times with2x SSC(lx SSC is 0.15M NaCl
plus 0.015Msodium citrate), treated with RNase,
washedseveral moretimes with2xSSC, dried, and
counted withaliquid scintillation counter. The
back-ground radioactivity that hybridized to a control
filter whichwaspresent ineach vialand contained
only pBR322 DNAwassubtracted. This value ranged
from 99 cpm in fraction 10 (the peak of total
radio-activity elutedfromthegel) to 30cpm in fractions
containing high-molecular-weight RNA (molecular
weight,>106). The T4 recombinant plasmids used in
,Bgt, and 30 migrated
heterogeneously (Fig.
9ethrough h). The molecularweights of the RNAs
thathybridized tothese genesranged from2.5 x 104to about 1.5 x 106. There were somehigh
points in the distributions ofspecificRNAsfor
gene 39, gene 40plusgene 41, gene 42plusgene
,Bgt,
andgene30, butmostof thepeaks
were notreproducible whenadifferent RNA
preparation
was analyzed. The peak of mRNA that was
complementarytogene 42plusgene
Blgt
in frac-tions18through20,whichrepresentedanRNA molecular weight of0.6x106,
wasthesamesizeasthe mRNA that coded for
f3-glucosyl
trans-feraseinvitro (Table3;
unpublished data).
Sizes of
pulse-labeled early
transcripts
after
denaturation
withglyoxal.
Electropho-resisofRNAscompletelydenatured by
glyoxal
(21) provided betterestimates of RNA molecu-lar weights than analysis of native RNAs and should have diminished artifacts caused by RNA-RNA aggregation.
Radioactively
labeledT4 RNA wastreated withglyoxal and fraction-atedbyelectrophoresisasdescribed
previously.
Theeluted RNAwas
hybridized
tofour differentplasmids containing restriction fragments of
genesrIIA,
nIB,
43, and 52.Controlexperiments showed thatglyoxalated RNA hybridized with the same efficiency as non-glyoxalated RNA, probably because the glyoxal adductswereun-stable and came offthe RNA at the elevated
temperatureusedforhybridization (Fig. 10). Glyoxalated RNAs complementary to genes rIIA,rIIB, 43, and 52 migratedmore heteroge-neously than the native RNAs. However, we
observed the same range ofmolecular
weights
with the denatured RNA as with the native RNA (whichwasdenatured with
dimethyl
sulf-oxide beforeelectrophoresis). Gene43RNA mi-gratedveryheterogeneously. The distribution ofRNAthathybridized togene 43 extendedto a
maximum molecular weight of approximately l
ax
106: this wassimilar
to the value of 1.1 x106 obtained whennative RNAwasused. Denaturation with
glyoxal
increased thehet-erogeneity of the migration rates of all four
RNAs
examined;
rIIB RNA was the leastaf-fected.Inother
experiments
weobserved similar increasedheterogeneity with otherearlyRNAs(complementarytoplasmids
containing
gene30,gene 40
plus
gene41,gene 42plus
gene8gt, andgene 39).Nevertheless, thesame discrete RNA
thehybridizations and the T4 genes whichthey
con-tainedwere asfollows: p622 containing gene43(a);
p621 containing gene52(b);containing pABIV
con-taining generIIB(c);pABI containing gene rIIA(d);
p626Acontaining gene39(e); p627containinggenes
40and41
(t);
p624containing genes42and/8gt (g);p625containing gene30(h).
VOL. 40,1981
on November 10, 2019 by guest
http://jvi.asm.org/
[image:10.500.50.245.54.345.2]782 YOUNG AND MENARD
S-Z
16s 23s generllAa
b
A4 gere52
6 F
2
'7 C: 11 '2. l^ A7 C7. 1 1
7 ID -523 31Y3 47 5 7 1'
Fractionnumber
FIG. 10. Fractionation of
pul
RNA denatured by glyoxal andT4 RNAsbyhybridizationtorec
TheRNA preparation described
9wasanalyzed inanidenticalZ
the RNAwastreated with1M
gl
sulfoxidein 10 mMKPO4 buffer
at50°Cbeforeelectrophoresison
0.5%agarosegel. (a)pABI (gene
52). (c)pABIV(generIIB). (d) p6
species was often discerned
RNAswiththesemolecularw
werenotdueto aggregation.
CotranscriptionofrIIAa
etal. (31) detected cotranscr
rIIBbyusingdeletion hybridi
Wedetectedcomigration ofn
rIIA and rIIB polypeptidesbu duciblydetect radioactiverII
this molecular weight. The fa active rIlA orrIIB RNA tha
expected position of a polyc
message could have been du
down of full-length transcript detectlinked rIIA-rIIBtransc:
mids containing restriction fi
rII region (Table 2) (35, 39
early RNA was hybridized
containsarestriction fragmen
3' end ofrIIB, the 5' end of
region in between which isni
(7). After hybridization to
pTB17DNA, the RNA was E
izedtoeitherapTB17filteroi
pTB35DNA. pTB35containE
restrictionfragment. Wecoulo
detectlinkedrIIA-B transcri] cedurewasused, although th
bridizedefficientlyto
pTB17;
possible explanation for thi
small amountof polycistroni
comparedwith thesmaller b
rIIBtranscripts(Fig. 6),whicl
pleted more effectively forti
16s 23s DNA in theinitial hybridization.However, when gene r
lB
wehybridized
the RNA firsttopTB35
and then to pTB17, we also failed to detect linkedrIIAB
transcripts.The most likely reason for the failure to detect
linked
rIIAB
transcripts with pTB35 and pTB17 was that there were very few intact polycistronicd transcripts. The T4 DNA fragments in pTB35
gene43 andpTB17 are separated in the T4 genome by about 0.9 kilobase. A cleavage of the polycis-tronic rIIAB RNA anywhere in this region, either in vivo, during isolation, or during the first
hybridization,
would have unlinked the 5 23 31 55rIIA-
andrlIB-specific
RNAs. If this reasoning se-labeled T4 early iscorrect,
linkedrIlAB
transcripts might
have detection of specific been detected moreeasily
by using
rIIA-
andombinantplasmids.
nIB-specific
DNAfragments
that were morein the legend to Fig. closely linked on the T4 genome.
Selzer
(unpub-manner, except that lished data) obtained recombinant DNA clones1yoxal-50%dimethyl from the rIIAB junction region by partial RI*
(pH
7.0) for 60min digestion of a HindIIIfragment
spanning thea2.0%acrylamide-
rIIAB
junction region. The recombinant clonesrIIA).
(b)
p621 (gene obtainedby
this procedure included a clone(gene 43). which had only
rIIA
sequences (pABI) and an-other clone which had onlyrIIB
sequences1,
suggesting that (pABIV) on the plasmid. Recently, this region eights existed and was sequenced (26a), and a promoter sequence (Pribnow box) and the presumed start site ofmd
rIIB.
Schmidt therIlB middle-mode RNA were located in a iption ofrIIA and DNA region ofrIIA
that lie between the two ization techniques. fragments cloned in pABI and pABIV. These nRNA activity for two plasmids should hybridize only to anrIIA-itcould not repro- specific RNA and an
rIIB-specific
RNA, respec-'A orrIlB
RNA of tively, because pABI lies to the left (5') of the ilure to find radio-rIIB
middle-mode promoter.it migrated at the Cotranscription of
rIIA
andrIIB
was detected :istronic rIIA-rIIB by rehybridizing RNA first hybridized to pABIie to rapid break- or pABIV and then subjecting the preparation s. In an attempt to to a second annealing. In the second hybridiza-ripts, we used plas- tion reaction radioactive RNA that had been ragments from the eluted from pABI was able to hybridize to ). Radioactive T4 pABIV, and RNA that had been eluted from to pTB17, which pABIV was able to hybridize to pABI (Table 4). It that includes the The control experiment with RNase-treated hy-Fgene 52, and the brids showed that the abilities of RNAs re-ot deleted by saA9 covered from pABI and pABIV to hybridize to filters containing pABIV and pABI, respectively, depended on eluted and hybrid- RNA sequences that were not in a DNA-RNA a filter containing hybrid form during the first annealing. None of ed aninternal rIIA the radioactive RNA that was eluted from pABI d not reproducibly and pABIV was able to hybridize in the second pts when this pro- annealing to pABIV and pABI, respectively, if Le eluted RNA hy- the first hybrids were treated with RNase,
al-asecond time. One though the RNase-treated hybrids did yield
is failure was the RNAs that rehybridized to plasmids containing c rIIAB transcript sequences identical to those used in the first
out
more abundant annealing. An additional control showed the hwould have com- specificity of hybridization. The same low level he complementary of background radioactivity was observed withJ.
VIROL,
on November 10, 2019 by guest
http://jvi.asm.org/
[image:11.500.59.253.51.213.2]TABLE 4. Cotranscription of rIIA and rIIB
RNasetreat- Inputfor
Amt
(cpm)annealed in secondhybridization %ReannealedtofilterRNasetreat-Input for
~~with:
containiing:C
Plasmidusedin mentafter secondan-
with:_______ning:'
firstannealing0 first anneal- nealing pAI ABV p6(gn pB22 rA nB
ing (cpm) (priAI) (IIB) 12) pBR322 rIIA rlIB
pABI(rIIA) + 416 134 30 21 32 25 0
pABI(rIIA) - 1,460 551 275 30 46 36 17
pABIV(rIIB) + 290 30 110 26 24 0 27
pABIV(rIIB) - 408 116 224 24 35 21 48
aThe initial input radioactivity was approximately
106
cpm of[3H]RNA
labeled from 3 to 6 min afterinfection by wild-type T4 at300C,andhybridization was for 30min at 40°C in a final volume of 150ul containing
50%formamide, 0.03 M sodium citrate, and 0.3 M sodium chloride.
bEach second hybridization reaction mixture was incubated for 72 h under the conditions described in
footnoteafor theinitial hybridization. The hybrids were not treated with RNase after the second reaction.
'Abackground of 30 cpmwassubtracted from
all
rawdata.filters containing either the vector alone (pBR322)ora
plasmid
containing unrelated T4gene 12.
The relative amounts of the
polycistronic
rIIABtranscript and the non-polycistronic rIIB transcript could be estimated from the data in Table 4. It appeared that at least 50% of the elutedrIlA transcripts extended into rIIB and
that about 50% of the
rIlB
transcripts labeled between 3 and 6 min after infection includedrIIA sequences; that is, they were transcribed from therIIA promoter.Thesevalueswerevery
rough estimates since the
hybridization
effi-ciency inthe firstannealing
wasunknown and in the secondannealing
not all of the RNApresent wasrecoveredashybrid.
Sizes
of
immediateearly transcripts
syn-thesized in the presence and absence of
CAM.T4
protein synthesis
isrequired
in vivofor the
synthesis
of allprereplicative transcripts (12, 30). The average sizes of IPII and IPIII transcripts made in thepresence of CAMwere less thanthe sizes of thesetranscripts
madeinthe absenceofCAM(1).Thiswas
interpreted
to mean that CAMprevented
these transcriptsfrom
being
elongated
intoneighboringgenes. Analternative
interpretation
wasthatCAM causeda breakdown ofthe RNA so that transcripts shorter than
full-length
transcripts were de-tected. The data discussed above indicated thatmany T4 RNAswerepresent
primarily
intran-scriptsshorter thanfull-length transcripts even in the absenceofCAM.
We examined the sizes ofnormal transcripts and transcripts from cells (CAM RNA)
CAM-treated from immediateearlygene 30, gene 39, gene 52, gene42plusgene
,fgt,
andgene40plusgene41byhybridizingfractionated RNAstothe
appropriate
plasmids.
Other experiments (41)had shown thatplasmids containingthesegenes
hybridized to T4 RNAsynthesized inthe pres-ence ofCAM. Our results are shown in Fig. 11.
The RNAs wereglyoxylated before
electropho-resistoprevent
aggregation
and toprovide
morereliable
estimates
of molecularweights.Asnoted above,denaturation ofnormalT4RNAledto a moreheterogeneousmigrationpattern.
Themo-lecular weight distributions of the
complemen-tary normal and CAM RNAs were distinctly different only for gene 40 plus gene 41. The
molecularweight of the CAM RNA forgene 40
plusgene41waslessthan the molecularweight of the normal RNA for gene 40 plus gene 41. However,the sizes ofCAM RNAs synthesized
fromgene30, gene 39,andgene 42plusgene,Bgt were notdifferent than the sizes of the normal
RNAs
complementary
tothese genes. Themax-imum
molecularweights,
aswell
astheaveragemolecular
weights,
wereapproximately
thesame in normal and CAM RNAs complementary tothese genes. For genes 30 and 39 only
small
fractions of the radioactive RNAs made either
in the presence or the absence of CAM were
largeenoughtobefull-length transcripts.
We also studied the size distribution of
func-tionalT4mRNAinapreparation of CAMRNA.
Nonradioactive RNAwasisolated 10minafter infection by T4, and CAM was added 5 min
before infection. The RNAwaselectrophoresed
on a preparative RNA gelandanalyzed as
de-scribed in the legendto Fig.2. No polypeptide with a molecular weight greater than about 21,000was
synthesized
(41). All of theT4 CAMRNA
polypeptides
weresynthesized
by mRNA'shaving molecularweights of0.1 x 105to0.2 x
105 (datanotshown).Althoughnospecificgene
productswereidentified,many of thesmall
poly-peptides codedforbyCAM RNAs had thesame
mobilities as the
small
polypeptides coded forby normal RNAs. However, in normal RNAs someof the
small
polypeptideswereencodedby large mRNA's, as shown in Fig. 2, whereas in CAM RNAs no largeT4 mRNA'swereidenti-fied bytranslation offunctional mRNA's.
VOL. 40,1981 T4 EARLY mRNA SIZES
783
on November 10, 2019 by guest
http://jvi.asm.org/
I,
0c
Fraction number
FIG. 11. Analysis of fractionatedT4CAM-treated
RNAsby hybridizationtorecombinantplasmids.
Ra-dioactive RNAwaspreparedbylabelingcells with
[3H]uracilbetween 3 and6min after infection byT4
52amEA118 either in the absenceorpresenceof200 pgofCAMpermladded5min before infection.The
RNA wasisolated andanalyzed byelectrophoresis
andhybridizationasdescribedinthelegendtoFig.
9 and in the text. The RNA was denatured with
glyoxal before electrophoresis as described in the
legendtoFig.10 andfractionatedbyelectrophoresis
on a 2.0% acrylamide-0.5% agarose gel; 0.9 ml of
every other 1-mlfraction was hybridized tofilters
containing the following plasmid DNAs: (a) p625
(gene 30),noCAM; (b) p625 (gene 30), plusCAM; (c)
p626A (gene 39), no CAM; (d) p626A (gene 39), plus
CAM; (e) p627 (genes40and41), no CAM; (D)p627
(genes40and41), plusCAM; (g) p624 (genes42and
,8gt),noCAM; (h) p624 (genes42 andfigt), plusCAM.
DISCUSSION
With a few notable exceptions, T4 mRNA's
are unstable, as are mostprocaryotic mRNA's.
Nevertheless, estimatingthemolecularweights ofspecific T4transcriptsunder different
condi-tions, such as different times after infection, duringinfections with putative regulatory mu-tants(tsGl,regA),andduringinfections of
mu-tant host cells (RNase I-, RNase III-), should
provide information important to an
under-standingofthe arrangement of T4transcription
units and
synthesis,
processing, anddegradation
ofT4 mRNA.
Functional T4 mRNAwas detected by
frac-tionating
total RNA from T4-infected cells onpreparative polyacrylamide
gels, followedby
translationinvitro toidentify specific
polypep-tides
and, indirectly,
specific mRNA's. Sincewe used autoradiography of one-dimensionalso-dium
dodecyl
sulfate-polyacrylamide gels toidentify
T4polypeptides,
only themostpromi-nentT4early proteins could be identified easily. Theminimumsizeofthe mRNAdetected had
tobesufficienttocode for theintact
polypeptide
chain,
since that was our means ofidentifying
themRNAactivity.Oursizeestimates of the T4
RNAswerebasedon comparisonsoftheir
mo-bilities
(more
properly, their mobilities andelu-tion rates) with the mobilities of a series of RNAs with known molecular weights. For T7
early
andlatemRNA's, this method providedafairly accurate measurement of molecular weights (26).Wehavenoreason to believe
other-wise for T4RNAs. For four of thenine mRNA's
whose sizes are showninTable 3, we observed
asingle messenger peakofapproximately
mon-ocistronicsize. In each casetheapparent
molec-ularweight exceeded the coding capacity.Some
extra nucleotides were presumably present in
theflankingregions, but the apparentexcessof
200 to600nucleotideseven forthose RNA
spe-ciessuspectedtobemonocistronic (suchasthe gene 43 message) might indicate that our size estimateswere toolargeforall ofthemRNA's.
Forexample, the onlygene 43 message activity
that was detected had a molecular weight of approximately 1.2 x 106. Sincegp43 hasa
mo-lecularweightof 1.15 x
105,
amessengerwith amolecularweightofapproximately 1.0 x 106 is requiredtocode for thegp43polypeptide. There
wasverylittle ifanygene 43 mRNAactivityof
higher molecular weight, as might have been
expected if this RNA had been derived from a
larger precursor. However, rapid processing to
thesize observed couldhave prevented the iden-tification ofalargegene 43transcript.
Severalearly messagesdisplayedtwo ormore
peaksofactivity. Gene52messageactivityhad
peaks representing RNAs with molecular weightsofapproximately0.65 x 106, 1.0 x 106, and1.3 x 106. Thesmallestofthesecouldhave
beenamonocistronicgene 52message since gp52
has amolecularweight of about55,000.The two
largergene 52 messagescould have been poly-cistronic messages. The bestway to determine this and to determine which other genes are
linkedontheputativepolycistronic messages is
to useplasmids containinggene 52 andadjacent
regions. Only thegene 52 mRNA with a
molec-ular weight 0.8 x 106 was observed in RNA
on November 10, 2019 by guest
http://jvi.asm.org/
[image:13.500.63.256.54.355.2]VOL. 40,1981
extracted after infection with the rII deletion
mutant del1241. The deletion endpoint nearest gene 52isveryfar fromgene52. This
observa-tion isinterestingbuthas not beenreproduced.
Itis alsointeresting that another deletioninthis
sameregion affects gp52synthesis. The deletion
saA9 (7) causes overproduction ofgp52 specifi-cally (Mattson and Bolle,personal communica-tion). Inthis casethe effect of the deletion on gp52 expression could be due to transcription
from therIIBpromoter(Mattson,personal
com-munication),although itmaybe thatsequences
in betweenrIIB and gene 52 have an effecton gene 52
transcription.
Gene32isrepresented bytwoactive mRNA species. These have
approximately
thesamemo-lecularweightsas two
peaks
of stable radioactive RNA activity that were observed after along
chase in the presence of
rifampin
(H. Krisch,personal communication).
In thiscase the twopeaks of radioactive RNAwereassociated with
gp32 message activity. In both our studies and those of
Krisch,
most ofthegp32-synthesizing
activitywas associated with the lower-molecu-lar-weightgene 32mRNA.Itisalso noteworthy thattheRNA extracted fromcellsinfected with
agene 32amber mutant contained a full-sized
gene 32 message.
Unlike the messages just described, several mRNA activities migrated very heteroge-neously. One such mRNA coded for gp3O; the
minimum molecular
weight
ofthis mRNA wasabout 0.60 x
106,
and its maximum molecular weightwas1.5x 106to2.0 x 106. Severaldiffer-entmechanisms couldgive risetomRNA's that display heterogeneous
electrophoretic
mobili-ties. They could be derived fromlarger precur-sors which aredegraded
randomly
intofrag-ments, someof which canstill functionas
tem-plates, or they could be transcription products of genetic regions lacking strong termination signals. Therearealso otherpossibilities.
Hybridizationtoclonedrestrictionfragments containing known T4 genes was
performed
in ordertoidentify specific
T4 mRNA'sdirectly.
Pulse-labeled RNA was used to increase the fraction ofnascentand
newly synthesized
tran-scripts. Forseveral
early
T4 genesafunctionalmRNA had been identified unambiguously by
translation in vitro (genes 43, 52, 39,
rnIB,
andrIIA),and we had ahybridization probetoassay
specificallyanddirectlyfor theRNA (thegene 52probemay have contained asmallamountof non-gene 52 DNA). For genes 43, 52, 39, and rIIB the radioactive RNAs that were able to
hybridizetothecorrespondingplasmidshad size distributions that included peaks representing RNAswith the same molecular weights as the
functional mRNA's detected. However, there
T4 EARLY mRNA SIZES 785
wasalsoa considerableamount of
lower-molec-ular-weight RNA, presumablyrepresenting
de-graded mRNA. In thecaseoftherIIAmessage,
essentially allof thehybridizing RNA that was
detected was smaller than the functional
mRNA.However, most significant was the fail-ure toobserve pulse-labeledRNA with a higher
molecularweight than the molecular weight of
thefunctional mRNA's fromthe same gene. It isunlikelythat this failure was due to the pres-enceofonly incomplete transcriptsin the
pulse-labeled RNA since a 3-min pulse should have
beenlong enoughto
allow
synthesis of an RNAwitha molecular weight of 2 x
106
to 3 x106
(considerably larger than the transcripts
de-tected). However, to test this hypothesis
di-rectly,RNA waslabeled from0 to 10 and from 2to 4minafter infectionwith T4, the RNA was
isolated immediately afterlabeling was
termi-nated, and the distribution ofgene 43message was measured by hybridization to a gene
43-containingplasmid (datanot shown). The RNA
labeled from 0 to 10 min after infection had a
distribution ofgene
43-annealing
RNAvery sim-ilartothat showninFig. 9forRNA labeled from 3to 6min. Inthe RNApulse-labeled from2 to 4 minafter infectionthere was relatively more gene 43 message in the peak with amolecular weight of 1.2 x106.
Higher-molecular-weightgene 43 RNA was notdetected in either RNA
preparation. This suggested that the gene 43 message with a molecular weight of 1.2 x
106
represented thetranscription unit forgene 43 or
thatprocessing occurred
during
transcription,
asappears tobe thecasefor
bacteriophage
T7.Several different mechanisms could have
pro-duced the
multiple transcripts
from othergenes,bothidentified (52,
nIB,
32,IPIII)
and uniden-tified. Thesetranscripts
could have arisenby
processing ofa larger
transcript,
as is the casewith theT7
early region primary transcript
and several T7 latetranscripts. Alternatively, they
couldhave been
synthesized
fromindependent
promoters and a common termination site. There arewell-documented
precedents
for this possibility. Early T7 transcription by E. coliRNA
polymerase
andlate T7transcription by
T7 gene 1RNA
polymerase
utilizemultiple
pro-moters and a common termination signal to generate all of theearly
messages and severallatemessages,respectively (9, 11,26).The
mul-tiple transcripts of the small DNA
phages
arealso derivedfrom
multiple
promoters anda com-mon termination site(35).
Synthesis
of RNAtranscripts froma
single
promoter andmultiple
termination sites could also generate multiple
mRNA
species.
Anexample
of suchatranscrip-tion unit occurs in
bacteriophage
lambda. Thetranscripts
promoted
fromPI
and Pr terminateon November 10, 2019 by guest
http://jvi.asm.org/
at ti and
t,
unless the lambd,- N protein is present.InthepresenceofNprotein,transcrip-tion continues into more distal genes. No T4 proteinwith N-likeactivity has been identified,
althoughanti-terminationhas beensuggestedas
anexplanationfor thesynthesisofdelayed early
RNAbyreadthroughfromadjacentimmediate early regions (30, 31).
Thesize of the RNA from the rIIregion is of
particular interest. rIIB message activity was
detected by thesynthesis of either thewild-type
rIIB polypeptide or the synthesis of a shorter rIlB polypeptide coded for by a mutant rIIB gene containing an internal in-phase deletion
which removed approximately 300 base pairs.
rIIA message activitywas more difficultto de-tect,butwasidentifiedunambiguously by using
anrIlAmutantcontaininganinternal in-phase
deletionwhich reducedthemolecularweight of the rIIA polypeptide by about 30,000 (from 85,000to 55,000) in ourgel system. A multiple
mutant containing both rII in-phase deletion
mutations, a gene 32 amber mutation which
producedanidentifiableamberfragment,anda
gene 44 amber mutation allowed us to detect
and measure both rIIA mRNA and rIlB mRNA,
as well as gene 32 mRNA in the same RNA
preparation. Threepeaks of rIIBdeletion mes-sageactivityweredetected;thesecorresponded
to molecular weights of approximately 0.45 x
106,
0.78x 106,and 1.3x 106 (designatedmrIIBa, mrIIBb, and mrIIAB, respectively). The twosmaller rIlB messages were alsoidentified
un-ambiguously in wild-type rII RNA (T4 32amA453 44amN82) (Fig. 2) and in RNA
con-tainingwild-type rIlAbutrlIB deletionmessage (T4 32amH1844amN82) (Fig. 5). The high-mo-lecular-weight rIlB message (mrIIAB)
comi-grated with message activity forrIIAdelEM66. Asimilar comigration ofrIIA andrIIB message activitieswasobservedwhenweused RNA
con-tainingeitherwild-typerIIAandrIlB messages,
rIIB
deIrlIA
wild-type message, orrnIB
wild-typerIIAdeletionmessage. Inthecase of wild-typeRNA,the message activitiesforgprIIBand for gprIIA migrated more slowly, representing
higher molecularweights than when RNAs from thedeletionmutants were examined. The find-ing that the molecularweight of thelargest
rnIB
message was decreased by a deletion in rIIA proved that thetwomessage activitieswere on
thesamemolecule.
Hybridizationto aclonedrestrictionfragment
of
rnIB
confirmed the existence of two of the functional messages just described above. An rIIBprobe detected peaks of rIlB RNAatmo-lecularweights ofapproximately 0.5 x 106 and
0.8 x
106,
approximately the same size as ob-served forfunctional rIIB message. There wasalso aconsiderable amount of lower-molecular-weight rIIB RNA,presumablyrepresenting de-graded rIIB RNA. No high-molecular-weight
polycistronic rIIAB RNA was detected
repro-ducibly with either the rIIA-specific probe or
the nIB-specific probe. Essentially all of the rIIA-specific RNA was present in a heteroge-neousdistribution withmolecular weights rang-ing from 2.5 x 104 to 1.0x
106,
suggesting that rIIAmRNA wasbroken down more rapidly than rIIB mRNA. Polycistronic rIIAB transcripts weredetectedbyrehybridizingradioactive RNA first hybridized either to a plasmid containing the 5'-terminal end of rIIB or to a plasmidcontainingthe 3'terminusof rIIA.
Schmidt and co-workers (31) originally
re-ported apolycistronic rIIAB transcript, aswell
as a monocistronic rIlB transcript, based on
studies in which hybridizationto phage DNAs
containing various rII deletions was used. Our
results confirm the existence of both a polycis-tronicrIlAB transcript and a smaller, possibly
monocistronic rIIB transcript. In addition, we
foundathirdrIIB message thatwas not
antici-pated. Other investigators havestudiedthesize of rII RNAbyusing deletion hybridization tech-niques. Sederoffetal. (33) foundrII RNA that
sedimentedinabroad distributionrepresenting
molecularweights rangingfrom 2.5 x 104to 1.2
x106.These authorsdid not distinguish between
rIIA-specificandrIIB-specific sequences. This is the same molecular weight range that we
ob-served when we used hybridization to specific rIlA and rIIB probes. On the other hand,
Wit-mer(38) claimedthat he detected an
rnIB
tran-scriptwithamolecularweight of0.43 x 106, an
rIlA transcriptwith a molecular weight of 0.71 x 106,andapolycistronicrIIABtranscript with
amolecular weight of1.1 x
106.
Onlythe poly-cistronicrIIABtranscriptwasdetected when T4 RNAsynthesizedinvitrowasanalyzed byWit-mer (38). Therewas nolower-molecular-weight
RNA present in the RNApreparations of
Wit-merdespite repeatedincubation of the RNA at
an elevated temperature during hybridization
beforeanalysisof the RNAbysucrosegradient
sedimentation. We cannot account for the dis-crepancies betweenourresults and those of Wit-mer (38). In particular, we never detected a
monocistronic rIlAtranscript,norhavewe ever examinedanRNApreparationthatdidnot
con-tain significant amounts of lower-molecular-weight rIIA-specific RNA and rIIB-specific
RNA.
Itis thought that CAM inhibitssynthesis of
delayed earlyRNAby preventingelongation of
transcriptsfrom immediateearlygenes into
de-layed early genes. If thiswere the case, CAM-treated RNA transcribed from immediate


![FIG. 4.[3S]methioninetocoliwt-16-20']).electrophoresistractedtypes(infectioning the T4 polypeptides synthesized in vivo by us- wild-type (wt) and mutant phage](https://thumb-us.123doks.com/thumbv2/123dok_us/1469875.99608/6.500.47.238.76.529/fig-methioninetocoliwt-electrophoresistractedtypes-infectioning-polypeptides-synthesized-mutant-phage.webp)