0022-538X/79/07-0017/08$02.00/0 Vol. 31, No.1
Recombination
in
Bacteriophage
Ti in the Presence
of Host
Restriction
JEROO S. KOTVAL,tTHOMAS V. POTTS,4 ANDJ. R. CHRISTENSEN'
Department of Microbiology, University of Rochester School of MedicineandDentistry,Rochester, New York 14642
Received for publication11December1978
Whenunmodified phage Ti infects restricting hostcellsathighmultiplicities ofinfection, there isanincrease in recombination frequency inallregions of the
Ti mapcomparedtothe level of recombination in standardcrosseswhenshort
distances are examined. The enhancement of recombination frequency is not
uniformfor all regions but isgreatestformarkersnearthecenterof themapand notsogreatfor markersnearthe ends. Crosses between markersattheextremities of the map show that there is no increase in recombination frequency under restriction conditions. An examination of phage Ti he'terozygotessuggeststhat anincrease of ends created by theprocessof P1 restriction increases recombina-tion. When Ti crosses aredone in the absence of host restriction, recombination defects inthe host havenoeffectonphage recombination andweconclude that phage Ti codes for itsownrecombinationgenes. Host recombinationfunctions
arealsodispensable for the recombination occurring during infection of restricting hostcellsby unmodified phageathigh multiplicities of infection.
BacteriophageTiissubjecttotherestriction and modification system of the prophage P1 (12).PhageTi grown on P1lysogens (T1lP) is
modified,and itplates onP1 lysogens withan efficiency of plating (EOP) of 1.0. Phage Ti grown onhostsnonlysogenic for P1(Ti .0)lacks the Pi modification andplatesonPi lysogens witha
markedly'reduced
EOP.Ithas beenproposed(17) that the P1 restric-tionenzymebindstounmodified DNAat asite called the mediation site and cleaves at distal sitescalled cleavage sites. For each mediation
site,agivenDNAmolecule iscleavedonlyonce atany one of itscleavagesites. The restriction
enzymedoesnotappearto turn over(17). Despite the inability of Ti .0 to infect P1
lysogens successfully, under certain conditions T1.0cansuccessfully infect asubstantial frac-tion of a population of restricting cells. Thus,
under conditions ofstrongaeration and in the
presenceof nutrient medium withanincrease in
multiplicity of infection
(MOI),
there is morethan a proportional increase in the fraction of
successfully infected cells (9). If genetically
marked phage strains are used in the experi-ment, amarkedly enhancedfrequencyof recom-bination betweenmarkerscanbe observed (9).
In this paper, the successful infection of a t Present address: Department of Radiation Biology and Biophysics, University ofRochester, School of Medicine and Dentistry,Rochester, NY14642.
tPresent address: 2123Bailey Ave., Buffalo, NY 14211.
substantial fraction ofrestricting host celLs at
high MOIs of unmodified phage will be called cooperative infection, and the recombination thatis seenin theseexperiments will be called
cooperativerecombination.
TheTi geneticmapdoesnotshowauniform
recombinationfrequencyperunitlength ofDNA (15).There isahigh frequency ofrecombination perunit lengthof DNA atthe endsof themap
and alowfrequency of recombinationperunit lengthof DNA in thecenterof themap.Electron microscopic autoradiography has displayed
qualitatively similar results for Ti recombina-tion and further reveals the insertion of short pieces of labeled DNA intounlabeledchains(2).
The inferencedrawnfrom these data has been thatfree endsarerecombinogenicforTi
recom-bination (2, 15).
Phage heterozygotes have been proposed to
berelatedtointermediates inbacteriophage
re-combination (19). In three-factor crosses with
looselylinkedmarkers,ifonestudies the
segre-gation pattem of plaques
arising
fromphage
that are heterozygous for the central marker,
twokinds ofresultsarefound.Oneisthe
segre-gation oftwo genotypes which have the same
recombinantconfiguration for thetwoflanking markers. Theinferredstructurefor the
hetero-zygote producing this result can be called an
"overlap"heterozygote. Theothercommon
seg-regation resultis tofindoneparentaltypeand
a double recombinant, from which an
"insr-17
on November 10, 2019 by guest
http://jvi.asm.org/
18 KOTVAL, POTTS, AND CHRISTENSEN
tional" type ofheterozygoteisinferred. Except for very closely linked markers, heterozygosis for either of the unselected markers is
uncom-mon(19-21). A study of phageTIheterozygotes is thereforeavalidgenetictool forexploring the natureof recombinationintermediates.
In the present paperwe report studies done under conditions where unmodified Ti infects restricting hosts at high MOLs. We show that under these conditions: (i)recombinationin the
centerof the Ti map is enhanced compared to
normal Ti recombination; (ii) despite the
en-hanced recombination under these conditions, the ends of the TI molecule are notgenetically
dislinked and the enhancement of recombina-tion is due to the insertion of short,
single-stranded fragments; and (iii) these
recombina-tion events, like normal Ti recombination, do
not require any host recombination pathway requiring the recA gene product.
MATERLALS AND METHODS Phage strains. The Tlam strains and the prepa-ration ofTlam stocks have beenpreviouslydescribed (8).All of the three-factorcrossesreportedherewere betweenTi Sb HrKn+ and Ti Sb+ Hr+Kn. These markersareplaquemorphologymarkers andarewell separatedonthegenetic map ofTi soas toavoidthe problem of high negative interference (1). All eight combinations of the three markerscanberecognized ondye-containing agar plates (TDM-2) (9).
Umnodi-fiedTi stocks (Ti-0) werepreparedon Escherichia coli B andmodifiedTistocksonE. coliB(Pi) by the plate lysate method, using LB agar plates (7). All phage stocks were sedimented by centrifugation at 30,000 rpmfor90min andsuspended in nutrient broth (NB), whichhasanion concentrationfavorable forTi
attachment.
Bacterial strains. E. coli strainsB,B/i,t, B(Pi),
and B/i,t(P1) are nonpermissive for Tlam mutants andhave been described (7),ashave the amber per-missive strains KB3 and CS100 (5). KB3(P1) and
CS100(P1) were derived in this laboratory. Strain JC4583wasthewild-type strain for comparison with the various recombination-deficient strains. JC4588 (recA56), JC4584 (recB21 recC22), and SDB1006 (recA56 recB21 recC22) are isogenic with JC4583. Theyarereferred to in the text, respectively, as rec+, recA-, recB-, and recA- recB-. These were gifts of S. D.Barbour and are describedfurther byCapaldo et al. (3). These strains were madelysogenic for phage P1CM (11), which is phageP1carrying a DNA inser-tionconferringchloramphenicol resistance and was a gift fromJ. L. Rosner. TheP1CM lysogens of these strains were found togradually lose their ability to restrictTi .0during serialpropagation. Therefore, for eachstrain,asingle freshly isolated lysogenic colony wasinoculated into LB containing 12,tgof chloram-phenicol permland incubated at 37°C with aeration till aturbidculture was achieved(approximately 12 to 18 h). Those cultures which showed at least 96% of thecells capable ofrestriction were distributed in 1-mlquantities inseveral vials and stored at -70°C in 10%dimethylsulfoxide.
Media. Luria broth (LB) and NB have been de-scribed(7),ashas TDM-2(9). LB agar waspurchased from GIBCODiagnostics. NB + NaCl is NB+ 0.5% NaCl. Cyanide broth is NB + 0.5% NaCl + 10-2 M KCN.
Preparation oflog-phase cells for Ti infection. In the case ofP1CM lysogens of the isogenic set of recombination-deficient strains, one vial of the frozen stockwasthawed at 37°C, and 0.2 to 0.4 ml of the culture was diluted into 10 ml of LBcontaining 12 ,ug ofchloramphenicol per ml. This was grown at 37°C with aeration until the cells were growing exponen-tially andatiter of1x 108 to2 x108 cells per mlwas achieved. The log-phase cells were centrifuged and suspended in an equal volume ofhalf-strength NB. These cells were used for attachment of Ti for the execution ofcrosses.The preparation of log-phase B and B(P1) for Ti attachment have been previously described(16).
Procedure used for phage crosses. The tech-niques usedweresimilartothose usedby Drexler and Christensen (7). All infectionsweredone in NB, and thephagewereallowed toattach at37°C for5min. The phage-infectious cyclewas thenarrestedby the addition ofpreviously chilledcyanide broth, and the tube was centrifuged. The KCN inhibition was re-versed either by washing with NB + NaCl or by diluting out the KCN with NB + NaCl before any titrationorbyacombination of the two procedures. Thesupernatant was titrated for unattached phage by plating on KB3 in the case of Tlam crosses or on a mixture of B+B/1,t (4:1, vol/vol) in the case of three-factor crosses,usingplaque morphology markers. The pellet, which is henceforth called the infected-cell preparation, was washed by centrifugation and titrated for infectious centers on CS100(P1) in the case of cooperative crossesto measurethe modification ratio, which monitors the extent of cooperative infection. The modification ratio is the proportion of cells that release at least one modified progeny phage (9).
Above an MOI of 5 in cooperative crosses, the modification ratios are the same as yield ratios (9). We verified this for all crosses, but the data shall not be presented. For standardcrosses innonlysogenic hosts, theinfected-cell preparationwastitrated on KB3 for theTlamcrossesand on Bfor thethree-factor crosses. A portion of the infected-cell preparation was then chloroformed and titrated on appropriate hosts to monitor the number of free phage particles in the infected-cell preparation.
The washedinfected-cell preparation was then di-lutedinto atube ofNB +NaCl to giveafinal concen-tration of102 to103 yielder cells per ml.Thisgrowth tubewasincubated at 37°C for 30 min to allow phage bursttooccur. Itwas thenchilled,and1to 2drops of chloroformwereadded. Thetube was then titrated on theappropriate nonlysogenic host to measure the total progeny released upon lysis. For cooperative crosses, the growth tube was also titrated on CS100(P1) to measurethetotalyieldofmodified phage.
InTlam crosses, the extent of recombination was monitored by measuring the proportion of progeny phage thatweream'. This should measure one-half ofthe totalrecombinantfrequency, since double am recombinants are not measured. To rule out the pos-sibility that thenonpermissive character of the host J. VIROL.
on November 10, 2019 by guest
http://jvi.asm.org/
RECOMBINATION IN BACTERIOPHAGE Ti
would influence the results of these crosses, several experimentswerecarriedoutin theamber-permissive
hosts, KB3 andKB3(P1). Resultsweresimilartothose obtained in B and B(P1) (datanotshown), butmost ofourambercrossesweredone in B and B(P1).
Inthecaseofthree-factorcrossesbetween plaque morphology markers, the progeny were scored on TDM-2plates. Thebacterial cultures usedweregrown tolog phasein NB+NaCl. A mixture of4volumes of Bto1volume of B/1,twasusedtoscoretheprogeny ofstandardcrosses in nonlysogenic hosts. Four vol-umesof B(P1) + 1volume of B/1,t(Pl) wasusedto
scoretheprogenyofcooperativecrossesandstandard
crossesinP1lysogens,toavoid thescoring of unad-sorbedphageparticles. Itwasverified that theprogeny
fromcrossesin P1lysogenswereessentiallyall modi-fied, and thus would form plaques on this plating mixture.
TheTDM-2plateswerespread with 0.2 ml of the
appropriatebacterial mixture and0.1ml ofthelysate from the growth tube, diluted to give 200 to 300 plaquesperplate. The plateswereincubatedat37°C for10to12h, andtheplaqueswereidentified under adissecting microscopewith strongillumination.
Heterozygotes. To screen for heterozygotes for the Hrmarker, the growth tube from thecross was platedonamixtureof B(P1)+B(Pl)/l,t (4:1,vol/vol)
on anLBplateatadilution which wouldgive only50
to60plaquesperplate. Those plaques which showed
a sectored morphology (heterozygotes for the Hr marker) were picked into sterile NB, diluted, and spreadonTDMplates with 0.2mlofamixtureof B
+ B/1,t (4:1, vol/vol). After incubation, the plates were examined for the progeny segregated by the
heterozygousplaque.
RESULTS
Distribution of exchanges along the Ti
geneticmap.Freshmanetal.(9) observedthat forthe markers Hrand Sb,recombination
fre-quencyincontrolcrossesvaried littleasa func-tion of theMOI,whereas in therestrictinghost the recombination frequency increased
mark-edlywithMOI,andatsufficiently highMOI(10
ormore),recombinationwasabout four timesas
frequent as in control crosses. We have done
similar experiments with markers spanning var-iousregions of theTi mapand findthat there isagreatvariation fromregiontoregion. Some
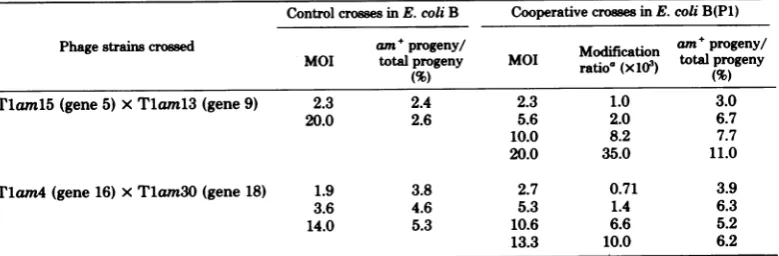
typical resultsareshown in Table 1. It canbe
observed thatinthecaseofTlaml5 (gene 5) x
Tlaml3 (gene 9), which are markers in the
central region ofthe Ti map, the MOI hasno
effect onrecombination frequency in standard crosses, whereasthere isan almostfourfold in-crease in recombination frequency with MOI under cooperative conditions. On the other
hand, in thecaseofTlam4 (gene 16) xTlam30
(gene 18), whicharemarkers atone endof the
map,there isless thanadoubling of the
recom-bination frequency withincreasing MOI,under
cooperative conditions, andatmostaslight dif-ference between therecombination seen in
re-stricting andnonrestrictinghosts. Thissuggests
that theextentof the increase in recombination observedtooccurin cooperative infectionmay
berelatedtomapposition.
Tofurther investigatethedistribution of the
effect,wehavecarriedoutanextensive series of
amxamcrosses,withpairsofmarkersspanning
variousregions of themap.Somerepresentative
resultsaredisplayed in Fig.1,whereanincrease
inrecombination in cooperativecrossesis seen
as a dilation of the genetic map. It isevident
thatpairsof markersneareither endofthemap
show little or no increase in recombination,
whereaspairs of markersnearthecenterof the
mapshowamarked increase in recombination incooperativecrosses.
Crosses between amber markersnearthe
ex-tremeends of theTi mapshowthat there isno increase inrecombination frequencyin
cooper-ativecrossescomparedtostandardcrosseswhen alargeregionofthe Ti mapisexamined. Typ-icalresultsareshowninTable 2.
TABLE 1. Geneticrecombinationaccompanying cooperative infectionindifferent regions of the Tl map Controlcrossesin E.coli B Cooperative crossesinE.coliB(P1)
Phage strainscrossed am+progeny/!am progeny/
MOI total progeny MOI ratioa(x103) totalprogeny
Tlam15(gene 5) xTlam13(gene 9) 2.3 2.4 2.3 1.0 3.0
20.0 2.6 5.6 2.0 6.7
10.0 8.2 7.7
20.0 35.0 11.0
T1am4(gene 16) xT1am3O(gene 18) 1.9 3.8 2.7 0.71 3.9
3.6 4.6 5.3 1.4 6.3
14.0 5.3 10.6 6.6 5.2
13.3 10.0 6.2
aThemodification ratio is theproportionof
dually
infectedcellsthatyieldatleastoneP1-modifiedparticle. Where theMOI isinsufficienttoinsurethat all cellsareduallyinfected,the Poisson distributionwasusedto correct forcells that donotreceive bothphage parents.19
VOL. 31,1979
on November 10, 2019 by guest
http://jvi.asm.org/
[image:3.508.62.452.509.637.2]20 KOTVAL, POTTS, AND CHRISTENSEN
Multipleheterozygosity among progeny
of Ti crossesdoneunder conditions of
co-operative infection. An increase in
recombi-nation frequency overshort distances that
de-pendsonthemapposition,whichweobservein crossescarriedoutundercooperativeconditions,
cannotbeexplained by merelyassumingan
in-creasein the roundsofmating,since thiswould accountforonlyauniform increase inallregions
of the map. However, an increase in the fre-quency of initiation of single-strand transfers
(19) in regions of the genome in which such transfersarenornallyuncommonmight provide an explanation. Such transfers produce
struc-tures containing heteroduplex DNA, and we
soughtevidence for their existencebyexamining
the frequency and genetic structure of phage
heterozygotes.
Table3 presentsthe results ofananalysisof
heterozygotes from three-factor crosses,
be-tween well-separated markers in the center of
the map, done under standard conditions. The
frequencyofheterozygous particlesis0.1% and is comparable to the frequency reported by Trautner (20).The pattern of strandsegregation
is consistent with thehypothesisofDNAstrand
T IMAP INEco/liB
GENE NOI 2 3 5911 161I
AMBER NO 16 3 4115132910 4 30
13 .4
15 110.5 11.1 1 13.4 79 124
16 3 41 15 13 29 10 4 30
TiMAPINfco/iB(PI)
FIG. 1. Comparison ofthegeneticmapsobtained
from two-factor amx am crossesofunmodified Ti carriedoutin E.coli strains BandB(PI). Allcrosses werecarriedout athigh MOI, underconditions that promote cooperative infection. The numbers below thelineoneachmaprepresentthe proportion ofam' progeny amongthe totalprogenyproduced in each
cross.
equivalence for resolution (18), whichaccounts
forapproxinately equal numbers of overlap and insertional heterozygotes.Heterozygotes which
segregate one recombinant and one parental
strand could be the products of short single-strand insertions in the region of the Hr marker. Those heterozygotes whicharerecombinant for
the outside markers reflectheteroduplex
over-laps for the central marker (19). We noticed the inequality between the two types of possible insertional heterozygotes but have no
explana-tion for it.
The data for heterozygotes selected for the central marker from cooperative crosses are
shown in Table4.Thefrequency of heterozygous particles is increasedto0.3% ofthe total
popu-lation, consistent with an increase in
recombi-nationseeninthesecrosses.Heterozygotes
seg-regating the two patterns seen in standard
crossesaccountforonly 55% of the total
heter-ozygotes. The other 45% are heterozygous for
twoloci(41%)orallthree loci (4%) anddeserve
TABLE 2. Control andcooperative crosses between Ti markers thatarewidelyseparated onthe map
Control crosses in E. Cooperativecrosses coli B in E.coli B(PI)
Phage am+ +
strains
progeny/
progeny/
crossed MOI total MOI total
progeny progeny
(%) (%)
Tlaml6 16 20 20 20
(gene 1) x Tlam30
(gene18)
Tlaml6 15 18 19 19
(gene1)x Tlam4 (gene 16)
Tlam3(gene 14 15 18 18
2)x Tlam4
(gene16)
TABLE 3. Segregatingphagetypesfound in plaques, selected asHr/Hr+heterozygotes,fromacross of modified TlSb HrKn+xmodified 5b+ Hr+ Knperformed inB(Pl)'
Segregation like that expected from overlap heterozy- Segregation likethat expectedfrom insertional
heterozy-gotes gotes
Exptno., Sb HrKn and + + +and Totalfrequency Sb Hr + and + +Kn and Totalfrequencyof
Sb+Knb + Hr + of overlap seg- Sb + + + HrKn insertional
segre-regation gation
1 3 10 1 15
0.45 0.55
2 3 4 0 8
aAtotal of 29,783 plaques from cross 1 and 15,120 from cross 2 were inspected. Frequency of Hr/Hr+ heterozygous plaques: 0.12%. Three plaques segregated the two parental genotypes and are considered to have arisenby accidental superimposition of plaques of the two parental types.
bAll
genotypesareexpressed in the order Sb Hr Kn, and the wild-type allele is designated by a "+" in the appropriate position. Thesameconvention is used in all tables thatfollow.J. VIROL.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:4.508.64.459.327.552.2]TABLE 4. Segregating phage types found in plaques, selected as Hr/Hr+ heterozygotes, from a cross of unmodifiedTl Sb Hr Kn+xunmodified Sb+ Hr+ Kn performed inB(Pi)'
Segregationlikethat expected from overlap Segregation like thatexpected
heterozygotes from insertionalheterozygotes
Total
fre-Heterozygosis ~~~~~
~~~~~~~~~~~~Fre-
quencyoffeeor:gsi
Exptno. Frequency Sbr +n quency thisclas
for: Sb Hr Kn + + + and + amongsingle and SbH+ +HK
among
anionHr-andSb+and+Hr 9 Hr+ hetero-andSb+Kn Hr+ heterozy- + Kn single zygotes
gotes
heterozy-gotes
Selected marker 0.55
only 1 0 8 5 5
0.53 0.47
2 2 17 6 8
Segregation of one parent and one single recombinant, suggesting heterozygosis in
cis
Frequency among double heter-Sb Hr Sb + Kn + ++ + Hr + ozygotesKn and and Sb and Sb and + +
+ +Kn Hr + Hr + Kn
Twomarkers 0.41
1 1 1 5 3
0.47
2 1 1 2 4
Segregation of one single recombinant and onedoublerecombinant, suggesting
heter-ozygosis in trans
SbHr Sb+ Kn + + + +Hr+ Knand and+Hr and+Hr and Sb +
Sb+ + Kn Kn +
1 0 2 2 6
0.53
2 0 3 1 6
Segregationoftwosinglerecombi- Segregationoftwo Segregationoftwo
nants double recombi- parents nants
SbHrKnand+ Sb+Knand+ Sb++and+Hr SbHr+and++
++ Hr+ Kn Kn
Threemarkers 0.04
1 0 0 1 3
2 0 2 1 3
aAtotal of12,162plaquesfromcross 1and17,621plaquesfromcross2 wereinspected.FrequencyofHr/Hr+
heterozygotes: 0.31%. The sixplaques that segregated thetwoparentaltypeswereassumedtohave arisenby
the accidentalsuperimpositionoftwoplaques,andtheywere notincluded incalculatingany of theproportions
listed here.Since theparentaltypeswerethemost common twoclasses ofplaquesin the progenypopulation,
themost commonclassofsuperimposedplaqueswouldgivethissegregationresult.Therefore,few if any of the otherplaquesrecorded here could have arisen this way, except thatavery few of theplaquesthatsegregated
oneparentaltype andonesinglerecombinantcould have been of this kind.
further consideration. Of the38particles which
areheterozygousfortwoloci (double
heterozy-gotes), 18 areheterozygotesfortwolociderived
fromthesamestrand(incis).Thesecould arise
if the entire length of material encompassing
these markers were obtained from one parent
(i.e., long overlap heterozygote) orbytwoshort noncontinuous single-strand insertions at the
two regions of heterozygosity. However, 20 of
these double heterozygotes have flanking markersderived from differentparentalstrands
(in trans). These canbe bestaccountedforby assumingthattwosingle-strand insertionsgave rise tothesestructures. If the doublecis heter-ozygotesareinterpretedtoarise from
long
over-laprecombinant structures,theywould standas ananomalousclassthat isnotseeninstandard crosses, and they would not contribute to the
31, 21
on November 10, 2019 by guest
http://jvi.asm.org/
[image:5.508.58.447.83.495.2]22 KOTVAL, POTTS, AND CHRISTENSEN
disproportionateincreasesof recombination
fre-quencyincentralmappositionsthatare seenin
cooperativecrosses.Since the double cis
heter-ozygotes arealmost equal tothe double trans
heterozygotes, it islikelythatthe cis
heterozy-gotes are also produced by two independent
insertionsattheregionsofheterozygosity.
Fur-thermore, of the 93 Hr/Hr+ heterozygotes, 30 areheterozygousalso forSb,and 16are hetero-zygousforKn. If theseregionsofheterozygosity
wereindependentamongHr/Hr+heterozygotes,
about five triple heterozygotes would be
ex-pected,and in fact fourwerefound.
Effect of recombination defects of the
host on Ti crosses under normal and
co-operative conditions. Toinvestigatewhether
phage Ti codesfor itsownrecombinationgenes,
three-factorcrosseswerecarried out under
nor-mal conditionsinisogenic wild-typeand
recom-bination-deficient hosts. The results (Table 5) show thatthere isnodifference in recombination
frequency whetherthecrosses arecarriedoutin
hoststhatarerec+, recA-, recB-,orrecA- recB-.
Since recA mutations in E. coli are known to
abolishlegitimaterecombination in the host(6),
the data suggest thatphageTi codes for itsown
general recombination functions. (The possibil-ity ofan undiscovered recA-independent host
recombinationpathway allowingnormal Ti
re-combination isnotformally excluded.)
Cooperative infection differs from standard recombination not only in the conditions
re-quired for it to be observed, but also in the
resultsthat areobtained incrossesdone under
cooperative conditions. Itwastherefore possible that the host's recombination systemmight play
a role in cooperative crosses, even ifit is not
required during ordinaryTi crosses (Table 6). Inthe first sectionaretheresultsofcrossesof
T1iP in the rec+(PlCM) host, to serve as a control forthecooperative crossesinthe same
J. VIROL.
host.The resultsareverysimilartotheresults presented in Table1forT1-0 crossed in nonly-sogenic hosts.Additionalexperiments with Ti1 P crossed in thevarious recombination-deficient
PlCM lysogens gave similar results (data not
shown).Thus,Pi lysogenyper sedoesnot influ-ence the results of the crosses, so longas the
incoming phage isnotsubjectedto restriction. Crosses of Tl*0 at high MOIs in the
rec+(PlCM) hostare presented in Table2; the
enhanced recombinationassociatedwith
coop-erative crosses (9) is clearly seen. Taking into account both the single and double
recombi-nants, the totalrecombination in the Sb toHr
region is at least three times as high in the
cooperative crosses as in the standard crosses,
and thatintheHrtoKnregionisatleasttwice
ashigh.
Recombination frequencies in cooperative
crosses in the recA-(PlCM) host are
interme-diate between those observed in standard
crosses and in cooperativecrosses done in the
rec+(PlCM)host. However,anyhypothesis that
recA-dependent host recombination pathways
are required for cooperative recombination is
refuted by the results in the
recB-(PlCM)
and recA- recB-(PlCM) hosts, which give resultssimilartothose obtained in cooperativecrosses
in therec+(PlCM)host. These resultsare
con-sidered furtherbelow.
DISCUSSION
Astudy of recombination in phageTi shows that thedistribution of exchange eventsin
co-operative crosses is clearly different from that seen instandardcrosses (Fig. 1). Michalke (15)
has concluded that in T1 crosses most of the recombinationoccurs nearthe ends of the mol-ecule. Fromourresults itappearsthat in
coop-erativecrosses recombinationeventsaremuch
[image:6.508.67.463.512.654.2]more uniformly distributed. In regions where
TABLE 5. Crosses of unmodified Ti Sb HrKn'xTi Sb Hr+ Kn in nonlysogenic hosts with differing
recombinationproficiencies
Recombinant progeny/totalprogenycounted Hostgenotype TotalMOI Burstsize Total phage
counted Sb+Knand Sb Hr Kn and Sb+ +and+
+Hr+ +++ HrKn
reck 18 41 2,930 0.051 0.13 0.016
15 137 2,048 0.071 0.12 0.033
recA 12 11 2,408 0.056 0.12 0.016
15 36 1,391 0.062 0.12 0.029
recB- 11 72 1,971 0.049 0.11 0.016
17 100 1,164 0.052 0.12 0.027
recA-recB 9 45 2,178 0.057 0.10 0.022
11 70 2,038 0.071 0.11 0.027
on November 10, 2019 by guest
http://jvi.asm.org/
RECOMBINATION IN BACTERIOPHAGE Ti 23 TABLE 6. Crosses of Tl Sb Hr Kn+x TlSb+Hr+ Kn inP1CMlysogenswithdiffering recombination
proficiencies
Recombinantprogeny as a proportion of to-Host genotype Total MOI Burst size Total phage
counlted
tal progenySb + Kn and Sb Hr Kn Sb + + and
+Hr+ and+++ +HrKn
Cros8se with modifiedphagea
rec+ (P1CM) 25 20.0 2,258 0.081 0.14 0.022
23 8.5 1,597 0.054 0.12 0.024
Crosses with unmodified
phageb
rec+ (P1CM) 16 3,006 0.18 0.27 0.095
23 2,704 0.19 0.24 0.12
recA-(P1CM) 19 3,872 0.077 0.17 0.051
21 1,861 0.077 0.18 0.068
recB-(P1CM) 22 2,069 0.20 0.23 0.15
24 2,903 0.21 0.24 0.13
recA- recB- (P1CM) 15 2,066 0.22 0.27 0.12
20 2,845 0.21 0.26 0.13
a
Control
for effect ofP1
prophagein theabsence ofrestriction.
b
Restriction
present.
recombination is already high, cooperative crossesgivethe same results as control crosses, but inregions where standard crosses give less recombination, the recombination under coop-erative conditions is markedly enhanced. In
ad-dition,whenpairs of markers at the extremities of the Timap are crossed under standard and cooperative conditions (Table 2), there is no difference in recombination frequency and it appears thatcooperativerecombination does not
involvethereassembly of double-stranded frag-ments any more than standard recombination does. It is conceivable that the process of P1 restriction creates recombinogenic ends which enhance recombination inintemal regions of the
Timap.
Analysisofheterozygousparticles from stan-dard and cooperative crosses (Tables3 and 4) indicatesthatduring cooperativerecombination the majority of recombination intermediates
(55%) are of the typeseen instandard crosses.
However,multiple regions ofheteroduplexDNA where the flanking markersarein theparental configuration (double trans heterozygotes and
triple heterozygotes) are enhanced under re-stricting conditions
compared
torecombination instandard crosses. Such insertionalheterozy-gotesarehypothesizedtobeduetothe insertion
ofsingle-strandedDNA intotheparentalDNA
moleculetoyieldtherecombinant (19).In these
experiments, wehave selected forunreplicated
recombinant moleculesby selectingforasingle
heterozygous marker and then examined two
unselected sitesforheterozygosity.
Heterozygos-ity at thesesites wasfound frequently enough tosuggestthat, hadwebeen abletoexaminea
larger number of sites for heterozygosity, we would have foundmultiple heterozygosis in an evenlargerproportion of the plaques examined. Inaddition, we recallthat the frequency of
het-erozygosissubstantiallyunderestimates the fre-quency ofheteroduplex regions of DNA. First, one-half of the heteroduplex regions will be homozygous, because the two DNA strandswill
befromparents of the same genotype.Second, inseveralphage systems (21),there is a signifi-cant occurrenceof correction of any base pair
mismatches thatoccurwithinregions of
heter-oduplex DNA. We also note that there is a pronouncedincrease inrecombination frequen-ciesbetween markers in thecentral part of the genetic map, but that this effect isnot accom-panied by a similar increase in recombination
fiequencybetween moredistant markers. This
resultis whatwould be expectedfrom the rep-lication of the kind of recombination
intermedi-ateproposed above.
Recombination ofphage
Ti
inthe absence of host restriction appearscompletelynormal in all of therecombination-defective hosts(Table 5).
Since recA mutations in E. coli are known to
completely abolish the recombination
profi-ciency ofthe host (6), phageTi mustcode for its ownrecombinationgenes.
TheresultspresentedinTable6demonstrate
thatcooperativerecombinationisalsounaltered
in therecA-
recBi(PlCM)
host.Therefore,
nei-thertheRecB,C pathwaynoranyother E. coli
pathway that depends on the recA function is
required for cooperative recombination. It is known thatgenes carriedonunmodifiedTi can
express themselves under
restricting
conditionsVOL. 31,1979
on November 10, 2019 by guest
http://jvi.asm.org/
24 KOTVAL, POTTS, AND CHRISTENSEN
(4). Therefore, it is most likely that phage T1
recombination functionsare responsiblefor
co-operative recombination.
Inthe recA- (P1CM) host,the usual enhance-mentin recombination seen under cooperative conditions is much reduced(Table 6).Thelower level of cooperative recombination may occur
because someintermediate ofrecombination un-der these conditions issensitive tothe RecB,C nuclease, exoV, which is known to be particu-larly active in recA- strains (6). The P1 restric-tionenzyme isatype I restrictionenzyme which
acts onunmodifiedDNAbycreatinganickand thenconverting the nick intoadouble-stranded break, via two steps (13).
Cleavages
are notatuniquesites,since there isnoevidence for unique sizeclasses(13, 17). ExoVacts onnicked double-stranded DNAor on single-stranded DNAin a
manner which creates single-stranded DNA fragments (14). Hollomanetal. (10) have dem-onstrated that invasion of a double-stranded
DNA molecule by homologous single-stranded
DNAmayinitiate theprocess ofgenetic
recom-bination. DNA subjectedtoP1restriction could conceivably provide the free single-stranded DNA which Holloman etal. show to stimulate
recombination. The free single-stranded DNA maybe astranddisplaced fromaDNAmolecule nicked by the P1 restriction enzyme or it may be a product ofpostrestriction degradation, as
for example by exoV. Since exoV degrades nicked double-stranded DNA and single-stranded DNAin astepwise manner (14),these recombinationintermediateswouldbedegraded
in a recA- (P1CM) host where the activity of
thenuclease isknown tobe increased (6) com-pared to wild type. The suggestion has been made that the complexity of type Irestriction enzymes suggests that they may have arisen through evolution from a DNA recombination
enzyme (13). Our data lend credence to this plausible role for the P1 restriction enzyme in
vivo.
ACKNOWLEDGMENTS
We thank David Figurski for helpfuldiscussions in the courseofthis workand A. J.Clark, S.Barbour, and J. Rosner
forsome of thebacterialandphagestrains used.
Supportwasprovided by Public Health Serviceresearch grantAI-02781andtraining grants DE 00003 and GM 00592.
LITERATURE CITED
1. Bresch, C., and H.Mennigmann.1954.Weitere Unter-suchengen zur Genetik von TIBakteriophagen.Z.
Na-J. VIROL. turforsch.9B:212-215.
2. Bresler,S.E.,L. P.Dadivanjan,and M. I.Mosevitsky.
1970.Electronmicroscopic autoradiographyof recom-binant DNA molecules ofbacteriophage Ti.Biochim.
Biophys.Acta224:249-252.
3. Capaldo,F.N.,G.Ramsey,and S. D.Barbour.1974.
Analysis of the growth of recombination-deficient strains ofEscherichiacoliK-12. J.Bacteriol. 118:242-249.
4. Christensen,J. R.1974.Complementation byrestricted
phageTi.J.Virol.14:1411-1418.
5. Christensen, J. R., andJ. M. Geiman. 1973. Anew effect of therexgeneofphageA:prematurelysisafter infectionby phageTI.Virology56:285-290.
6. Clark,A. J. 1973.Recombinationdeficientmutantsof E. coli andotherbacteria. Annu. Rev. Genet. 7:67-86.
7. Drexler, H., and J. R. Christensen. 1961. Genetic crossesbetween restricted and unrestrictedphageTIin
lysogenicandnonlysogenichosts.Virology13:31-39.
8. Figurski,D.,and J. R.Christensen. 1974.Functional characterization ofthegenesofbacteriophageTI.
Vi-rology59:397-407.
9. Freshman, M.,S. A.Wannag,and J.R.Christensen. 1968.Cooperativeinfection ofPI-lysogenicbacteriaby
restrictedphageTi. Virology35:427-438.
10.Holloman, W. K., R. Wiegand, C. Hoessli, and C.
Radding.1975.Uptakeofhomologoussingle-stranded fragments by superhelicalDNA:apossiblemechanism forinitiation ofgeneticrecombination. Proc. Natl. Acad. Sci.U.S.A.72:2394-2398.
11. Kondo, E.,D. K.Haapala,and S.Falkow. 1970.The
production of chloramphenicol acetyltransferase by
bacteriophageP1CM.Virology40:431-440.
12. Lederberg,S. 1957.Suppressionofthemultiplicationof
heterologousbacteriophageinlysogenicbacteria.
Virol-ogy3:496-513.
13. Linn, S., J. A. Lautenberger, B. Eskin, and D.
Lackey. 1974.Host-controlledrestriction and modifi-cationenzymesofEscherichiacoli B. Fed. Proc.35:
1128-1134.
14. Mackay, V., and S. Linn. 1974. The mechanism of
degradation of duplex deoxyribonucleic acid by the recBCenzymeofEscherichiacoliK-12. J. Biol. Chem. 249:4286-4294.
15. Michalke,W. 1967.ErhohteRekombinationshaufigeitan den endenTI-Chromosoms. Mol. Gen. Genet. 99:12-23.
16. Potts,T.V.,and J. R.Christensen.1974.Physiological study ofcooperative infection byrestricted
bacterio-phageTi.J.Virol.14:1319-1325.
17. Risser, R.,N.Hopkins,and R. W.Davis.1974.Action ofEscherichiacoliP1restrictionendonucleaseon Sim-ianVirus 40DNA. J. Mol. Biol.89:517-544.
18. Sigal,N.,and B.Alberts.1972.Geneticrecombination: thenatureofthe crossed strandexchangebetweentwo homologousDNA molecules. J. Mol. Biol. 71:789-793.
19. Signer,E. 1971.Generalrecombination,p. 139-174. In A.
D. Hershey (ed.), The bacteriophage lambda. Cold SpringHarborLaboratory, ColdSpringHarbor, N.Y.
20. Trautner,T. A.1958.UntersuchungenanHeterozygoten
desPhagenTi.Z.Vererbungsl.89:264-271.
21. Wildenberg,J.,and M.Meselson.1975.Mismatch
re-pairinheteroduplexDNA. Proc. Natl. Acad. Sci. U.S.A. 72:2202-2206.
on November 10, 2019 by guest
http://jvi.asm.org/