0022-538X/92/063306-10$02.00/0
Copyright©D1992, American SocietyforMicrobiology
Effects of Deletions in the
Cytoplasmic Domain
on
Biological
Functions of Human
Immunodeficiency Virus Type
1Envelope
Glycoproteins
DANA H. GABUZDA,1ANDREW LEVER,2t ERNESTTERWILLIGER,3 ANDJOSEPH SODROSKI2,3* Division of Human
Retrovirology,
Dana-Farber CancerInstitute, 44Binney Street,2 and Departments of Pathology3 andNeurology,
1 HarvardMedicalSchool, Boston,
Massachusetts 02115Received 31December1991/Accepted 25 February1992
The role of the cytoplasmic domain of the human immunodeficiency virus type 1 (HIV-1) envelope glycoproteinsinvirusreplicationwasinvestigated. Deletionofresidues 840to856atthecarboxylterminus of gp4l reducedthe efficiencyof virusentryduringanearlystepinthevirus lifecyclebetweenCD4 binding and formation oftheDNAprovirus without affecting envelope glycoprotein synthesis, processing, or
syncytium-formingability. Deletion of residues amino terminaltoresidue846wasassociatedwith decreased stability of envelope glycoproteins made in COS-1 cells, but this phenotype was cell typedependent. Thecytoplasmic domain ofgp4lwasnotrequiredfor theincorporationof theHIV-1envelopeglycoproteinsintovirions.These resultssuggest thatthecarboxylterminus ofthe gp4l cytoplasmic domain playsa role inHIV-1 entryother than receptor bindingormembrane fusion. Thecytoplasmic domain ofgp4l also affects the stability of the
envelope glycoproteininsome cell types.
The presence of a long intracytoplasmic domain at the carboxylterminusof thegp4l transmembraneproteinof the humanimmunodeficiencyvirustype1(HIV-1) distinguishes the HIV-1 envelope glycoproteinfrom that ofmost
retrovi-ruses(4, 12, 34). The transmembrane proteinsof thetypeC
and type D retroviruses terminate within 50 amino acids carboxy terminal to the transmembrane domain (34). The
presence ofacytoplasmic domainextendingmorethan 100
amino acidsbeyondthe transmembraneregioninHIV-1,the simian immunodeficiency virus (SIV), visna virus, and the equine infectious anemia virus suggests a specific function
for thisregion in the lifecycleofthe lentivirussubfamilyof retroviruses (12, 34). HIV-2 and SIV are frequently trun-cated just after the membrane anchor domain, but these truncated formsappeartoresult from selectionduringtissue culture propagation in human cells (4, 19, 20, 22). The natural form of the SIV transmembrane protein is the full-length41-kDaprotein (20, 22).
The HIV-1 envelope glycoprotein is initially made as a 160-kDa glycosylatedprecursor that is cleaved to form the gpl20exteriorandgp4l transmembrane subunits(7, 8, 40). Thegpl20 glycoprotein determines the tropism of HIV-1 for specifictarget cells by bindingto the CD4molecule (6, 21, 27). The gp4l transmembrane glycoprotein contains an
extracellular domain that is required for membrane fusion and noncovalent association withthegpl20 glycoprotein, a hydrophobic transmembrane region that anchors the gpl20-gp4l complex in the cell or virion lipid bilayer, and an intracytoplasmic domain of unknown function (2, 11, 14, 24, 35).Thecytoplasmic domain containstwohighly conserved segments with the potential to form amphipathic ao helices
andmayform asecondaryassociationwith the lipid bilayer
(1, 14, 15,39).
The cytoplasmic domain ofgp4l is required for efficient HIV-1 replication, but the function of this domain in the
*Correspondingauthor.
tPresent address:DepartmentofMedicine,Addenbrooke's Hos-pital, Cambridge, CB2 2QQ United Kingdom.
virus life cycle is unknown (9, 26, 37). The cytoplasmic domain isnotrequiredforsyncytiumformationorbindingto CD4(7, 8, 24)butmaymodulate intracellular transportand processing of gpl60 in some cell types (16). Whether the cytoplasmic domain is involved in virion assembly, for example, by facilitatingtheincorporationofenvelope glyco-proteinsintovirions, isunknown.
Toinvestigatethe roleof thegp41 cytoplasmicdomain in virusreplication,the effect ofdeletionsin thisregiononthe synthesis, processing,cellsurfaceexpression, incorporation into virions, and biologicalfunction of the HIV-1 envelope glycoproteinswas investigated.
MATERIALSANDMETHODS
Plasmids. The pSVIIIenv plasmid expresses the HIV-1
envandrevgenesof theHXB2 strain(31)underthe control
of the HIV-1 long terminal repeat (LTR) (17, 24). The pHXBAenvCAT plasmidcontainsanHIV-1provirus withan
in-frame deletion from theBglII to BglIIsites (nucleotides 6620 and7200)of thesequenceof Ratneretal.(32)intheenv
gene and a chloramphenicol acetyltransferase (CAT) gene replacing the nefgene (17, 36). The pHXBAenvAEcoCAT
plasmid, derived from pHXBAenvCAT, contains an
addi-tional deletion from EcoRI to EcoRI (nucleotides 4231 to 5325) overlapping thepol gene. These plasmids contain a
simianvirus 40originofreplication. Envelope glycoprotein deletionmutantsweremadeby Bal 31 exonuclease digestion ateitherthe BamHI siteorthe XhoI site(nucleotide 8053or 8476, respectively) (24, 32, 37). Theenvopenreading frame wasrestored when necessaryeither by creatingblunt ends prior to religation or by inserting ClaI linkers. The A(767-856) mutant was made by inserting an EcoRI linker at an
MnlI site to create aframeshiftatamino acid 767. Oligonu-cleotide-directedmutagenesis wasusedtocreatethe A(796-803), A(796-804), A(846-856), A(851-856), and A(854-856) deletions in an EcoRI-to-XhoI (nucleotides 5325 to 8476) fragment of HXB2 subcloned into the pBluescript plasmid (Stratagene) (25).Allplasmidsweresequencedin theregion
3306
on November 9, 2019 by guest
http://jvi.asm.org/
of the introduced mutation. ThepHXB2A(840-856) provirus
was made by subcloning the deleted BamHI-to-XhoI frag-ment into the pHXB2 plasmid (31). The rev expressor plasmidpSVIIIenvAKS contains a
KpnI-to-StuI
(nucleotide 5926 to6411) out-of-frame deletion in the envelope gene and expressesHIV-1 rev but not theenvelope glycoproteins.Virus replication studies in Jurkat lymphocytes. Jurkat T lymphocytesweretransfected by the DEAE-dextran method
(30) with 10 ,ug of pHXB2 orpHXB2A(840-856). Following
transfection, cultures were maintained in RPMI 1640 plus
10% fetal calf serum with daily medium changes. Reverse transcriptase activity of pelleted virions was measured as
previouslydescribed(33).
Radioimmunoprecipitation. For measurementof viral pro-tein expressionin infected Jurkat cells, 5 x 106 cells were metabolically labeled with 100 ,uCi (each) of [35S]cysteine
and [35S]methionineperml for 16 h. Virionswereharvested
for immunoprecipitation by centrifugation of supematants for 10 min at 1,500 x g to remove cell debris and
ultracen-trifugation at 12,000 xgfor 1 h. Cellsorvirionswerelysed
in RIPA lysis buffer, and HIV-1 proteins were immunopre-cipitated with AIDS patient serum and analyzed by sodium dodecyl sulfate-polyacrylamide gel electrophoresis
(SDS-PAGE) (23). The human monoclonal antibody (50-69) to
gp4l was obtained from the National Institutes of Health AIDS Research and Reference ReagentProgram andwas a donation of Susan Zolla-Pazner. For measurement of enve-lopeglycoprotein expression in transfected COS-1 cells, 3 x 106 cellsweremetabolically labeled with 100 ,uCi of
[35S]me-thionine per ml for 16 h at48 h posttransfectionwith 5 ,ug each ofanenvelopeexpressorplasmidand therev expressor
plasmid bytheDEAE-dextran method(5).The labeled cells were lysed in 10 mMTris-HCl (pH 7.4)-150 mM NaCl-5%
Triton X-114 at 0°C, and the detergent phase containing integralmembraneproteinswasisolatedby phaseseparation
at30°C,dissolved in RIPAlysis buffer,
clarified,
andimmu-noprecipitated aspreviously described (3, 11). Virions pro-duced in COS-1 cells were pelleted as described above
following labeling of transfected COS-1 cells for 16 h and
chasing for 3 h with medium containing excess unlabeled methionine and cysteine.
PCR assayfordeterminingtheefficiency of provirus forma-tion following acute infection. Fresh virus stocks were
pre-pared from supernatants of infected Jurkat cell cultures by centrifugation at 1,500 x gfor 10 min and filtration
(0.45-,um-pore-size filter)to removecell debris. Virus stockswere treated with DNase(Worthington) (2
,ug/ml)
for 20 min at roomtemperaturetoeliminateplasmidDNAcontamination(41). Heat-inactivated virus control supernatantswere incu-bated for 90 min at 60°C. Jurkat cells
(107)
were incubated with HXB2orHXB2A(840-856)
for 7 hat37°C. After 7h,2 x 106cells were harvestedbycentrifugation,washedtwice, lysedin 50,ul of0.2% NonidetP-40, and boiled for 15 min. Thecell lysatewasclarifiedbycentrifugation
at12,000
xg for 2 min and stored at -20°C. For polymerase chain reactions(PCR),theDNAin 2.5,ul of the celllysate(100,000
cell equivalents) was
amplified by using
the HIV-1 LTR R/U5 primers 5' GGCTAACTAGGGAACCCACTG 3' and 5'CTGCTAGAGAT'lTCCACACTGAC
3', whichare sim-ilartotheAA55 andM667primer pair previouslydescribed(41). PCRreactionswere
performed
according
tothemanu-facturer's instructions
(Perkin-Elmer
CetusCorp.)
for 33cyclesof94°Cfor 1min,
56°C
for1min,and72°C
for1min. ThePCRproductswereanalyzed by
electrophoresis
on2% agarosegels.Replication complementation assayformeasuring the
rep-licative potential of mutant envelope
glycoproteins.
Atran-sient
trans-complementation
assaywas used to assess thereplicative potential
ofmutantenvelope glycoproteins
inasingle round ofvirus
replication (17).
Cell-free virus trans-missionwasassessed bycotransfecting
COS-1 cellsby
the DEAE-dextran method(5)
with 5 ,ug ofanenvelope expres-sorplasmid
and 5 ,ug ofpHXBAenvCAT.
At48to72 h aftertransfection,theCOS-1 cell supernatantswerefiltered
(0.45-pum-pore-size
filter)
and the reversetranscriptase
activity
wasmeasured
(33).
Equivalent
reversetranscriptase
units of cell-free supernatants were added to 5 x 106 Jurkat Tlymphocytes.
The Jurkatcellswereincubated for 48to72h and thenassayed
for CATactivity.
In a similartrans-complementationassay, both cell-free and cell-to-cell virus transmissionweremeasured
by
direct transfection of Jurkat Tlymphocytes
aspreviously
described(17).
Syncytium
formation assays. Theability
of the mutantglycoproteins
to mediate the formation ofsyncytia
was assessed in COS-1 cellsby
cotransfection of 5p.g
of theenvelope
expressorplasmid
and 5p.g
of the rev expressorplasmid
followedby
cocultivationwithCD4-positive SupTl
lymphocytes
for 6 h at 60 hposttransfection (38).
Thesyncytium-forming
abilities of the mutantenvelope
glyco-proteins
wereassessed inJurkat-tat cellsby
cotransfecting
8 ,ug of theenvelope-expressing
plasmid
and 8p.g
of therevexpressor
plasmid
into Jurkattatcells andscoring
syncytia
at 60 h
posttransfection (11).
Expression of envelope glycoproteins on the cell surface. COS-1 cells cotransfected with an
envelope
expressorplas-mid and the rev expressor
plasmid
weremetabolically
la-beledwith[35S]methionine
for 16 hat48hposttransfection.
The intact labeled cells were washed twice with ice-cold
phosphate-buffered
saline(PBS) containing
2% heat-inacti-vated fetal calf serum, incubated with a 1:100 dilution of AIDSpatient
serum reactive with theenvelope
glycopro-teinsat
4°C
for 30min, rinsedtwice with PBScontaining
2%fetal calf serum,and
lysed
in RIPAlysis
buffercontaining
2.5p.g
of unlabeledgpl20
(American
BioTechnologies, Inc.)
at4°C.
The celllysates
were clarifiedby
ultracentrifugation,
and bound
envelope
glycoproteins
wereimmunoprecipitated
by
incubation withprotein A-Sepharose
asdescribed above. Soluble-CD4 inhibition ofsyncytium
formation and virusreplication. The effect of soluble CD4 on virus entry was
measured
by
producing
therecombinant HXBAenvCAT pro-viruses in COS-1 cells asdescribed above andpreincubating
equivalent
amounts of virus as determinedby
measuring
reverse
transcriptase activity
with different concentrations offull-length
soluble CD4(American
BioTechnologies)
for 1 hat37°C
prior
toinfection of Jurkatlymphocytes
(38).
RESULTS
Effects ofa deletion at the
carboxyl
terminus ofgp4l
onHIV-1
replication
in Jurkatlymphocytes.
Amutation which deletes residues 840 to 856(28)
atthecarboxyl
terminus ofgp4l
was introduced into an infectious HIV-1provirus
onplasmid pHXB2.
To compare thereplication
rate of this mutantviruswith that of thewild-type
virus,
thepHXB2
andpHXB2A(840-856) plasmids
were transfected into Jurkat cells and virusreplication
was monitoredby
measuring
reverse
transcriptase
activity
in the culture supernatants.The Jurkat cultures transfected with the
pHXB2A(840-856)
plasmid
demonstrated slowed virusreplication,
witha5-day
lagin thetime
required
toreachpeak
reversetranscriptase
activity
compared
with that ofthewild-type
virus(Fig.
1),
which is consistent withprevious
studies(9,
26,
37).
When
on November 9, 2019 by guest
http://jvi.asm.org/
250
C,,
0 X
E 200 E
C)
00
C-,
0) 150 co
.Q
C 100
cn
a) 5
CC
4
Day
FIG. 1. Replication of HXB2 and I Jurkat cells. Curves showreverse transc tants of Jurkat cell cultures transfected
pHXB2A(840-856)DNA.The results are least threeindependent experiments.
similar levels of virusproductionwe ofsyncytia formed in the
pHXB2-transfected cultures and the
viabilit'
indistinguishable (datanot shown).To examine the effects of a de terminus ofgp4longagandenvprc ing, secretion, andincorporationint steady-state andpulse-chaselabelin formed at a time when virus pro
HXB2A(840-856)-infected culturesv days aftertransfection, equal numb
metabolicallylabeled with
[35S]cyste
for 16 h and viral proteins were in celllysates, supernatants, andpelle
state levels of viral gag andenvproteinsweresimilar in the
pHXB2-
andpHXB2A(840-856)-transfected
celllysates,
su-pernatants,andpelleted virions (Fig.2A, B, and C and data notshown), althoughslightlyless cell-associatedgpl60was observed in the pHXB2A(840-856)-transfected culture (Fig.
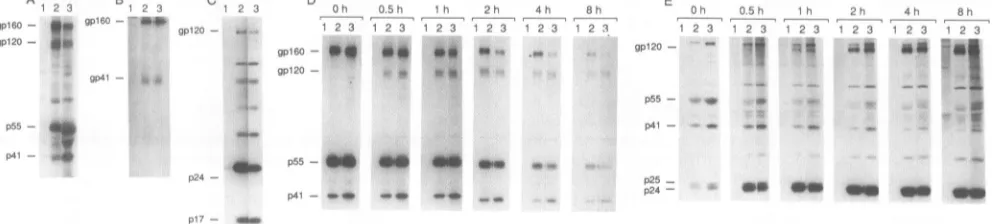
2A). Pulse-chase analysis of infected Jurkat cell cultures demonstrated similar synthesis, processing, and release of envelope glycoproteins(Fig. 2D and E). However, the level of themutantgpl60 in celllysateswasapproximately50% of thewild-typelevel aftera2-horlongerchase(Fig.2D).This decrease in the level ofgpl60at thelongerchase timeswas not accompanied by an increase in the level of
cell-associ-ated gpl20 glycoprotein. The gpl20 glycoprotein was re-HXB2 leased into the cell
supernatants
aftera1-h chasein both the wild-type and mutant cultures (Fig. 2E). These results sug-HXB2A(840-856) gest that deletion of residues 840 to 856 at the carboxyl terminus ofgp4l results in decreased stabilityof thegpl60 glycoprotein in infected Jurkatcells. Nonetheless, the syn-12 16 20 thesis, processing, secretion, andincorporationinto virions of the mutant HXB2A(840-856) envelope glycoproteins ap-pearto becomparable to those of the wild-typeglycopro-[XB2A(840-856)
viruses in teins.riptase activityinsuperna- Deletion of the carboxyl terminus of gp4l reduces the with 10
pLg
ofpHXB2 or efficiency of virusentry.To determine whether the cytoplas-typical of those seenin at mic domain ofgp4l
has an effect on the efficiency of virus entry, HXB2or HXB2A(840-856)virus stockswereused to infect Jurkat cells and the efficiency of HIV-1 provirusformationat7h afteracuteinfectionwasmeasuredbyPCR reachieved,the number amplification of the proviral DNA. Uninfected Jurkat cells and pHXB2A(840-856)- and Jurkat cells incubated with heat-inactivated virus con-y of these cultureswere trols were used as controls
(41).
The HIV-1 LTRR/U5
primer pair used for PCR amplification detects the first
oletion
at the carboxyl region of the viral DNA madeduring reverse transcription)teinsynthesis, process- and should detectvirtuallyall HIV-1 DNAmade,
including
ovirions in Jurkatcells, partial DNA transcripts
(41). Mapping
of restriction sites gexperimentswereper- within the amplified DNA product withHindIll,
HinflI,
duction in HXB2- and XhoII,and MboI restriction enzymes, whichcut atinternal xas similar. At 14 to 21 sites,confirmed that the140-bp fragment
amplified
bythese ersof viable cells were primers was the correct HIV-1 LTRfragment
(data
not -ine and[35S]methionine
shown). Followingacuteinfection ofJurkatcells,
formation nmunoprecipitated from oftheHXB2A(840-856)
provirusasdetectedbyPCRampli-ted virions. Thesteady- fication of viral DNA was
significantly
reducedcompared
A1 2 3 B1 2 3 C 1 2 3
gpl60- 9
g1gP02 -
-gpl2O- _ .2
gp54, - A1
D
0 h 0.5 h
2 3 1 23
l h 2h 4h Sh
12-3
12 3 1 2 3 1 2 3 1 2 'I
gpl60- Sol* *i 4
gpl2O- .. es
E
0 h 0.5 h 1h
1 2 3 1 2 3 1 2 3
gpl20- - t
I
2h 4h 8 h
1 2 3 123 12 3 -4Ma i i
p55- W 4w
p41- -. -_
a_, p55- _ d-_,_
p41- m .-i - _
-p25
-p24 - eS mm
NM
OM meFIG. 2. HIV-1envandgagproteins in HXB2- and HXB2A(840-856)-infected Jurkat cells and incorporated into virions.
Immunoprecip-itated celllysates (A, B, D), virions (C), andsupernatants(E)wereassayedatdays14to21followingtransfection of Jurkat cellswithnoDNA
(lanes 1), 10 ,ug of pHXB2 (lanes 2), or 10 ,ug ofpHXB2A(840-856) (lanes 3). Cells were metabolically labeled with [35S]cysteine and
[35S]methioninefor 16 h(A, B, C)orbyapulse-chase protocol (D, E). The gpl60, gpl20, and gp4l envelope glycoproteins and the p55, p24, andp17gagproductsaremarked.(A) Immunoprecipitatedcelllysates with AIDS patientserumanalyzed by4to12%gradient SDS-PAGE; (B) immunoprecipitated cell lysates with a human monoclonal antibody (50-69) that recognizes gp4l analyzed by 12.5% SDS-PAGE; (C)
immunoprecipitated pelleted virions with AIDS patientserumanalyzed by12.5%SDS-PAGE; (DandE) pulse-chase analysisof celllysates
(D)and supernatants(E) immunoprecipitated with AIDS patient serumandanalyzed by4to 12%gradient SDS-PAGE. Cellswerelabeled
for 10minand chased for the indicated times(in hours)with mediumcontainingexcess unlabeledcysteineandmethionine.
p55
-p41- e2,o
p24
-p17- _w
on November 9, 2019 by guest
http://jvi.asm.org/
[image:3.612.63.296.75.286.2] [image:3.612.62.559.529.641.2]A
B
C
M.,.9
M 1 2) I 4 S 1 9 q A 9;1 9 I A r,
HIV-1 R/U5
[image:4.612.83.273.80.197.2]-140bp
FIG. 3. Detection ofHIV-1provirusformation inJurkatcellsby PCR amplification of viral DNA following acute infection with HXB2 or HXB2A&(840-856) viruses. PCR amplification was per-formed byusingthe HIV-1 LTRR/U5primerpair M667-AA55 (41) for 33cycles toamplifyDNAfrom 100,000Jurkat cellequivalents harvested 7 h after acute infection with 4,000(A), 20,000 (B), or 100,000(C)cpmofreverse transcriptaseunits of novirus (lane1), heat-inactivated HXB2 (lane 2), HXB2 (lane 3), heat-inactivated HXB2A(840-856) (lane 4),orHXB2A(840-856) (lane 5)per ml. The 140-bp PCR product wasanalyzedbyelectrophoresison2%agarose gelsandvisualizedwith ethidium bromide. A DNA standard (PhiX-HaeIII digest)is shown in lane M.
512 NH2
wild
type
A
701-771
A704-751A
726-751
A
726-856
A
728-745
A
728-745+814-856
A
754-769
A
754-776
A
754-788
A
754-797
A
754-856
A
767-856
A
796-803
A
796-804
A
814-856
A
840-856
A
846-856
A
851-856
A
854-856
withthat of wild type (Fig. 3). Asshown in Fig. 3,the effect of the mutation on the efficiency of virus entry was most marked at the lower multiplicities of infection (4,000 and 20,000 cpm/ml). These results suggest that the carboxyl terminus of gp4l hasaneffectontheefficiencyof virus entry during anearly stage in the virus lifecyclepriortoformation of the DNAprovirus.
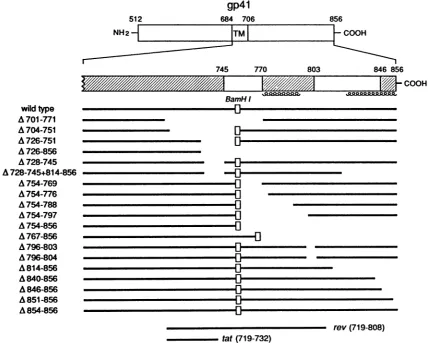
Mutationalanalysisofsequencesin thecytoplasmicdomain ofgp4l required forreplication. Thecytoplasmic domainof gp4l contains a stronglyhydrophilicregion (amino acids 724 to 745) followed by a region of alternating hydrophobicity
andhydrophilicity (amino acids 746 to856) (4,25). The latter region alsocontains two segmentswith the potentialtoform amphipathicothelices(amino acids 770 to 794 and 824to856)
(1, 39). A series of mutants containing deletions in the cytoplasmic domainwas constructed in the pSVIIIenv en-velope expressor plasmid (Fig. 4). Because many of these mutantsoverlap thetat or revsecond openreading frames,
rev orrev plus tat wassupplied in trans by cotransfecting
either the rev expressor plasmid or pHXB2AenvCAT, re-spectively. Of note, the truncation in theA(726-856)mutant issimilartothe truncation found insomeisolates ofSIV and HIV-2 (4, 10). The effect of theA(814-856) mutation in the
gp4l
684 706
TTMT
856 F COOH
745 770 803 846 851
'
V/////////A
/
Li-BamH I n
u
-0
n
r
n
rl
E.
El~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
_
ElEli
El
El--Il_
El
El
El--El
El
j6
-COOH
rev(719-808) tat(719-732)
FIG. 4. Diagram of the mutantHIV-1 gp41 transmembrane glycoproteins. Thewild-type HIV-1
gp4l
transmembraneglycoprotein
is depicted at thetop, with the hydrophobictransmembrane region designated TM. Thecross-hatchedregions
arehighly
conserved amongHIV-1isolates(28). Regionsofpotential amphipathicahelicesatresidues 770to794 and 824to856
(39)
areindicatedby
coils.Thepositions
ofthe secondopenreadingframes oftatandrev areshown.
on November 9, 2019 by guest
http://jvi.asm.org/
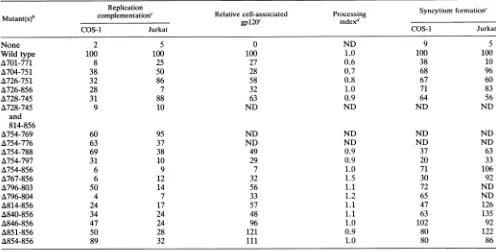
[image:4.612.96.524.349.692.2]TABLE 1. Phenotypes of gp4lcytoplasmic domain mutants'
Replication Snyimfrain
Mutant(s)b
complementation'
Relative cell-associated Processing Syncytium formation'COS-1 Jurkat COS-1 Jurkat
None 2 5 0 ND 9 5
Wild type 100 100 100 1.0 100 100
A701-771 8 25 27 0.6 38 10
A704-751 38 50 28 0.7 68 96
A726-751 32 86 58 0.8 67 60
A726-856 28 7 32 1.0 71 83
A728-745 31 88 63 0.9 64 56
A728-745 9 10 ND ND ND ND
and 814-856
A754-769 60 95 ND ND ND ND
A754-776 63 37 ND ND ND ND
A754-788 69 38 49 0.9 37 63
A754-797 31 10 29 0.9 20 33
A754-856 6 9 7 1.0 71 106
A767-856 6 12 32 1.5 30 92
A796-803 50 14 56 1.1 72 ND
A796-804 4 7 33 1.2 65 ND
A814-856 24 17 57 1.1 47 126
A840-856 34 24 48 1.1 63 135
A846-856 47 24 96 1.0 102 92
A851-856 50 28 121 0.9 80 122
A854-856 89 32 111 1.0 80 86
aDatareportedrepresentthe meansofatleast two or threeindependent experiments.ND, not done.
bThe number of the mutant refers to theaminoacid residues of the HXB2strainof HIV-1, where 1 is the initialmethionine (31).
cPercentage of thewild-type value.Values forreplication complementation represent the CAT conversion in target Jurkat cells for the mutantenvelope
glycoproteinrelative to the value for the wild type. Values shown for cell-associatedgpl20weredeterminedby gel densitometryofradioimmunoprecipitates
analyzed bySDS-PAGE andautoradiography.
dAmeasureof theconversionof mutantgpl60togpl20relative to that of the wild-typeglycoprotein. The amounts ofgpl6()andgpl20weredetermined
bygel densitometryofautoradiograms of SDS-PAGE gels.Theprocessingindex wascalculatedbythefollowingformula:processingindex=[(totalgp12O)mUta.,
x
(gp16O),,id
typej/[(gp16O)mutant
x (totalgp120),ildtypel-context of an infectious HIV-1 provirus has already been described elsewhere (37).
Toidentify sequences in the cytoplasmic domain thatare required for efficient viral replication, the ability of the mutant envelope glycoproteins to support cell-free virus transmission was determined by using a transient comple-mentation assay (17). In this assay, COS-1 cells cotrans-fected withanenvelope expressorplasmid and an envelope-defective provirus expressing the CAT gene produce recombinant virions thatareused to infect Jurkat T lympho-cytes (17). The efficiency of cell-free transmission of the recombinant viruses is determined by measuring CAT
activ-ityin the infected Jurkat culture. Most deletions within the carboxy-terminal 160 amino acids ofgp41 reduced replica-tioncomplementation of cell-free virus transmission to 4 to
69%of thewild-typevalue(Table 1). Todetermine whether similar regions are required for replication in Jurkat cells under conditions in which most of the virus transmission occurs by cell-to-cell spread, the abilities of the mutant glycoproteins to support a single round of virus replication wereexamined by using a similar transient complementation assayintransfected Jurkat cell cultures (17). The hydrophilic region extending from amino acids 726 to 751 and the immediately carboxyl region (amino acids 754 to 769) were
relatively dispensable for replication in Jurkat cells (Table 1). Otherwise, the regions in the cytoplasmic domain
re-quired to support the transmission of viruses produced in COS-1 cells and Jurkat cells were similar. These results suggestthat the function of some regions of the cytoplasmic
domain depends on the cell type in which the envelope
glycoproteins are expressed or onthe mode of virus
trans-mission.
Deletions in thecytoplasmicdomain ofgp4lareassociated withdecreased envelopeglycoprotein stabilityin COS-1cells. Todetermine whether mutations in the cytoplasmic domain ofgp41 alterenvelope glycoprotein synthesis orprocessing in COS-1cells, mutantenvelopeglycoproteins expressed in COS-1 cells were immunoprecipitated with AIDS patient serum and analyzed on SDS-polyacrylamide gels. The steady-state levels ofmutant glycoproteins with deletions within residues 846 to 856 at the carboxyl terminus were similar to those of thewild-typeglycoproteins (Fig. 5A and Table 1). The steady-state levels of mutant glycoproteins containing deletions amino terminal to residue 846 were reduced (Fig. 5A and Table 1). Two mutants containing deletions overlapping the transmembrane domain (amino acids 701 to 777 and 704 to 751)exhibited a processing defect (Fig. 5A and Table 1). The levels of wild-type and mutant
gpl20in the transfected COS-1 cell supernatants correlated with the levels of cell-associated gp120 (data not shown). When labeling COS-1 cells for 16 h wasfollowed by a 3-h chase with excess cold methionine and cysteine, the levels of mutantenvelope glycoproteins that were low in the steady statewerefurther reduced (Fig. 5B and Table 2). This result suggests that deletion ofresidues amino terminal to amino acid846 resultsin decreasedenvelope glycoprotein stability in COS-1 cells.
Thecytoplasmic domain ofgp4l is notrequired for incor-poration of envelopeglycoproteinsintovirions.Todetermine whetherthecytoplasmic domain ofgp4l is required for the
on November 9, 2019 by guest
http://jvi.asm.org/
[image:5.612.59.555.96.347.2]cC t3~QD~UD (.C Cn~st0U -Ln0,n00;n Ln00Lr u,C)400o o) sTr-
r-0 0: Co 0 0; 0 ;0 CD r "I
St D S D CD r«l 'I 'I St) Stn) C) CNO 000 '40000D:D -C 000
r-st-A 1 2 3 4 5 6 - 8 910 11 12
gpl60- e-. |gS b.
gpl2, -
is
S-a
-o \-C tn
-X~O UC 00
C)i -0
> OH H H H H H
un Lo Lo )tn u 0O coc0 cO 0 -4 O 0 - -4
; 'It 't LO U)
-
C.
0B 1 2 3 4 5 6 7 8 9 10 11 12 13 14 gpl60 - - '-m
gpl20
-4_*w w
00
O _
Lo
n) - N Lo 0 0 0? CO co 0c
°o 2Ozi o; (, "t Ct OD s st
CD s nt If
C 1 2 3 4 5 6 8 9 10 1132t314
gplO20
p55
-1_-p41
-gp4l
-gp4l - l* s
p24
[image:6.612.59.556.82.260.2]-FIG. 5. Wild-type and mutant HIV-1 envelope glycoproteins in COS-1 cell lysates (A, B) and virion pellets (C). Cell lysates and virions wereimmunoprecipitatedwith AIDS patient serum and analyzed by 10% (A, B) or 12.5% (C) SDS-PAGE. (A) Steady-state levels of mutant envelopeglycoproteinsexpressed inCOS-1 cells transfected with 5 ,ug of the rev expressor plasmid plus no DNA (lane 1), 5pgofwild-type envelopeexpressorplasmid(lane 2), or 5 ,ug of the mutant envelope expressor plasmids (lanes 3 to 12). (B) Expression of wild-type (lanes 1 and 3 to 8) and mutant (lanes 9 to 14) envelope glycoproteins in COS-1 cells after 16 h of labeling and 3 h of chase. COS-1 cells were transfected with5,ugof thewild-type envelopeexpressorplasmid (lane1),pHXBAenvAEcoCAT(lane2), or pHXBAenvAEcoCAT plus 0.1, 0.25,0.5, 1.0,2.5,or5.0,ugofthewild-typeenvelopeexpressor plasmid(lanes3 to 8,respectively) or 5p,gof the mutant envelope expressor plasmids (lanes 9 to14). (C) Virion-associated envelope glycoproteins. The lanes are the same as in panel B. The positions of the gpl20 envelope glycoprotein andp55,p41,andp24 gagproductsaremarked.
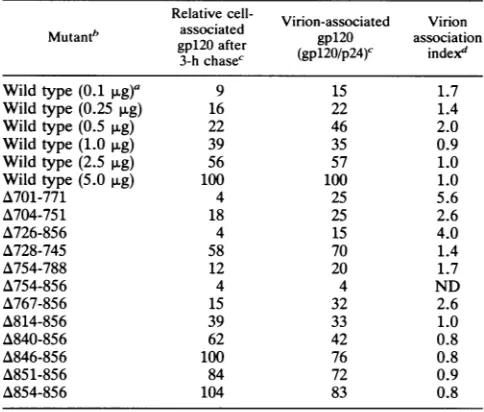
TABLE 2. Effectofmutations in thegp4lcytoplasmic domain onvirion-associatedenvelopeglycoproteinsa
Relative cell- Virion-associated Virion
Mutant" associated gp12a ascton gpl20after
gpl2O
association3-h chase' (g2Op)' idx
Wild type(0.1 pug)a 9 15 1.7
Wild type(0.25 ,ug) 16 22 1.4
Wild type(0.5 ,ug) 22 46 2.0
Wild type(1.0 ,ug) 39 35 0.9
Wild type(2.5 ,ug) 56 57 1.0
Wild type(5.0 SLg) 100 100 1.0
A701-771 4 25 5.6
A704-751 18 25 2.6
A726-856 4 15 4.0
A728-745 58 70 1.4
A754-788 12 20 1.7
A754-856 4 4 ND
A767-856 15 32 2.6
A814-856 39 33 1.0
A840-856 62 42 0.8
A846-856 100 76 0.8
A851-856 84 72 0.9
A854-856 104 83 0.8
aDatareported representthemeansofat leasttwo independent
experi-ments.
bThe numbers inparentheses refertotheamountofwild-type envelope
expressorplasmid DNAused for transfection. For all of themutant
glyco-proteins, 5 pugofplasmidDNAwasused.
cPercentageof thewild-typevalueat5 ,ug ofwild-typetransfectedplasmid
DNA.The values shownweredeterminedby geldensitometryof
radioimmu-noprecipitatesanalyzed bySDS-PAGE andautoradiography.
dCalculatedtonormalize the level ofvirion-associatedgpl20for the level
of cell-associatedgpl2O.Thevirion-association indexwascalculatedby using
the formula[(virion-associatedgpl2)0n,IanIX(p24)widtyp x (cell-associated
gpl2O)wjId
tyI]/[(p24)mun,n
x(virion-associatedgpl2O)wildtypex (cell-associ-atedgp12O)m.t.t]. ND,notdone.efficientincorporation ofHIV-1 envelopeglycoproteinsinto virions, the effect of deletionson the level of virion-associ-ated envelope glycoproteins was examined. COS-1 cells cotransfected with an envelope expressor plasmid and pHXBAenvAEcoCAT were used to produce recombinant virions. Pelleted virions were immunoprecipitated with AIDS patient serum and analyzed on SDS-polyacrylamide gels. Theamountoftransfectedwild-typeenvelope expres-sor plasmid DNAwas incrementally increased from 0.1 to 5.0 ,ug to allow comparison of the level of wild-type and mutant virion-associated gpl20 under conditions in which the levels of cell-associated envelope glycoproteins were similar (Table 2 and Fig. 5B and C). A virion association indexwascalculatedtonormalize the level of virion-associ-atedgpl20 for the level of cell-associated gpl20 (Table 2).
Deletions amino terminaltoresidue 846 caused reductions in the level of virion-associated gpl20 to 70% or less of the wild-type level (Fig. 5C and Table 2). However, the virion association index of themutantglycoproteinswassimilarto orgreaterthan that of thewild-type envelope glycoproteins
under conditions inwhich the levels of cell-associatedgpl20 weresimilar(Table 2).Moreover, the relativeincorporation
ofgpl20 into virionswasgenerally more efficient when the levels of wild-type or mutant cell-associated gpl20 were
relativelylow(Table 2). These results demonstratethat the
cytoplasmicdomain ofgp4l is notrequiredfor theefficient
incorporation of the gpl20 envelope glycoprotein into
viri-ons.It canalso beinferred that theincorporationof thegp4l
glycoprotein isnotdefective, since the cell orvirus associ-ation of the gpl20 glycoprotein depends on an interaction with gp4l(24).
Todetermine whether the observed decrease in the abso-luteamountofvirion-associated mutantenvelope
glycopro-tein was the sole cause of the replication defect in virions made in COS-1 cells, replication complementation
by
thewild-typeandmutantenvelope
glycoproteins
wascomparedunder conditions in which levels of virion-associated
gp120
A
on November 9, 2019 by guest
http://jvi.asm.org/
[image:6.612.56.298.424.630.2]B
100
/
~~~~~A
(854-856)A(754-788)
A(851-856)
l- A(846-856)
A(704-751) A(840-856) 0
A 0 A (728-745)
0 A(814-856) A(726-856)(
-0-wild-type(0.1-5.Ojg) (701-771) A(767-856) mutants
(5.Ojg)
'A(754-856)
*25 50 75 100
Virion-Associated
Envelope (%)0
0-0 U-6co
E
i-)._.
C:
cn
aI)
.)
75
F
50
F
25
o0
A(726-856) A(840-856)
0
A(814-856) 0
-0-wild-type (0.1-5.0 jg)
0 mutants
(5.Ojg)
III
25 50 75 100
Relative Envelope
GlycoproteinExpressed
onthe Cell Surface (%)FIG. 6. (A)Levels of virion-associated envelope glycoproteins andreplication complementation. Values for virion-associatedenvelope
glycoproteinswerederivedfrom Table 2. Values forreplication complementationweredeterminedby usingthereplication complementation assay in COS-1 cells(Table 1). All values areexpressed asa percentageof the value for thewild-type envelope glycoprotein at 5 ,ug of
transfectedplasmidDNA. Opencirclesrepresentthevalues, in consecutiveorder,for thewild-type envelope glycoproteinswhen0.1, 0.25,
0.5, 2.5,or5.0 p.gofenvelopeexpressorplasmidDNAwastransfected. For themutantenvelope glycoproteins,5 ,ugofenvelopeexpressor plasmid DNAwas transfected. (B) Levels of envelope glycoproteins expressed onthe cell surface and syncytium-forming ability. The
expression of envelope glycoproteins on the cell surface was determined by immunoprecipitation ofcell surface-accessible envelope glycoproteins following incubation of intact COS-1 cells with AIDS patient serum, analysis by SDS-PAGE, and quantitation by gel densitometry.All valuesareexpressed as apercentageof the value for thewild-type envelope glycoproteinsat5 ,ugoftransfectedplasmid
DNA.Opencirclesrepresentvalues, in consecutiveorder,for thewild-type envelope glycoproteinswhen0.1, 0.25, 0.5, 1.0, 2.0,or5.0 p.g
ofplasmidDNAwastransfected. For themutantenvelope glycoproteins, 5.0 pugofplasmid DNAwastransfected.
weresimilar. The relationshipbetween the levels of virion-associated wild-type glycoprotein anddegree ofreplication complementationwasnonlinear,with saturation of the latter
value when the level of virion-associated glycoproteinwas only 35%of themaximum value(Fig. 6A).As shown inFig. 6A, relatively low levels ofwild-type virion-associated en-velope glycoprotein can mediate virus entry efficiently. In contrast, most mutant envelope glycoproteins containing deletions within thecytoplasmic domain exhibiteda
replica-tion defectthat couldnotbesolely attributedtothe reduced levelof virion-associated envelope glycoproteins (Fig. 6A). Thegp4lcytoplasmic domain isnotrequiredfor syncytium formation. To determinewhetheradefect infusion wasthe causeofthe reduced replication complementation ability of
themutantenvelope glycoproteins produced in COS-1cells,
COS-1 cells were cotransfected with a wild-type ormutant
envelope expressorplasmid andcocultured with
CD4-posi-tive SupTl cells, and syncytium-forming abilitywasscored.
The syncytium-forming abilities ofmost of the mutant gly-coproteins expressed in COS-1 cellswereless than those of
thewild-type glycoproteins(Table 1). Since previous studies had demonstrated that syncytium formation in other cell types did not require the cytoplasmic domain (7, 24), the syncytium-forming abilities of the mutant envelope glyco-proteinswere also examined in Jurkat-tat cells (11, 18). In
Jurkat-tat cells, the syncytium-forming abilities ofthe mu-tant glycoproteins were usually equal to or greater than
those of thewild-type glycoproteins (Table 1).Theseresults suggest either that the syncytium-forming ability of the
mutantglycoproteinsisdependenton the celltype in which
theenvelope glycoproteinsareexpressedorthat syncytium
formation in COS-1 cellsmightbe reducedasaconsequence of decreased expression of the mutant envelope glycopro-teins onthe cell surface.
To determine whether the decrease in syncytium forma-tion observed in COS-1 cells could be solely attributed to reduced expression of envelope glycoproteins on the cell surface, the expression of selectedmutant envelope glyco-proteinsonthe cell surfacewasdetermined by immunopre-cipitation, analysison SDS-polyacrylamide gels, and
quan-titationby gel densitometry.Thesyncytium-forming abilities of the wild-type and mutant envelope glycoproteins were compared under conditions in which similar levels were expressed on the cell surface. For the A(726-856), A(814-856), and A(840-856) mutant glycoproteins, the level of envelope glycoproteins expressedonthe cell surfacewasnot
significantly different from the wild-type level when the levels of intracellular envelope glycoproteins were similar
(Table2 andFig. 6B). Furthermore, the syncytium-forming abilities of the mutant envelope glycoproteins made in COS-1 cells were similar to that of wild type when similar levels ofenvelope glycoproteins wereexpressedon the cell
surface (Fig. 6B). These resultsdemonstrate that the cyto-plasmic domain is notrequired for the cell surface
expres-A
-00.
c 0
.O
-c
CD
E
0
C)
E
0 0 0._o
'a
10075
50
25
0
on November 9, 2019 by guest
http://jvi.asm.org/
100
-\- A(814-856)
-\U
A(840-856)--\ A(846-856)
80 - -*- wildtype
-0 60
w co >
40-
20-0.001 0.01 0.1 1 10
[sCD4]
y/mI
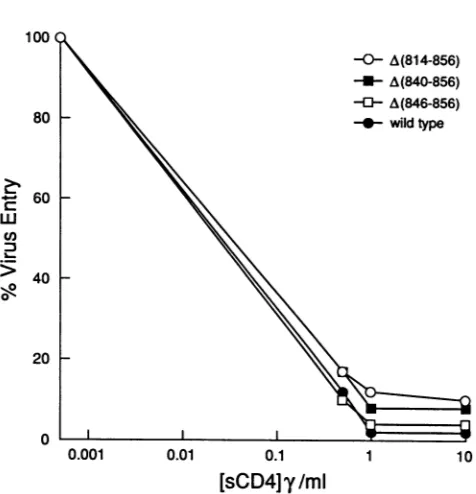
FIG. 7. Effect of mutations in the cytoplasmic domain on the sensitivity of replication complementation to soluble-CD4 (sCD4) inhibition. The effects of different soluble-CD4 concentrations on the CATactivity transferredtoJurkatlymphocytesby recombinant virionscarrying the wild-type or mutant envelope glycoproteins are shown. Each value represents the percentage of CAT activity observedforagivenmutantin the presence of soluble CD4 relative to the activity observed for the mutant in the absence ofsoluble CD4.-y,micrograms.
sion or the fusion function ofenvelope glycoproteins
ex-pressedin COS-1 cells.
Soluble-CD4 sensitivity of the mutant envelope glycopro-teins. HIV-1envelope glycoproteins that exhibit more than a twofold decrease in relative CD4-binding ability are less sensitive than the wild-type glycoprotein to soluble CD4 inhibition of virus entry(38). To investigate the possibility that deletions in thecytoplasmic domain ofgp4l affect the
CD4-binding abilityof the functional envelope glycoprotein
multimer, theeffect of mutations in thecytoplasmicdomain on soluble CD4 inhibition of virus entry was examined.
Equivalent amounts of recombinant virions produced in COS-1 cells as determined by measuring reverse
tran-scriptase activity were incubated with different concentra-tions of soluble CD4 for 1 h at 37°C prior to infection of Jurkat target cells. The A(814-856), A(840-856), and A(846-856)mutants exhibited the same sensitivity to soluble-CD4 inhibition of virus entryasthewild-type envelope
glycopro-tein (Fig. 7). From these results, it can be inferred that deletions in the cytoplasmic domain of
gp4l
do not signifi-cantlyalter theCD4-binding abilityof thenative,multimericenvelope glycoproteins. This conclusion is consistent with
previous observations that alterations in the
gp4l
cytoplas-mic domain do not affect the CD4-binding ability of the monomeric soluble form of thegpl20 glycoprotein (2, 7, 8, 24).
DISCUSSION
This study demonstrates that the
cytoplasmic
domain of the HIV-1 transmembrane glycoprotein has at least two effectsonviralreplication.Ahighlyconservedregionwithinresidues 840 to 856 at the carboxyl terminus of gp4l is important forefficient virus entry during an early step inthe virus life cycle between CD4 binding and formation of the DNAprovirus. The entire cytoplasmic domain amino termi-nal toresidue 846 affects the stability of envelope glycopro-teins made in COS-1 cells. The entire cytoplasmic domain appears to affect the efficiency of virus entry during the cell-free transmission of viruses made in COS-1 cells, whereas residues 726 to 769 are dispensable for cell-to-cell transmission inJurkat cells.Whether these differences area consequence of the cell type in which the envelope glyco-proteins are made or reflect differences in the requirement for this domain during cell-free versus cell-to-cell transmis-sion remainsto be determined.
The reduced efficiency of HIV-1 provirus formation fol-lowing acute infection with HXB2A(840-856) demonstrates that the carboxyl terminus of the HIV-1 transmembrane glycoprotein has an effect on the efficiency of virus entry duringanearly stage in the virus life cycle prior to formation of theprovirus. The steps in the virus life cycle required for virus entry includeCD4 binding, fusion of the virus and host cell membranes, and penetration anduncoating of the viral core (13). Deletion of the carboxyl terminus of gp4l is unlikely to affect CD4 binding, since the CD4-binding do-mainsareall contained within thegpl20 subunit (24, 27). The lack ofrequirement for the cytoplasmic domain for syncy-tium formation and the lack of alteredsensitivitytosoluble CD4providefurther evidence thatdeletionsin thisregiondo notsignificantly affect the CD4-binding affinity of the multi-meric glycoprotein complex. Deletions in the cytoplasmic domain of gp4l are also not likely to affect the fusion
function, since this domain is not required for syncytium
formation, asdemonstrated here and inprevious studies (7, 24). The possibility that the membrane fusion function of only the virion-associated envelope glycoproteins is defec-tive has not been excluded, but reliable methods for mea-suring the fusion function of the small fraction ofvirions relevant to infection are not available. The competence of the mutant envelope glycoproteins for CD4 binding and membrane fusion inconjunction with inefficient formation of the DNAprovirus raises the possibility that these truncated
envelope glycoproteins aredefective for astep involved in theuncoatingorpenetration of the viral core. The location of the gp4l carboxyl terminus within the virion close to the viral coreis consistent with the possibility that this region
plays arole in virusuncoatingorpenetration.
Independent ofarole in virusreplication, the integrity of thegp4lcytoplasmic domain amino terminal toresidue 846 is important for the stability of envelope glycoproteins expressedinCOS-1 cells. The observation that thestability
ofgpl60inHXB2A(840-856)-infected Jurkat cell cultures is decreased suggests that thecytoplasmicdomain also hasan effectonthestabilityof theenvelopeglycoproteins in other cell types.The effect of thecytoplasmicdomainonenvelope glycoprotein stability appears to be cell type dependent, judging bythe observation that the effect isrelativelyminor in Jurkat cells and the finding that some mutant
glycopro-teins that showed decreased stability in COS-1 cells were
fully functional, and thuspresumably stable, inJurkatcells. These results are consistent with previous studies
demon-strating that the stability and processing of the HIV-1
envelopeglycoproteins dependonthe cell typein which the
envelopeglycoproteinsareexpressed (8, 18).InCOS-1cells, approximately75% ofgpl60is cleavedto
produce
thegpl20
subunit,whereas inJurkatcells,only
asmallfraction(10
to20%) ofgpl60is cleaved.This
dependence
of theefficiency
on November 9, 2019 by guest
http://jvi.asm.org/
[image:8.612.61.299.62.309.2]ofgpl60
cleavage
oncell typemight,atleast in part, account for some of the celltype-specific
effects observed in thisstudy,
since a decrease in the steady-state level of gpl60 would be less deleterious in cells which process gp160inefficiently.
Inaprevious
study, Willeyetal. demonstrated that intracellular sorting results in the transport of most uncleaved gpl60 to lysosomes, where it is degraded (40).The mechanism by which the cytoplasmic domain affects
envelope
glycoprotein stability is unknown. One possibilityis that deletions in thecytoplasmic domain result in altered transport of gpl60, as observed in one study (16), which
might
affect localizationtosubcellular compartments wheredegradation
occurs.The cytoplasmic domain of the transmembrane protein doesnotappeartobe requiredfor the efficient incorporation
of the HIV-1
envelope glycoproteins
into virions. Thesefindings
are consistent with those ofprevious studiesdem-onstrating
that SIV and equine infectious anemia viruscontaining truncated envelope glycoproteins are still infec-tious
(4,
34). Similar studies of deletions of the shortercytoplasmic
domain of the Rous sarcoma virus envelopeglycoprotein
also demonstrated that thisregionwasdispens-able forvirionincorporation, although in this case,
infectiv-ity
was not affected(29).
The studies reported herein, combined with systematic mutagenic analysis of the HIV-1gp4l
transmembrane region (11), essentially eliminate thepossibility
that interaction ofspecific gp4l sequences with gagproteins
is a necessary step forincorporation ofenve-lope glycoproteins
into virions. The incorporation of the HIV-1 envelope glycoproteins into virions occurred moreefficiently
when levels ofwild-typeormutantcell-associatedenvelope glycoproteins
werelow. This result suggests thata mechanism for concentrating the envelope glycoproteinsinto virions is involved in HIV-1 virion assembly. Whether the
cytoplasmic
domain isrequiredfor another step invirusassemblyisnot known, but the examination of
HXB2(840-856)
virionsbyelectronmicroscopyand determinationofthe rate of gag precursor processing (Fig. 2D and E) did not reveal any gross defects in virus assembly or maturation(lla).
An HIV-1provirus containinga4-amino-aciddeletionand an additional 15 random residues in the gp4l cytoplasmic
domain,
X10-1, was reported to result in a replication-competent HIV-1exhibitingtheabilitytoform syncytia but no ability toefficiently
lyse single cells (9). A subsequentstudy
also proposed a role for the cytoplasmic domain ofgp4l
inviralcytopathiceffects(26).Contrarytothese earlierstudies,
Kowalski et al. demonstrated that the replication andcytopathogenicity
of the X10-1 mutant in Jurkat Tlymphocytes
wereindistinguishable from thoseof the wild-type virus(23).
The results of the present study provide further evidence that the cytoplasmic domainofgp4l does notplay
amajorrole in viralcytopathiceffectindependent of theeffectsofchanges inthisregiononviral replication rate.ACKNOWLEDGMENTS
We thank Kitty Lawrence for expert technical assistance, the AIDSResearch and Reference Reagent Programand Bruce Walker forreagents, and William Haseltine for helpful discussions.
This workwassupported bygrantsfrom the Leukemia Societyof America (toJ.S.) and the National Institutes of Health (AI01017,
A124755,
A128193, andA107386).We alsoacknowledge theCenter for AIDS Research grants (AI017386 and A128691) for supportingnecessarycorefacilities and forprovidingsalarysupportfor D.G.
REFERENCES
1. Andreassen, A.,H. Bohr,J. Bohr, S. Brunak, T. Bugge, R. M.J. Cotterill, C. Jacobsen, P. Kusik, B. Lautrup, S. B. Petersen, T. Saermark, andK.Ulrich. 1990.Analysis of the secondary struc-ture of thehuman immunodeficiency virus (HIV) proteins p17, gpl20, and gp4lbycomputermodelingbasedonneural network methods. J. Acquired Immune Defic. Syndr. 3:615-622. 2. Berman,P.W., W. M. Nunes, and 0. K. Haffar. 1988.
Expres-sionofmembrane-associated and secretedvariantsofgpl60of humanimmunodeficiencyvirus type1in vitro and incontinuous cell lines. J. Virol. 62:3135-3142.
3. Bordier, C. 1981. Phase separation of integral membrane pro-teins in Triton X-114solution. J. Biol. Chem. 256:1604-1607. 4. Chakrabarti,L., M. Emerman, P. Tiollais, and P.Sonigo. 1989.
The cytoplasmic domain of simian immunodeficiency virus transmembraneprotein modulatesinfectivity. J.Virol. 63:4395-4403.
5. Cullen, B. R.1987. Use of eukaryoticexpression technologyin the functional analysis of cloned genes. Methods Enzymol. 152:684-703.
6. Dalgleish, A. G., P. C. L. Beverly, P. R. Clapham, D. H. Crawford,M. F.Greaves, and R. A. Weiss. 1984.The CD4(T4) antigen isan essential componentof the receptor for the AIDS retrovirus. Nature (London) 312:763-766.
7. Earl, P. L., S. Koenig, and B. Moss. 1991. Biological and immunological properties of human immunodeficiency virus type 1 envelopeglycoprotein: analysisof proteins with trunca-tions and deletrunca-tions expressed by recombinant vaccinia viruses. J. Virol. 65:31-41.
8. Earl, P. L., B. Moss, and R. W. Doms. 1991. Folding,interaction withGRP78-BiP, assembly, and transportof the human immu-nodeficiency virus type 1 envelope protein. J. Virol. 65:2047-2055.
9. Fisher, A. G., L. Ratner, H. Mitsuya, L. Marselle, M. E. Harper, S. Broder, R. C. Gallo, and F. Wong-Staal. 1986. Infectious mutants of HTLV-IIIwith changes in the 3' region and mark-edly reduced cytopathiceffects. Science 233:655-659. 10. Franchini, G., C. Gurgo, H. G. Guo, R. C. Gallo, E. Collati,
K. A. Gargnoli, L. F. Hall, F. Wong-Staal, and M. S. Reitz. 1987. Sequenceof simian immunodeficiency virus and its rela-tionship to the human immunodeficiency viruses. Nature (Lon-don) 328:539-543.
11. Gabuzda, D., U. Olshevsky, P. Bertani, W. A. Haseltine, and J. Sodroski. 1991.Identification of membrane anchorage domains of the HIV-1 gpl60 envelope glycoprotein precursor. J. Ac-quired Immune Defic. Syndr. 4:34-40.
11a.Gabuzda, D., and J. Sodroski. Unpublished data.
12. Gallaher, W. R., J. M. Ball, R. F. Garry, M. C. Griffin, and R. C.Montelaro. 1989. A general model for the transmembrane proteins of HIV and other retroviruses. AIDS Res. Hum. Retroviruses 5:431-440.
13. Grewe, C., A. Beck, and H. R.Gelderblom. 1990. HIV: early virus-cell interactions. J. Acquired Immune Defic. Syndr. 3:965-974.
14. Haffar, 0. K., D. J. Dowbenko, and P. W. Berman. 1988. Topogenic analysis of the human immunodeficiency virus type 1 envelope glycoprotein, gpl60, in microsomal membranes. J. Cell Biol. 107:1677-1687.
15. Haffar,0. K., D. J. Dowbenko, and P. W. Berman. 1991. The cytoplasmic tail of HIV-1gpl60 contains regions that associate withcellular membranes. Virology 180:439-441.
16. Haffar, 0. K., G. R. Nakamura, and P. W. Berman. 1990. The carboxy terminus of human immunodeficiency virus type 1 gpl60 limits its proteolytic processing and transport in trans-fected cell lines. J. Virol. 64:3100-3103.
17. Helseth, E., M.Kowalski, D. Gabuzda, U.Olshevsky,W. Hasel-tine, and J. Sodroski. 1990. Rapid complementation assays measuring replicative potential of human immunodeficiency virus type 1 envelope glycoprotein mutants. J. Virol. 64:2416-2420.
18. Helseth, E., U.Olshevsky, D. Gabuzda, B. Ardman, W. Hasel-tine, and J. Sodroski. 1990. Changes in the transmembrane region of the human immunodeficiency virus type1gp4l
on November 9, 2019 by guest
http://jvi.asm.org/
lope glycoprotein affect membrane fusion. J. Virol. 64:6314-6318.
19. Hirsch, V., N. Riedel, and J. L. Mullins. 1987. Genome organi-zationof STLV-3 is similar to that of the AIDS virus except for atruncated transmembrane protein. Cell 49:307-319.
20. Hirsch, V. A., P. Edmondson, J. Murphey-Corb, B. Arbeille, P.R.Johnson, and J. L. Mullins. 1989. SIVadaptation to human cells. Nature(London)341:573-574.
21. Klatzmann, D., E. Champagne, S. Chamaret, J. Gruest, D. Guetard, T. Hercent, J. C. Gluckman, and L. Montagnier. 1984. T-lymphocyteT4moleculebehaves as the receptor for human retrovirusLAV. Nature(London) 312:767-768.
22. Kodama, T., D. Wooley, Y. Naidu, H. W.Kestler, M. D. Daniel, Y. Li, and R. Desrosiers. 1989. Significanceof premature stop codons in env of simian immunodeficiency virus. J. Virol. 63:4709-4714.
23. Kowalski, M., L. Bergeron, T. Dorfman, W. Haseltine, and J. Sodroski. 1991. Attenuation ofhumanimmunodeficiency virus type 1cytopathiceffectbyamutation affectingthe transmem-brane envelopeglycoprotein. J.Virol. 65:281-291.
24. Kowalski, M., J. Potz, L.Basiripour, T. Dorfman, W. C. Goh, E. Terwilliger, A. Dayton, C. Rosen, W.Haseltine, and J. Sodroski. 1987.Functionalregions oftheenvelopeglycoprotein ofhuman immunodeficiencyvirus type 1.Science237:1351-1355. 25. Kunkel, T. A., J. D. Roberts, and R. A. Zakour. 1987.Rapidand
efficientsite-specific mutagenesiswithoutphenotypicselection. Methods Enzymol. 154:367-382.
26. Lee,S.J., W. Hu, A. Fisher, D. Looney, V. Kan, H. Mitsuya, L. Ratner, and F.Wong-Staal.1989. Roleof thecarboxyl-terminal portion ofthe HIV-1transmembraneprotein in viral transmis-sion and cytopathogenicity. AIDS Res. Hum. Retroviruses 5:441-449.
27. McDougal, J. S., M. Kennedy, J. Sligh, S. Cort, A. Mowie, and J. Nicholson. 1986. Binding of the HTLV-III/LAV to T4+ T cells by a complex of the 110 K viral protein and the T4 molecule.Science231:382-385.
28. Myers, G., S. F. Josephs, A. B. Rabson, T. I. Smith, and F. Wong-Staal (ed.). 1988. Human retroviruses and AIDS. Los AlamosNationalLaboratory, Los Alamos, N.Mex.
29. Perez, L. G., G. L.Davis,and E.Hunter. 1987. Mutants of the Rous sarcoma virus envelopeglycoprotein thatlack the trans-membrane anchor and cytoplasmic domains: analysis of intra-cellular transport and assemblyinto virions. J. Virol. 61:2981-2988.
30. Queen,C., and D. Baltimore. 1983.Immunoglobulingene tran-scriptionis activated bydownstream sequence elements. Cell 33:741-748.
31. Ratner, L., A. Fisher, L. L. Jagodzinski, H. Mitsuya, R. S. Liou, R. C. Gallo, and F. Wong-Staal. 1987. Complete nucleotide sequenceof functional clones ofthe AIDS virus. AIDS Res. Hum. Retroviruses3:57-69.
32. Ratner, L., W. Haseltine, R. Patarca, K. J. Livak, B. Starcich, S. F. Josephs, E. R. Doran, J. A. Rafalski, E. A. Whitehorn, K. Baumeister, L. Ivanoff, S. R. Petteway, M. L. Pearson, J. A. Lautenberger, T. S. Papas, J. Ghrayeh, N. T. Chang, R. C. Gallo, and F. Wong-Staal. 1985. Completenucleotide sequence of the AIDS virus, HTLV-III. Nature (London)313:277-284. 33. Rho, H. M., B. Poiesz, W. Ruscetti, and R. C. Gallo. 1981.
Characterization of the reversetranscriptase froma new retro-virus (HTLV) produced by a human cutaneous T-cell lymphoma cell line. Virology 112:355-360.
34. Rice, N. R., L. E.Henderson, R. C. Sowder, T. D. Copeland, S. Oroszlan, and J. F. Edwards. 1990.Synthesis andprocessing of the transmembrane envelope proteinofequine infectious ane-miavirus. J. Virol.64:3770-3778.
35. Stein, B. S., S. D. Gouda, J. D. Lifson, R. C. Penhallow, K. G. Bensch, and E. G. Engleman. 1987.pH-independentHIV entry intoCD4-positiveTcells viavirusenvelopefusion to theplasma membrane.Cell 49:659-668.
36. Terwilliger, E., B.Godin,J. Sodroski,andW. Haseltine. 1989. Constructionanduseofreplication-competent human immuno-deficiencyvirus that expresses the CAT enzyme. Proc. Natl. Acad. Sci.USA 86:3857-3861.
37. Terwilliger, E., J. Sodroski, C. Rosen, and W. Haseltine. 1986. Effectsof mutations within the3'orfopenreadingframeregion ofhumanT-cell lymphotropicvirus type III(HTLV-III/LAV) onreplication andcytopathogenicity. J. Virol. 60:754-760. 38. Thali,M., U.Olshevsky, C.Furman, D.Gabuzda, J. Li,andJ.
Sodroski. 1991.Effects ofchangesingpl2O-CD4bindingaffinity onhumanimmunodeficiencyvirus type 1envelopeglycoprotein functionand solubleCD4sensitivity.J. Virol.65:5007-5012. 39. Venable, R. M., R. W. Pastor, B. R. Brooks, and F. W. Carson.
1989.Theoretically determined three-dimensionalstructuresfor amphipathic segments of the HIV-1 gp4l envelope protein. AIDS Res. Hum. Retroviruses 5:7-22.
40. Willey, R. L., J. S.Bonafacino,B.J.Potts,M.A.Martin,and R. D.Klausner. 1988.Biosynthesis, cleavage, and degradation ofthe humanimmunodeficiency virus1 envelopeglycoprotein gpl60.Proc. Natl.Acad. Sci. USA 85:9580-9584.
41. Zack,J. A., S. J.Arrigo,S. R. Weitsman, A. S.Go,A.Haislip, and I. S. Y. Chen. 1990. HIV-1 entry into quiescentprimary lymphocytes: molecular analysis reveals a labile, latent viral structure. Cell 61:213-222.