JOURNAL VIROLOGY, Mar. 1981, 922-930 0022-538X/81/030922-09$02.00/0
Analysis
of the
Virogenes
Related
to
the Rhesus
Monkey
Endogenous
Type
C
Retrovirus in
Monkeys
and
Apes
MICHAEL A.TAINSKY
BiologicalCarcinogenesisProgram,Frederick Cancer ResearchCenter,Frederick, Maryland21701
Molecularhybridization studieswerecarriedout
by
using
a[3H]complementaryDNA (cDNA) probe to compare the endogenous type C retrovirus of rhesus
monkeys (MMC-1) with other known retroviruses and related sequences in
various primateDNAs. The genomicRNA of theendogenous typeC retrovirus
ofstumptail monkeys (MAC-1) was found to be highlyrelated to the MMC-1
cDNA probe, whereasthe other retroviral RNAs tested showed no homology.
Relatedsequences were found in Old WorldmonkeyDNAs and to a lesserextent
in gorilla and chimpanzee DNAs. No homology wasdetected between MMC-1
cDNA and DNA ofgibbon,orangutan,orhumanorigin.Restriction endonuclease
analysisofgenomicDNAindicated thatmany oftheseveralhundredsequences
related to MMC-1 in rhesus monkey DNA differed from that
integrated
intoDNA of infected canine cells. Gorillaand
chimpanzee
DNAs containedaspecificrestriction endonucleasefragmentofthe MMC-1genome.
Endogenous retroviruses have been isolated
from tissue of many sources by cocultivation
withcelllines fromheterologous species
(16)
orby induction withhalogenated pyrimidines (1,
8). ThefirstreportedendogenoustypeC
retro-virus from a rhesus monkey was obtained by
cocultivation ofarhesus
monkey
(Macacamu-latta)
esophageal
carcinomacellline(816A)(11)
withacanineosteosarcomacell line
(D17) (10).
Sequences relatedto the genomeofthis
virus,
called MMC-1, were shown
by
molecularhy-bridizationtobepresent inthe DNA of rhesus
monkeys. MMC-1 has the
morphology
ofatypeC virus and contains the
expected
RNA-depend-ent DNA polymerase activity. In
addition,
MMC-1reactswitha
broadly
reactiveinterspe-cies antiserumtothep30protein ofmammalian
typeCviruses (10). Ahighly relatedretrovirus
hasbeenisolatedfrom M. artoides (15).
Ireporthere thepresenceandcharacteristics
ofMMC-1-relatedsequencesinthegenomesof
otherprimates,aswellastherelationship of the
MMC-1genome to that of otherprimate
retro-viruses. Molecular hybridization showed that
MMC-1-related sequences were present in
DNAsisolatedfromOldWorldmonkeys and to
alesser extent in DNAs ofgorillasand
chimpan-zees. In addition, MMC-1-related sequences werefound in catliverDNA.
To better understand the observed hybridi-zation of MMC-1 cDNA withgorillaand
chim-panzeeDNAs,restrictionendonuclease analyses
were carried out. MMC-1 genomes present in the chromosomal DNAof infectedcanine cells
were mapped by hybridization of
[32P]comple-mentary
DNA (cDNA) tonitrocellulose filters bearing immobilized DNA fragments produced by seven restriction endonucleases. This patternwasthen comparedwith cleavage patterns
ob-tained with DNA from normalrhesus monkey
(which contains several hundred MMC-1-re-lated sequences [10]); significant differences were found. A similar analysis of DNAs from gorillas andchimpanzees,whichcontain
MMC-1-related sequences, indicated that a specific
restriction fragment from the MMC-1 genome
had been conserved in these apes. This
con-servedfragmentwasdistinct from that observed
when restriction fragments from gorilla and
chimpanzeeDNAs werehybridizedto ababoon
endogenousvirus(BaEV) cDNA probe.Nosuch
related region was found in human DNA.
MATERIALS AND METHODS
Virusesandcells.MMC-1-infected canine thymus cells (Cf2Th) were propagated in Earle modified Eagle medium containing 10% fetal calf serum, 2 mM gluta-mine, 100
/tg
of streptomycin per ml, and 100 U of penicillinperml. Cellswerepassagedat a 1:10dilution weekly. Virus was harvested at 24-h intervals after the cells had reached confluency.Viruswaspurifiedasdescribed by Benton et al. (2). Other viruses usedwere:Rauschermurine leukemia virus grown inJLS-V9 BALB/c mouse bone marrow cells;Mason-Pfizer monkey virus grown in the human rhabdomyosarcoma cellline A204; BaEV strain M7 grown in the human osteosarcoma cell line HOS; RD114 grown inRDcells; equine infectious anemia virus grown inequine fetal kidney cells; bovine leu-kemiavirus grown in fetal lambkidney cells; reticu-loendotheliosis virus grown in transformed chicken bonemarrowcells; gibbon apeleukemiavirusgrown
922
on November 10, 2019 by guest
http://jvi.asm.org/
in 6G-1 gibbon T lymphoblasts; feline leukemia virus grown in FEAfeline cells; rat leukemia virus grown in Fisher rat embryo cells; squirrel monkey retrovirus grownin Cf2Th canine thymus cells; and M. artoides endogenous virus grown in A549 human cells.
Synthesis of MMC-1 [rHcDNA. MMC-1 [3H]-cDNA wassynthesized in a 0.250-ml reaction mixture containing1mMdATP, dCTP, and dGTP, 0.15 mM [3H]dTTP(44Ci/mmol), 0.04% Nonidet P40,0.05M
Tris-hydrochloride (pH 8.3), 5 mM magnesium ace-tate, 0.05 MNaCl, 0.01 Mdithiothreitol, 0.100 mg of actinomycin D perml, and approximately 2.3 mg of virus. After 2 h, sodium dodecyl sulfate (SDS) was addedto1%, and the mixturewascentrifuged at 40,000 rpmfor 2.5 h at 10°C in an SW41 rotor through a 15
to 30%sucrose gradient. A sharp peak of trichloro-aceticacid-insoluble radioactivity was observed in the middle of the gradient. The pooled fractions were incubated in 0.3 N NaOH for 3hat37°C,neutralized, and extracted with phenol followed by chloroform. The[3H]cDNAwascollectedby ethanol precipitation. Theseconditions generate probes with a high degree ofrepresentation andaspecific radioactivity of 3x107
dpm/iLg
(12).Preparation of nucleic acids for molecular hy-bridization. Viral RNAwaspreparedasdescribed by Schlometal.(13), and DNAwasprepared according
to the urea-phosphate method (5). Hybridizations
wereperformed with 1x103to 3x 103 cpmof [3H]-cDNA and100 to 500 ugofDNA or 0.5/g of RNA in
1Msodium phosphate buffer, pH 6.8, at650C.After appropriate times, thesamplesweredilutedto0.14 M phosphate buffer and 0.01%SDS, appliedto hydrox-yapatitecolumnsat500C, andeluted at 100C incre-ments.Fractionswereassayed for 3Hcountsper
min-ute after addition of2 volumes ofAquasol II. The
extentofhybridizationisdefinedasthe percentage of total radioactivity eluting from the column above 500C, andCot values are corrected to the standard conditions of0.12Mphosphatebuffer(5).
Preparation of high-molecular-weight DNA forSouthern blotanalysis. Packedcells were di-luted1:10in50mMTris-hydrochloride (pH 7.4)-100 mM NaCl-1 mM EDTA (TNE), and proteinase K (100
jg/ml).
SDSwasaddedto 1%,and the mixturewasincubatedat420Cfor2h. Aftertwoextractions
withphenol-chlorofoym(1:1), the aqueousphasewas
extracted once with chloroform and dialyzed three times against TNE. RNase A (boiled for 2min to
inactivate DNase) was added to 25 ug/mL, and the solutionwasincubatedat420Cfor2h.Theresulting mixture wasextractedasbefore with
phenol-chloro-form anddialyzedagainst10mMTris-hydrochloride,
pH 7.4, and1mMEDTA. The abovepreparationwas
modified for tissue DNA by firsthomogenizing the mincedtissue in aBrinkmannPolytronhomogenizer
at00C,using1g of tissueto10ml ofTNE.
Probe synthesis for Southern blot analysis.
MMC-1[32P]cDNAwaspreparedinareaction mixture containing80jgof 70SvirionRNAperml,400jgof calf thymus DNA primers per ml, 0.1 mM dGTP,
dATP, anddTTP,50mMTris-hydrochloride, pH8.3,
50mM KCl, 8 mMMgCl2, 8 mMdithiothreitol, 10
mCi of[a-3P]dCTPper ml(-400Ci/mmol),and1,000 Uof avianmyeloblastosisvirusreversetranscriptase
perml. After 8 h at370C, the reaction mixture was phenol extracted and the aqueous phase was treated with 0.3 MNaOH for 5 min at1000C.ThecDNA was purified by Sephadex G-25 chromatography in 3x SSC (SSC = 0.15 NaCl plus 0.015 sodium citrate). The specific radioactivity of this cDNA probe was 3 x 108 dpm/yg.
Restriction endonucleasedigestion.Restriction enzymes, obtained from New England Biolabs and Bethesda ResearchLaboratories, were used under re-action conditions as specified by the manufacturer. Completeness of digestion was monitored by including phage A DNA in each reaction and analyzing the fragmentation patterns on gels. Electrophoresis was carried out in 40 mM Tris acetate, pH 7.5, 5 mM sodiumacetate, and1mM EDTA, using 1% agarose (Seakem ME grade). Phage A DNA or4X174 replica-tive-form DNAcutwith various restriction enzymes was used as a size marker. Eucaryotic DNAs were used at 1 to 7
jig
perlane. Blot analysis was carriedoutby the method ofSouthern (14). Nitrocellulose filters (Schleicher &SchuellCo.) were hybridized for 16to24hinasolution containing 5x SSC, 0.2% bovine
serumalbumin, 0.2%Ficoll 400, 0.2% polyvinylpyrrol-idone, 20ugofcalf thymus DNA per ml (sonicated anddenatured),20ygof Cf2Th caninecell total RNA per ml, 0.1% SDS, and 20ngof
[nP]cDNA
per ml. Washed nitrocellulose filters were analyzed byra-dioautography, using Kodak XR-5 film and du Pont Lightning Plus intensifying screens.
RESULTS
Molecular hybridization. The relationship
ofMMC-1 toother known viruseswasstudied
by nucleic acid reassociation, using MMC-1
[3H]cDNA
and RNA extracted from purifiedvirions. When
assayed
byhydroxyapatite
chro-matography, the MMC-1 [3H]cDNA probewas
homologoustoMMC-1andMAC-1 viral RNAs
(10). Undertheseconditions, no hybrid
forma-tion was observed with RNA extracted from
other retroviruses, including bovine leukemia
virus, BaEVstrainM7, equineinfectiousanemia
virus,Mason-Pfizer monkey virus,feline
leuke-mia
virus,
feline sarcomavirus,
rat leukemiavirus,gibbonape leukemiavirus,Rauscher
mu-rine leukemia
virus,
and thesquirrel
monkey
retrovirus.
To assay for the presence of viral gene
se-quencesrelatedtoMMC-1 in other animal
spe-cies,
the[3H]cDNA transcript
wasannealedtohighly
purified
DNApreparations.
TheMMC-1[3H]cDNA hybridized
essentiallycompletely
to canine cells infected with MMC-1(Table
1),
whereas thesecells containednorelatednucleic acid sequencesbeforeinfection. In
addition,
high
levelsof
hybridization
wereobservedwithDNAsfrom various Old World
monkeys
(i.e.,
rhesusmonkeys,
stumptail
monkeys, baboons,
andSykes
monkeys).
Theseresultsarequalitatively
similar tothoseof Todaroetal.
(15)
in thestudy
on November 10, 2019 by guest
http://jvi.asm.org/
924
TAINSKYTABLE 1. HybridizationofMMC-I[3HJcDNAto
cell and tissue DNAs
DNAsource %
Hybridi-
zation OldWorldmonkeysMacaca mulatta liver(rhesus
monkey) ... 100
M. mulattaspleen (rhesus monkey) 99 M.mulattakidney (rhesus monkey) 90 M.arctoides MA250 cells(stumptail macaque)... 89
Papiohamadryas kidney(Russian baboon) ... 90
Cercopithecusalboqulariskidney (Sykes monkey) ... ... 81
NewWorldmonkeys Lagothrix logothricha kidney(woolly monkey) ... 4
Aotustrivirgatuskidney(owl monkey) ... 4
Apes Gorillagorillamachicells(gorilla) 38 Pantroglodyteskidney(chimpanzee) 31 HomosapiensA204cells(human) ... 3
Hylobates larkidney (gibbon ape) ... 4
Orangutan CP81 cells ... ... 4
Nonprimates Musmusculus kidney (BALB/c mouse) ... 8
M. musculus liver(AKR mouse) ... 8
Canisfamiliarisliver(dog) ... 4
Feliscatus(cat) ... ... 42
Bostaurus(calf) ... ... 4
Salmon ... 4
Control cell lines D17caninecells ... ... 3
D17caninecells producing MMC-1 .. 97
Cf2Th canine cells ... ... 3
Cf2Th caninecellsproducingMMC-1 90
of another macaque endogenous virus isolate, MAC-1. DNAs from NewWorld monkeyswere
also tested for sequence relatedness to the MMC-1 genome, but neither woolly nor owl
monkey DNAs showed hybrid formation. MMC-1 [3H]cDNA reassociated with DNA purified from some ape tissues and cells. Chimpanzee
andgorilla DNAshybridizedtoasimilarextent (31 and 38%, respectively) with the MMC-1 probe. No hybrid formationwas observed with
DNAfromhumans,orangutans,orgibbonapes,
evenunderthese conditions of low stringency.
DNAs extracted from various nonprimates, i.e., mice, dogs, calves, and salmon, were also
examined forsequencerelatednesstothe MMC-1genome,butnohomologywasobserved.
Sim-ilar to MAC-1 (4), asurprisingly high level of
homology was found in DNA extracted from
domestic
catliver(42%).
Nucleic acid reassociation kineticswereused
todetermine thereiterationfrequency of
MMC-1-relatedsequences intheDNAof
primates
andMMC-1-infected
cells. These sequences were present inmultiple copies
(200 perhaploid
ge-nome) in the genome of the species of
origin,
rhesusmonkeys (10).Inaddition, DNA samples
from other Old World monkeys were tested;
baboons, Sykes monkeys, and African green
monkeys
all contained MMC-1-related se-quences inmultiple
copiesranging fromapprox-imately40 to 300copiesperhaploidgenome.In
the
MMC-i-infected
Cf2Th canine cellline,
MMC-1
sequences occurred at afrequency
ofabout30
copies
perhaploid
genome.DNAfromhuman HOS cells
productively
infected withMMC-1 virus were found to contain about 20
copies of MMC-1-related viralgenes perhaploid
genome.Inagreementwith Bonner and Todaro
(4), who found MAC-1-related sequences
pres-entinmultiple copiesin catDNA, MMC-1 DNA
sequences werefoundtobe reiterated incatliver DNA at
approximately
150 copies per haploidgenome.
As reported previously (10), the MMC-1
ge-nomesharedextensivesequencehomology with
the endogenous virus of stumptail macaques,
MAC-1. The thermal stabilities of the hybrids
formed betweentheMMC-1 probe andgenomic
RNAsfrom thesemacaqueendogenous viruses
were foundtobeidentical within experimental
variation (data not shown). This is consistent
with the observedsimilarity of the proteins of
the two viruses asanalyzed onpolyacrylamide
gels
(10).Genomic
organization
of integrated
MMC-1
proviruses. High-molecular-weight
DNA isolated from canine
cells
chronically
in-fected with MMC-1 was fragmented by using
restriction enzymes andanalyzedby
hybridiza-tiontoSouthem blots, usinga
[32P]cDNA
probe.Thespecificity of the MMC-1
['P]cDNA
probewas tested by comparison of the hybridization
to DNA
from
uninfected and infected caninecelis
cleaved witheither EcoRI or HindIII. Nobandswereapparentwhen uninfected
cell
DNAwas analyzed (Fig. 1). In contrast, analysis of
infected cellDNAyieldedasingleintense band
for eachoftheseenzymes. The EcoRI
fragment
was 2.2 kilobase pairs (kbp), and the HindIII
fragmentwas 6.4kbp.
HindIII,
EcoRI,
andXhoIcleaved each of theintegrated MMC-1 genomes twice, generating
single intense bandswithmolecular sizes of 6.4,
2.2, and 1.3 kbp, respectively (Fig. 2).
When
EcoRI and
HinduII
were used in combination,on November 10, 2019 by guest
http://jvi.asm.org/
bands of2.4, 2.2,and1.8kbpwereobserved.The
2.2-kbp
band represented the EcoRI cleavageproduct band, and the 2.4- and 1.8-kbp
frag-ments were on either side of it, derived from the
HindIII 6.4-kbp band. When XhoI and HindIII
were usedin double-digestion experiments,the
1.3-kbp XhoI bandwasobserved alongwith two
other fragments of3.55 and 1.55 kbp (Fig. 2), which were oneither side ofthe 1.3-kbp
frag-ment within the HindIII 6.4-kbp fragment.
When EcoRI and XhoI were used in
double-digestion experiments,adoublet band of1.1kbp
was observed. Therefore, the XhoI had one
cleavage siteinthe middle of the EcoRI 2.2-kbp
fragment and one site very close to the one
EcoRI cleavage
site,
yielding a fragment toosmalltobeobserved. When EcoRI,HindIII, and
XhoIwereused in
triple-digestion experiments,
bandswereobservedat 2.4and1.55kbp,aswell
as the
EcoRI-XhoI
1.1-kbp doublet. The1.55-kbpfragmentarosefromXhoI-HindIIIcleavage.
The2.4-kbp fragmentwasgenerated by cleavage
by
EcoRI
and HindIII and must have comefrom the region of the
XhoI-HindIll
3.55-kbpfragment.
The restriction enzyme BamHI cleaved the
genomethree timestoyieldtwo
fragments,
2.65and0.95 kbp (Fig. 3). Indouble-digestion
exper-iments
withHindIIIandBamHI, twoadditionalfragments,1.15and1.65kbp,wereobserved(Fig.
3). These arose from sequences between the
outside BamHIsites and the two HindIII sites.
WhenBamHIandXhoI were used in
combina-tion (Fig. 3), three major bandswere detected,
at lengths 1.45, 1.25, and 0.95 kbp. Therefore,
theBamHI fragmentat2.65 kbp was cleaved by
XhoI intotwofragments, of lengths1.45and1.25
kbp. The 1.25-kbp band arose from a BamHI
site within theXhoI 1.3-kbp
fragment.
Thisfrag-ment(lane 3) had aslightly slowermobility than
the 1.25-kbpfragment.
SacI cleaved thegenomefourtimes,
generat-ingthreefragmentsoflengths5.6, 1.83, and 0.7
kbp (Fig. 4). WhentheSacI digestwasfurther
cleaved by XhoI, the0.7-kbp
fragment
remainedunchanged.A new0.95-kbp fragment appeared
fromcleavage of the 5.6-kbp by
XhoI.
There-maining0.38-kbp band resulting from cleavage
of the SacI1.83-kbp band into the 1.45-kbp band
wasonly faintly observed afterprolonged
expo-sure. When the SacI digest was treated with
SmaI,
the1.83-kbp fragment
was cleaved intotwo
fragments
of1.3and0.53kbp (Fig. 4). SmaIalso cleaved the
HindIII
6.4-kbp fragmentonce,yieldinga4.95-kbp anda1.35-kbp band. When
Infected Cells
Hind III RI
Uninfected Cells
Hind III RI
Hind III
+
Hind IllHindIII RI
RI
+ + + +
[image:4.488.70.214.356.615.2]Hindi11l Rl Xho I Rl Xho I Xho I Xho I
FIG. 1. Autoradiograph ofSouthernblotof
restric-tion endonucleasedigestion of MMC-1-infectedand
[image:4.488.249.442.369.608.2]uninfectedcanineCf2Thcells.RI,EcoRI.
FIG. 2. Autoradiograph ofSouthern blotof
restric-tion endonuclease digestion of MMC-1-infected
ca-nine
CQ2Th
cells. Kilobasepair size estimates areindicatedontheright.RI,EcoRI.
Kbp
-6.4
-3.55
-2.4 -2.2
-1.55
-1.3
-1.1
on November 10, 2019 by guest
http://jvi.asm.org/
926 TAINSKY
BAM H- I + BAM Hl1 Xho I
.: _
BAM H-Xho I Hind III
Kbp
-6.4 -4.95
-3.55
-2.65
[image:5.488.276.415.67.580.2]- 1.65 -1.3 -1.15 -0.95
FIG. 3. AutoradiographofSouthern blotof restric-tion endonucleasedigestion ofMMC-1-infected
ca-nine Cf2Th cells. Kilobase pair size estimates are
indicatedontheright.
theXhoI-HindIII double
digest
wastreated withSmaI, the1.55-kbp side bandwasreduced toa
doublet of1.35and 1.3kbp (Fig. 5),thus
confirm-ingtheorientation ofthe
Sacd
1.8-kbp fragment
asoverlapping the HindIII-XhoI
1.55-kbp
frag-ment. SmaI treatment of the BamHI-HindIII
doubledigest resultedincleavage ofthe
flanking
1.65-kbp band intoa1.35-kbpfragment identical
with theHindIII-SmaI 1.35-kbp band. The
po-sitionof theSacl 0.7-kbp
fragment
wasarbitrar-ily assigned to the right end ofthe map. The
presence ofSacIsitesatthe ends of the
MMC-1 genome was confirmed by Lovinger and Schochetman (7), who found the site in a se-quence of the MMC-1 strong stop DNA.
PstI treatment of the HindIII-XhoI double
digestresulted in loss of the3.55-kbp fragment
andthe appearance ofa newband of3.05 kbp
(Fig.6). Further evidence for the position of the
PstI cleavage site came from PstI and SmaI
doubledigestion,whichyieldedoneband of4.55
kbp (datanotshown). From thesedata,a
restric-tion enzyme mapwas constructed for much of the genomeintegratedininfectedcells(Fig.7).
Sacl
XhoI
SacI
Sacl + XhoI
SmaI
-5.6
-4.55
-3.05
-1.83
-1.45
-1.3
- .7
-1.3
-0.95
-0.70
-0.53 -0.38
FIG. 4. AutoradiographofSouthern blot ofrestric-tionendonuclease digestion ofMMC-1-infected ca-nine Cf2Th cells. Kilobase pair size estimates are indicatedon the right. Lower portion was exposed longertoobserve weaker bands.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:5.488.73.218.71.352.2]BAM-HI
Xho I
+ +
Hind III
Hind III
+ +
[image:6.488.63.216.53.490.2]SMA I SMA I SMA I SMA I
FIG. 5. Autoradiograph ofSouthernblotof restric-tion endonuclease digestion of MMC-1-infected
ca-nine Cf2Th cells. Kilobasepair size estimates are
indicatedontheright.
Structuralanalysisof MMC-1-related se-quencesinprimateDNAs. DNAwasisolated
from a rhesus monkey embryonic kidney cell
line (MA104) and digested with EcoRI and HindIII. The major restriction fragments
ob-servedpreviously in infected canine cell DNA were alsoobserved inSouthern blots of rhesus monkeycell DNA (Fig. 8).Additional hybridiz-ing high-molecular-weight fragments were ob-served which couldbeaccounted forbyseveral
FIG. 6. Autoradiograph ofSouthern blotof restric-tion endonuclease digestion ofMMC-1-infected
ca-nine Cf2Th cells. Kilobasepair size estimates are
indicatedontheright.
explanations (e.g.,multiple flanking bands,
het-erogeneityamong genomes, etc.). However,no
furtherexperiments were carried out toclarify theirorigin. When baboon DNAwasanalyzed,
theEcoRI 2.2-kbp bandwasobserved, but not
theHindIII 6.4-kbpband(datanotshown). In XhoI digests, the 1.3-kbp fragment
ob-Xho I
Hind
IIIPST-I
PST-I
Kbp
-3.55
-2.65
-1.65
- 1.3
- 1.1
- .9
Kbp
-3.55
-3.05
-1.55
-1.3
on November 10, 2019 by guest
http://jvi.asm.org/
[image:6.488.283.405.63.520.2]928 TAINSKY
H P B B R
Kbp aSa
2 3
X SR BX SMA
4 5
0
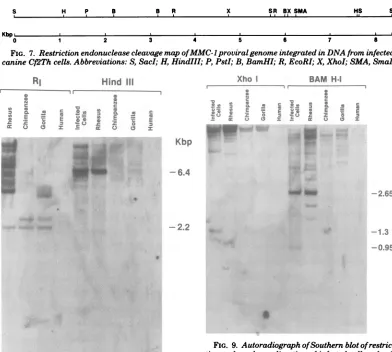
FIG. 7. Restriction endonucleasecleavagemapofMMC-lproviralgenome integrated in DNAfrom infected
canineCf2Thcells. Abbreviations:S,SacI;H,HindIII;P, PstI;B,BamHI; R, EcoRI; X, XhoI; SMA,SmaI.
RI
Hind III~~~~I
:
c c E EB * E
I
E.C C 2 z
cc 0 a ccE 0 0
Kbp
-6.4
[image:7.488.53.445.63.415.2]-2.2
FIG. 8. Autoradiograph ofSouthern blotof restric-tionendonucleasedigestion of infectedcelland
pri-mateDNAs.RI,EcoRI.
served in infected cell DNA was not seen in
rhesus monkeyDNA. WhenBamHIwasused,
theBamHI2.65-kbp fragmentwasseen,but not the 0.95-kbp fragment (Fig. 9). The additional BamHI fragments observed indicated the het-erogeneity of the MMC-1-related sequences in rhesus monkey DNA. No bandswereobserved
when rhesus monkey DNA was digested with
SacI (datanotshown).
Asreported here, DNAs fromtwoapes,
goril-las and chimpanzees, hybridized with MMC-1 [3H]cDNA; however, those from humans,
orang-utans,andgibbonapesdidnot.Restriction anal-ysisofapeDNAsindicated thattheEcoRI 2.2-kbp fragment found in rhesus monkey and in-fectedcellDNAwasalsopresentingorilla and chimpanzee DNAs (Fig. 8). In addition, an
Xho I
1, a
1= : E! cc0
I I BAM H-I
I
a C. 1. 1
. a
E
20 P.
a .5 0
-2.65
-1.3
-0.95
FIG. 9. Autoradiograph ofSouthernblotof restric-tion endonucleasedigestion of infected-celland
pri-mateDNAs. Kilobasepairsize estimates are indi-catedontheright.
EcoRI fragment of 2.5 kbp was seen that was
common to both ape DNAs. The EcoRI frag-ment at 3.9kbp (Fig. 8) found by using gorilla muscle DNA was also observed in analysis of DNA from aherpesgorilla-transformed gorilla
cell line (datanotshown) and thusappearedto begorilla specific. HindIII (Fig. 8), BamHI, and XhoI (Fig. 9) cleavageof theapeDNAsdidnot revealanyDNAfragments which could be
cor-related with the integrated genome from
in-fectedcaninecells(Table 2). It should be noted
that those ape DNAs which hybridized with
[3H]cDNA in solution hybridization experiments
werepositiveonSouthem blots also. Even with enzymesforwhich no internalbands were
ob-served,somehybridization with
high-molecular-weight DNAs was seen. In contrast, human DNAwasuniformly negative both in blots and
insolutionhybridization.
HS S
W. I I . . - II
--S
i
5 7 a
I
on November 10, 2019 by guest
http://jvi.asm.org/
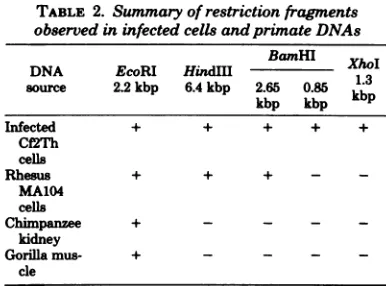
[image:7.488.244.446.117.401.2] [image:7.488.53.249.141.432.2]TABLE 2. Summary of restrictionfragments observed in infected cells andprinateDNAs
BamHI
DNA EcoRI HindM
1X
source 2.2kbp 6.4kbp 2.65 0.85
kb3p
kbp kbpkb
Infected + + + + +
Cf'2Th cells
Rhesus + + + - _
MA104 cells
Chimpanzee +
kidney
Gorilla mus- + - - -
-cle
Both MMC-1and the BaEV probeshybridize
to
African
apeDNAs (3). To determinewhetherthe
homology
istoidenticalapeDNAsequences,Southern blots of restriction enzyme DNA
di-gests of primate DNAs were annealed with
BaEV (M7) cDNA probe. Althoughthisprobe
hybridized to somefragments, the 2.2- and
2.5-kbp fragments were not detected in digests of
anyofthe
primate
DNAs. Therefore, theretro-virus-relatedsequences in the African apesfall
intoatleasttwosubsets:those which are related
toMMC-1 and those whicharerelatedtoBaEV.
DISCUSSION
Onepropertyofagenetically transmitted
ret-rovirus is thepresence of cross-hybridizing
nu-cleic acid sequences in closely related species.
MMC-1-relatedsequences werefound in
multi-ple copies in
all
Old World monkeys thus faranalyzed. In addition, MMC-1-related nucleic
acidsequences were presentinchimpanzees and
gorillas. No homology was observed in DNA
fromhumans, gibbonapes, or orangutans. Itstill
remainspossible thatamoderatelymismatched
retrovirus-related sequence present in human
DNA has eluded analysis because ofthe
insen-sitivity of thepresentmethodologies.Since
mis-matching affects the stability of nucleic acid
hybrids, onedoesnotknow whether the38and
31%
hybridization
found ingorillas
andchim-panzees,
respectively,
isrelatedtothe fact thatsomeof theregions ofMMC-1-relatedsequence
divergeextensively, with others being relatively
conserved, or tothe fact that the entire region
diverges to an intermediate extent (assuming
that these sequences and MMC-1 arose from a common progenitor). A cleavage map of the
integrated genome from infected canine
cells
was prepared to compare the MMC-1 genome
withrelatedsequences observed in various
pri-mates.
Theobservation that asmallbut discrete
re-gionof the MMC-1proviralgenome, theEcoRI
2.2-kbp fragment, was found in Southem blot
analysis of both chimpanzee and gorilla DNA
couldreflectaspecific regionalconservation in
these ape DNAs. This wasconsistent with
so-lution hybridization data with ape DNA. The
absenceofXhoIandBamHI
fragments
from thesame
region
of the viral genomeas the EcoRI2.2-kbp fragment indicates atleastsome
diver-gence within these
specific
sequences in DNAfrom these apes. Structural analysis of the
en-dogenous virogenes relatedtoMMC-1inrhesus
monkey DNA revealed some differences in
re-striction endonuclease cleavage patterns when
compared with integrated proviral genes from
infected canine cells. Theprecise determinants
for these differences are as yet undefined. A
possibility is that the endogenous virogene
cop-ies in rhesus monkey DNAare heterogeneous
with respect to at leastsome DNAsequences. These
small
differences manifest themselveswithin thisanalytical framework by alterations
inthe recognition sequences at restriction
en-donuclease cleavage sites such thattheyare no
longer cleaved. Therefore, one could postulate
that thecopyof thelarge number ofendogenous
virogenes (200) whichgaveriseto aninfectious
viruswasacopy presentinlowfrequency. If this
putative infectious virogene were even at the
level of a few percent of the total number of
endogenous
copies,
itwouldnotbedetected inthepresent
analysis.
Analternativehypothesis
for the difference incleavage
patterns
betweenMMC-1-infected canine cell and rhesus
monkey
cell DNAs is that
methylation-specific regions
inhibit endonuclease cleavage in some loci of
MMC-1-related sequences of rhesus
monkey
DNA.Cloning of these DNA
fragments
followedby sequence
analysis
will enable resolution oftheabovealtematives.
Hybridization of MMC-1
probes
to DNAsfrom
chimpanzees
andgorillas
andnot toDNAfromhumans,
gibbon
apes,ororangutansfollowsaninterestingpattern similartothatseenwith
the baboon viruses
(3).
These nucleic acidse-quencesfrom rhesus
monkeys
arehomologous
to DNAs from two African apes,
gorillas
andchimpanzees, butnot toDNAs from the Asian
apes, orangutansand
gibbon
apes. Humansarebelievedto have arisen in Africa
and,
withre-spect to the presence of MMC-1-related
se-quences,are different from the Africanapes. It
ispossiblethat lowlevelsof
homology
exist inallapesandthat thepresent
hybridization
meth-ods aretooinsensitivefor suchananalysis.
ACKNOWLEDGMENTS
Ithank Raymond V. Gilden, Nancy Rice, and Maurice Cohen for theirexcellent advice and discussion during the
on November 10, 2019 by guest
http://jvi.asm.org/
[image:8.488.46.239.70.213.2]course of thisstudy, MartinZweig for careful reading of the manuscript, and ToddAllen for technical assistance.
LITERATURE CITED
1. Aaronson,S. A., G. J. Todaro, and E. M. Scolnick. 1971.Induction of murine C type virus fromclonal lines ofvirus-freeBALB/3T3 cells.Science 174:157-159. 2. Benton, C. V., H. M. Hodge, and D. L. Fine. 1978.
Comparative large scale propagation of retroviruses from OldWorld(Mason-Pfizer monkey virus) and New World (squirrel monkeyvirus) primates. In Vitro 14: 192-199.
3. Benveniste, R. E., andG. J. Todaro. 1976. Evolution of typeC viral genes evidence foranAsianorigin ofman.
Nature(London) 261:101-108.
4. Bonner, T. I.,and G. J. Todaro. 1979.Carnivores have sequencesin their cellular DNAdistantly related to primate endogenous virus, MAC-1. Virology 94:224-227.
5. Britten,R. J., D. E. Graham,and B. R.Newfield. 1974.Analysis ofrepeating DNA sequences by reasso-ciation. MethodsEnzymol. 29:363-418.
6. Charman,H.P., M. B.Gardner,R. M.McAllister,N. Kim, and R. V.Gilden. 1976. Humoral immune re-sponses ofcats tomammalian type C virusp30s. Int. J. Cancer 17:98-108.
7. Lovinger, G. G., and G. Schochetman. 1980.5' Termi-nal nucleotide sequences of type C retroviruses: features common tononcoding sequences of eucaryotic messen-gerRNAs. Cell20:441-449.
8. Lowry,D.R.,W. P.Rowe,N. M.Teich,and J. W. Hartley. 1971. Murine leukemia virus: high frequency activation in vitro by 5-iododeoxy-uridine. Science 174: 155-156.
9. Oroszlan, S.,and R. V.Gilden.1980.Primarystructure
analysis of retrovirusprotein,p.299-343.InJ. R. Ste-phenson (ed.), Molecularbiologyof RNAtumorviruses. Academic Press,Inc., New York.
10. Rabin, H., C. V. Benton, M. A.Tainsky,N. R.Rice, and R. V. Gilden. 1979. Isolation and characterization of anendogenous virus fromRhesus monkeys. Science 204:841-842.
11.Rabin, H., R. H. Neubauer, M. A. Gonda, W. A. Nelson-Rees, H. P.Charman,and M.G. Valerio. 1978.Spontaneousesophageal carcinoma andepithelial cell lineofanadult rhesusmonkey.Cancer Res. 38: 3310-3314.
12.Rice,N. R., S.Simek,0. A. Ryder, and L.Coggins. 1978.Detection ofproviral DNA in horse cells infected withequine infectious anemia virus. J. Virol.
26:577-583.
13.Schlom,J., D.Colcher,S.Spiegelman,S.Gillespie, andD.Gillespie. 1973. Quantitation of RNA tumor viruses andvirus-like particles in human milk by hy-bridization topolyadenylic acid sequences. Science 179:
696.
14.Southern, E. M.1975. Detection ofspecific sequences among DNAfragments separated by gel electrophore-sis. J. Mol. Biol.98:503-517.
15.Todaro,G.J.,R.E.Benveniste,S. A.Sherwin,and C. J. Sherr. 1978. MAC-1, a newgenetically transmit-ted type C virus of primates: "low frequency" activation from stumptail monkey cell cultures. Cell 13: 775-782.
16. Todaro,G. J., C. J.Sherr,and R. E. Benveniste. 1976. Baboons and their close relatives are unusual among primatesintheirabilitytorelease nondefective
endog-enoustypeC viruses.Virology72:278-282.
J. VIROL.