0022-538X/84/07042-05$02.00/0
Copyright C) 1984, American Society forMicrobiology
Valyl-tRNA Synthetase
Modification-Dependent
Restriction of
Bacteriophage
T4t
NORMA J. OLSON* ANDGEORGE L. MARCHIN
Division of Biology(Microbiology), Kansas State University, Manhattan, Kansas66506 Received 7 November 1983/Accepted 25 March 1984
AstrainofEscherichia coli, CP790302, severely restricts the growth of wild-type bacteriophageT4. In broth
culture,mostinfections of single cellsareabortive, althoughafewinfected cells exhibit reducedburstsizes.In contrast,bacteriophage T4mutantsimpairedin the abilitytomodify valyl-tRNA synthetase developnormally
onthis strain. Biochemical evidence indicates thatthephage-modified valyl-tRNA synthetase in CP 790302 is
different from that previously described. Althoughtheenzyme is abletosupportnormalproteinsynthesis, a
disproportionateamountofphage structural protein (serum blocking power) fails to matureintoparticlesof the appropriate density. The results with host strain CP 790302 ate consistent with either a gratuitous
inhibition of phage assembly by faulty modification or abrogation of an unknown role that valyl-tRNA
synthetase might normally playinviralassembly. Shortly after bacteriophage T4 infects Escherichia coli, thehost valyl-tRNA synthetase is modified by the addition ofaphage-coded peptide calledT(12). The phagegenevsat
56 kilobases on the standard T4 genetic map is responsible for thismodification and presumably codes for theTpeptide (14). Bacteriophage T4 mutants that are deficient in the
abilitytomodify the host synthetasegrowwellontypicalE.
colilaboratory strains. The biological role for modification is thus difficultto define.
We have nevertheless continued tostudy the valyl-tRNA
synthetase modification becausewefeelthat itmight repre-senta unique kind ofregulatory eventtothebacteriophage T4, and that it might beamodel forsynthetase alterations in a number ofbiological systems(20).
Recentwork inourlaboratory has demonstratedthat
wild-type bacteriophage
T4vs'
does not formplaques ona host strain of E. coli called CP 790302 (11). A variety of phagemutantsdeficient inmodification of hostvalyl-tRNA synthe-tase, however, plaque normally onstrain CP 790302.
Previ-ous genetic evidence had indicated that the vaiSts and relA
alleles in E. coli CP 790302 probably were responsible for
thisrestriction phenomenon.
In this article, we present evidence that the inability of T4vs+ toformplaquesonstrain CP 790302 is duetoagreatly
reducednumber of infectiouscenters,aswellastoareduced averageburstsize. Inaddition, the biochemical propertiesof thephage-modified valyl-tRNA synthetasein CP 790302are
different than those previously described foraT4-modified
valyl-tRNA synthetase (19). Thus, we hypothesize that the
novel valyl-tRNA synthetase, while continuing to amino-acylate tRNAs and participate in protein synthesis,
some-howinterferes with viraldevelopmentatalatestageinviral assembly; wedonot knowwhether this postulated
interfer-enceis direct orindirect.
MATERIALSAND METHODS
Bacteria and bacteriophage. Several strains of E. coliand bacteriophage T4wereemployedin this studyandare listed in Table 1. Cells were routinely grown in
morpholinepro-* Correspondingauthor.
tContributionno.84-208-J,Division ofBiology,Kansas
Agricul-tural Experiment Station, Manhattan, KS66506.
panesulfonic acid TGYE medium prepared according to
Neidhardtetal. (18). High-titer lysates of T4wereprepared
onE. coli NP 4aspreviously described (13),exceptthat 20-ml lots were used. Lysates were titrated on strain NP 4
unless stated otherwise.
Preparation of cell-free extracts. Cell-free extracts were
preparedasdescribed previously (16), by sonicating cultures on ice in phosphate buffer (0.006 M potassium phosphate, 0.006 M 2-mercaptoethanol [pH 7.2]). The protein
concen-trationwas determined by the method of Lowryetal. (10), andextractswere stored at4°C; they werenotusedafter 5 days.
Enzyme assays. The specific activity ofvalyl-tRNA syn-thetase aminoacylation was measured by attachment of
[3H]valinetotRNA. Reaction mixtures contained 250 ,ug of tRNA, 100 ,ug of bovine serumalbumin, 2 .mol of dipotas-siumATP, 1 ,umol of glutathione, 10 p.mol of[2,3-3H]valine
at 15
liCi/VLmol,
5 xmol of KCl, 50,umol of Tris base, and5,umol of MgCl2, with afinalpH of 7.2. A limitingamountof valyl-tRNA synthetase (cell-free extract) was addedat 4°C
togiveafinalvolumeof 0.5 ml. Blanks contained everything butenzyme. After 5 min of incubationat 37°C, the reaction
wasstopped by cooling quickly inan ice bath and adding 3
mlofcold 5%trichloroacetic acid. After 30min,precipitates werecollectedonfiber glass filters and washed with 10 ml of
5%trichloroacetic acidand 5 ml of 67% ethanol. Filterswere
driedat 80°C for 30 min and counted in a Beckman LS-230
liquid scintillationcounterwith 5 mlof toluene base scintilla-tioncocktail. One unit ofsynthetase specific activity is the amount of enzyme that catalyze the addition of valine to
tRNAattherateof 1 p.mol/min underourassayconditions. Experiments measuring aminoacylation specific activity in CP 790302, when indicated, were performed as described
above, except that a "protective buffer" containing 0.1 M
Tris base, 0.001 M valine, 10% glycerol, and 4.2% 2-mercaptoethanol, with afinal pHof 7.3, wasused.
Serum blocking experiments. Serum blocking power was
determinedby using the standard endpointassayofDe Mars (6) and procedures thoughtfully provided by M. Snyder. Phage lysates wereprepared by adding T4vs+orT4vs2,ata
multiplicityof infection(MOI) of7,to20 ml ofE. coliNP 4
orCP790302 broth culturesgrownat30°Ctoadensityof 2x 108 cells per ml. After 90 min of incubation at 30°C with
aeration, cells were lysed by the addition of chloroform.
TABLE 1. Strains ofE. coli andbacteriophage T4
Strain Source Description
T4Bc+ Thislaboratory Wildtype; normalmodification of VRSa
T4vsl This laboratory Missense mutant; unusual modified VRS
T4vs2 This laboratory Amber mutant; nomodified VRS T4vs2 This laboratory Spontaneous revertant of T4vs2;
RFOl modified VRS
T4rII M. Snyder rdf deletion
E. coli This laboratory Wild-typeE.coli B; indicator strain NP 4
E. coli This laboratory Wild-typeE.coliKB NP 2
E. coli This laboratory valS"s NP29
E. coli CP G.Bjork F-pyrB hismetBrelAvalS'" rpsL 790302 ampA(vaiS"s by P1 fromNP
910212)
E. coli NP Thislaboratory F- recA strA valS'"pyrB trpA 910212
E. coli M.Snyder Lambdalysogen; restrictive host
G(A) for T4rII
E. coli G.Bjork leuvalS's relA+ NE 536
E. coli G.Bjork leuvalSisrelA NF 537
aVRS, Valyl-tRNA synthetase.
Lysate sampleswere incubated withT4antiserum in serum
blockingbufferat48°Cfor 12htoallowacompletereaction.
The residual neutralizing activity in each sample was
mea-sured by adding a known concentration of purified (22)
(T4vs+) tester phage and determining the inactivation after
48minof incubationat46°C. PFUproduced onE. coli G (X)
by surviving tester phage in the lysate samples were
com-pared with PFU produced by surviving tester phage in samples containing a known concentration ofCsCl-banded
T4rII (rdf deletion). Testerphage (T4vs+) form plaques on
indicator strain G (A), but T4 rII does not. Thus, serum
blockingpowerconcentrations in thelysatescanbe
convert-edtophage equivalents by using standard curvesthat relate
PFU produced by tester phage to phage equivalents ofT4 rll. Standard curves were run in eachexperiment.
RESULTS
RestrictionofT4vs+ byCP790302. Workers in our
labora-tory have demonstratedthatE. coli CP 790302 is restrictive for T4vs+ but permissive for T4vs2 (11). A pair of E. coli strains(NF536and NF 537),whichcontain adifferent valS's
allelethan CP790302,permitT4vs+ and vs-mutantstoform
plaques efficiently. Consequently, the restriction appears to
be dependent uponthe valS'sallele employed in CP790302.
We haveverified this byPlvirtransduction ofvalS+ intoCP 790302 (11), as well as by crosses with a number of Hfr
strains (P4X, KL25, KL16-99, and Hfr H). In all cases,
T4vs+ formed plaques on the valS+ recombinants at a high
frequency. In contrast, his+, metB+, and pyrB+ recombi-nants, either alone or in combination, continued to restrict T4vs+ plaqueformation (datanotshown).Theimportanceof
therelA alleletothe restriction phenomenonis more ambig-uous. Although we were unable to directly select relA+
recombinantseither via PlvirtransductionorHfrcrosses, a
related strain, E. coliNP 910212, which contains the same
valS'sallele asCP 790302 but isrelA+,allowed formation of
minuteplaques with T4vs+.
TABLE 2. Virus production in E. coli CP 790302 during single andmixedinfectionsa
Virus produced
Infection perinfectious %Reduction'
center (PFU)
T4vs+ 9
T4vs2 290
T4vs+ plus T4vs2 57 80
T6 139
T4vs+ plusT6 19 86
T4vs2 plus T6 340 0
T7 119
aCells were grown at 30°C to a density of5x107cellsper ml, and identical
cultures were infected with one or more strains of virus each at an MOI of 4. After 10min, unadsorbed phage were inactivated with phage antiserum. Five minutes later, each culture was diluted and plated on wild-type indicator strain E.coli NP 4 to determine infectious centers. Cultures were diluted, and after 90min,samples were removed and plated on the indicator strain. Experimen-taldetails were described by Adams (1).
bThe data are expressed as the percent reduction in virus produced (per infectious center)during mixed infection compared with the corresponding single infection with T4vs2 or T6.
Infection andvirus progenyproduction.Five minutes after T4vs+ or T4vs2 phage were added to actively growing
cultures of E. coli CP 790302, approximately 80% of the phage wereadsorbed, both cultures experiencedadecrease in optical absorbance, and fewer than 1% of the bacteria survivedeitherinfection (datanotshown). Adding T4vs+ to lawns of CP 790302, however, failed to produce plaques,
whereas T4vs2 formed plaques efficiently on the E. coli
strain (11). This phenomenon was further investigated by
using single-cell burst and one-step growth experiments.
When acultureof CP790302 wasinfectedwith T4vs+ and
unadsorbed phage were neutralized with antiserum, plating infectious centers on the permissive host E. coli B (NP 4)
resultedinapproximately 25% (15 of66) as manyplaquesas when the infection was conducted with T4vs2. Ninety
min-utes afterT4vs2 additionto CP 790302, an average ofmore than 200 PFU per infectious center were observed. Fewer than 10 PFU per infectious center were produced after infection with T4vs+ (Table2).Chloroformadditiontoeffect
cell lysis did not increase the burst sizes.
Furthermore,
T4vs+ phage produced on CP 790302 formed plaques with equal efficiency on CP 790302 and a standard indicator
strain, NP-4 (datanot shown).
Trans-dominant effect of infection with T4vs+. Virus
pro-duction was examined when T4vs+ and T4vs2, each at an
MOI of 4, were added
simultaneously
to cultures of CP790302. An 80% decrease in progeny virus
production
wasobserved at 90 min after this co-infection compared with virus production during T4vs2infection alone (Table 2).
Neidhardt and Earhart (19) reported the appearance of
heat-stable valyl-tRNA synthetase activity in extracts of bacteriophage T4vs+- orT6-infected NP 29 cultures. The T-odd bacteriophage were examined similarly and lacked the
ability to thermally stabilize the temperature-sensitive syn-thetase of NP 29.Theirdatasuggest thatmodifiedenzymeis formed after bacteriophage T4 or T6 infection of E. coli.
However, our results demonstrate that, unlike T4infection,
T6andT7infections of CP790302 result inplaqueformation (Table 2).
Bacteriophagemacromolecularsynthesisandassembly.The accumulation of macromolecules in CP 790302 cells
during
T4infection was analyzedbymeasuring the incorporationof
3H-labeled
precursors into trichloroaceticacid-precipitable
[image:2.612.312.553.93.192.2]TABLE 3. Proteinsynthesis andphage productionin E. coli strains at 30 and43.5°C
Relative rate of protein synthesis(%)' Burstsize on hostb
Strain 300C 43.5°C 300C 43.5°C
Uninfected T4vs+ Uninfected T4i's' T4vs+ T4vs2 T4vs+ T4vs2
CP 790302 100 96 12 10 4 254 NDC ND
NP910212 100 115 5 8 86 62 4 5
NP29 100 90 6 76 87 79 96 <.01
NP 2 ND ND ND ND 244 277 105 87
a Cultures were grown to a density of108cells per mlin a 30°C rotating water bath. Identical samples of each culture were left uninfected or were infected with T4vs+ at an MOI of 5. After 10 min, a portion of each sample (uninfected or phage infected) was either reincubated at 30°C or shifted to 43.5°C. After a10-min incu-bation,[3H]argininewas added, andits incorporation into trichloroacetic acid-precipitable material was measured by filtering and counting samples.
bBacteria weregrown at 30°C to 3x108cells per ml,diluted to 5x107 cells per ml, and infected at 30°C at an MOI of 0.1. After 10min,anti-T4antiserum was added and the cultures were diluted to tubes at 30 and43.5aC.After 5min, cultures were diluted and incubated for 100min. Samples were taken for determination ofphage production and are expressed as yield per infectious center.
cND, Not determined.
accumulated during restrictive infection (T4vs+,
T4vs2RF01) wascomparableto that measured during
infec-tions inwhich virus production was normal (T4vs2, T4vsl) (datanot shown).
In additional experiments, viral proteins were pulse-la-beled (under conditions that diminished labeling of host
proteins)andanalyzed byelectrophoresis in sodium dodecyl sulfate and byautoradiography (2, 3). Apair-wise compari-son of proteins from cultures of CP 790302 infected with T4vs+ or T4vs2 revealed nomajor differences in autoradio-graphic intensities.Thetemporalappearanceofproteinswas
similar inboth infections (datanot shown).
Phage antigen synthesis was determined by measuring
serum blocking power in
lysates
derived from E. coli cul-tures infected with T4vs+ or T4vs2. The ratio of PFU to serum blocking power measured in the lysate of T4vs+-infected CP 790302(2.4x108 PFU/3.3
x109
serumblocking
power [sbp] =
7.2%)
isonly one-fifth theratio measuredin thelysate of T4vs2-infectedCP 790302(2.2 x 109 PFU/6.0x109
sbp = 36.5%).Although
infectious virus concentrationwaslow intherestrictiveinfection,thelysate contained 55%
asmuchserumblockingpower as wasmeasuredin thelysate fromthepermissive infection.
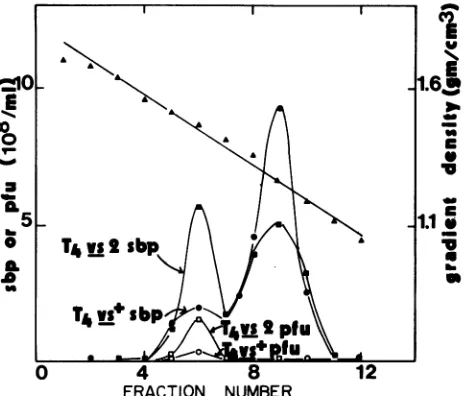
Lysates of CP 790302 cultures infected with T4vs+ or T4vs2 weresedimentedoncesium chloride
density gradients
(Fig. 1). Althoughtwopeaks ofserum
blocking
powerwereobserved for bothlysates,agreater
proportion
ofthemateri-al derived from the
permissive
infection sedimented at adensity typical ofassembled phage
particles. Plaque
assaysverified thepresenceof infectiousvirus. Mostofthe material
frotn the nonpermissive infection sedimentedat thedensity ofprotein. These studies suggest that
phage
maturation inT4vs+-infected CP790302cellsis restrictedatthepositionof
virusassembly. Thefollowing experiments weredone inan attempt toidentifythe cause of the restriction.
Aminoacylated
tRNA levels. The in vivo levels ofamino-acylatedtRNAweremeasured bytheperiodate method (5).
The fraction oftotal tRNAval that was aminoacylated was
87% in uninfected, 80% in T4vs+-infected, and 72% in T4vs2-infected cultures of CP 790302. The high level of
charging of
tRNAVal
in each culture is consistent withaminoacylationlevels
reported
by Comerand Neidhardt(5),
who used cultures ofuninfected and T4vs+-infected E. coli B.
Proteinsynthesis and virusproduction at high temperature inT4-infectedcultures. Table 3comparesproteinsynthesisin
uninfected and T4vs+-infected cultures of CP 790302, NP 910212, and NP 29 at30°Cand when thecellswereshifted to
43.5°C. The three uninfected, temperature-sensitive strains
were unable to synthesize protein at 43.5°C because they contained a thermolabile valyl-tRNA synthetase. Thermal stabilization of the temperature-sensitive enzyme occurred
after T4 infection of NP 29, and protein synthesis continued when the infected cells were shifted to43.5°C. In contrast, protein synthesis remained depressed at 43.5°C in T4vs+-infectedcultures ofCP 790302 and NP 910212. These E. coli strainsharbor the same vaiSts allele, which is different from that in NP 29 (8, 11).
These dataareconsistent withdata obtained by comparing averageburstsizes on these strains (Table 3). Virus produc-tion in T4vs+-infected NP 29cells was the same at both 30 and43.5°C. However, T4vs+-infected NP 910212 produced progeny virus at 30°C but not at 43.5'C.
Properties of valyl-tRNA synthetase in uninfected and
T4vs'-infected
temperature-sensitive and wild-type E. coli strains. We examined valyl-tRNA synthetase activity in extractsof uninfectedandT4vs+-infectedCP790302cellsto determine whether the host enzyme was actually modified. Extracts ofuninfected, temperature-sensitive cells arede-0
4
8
12
FRACTION NUMBER
FIG. 1. Serumblockingpower and PFU inlysates fractionated
oncesium chloridedensity gradients.Equivalentamountsofserum
blocking material from T4 vs+- and T4 vs2-infected CP 790302 lysateswerelayeredontocesiumchloridegradientsandcentrifuged for 1 hat 35,000rpm inaBeckman SW50.1rotor.Gradientswere
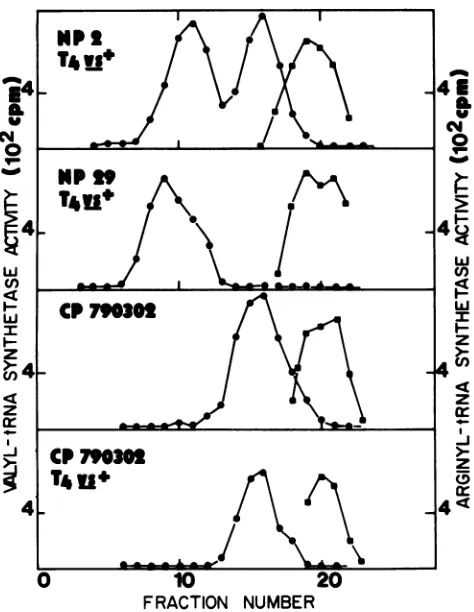
[image:3.612.328.561.462.660.2]FRACTION NUMBER
FIG. 2. Sucrose density gradient centrifugation of valyl-tRNA synthetases.E. coli culturesgrownat30°Cwereleftuninfected or were infected with T4 vs+ at an MOI of 7for 10 min. Cell-free
extracts were prepared from the cultures, and 0.2 ml of each preparation was layered separately ontolinear 5 to 20% sucrose
densitygradients.ThesewerecentrifugedinanSW50.1rotor at4°C for 10 h at 40,000 rpm. The gradients were collected from the
bottom, andalternatefractionswereassayedforspecific activityof
valyl-tRNA synthetase aminoacylation. Closed squares represent
arginyl-tRNA synthetase specific activity, usedas amarker in the assay. The high-molecular-weight formof themodifiedenzyme is duetoitsassociationwith tRNA.
void of thisenzymeactivity when examined by the standard
in vitro 3H-amino acid attachment assay (8; N. Olson and
G. L. Marchin,unpublished data). However, whenextracts wereprepared by usinga"protective buffer" and theassays wereperformedat30°C for 15 min, in vitro specific activity ofunmodified and modified temperature-sensitive enzyme could be observed and compared. Under these modified conditions, valyl-tRNA synthetase from T4vs+-infected CP 790302 (90.6 ,umol/minper,ug of protein) exhibited approxi-matelya two-thirds reduction inspecific activity compared with enzyme from uninfected cells. This reduction (2.0 ,umol/min per ,ug of protein) is characteristically seen with modifiedenzyme(4).
Figure 2 shows difference in enzyme sedimentation
pat-ternsonsucrosedensitygradients whenextractsof uninfect-ed and T4vs+-infected cells werecompared. Extracts from
T4vs+-infected NP 29 contained a high-molecular-weight
form of the modified enzyme due to its association with tRNA (13). In contrast, T4vs+-infected CP 790302 cell
extracts contained only the lower-molecular-weight form observedinuninfected E. coli cellextracts.Thetoppanel of Fig. 2 illustrates both forms of the enzyme, which were
observed after infection of wild-type E. coli NP 2. The
absence ofahigh-molecular-weightcomplex in infected CP 790302 further implies that modification afterT4vs+
infec-tion ofthesecells is unusual, and that, in turn, leads to an alteredinteraction of the modified synthetase with tRNA.
DISCUSSION
Apreviousreportfrom this laboratory demonstratedthat
bacteriophage T4 with the wild-type vs gene is unable to
formplaquesat30°C onE. coli CP 790302 (11). Phagewith
various amber, missense, and deletion mutations in the vs geneplate efficientlyon this strain. The restriction is
attrib-utabletoinfectiouscentersthatexhibitadramatic reduction in burstsizeorfailtoproduceevenasingleinfectious virus
particle.
Despite differences intheproduction of PFU,the
accumu-lation of DNA, RNA, and protein in the twoinfections is
similarasmeasuredbyprecursor accumulation. The kind of
protein synthesized alsoappears to besimilar in the restric-tive andpermissive infections; their temporalappearanceis also the same. However, although substantial amounts of
serum blocking power are synthesized in the restrictive
infection, only a small fraction is assembled intoinfectious virus particles. The remainder sediment with a density of proteinonCsClgradients. Theseresultsindicate,therefore,
that virus maturation is blocked at some position before complete assembly.
Interestingly, bacteriophage T6 forms plaques on CP 790302. Since the
phage
is thought tomodify
valyl-tRNA synthetase, and since simultaneous infection with T4vs2(Table2)doesnotresult insubstantially reduced burst sizes, then theinferencecanbe drawn that theT
peptide
in this T-evenphage is structurally different fromthe onein bacterio-phage T4. This conclusion is analagous tothe observations ofMoen et al. (15) that bacteriophageT4 and T6 specifyadifferent
repertoire
oftRNA molecules. Theability ofT7 toplate on CP 790302 is consistent with the observation of Neidhardt and Earhart (19) that the T-odd phage do not
modifyE.colivalyl-tRNA synthetase. Thus, theyare pheno-typically like T4vs2.
Earlier studies in this laboratory (17) had indicated that, during modification ofthevalyl-tRNA synthetase inE. coli,
the T peptide is buriedin the cleft betweenthe twoglobular domains of the enzyme. The aberrant modification ofthe enzyme inCP790302described in thispaper mayproducean enzyme with a different conformation and,
therefore,
analtered interaction with other elements in the infected cell.
The amino acid composition and molecular weight oftheT
peptide
(16) aresimilartothoseofanumberof nucleic acidbinding
proteins, suchashistoneH2A(9) and thecroprotein ofbacteriophage lambda (21).That themodifiedenzymefromT4vs+-infected CP 790302 fails to sediment as a
high-molecular-weight
complex on sucrosedensity gradients
isonedemonstrableexample ofanaltered interaction with a cellular component. This
alter-ation, however, was not detectable by any change in the
overall in vivocharging level of tRNAval in thesecells. Our
studies do not reveal more subtle alterations that may exist in the charging of different valyl-tRNA species or in the relative rates of tRNAcharging.
Theprecisemannerin whichvalyl-tRNA synthetase mod-ification arrests virus development in CP 790302 is not
understoodatthis time. The unusualmodification
apparent-ly occurs
during
T4vs+ infection of NP 910212, but tiny plaques are produced presumably because the stringentDonini (7) has shown that T4 recruits the host relA gene function during T4 phage DNA synthesis.
In summary, wehave demonstratedthat a normal, seem-ingly mild regulatory event, i.e., the modification of valyl-tRNA synthetase, goes awrywhenbacteriophageT4 infects CP 790302. Even though the modification is apparently nonessential and normally can be dispensed with by a variety of mutations, in this particular strain its operation severely hampers virus assembly.
LITERATURE CITED
1. Adams, M. H. 1959. Methodsofstudy of bacterial viruses, p. 443-522. InA. D. Hershey (ed.), Bacteriophages. Interscience Publishers, Inc.,NewYork.
2. Bonner, W. M., and R. A. Laskey. 1974. A film detection method for tritium-labelled proteins and nucleic acids in poly-acrylamide gels. Eur.J. Biochem. 46:83-88.
3. Chace, K. V., and D. H. Hall. 1975. Characterization ofnew regulatory mutants of bacteriophage T4. II. New class of mutants. J. Virol. 15:929-945.
4. Chrispeels, M. J., R. F. Boyd, L. S. Williams, and F. C. Neidhardt. 1968. Modification of valyl-tRNA synthetase by bacteriophage inEscherichia coli.J. Mol. Biol. 31:463-475. 5. Comer, M. M., and F. C. Neidhardt. 1975. Effect of T4
modification of host valyl-tRNA synthetaseonenzymeactionin vivo.Virology 67:395-403.
6. De Mars,R. I.1955.Theproduction of phage related materials when bacteriophage development isinterrupted byproflavine. Virology 1:83-99.
7. Donini, P.1970. Amino acid controloverdeoxyribonucleic acid synthesis in Escherichia coli infected with T-even bacterio-phage. J. Bacteriol. 102:616-627.
8. Eidlic, L.,and F. C.Neidhardt. 1965. Protein and nucleic acid synthesis intwo mutantsof Escherichiacoli with temperature-sensitive aminoacyl ribonucleic acid synthetases. J. Bacteriol. 89:706-711.
9. Iwai, K., K. Ishikawa, and H. Hiyashi. 1970. Amino-acid se-quence of slightly lysine-rich histone. Nature (London) 226:1056-1058.
10. Lowry,0. H.,N.J.Rosebrough,A. L.Farr,and R.J.Randall. 1951. Protein measurement with the Folin phenol reagent. J.
Biol. Chem. 193:265-275.
11. Marchin, G. L. 1980. Mutations in a nonessential viral gene permit bacteriophage T4 toform plaques on Escherichia coli valSts relA. Science 209:294-295.
12. Marchin, G. L., M. M. Comer, and F. C. Neidhardt. 1972. Viral modification of the valyl-tRNA synthetase of Escherichia coli. J. Biol. Chem. 247:5132-5145.
13. Marchin, G. L., U. R.Muller, and G. H. Al-Khateeb. 1974. The effect of transfer ribonucleic acid on virally modified valyl transfer ribonucleic acid synthetase of Escherichia coli. J. Biol. Chem. 249:4705-4711.
14. McClain, W. H., G. L. Marchin, and F. C. Neidhardt. 1971. Phage induced conversion of host valyl-tRNA synthetase, p. 191-205.InG.E. W.Wolstenholme andMaeveO'Connor (ed.), Strategy of the viral genome. ChurchillLivingstone, London. 15. Moen, T. L., J. G. Seidman, and W. H. McClain. 1978. A
catalogue of transfer RNA-like molecules synthesized following infection of Escherichia coli by T-even bacteriophages. J. Biol. Chem. 253:7910-7917.
16. Muller, U. R., and G. L. Marchin. 1977. Purification and properties of a T4 bacteriophage factor that modifies valyl-tRNAsynthetase of Escherichiacoli.J. Biol. Chem. 252:6640-6645.
17. Muller, U. R., and G. L. Marchin. 1977. Analysis of the structureofT4bacteriophage-modified valyl-tRNA synthetase by limitedproteolysisandisoelectric focusing. J. Biol. Chem. 252:6646-6650.
18. Neidhardt, F. C.,P. L.Bloch, and D. F. Smith. 1974.Culture medium forenterobacteria. J. Bacteriol. 119:736-747. 19. Neidhardt, F. C., and C. F. Earhart. 1966. Phage-induced
appearanceofavalyl sRNA synthetaseactivity in Escherichia coli. ColdSpring Harbor Symp. Quant. Biol. 31:557-563. 20. Spadafora, C., T. Igo-Kemenes, and H. G. Zachau. 1973.
Changes in tRNAs and aminoacyl-tRNA synthetasesduringsea urchindevelopment. Biochem.Biophys. Acta312:674-680. 21. Takeda, Y.,A.Folkmanis,and H.Echols. 1977. Croregulatory
proteinspecified by bacteriophage X. J. Biol. Chem. 252:6177-6183.