Copyright © 1997, American Society for Microbiology
Both T and B Cells Shed Infectious Mouse
Mammary Tumor Virus
JOHN L. DZURIS, TATYANA V. GOLOVKINA,
ANDSUSAN R. ROSS*
Department of Microbiology/Cancer Center, University of Pennsylvania
School of Medicine, Philadelphia, Pennsylvania 19103-6142
Received 28 January 1997/Accepted 18 April 1997
Mouse mammary tumor virus (MMTV) infected both B and T tissue culture cells and primary B and T cells
in vivo after milk-borne transmission of the virus. The infected tissue culture cells processed viral proteins, and
both these and primary B and T cells shed virus when cultured in vitro. Moreover, the infected B and T tissue
culture cells transmitted virus to uninfected mammary gland cells in vitro. The level of infection of these
different cell types in vivo was dependent on the strain of mouse, with C3H/HeN mice showing greater B-cell
infection and BALB/c mice greater T-cell infection after nursing on MMTV-infected C3H/HeN mothers.
Although their B cells were less infected, BALB/c mice developed tumors more rapidly than C3H/HeN mice.
These results indicate that both infected T and B cells are potential carriers of MMTV in vivo.
Mouse mammary tumor virus (MMTV) is a milk-borne
ret-rovirus that causes mammary tumors in mice (15). MMTV
encodes a superantigen (Sag) protein in its long terminal
re-peat (LTR) that is required for its transmission (1, 17). MMTV
first infects B cells in the Peyer’s patches of the gut, and these
infected B cells present the Sag to cognate T cells (14). B-cell
infection is required for stimulation of T cells, and infection
cannot be established in mice lacking B cells (2). Similarly, the
stimulation of cognate T cells by Sag is a requisite step in the
MMTV infection pathway, since mice lacking such T cells
cannot be efficiently infected (5, 11). Both B-cell and T-cell
amplification result from this stimulation, creating potential
reservoirs of infection-competent cells (5, 11).
Although both B and T cells are required for the milk-borne
transmission of MMTV, it is not known how this virus gets to
its target tissue, the mammary gland. Because it has been
shown that B cells are the most highly infected of all the
lymphocyte subsets, at least at early times after infection (10),
it has been suggested that these cells are ultimately responsible
for mammary gland infection (2, 10). However, lymphoid
or-gans rich in T cells also acquire integrated copies of MMTV
proviruses after milk-borne infection (4, 14) and transfer of
either B or T cells from an MMTV-infected donor to a naive
recipient results in transfer of the virus to both cell types of an
uninfected host (20, 21). Whether one or both of these cell
types carry virus to the mammary gland or even shed MMTV
virions has not been demonstrated.
We show here that both B and T tissue culture cells were
infected with MMTV and could transmit virus to other cells in
vitro. In vivo, both B and T cells were also infected with
MMTV and their level of infection was strain dependent, since
MMTV-infected C3H/HeN mice showed greater infection of B
cells than T cells, while BALB/c mice showed the opposite.
Splenic and thymic lymphocytes derived from transgenic mice
expressing a molecularly cloned MMTV and primary B and T
cells from C3H/HeN mice infected with MMTV(C3H) shed
virus when cultured in vitro. These results indicate that both B
and T cells are capable of carrying MMTV to the mammary
gland in vivo.
MATERIALS AND METHODS
Cell culture.The B-cell tumor line A20, B-cell hybridomas LBB.A, LBB.11, and LK4.5, and the T-cell tumor line BW5147 were cultured in RPMI 1640 (GIBCO/BRL, Gaithersburg, Md.) supplemented with 10% fetal calf serum and 0.5mMb-mercaptoethanol. The normal mammary gland cell line NMuMG and MMTV (HYB PRO)-transfected NMuMG (NMgCl 1) were cultured in Dulbec-co’s modified Eagle’s medium supplemented with 10% fetal calf serum and 10mg of insulin/ml. The rat hepatocyte XC cell line and the mammary gland tumor cell line Mm5MT were cultured in Dulbecco’s modified Eagle’s medium supple-mented with 10% fetal calf serum.
The NMgCl 1 cell line was generated by cotransfecting NMuMG cells with a molecular clone of MMTV [called hybrid provirus (HYB PRO), containing the
env and sag genes from MMTV(C3H)] and the MMTV-neo constructs
(de-scribed by Shackleford and Varmus [19]). Independent G418-resistant transfec-tants were isolated, and the NMgCl 1 clone was chosen because it shed high levels of MMTV virions (not shown). The MMTV-neo RNA was inefficiently packaged in this cell line (approximately 100 times lower than the HYB PRO RNA [not shown]).
Mice.C3H/HeN MTV2, C3H/HeN MTV1, and BALB/c mice from colonies of germ-free-derived, defined-flora animals were purchased from the National Institutes of Health, Frederick Cancer Research Facility, Frederick, Md. Trans-genic mice bearing the HYB PRO construct have been previously described (6, 7). MMTV-infected mice were examined weekly by palpation for mammary tumors.
Isolation of primary B and T cells.Primary lymphocytes were isolated from the spleen, thymi, and lymph nodes of MMTV(C3H)-infected C3H/HeN or BALB/c mice or from those of the HYB PRO transgenic mice. T and B cells were purified from the pooled lymphoid organs of two to three mice. Single-cell suspensions were prepared in Hanks balanced salt solution (HBSS) supple-mented with 20 mM HEPES. Cells were washed two times with HBSS. Eryth-rocytes were removed by 0.15 M NH4Cl lysis for 5 min at room temperature.
Cells were washed two more times with HBSS and then resuspended in RPMI 1640 medium supplemented with 20 mM HEPES. B cells were then isolated by negative selection during three rounds of panning 43108lymphocytes on
150-mm-diameter petri dishes coated with anti-Thy1 antibody (100mg/ml) to remove the T cells. T cells were isolated by similar panning on plates that had been coated with anti-mouse immunoglobulin antibody (100mg/ml) to remove the B cells. Panning was carried out at 4°C for 1 h during each round. In between each pair of panning steps, the petri dishes were agitated to resuspend nonad-herent cells, which were then transferred to the next dish. B- and T-cell popu-lations were washed two times with HBSS-HEPES. The purity of the cell pop-ulations was determined by fluorescence-activated cell sorter analysis with fluorescein isothiocyanate (FITC)-labelled anti-immunoglobulin or anti-CD4 and anti-CD8 antibodies (GIBCO/BRL). Fluorescence-activated cell sorter anal-ysis of the purified cells showed that the T- and B-cell populations were 93 to 99% pure (not shown).
Virus infection of lymphocyte cell lines.Infection of the lymphocyte cell lines was done by coculturing lymphocyte cell lines (104cells/plate) and NMgCl 1 (104
cells/plate) in 60-mm-diameter tissue culture plates for 5 days, followed by
* Corresponding author. Mailing address: Department of
Microbi-ology/Cancer Center, University of Pennsylvania School of Medicine,
415 Curie Blvd., Philadelphia, PA 19103-6142. Phone: (215) 898-9764.
Fax: (215) 573-2028. E-mail: ROSSS@mail.med.upenn.edu.
6044
on November 9, 2019 by guest
http://jvi.asm.org/
transfer to 100-mm-diameter plates for an additional 5 days. For some experi-ments, coculturing was carried out with 0.4-mm-pore-size filter inserts (Falcon 3090) to prevent cell-cell contact. The cultures were supplemented with 8mg of Polybrene/ml. The adherent NMgCl 1 cells were then removed from the cultures by passaging the nonadherent lymphocytes to new T-25 tissue culture flasks four times in succession. The lymphocyte cultures were supplemented with 1mM dexamethasone. Biological clones of the MMTV-infected lymphocytes were ob-tained by limiting dilution cultures in 96-well flat-bottom tissue culture plates.
Passage of virus from the newly infected lymphocytes to uninfected NMuMG and XC cells was done by similar coculturing in the presence of 0.4-mm-pore-size filters. After the 10 days of coculturing, the lymphocytes were removed. The NMuMG and XC cells were cultured for 10 more days in the presence of 1mM dexamethasone before DNA was harvested.
PCR and Southern blot analysis.Equal amounts of genomic DNA (0.25mg) from the different cell samples were used for PCR with Taq polymerase accord-ing to the instructions of the manufacturer (Promega, Madison, Wis.). Oligonu-cleotides used to amplify MMTV(C3H) (59primer and 39primer, nucleotides [nt] 268 to 289 and 894 to 871, respectively, from the 59end of the LTR) and endogenous MMTVs (59primer, nt 507 to 527 and 39primer, nt 1203 to 1184) were described previously (7). Semiquantitative PCR, in which the amplification was still in the linear range, was carried out by 31 cycles of 1 min at 55°C, 1 min at 72°C, and 1 min at 94°C (9). After PCR amplification, a sample of the product was digested with 2 U of MunI (NEB, Beverly, Mass.) for 2 h at 37°C, as previously described (2); MunI only cuts exogenous MMTV(C3H). Genomic DNA from the Mm5MT mouse mammary tumor cell line was used as a positive control for amplification of MMTV sequences.
Southern blot analysis to determine whether cells were infected with HYB PRO was carried out as previously described (8, 19). Briefly, genomic DNAs were digested with BglII and PstI, which cut internally within the proviruses, and the blots were hybridized with an env-specific probe. Copy numbers were esti-mated by comparing the relative intensities of hybridization of the exogenous (Fig. 1, band V) and endogenous (Fig. 1, band E) MMTV bands. All of the B and T cell lines have three to six copies of endogenous MMTV.
Virus purification and RNA analysis.The B and T cell lines were plated at 106
cells/ml in the presence of 1mM dexamethasone and cultured for 3 days. Virus was isolated from 50 ml of supernatant filtered through 0.2-mm-pore-size filters by adding 0.5 volumes of 33polyethylene glycol (PEG) 6000 (25% PEG 6000, 1.5 M NaCl) and mixing at 4°C for 2 h. Virus was pelleted by centrifugation at 12,0003g for 20 min at 4°C. RNA was then extracted from the virus pellets.
Cellular and viral RNA were extracted by guanidine thiocyanate extraction and CsCl gradient centrifugation (3).
Primary lymphocytes (107) isolated from 6-month-old C3H/HeN MMTV
(C3H)1, BALB/c MMTV(C3H)1, or HYB PRO transgenic mice were plated at 106cells/ml and cultured for 3 days in the presence of 10mg of concanavalin A/ml
(T cells) or 50mg of lipopolysaccharide/ml (B cells) (both from Sigma, Inc., St. Louis, Mo.). Virus was isolated from 10 to 20 ml of filtered supernatant by centrifugation at 156,0003g for 1.5 h at 4°C. RNA was extracted from the virus
pellets.
For RNase T1protection assays, labeled RNA probes were synthesized from
a pBluescript plasmid carrying the Sau3A fragment of the MMTV(C3H) LTR with T3 RNA polymerase, as previously described (8). Either 40mg of total cellular RNA or viral RNA isolated from 50 ml of culture supernatant was used for RNase T1protection analysis, as previously described (6).
Reverse transcription (RT)-PCR was accomplished by synthesis of cDNA from RNA isolated from the B- and T-cell culture supernatants as previously
described (7). After amplification and agarose gel electrophoresis, the products were transferred to nitrocellulose and hybridized with a probe specific to the MMTV LTR (5). Relative amplification was determined by densitometric anal-ysis of the autoradiographs.
Western blot analysis.Total cellular protein was extracted from cells or from 10 ml of pelleted cell culture supernatant with lysis buffer containing 50 mM Tris-HCl (pH 8.0), 150 mM NaCl, 1 mM EDTA, 0.1% sodium dodecyl sulfate, 1.0% Nonidet P-40, and 1.0% Triton X-100. Cellular protein (100 mg) was separated by sodium dodecyl sulfate–10% polyacrylamide gel electrophoresis, and Western blot analysis was performed, as previously described (6), with goat anti-MMTV gp52 (SU) or goat anti-MMTV p27 (Gag) polyclonal antibodies or normal goat serum (National Cancer Institute Biochemical Carcinogenesis Branch Repository, distributed by Quality Biotech Inc., Camden, N.J.).
RESULTS AND DISCUSSION
We used two approaches to determine whether T and B cells
could both be productively infected with MMTV. First, we
established T- and B-cell tumor lines that were infected with a
molecular clone of MMTV, called HYB PRO (19). We
in-fected the B-cell tumor line A20, B-cell hybridomas LK4.5,
LBB.A, and LBB.11, and T-cell tumor line BW5147 by
cocul-turing these cells with a HYB PRO-transfected mammary
gland cell line, as described in Materials and Methods. Clonal
cell lines from each of the infected populations were isolated
and examined for the presence of integrated viral DNA by
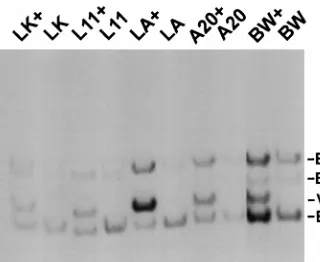
Southern blot analysis (Fig. 1). The infected cell lines had a
diagnostic 2.3-kb band (Fig. 1, band V) that was present after
digestion of the DNA with PstI and BglII. The clones selected
for further study had from 6 to 10 copies of newly integrated
MMTV proviruses, based on the relative hybridization to the
endogenous virus- and HYB PRO-specific bands (Fig. 1).
The clones from each infected cell line were then examined
for transcription of HYB PRO-specific RNA and the
produc-tion of processed MMTV proteins. RNase T
1-protection
anal-ysis with a probe specific for the U3 region of the HYB PRO
transcript was performed with RNA isolated from the various
cell lines. As can be seen in Fig. 2A, all of the infected clones
examined produced HYB PRO-specific RNA.
It had previously been reported that a T-cell lymphoma line
was not able to process virus proteins and therefore could not
produce infectious virus particles (16). To determine if all of
the cells were able to produce MMTV proteins, Western blot
analysis was performed on total cell extracts with monospecific
anti-SU (Fig. 2B, gp52) and anti-Gag (Fig. 2B, p27) sera. All of
the cell lines produced both processed viral proteins.
There-fore, there was no block to the translation and processing of
MMTV proteins in these B and T cell lines. In some cell lines
(i.e., LBB.11 and LK4.5), we detected unprocessed Gag but
not Env precursor proteins in the uninfected cells (Fig. 2B).
These most likely represent proteins produced from the
en-dogenous viruses present in these cell lines. The lack of
pro-cessed Gag and Env in the uninfected cells indicates that these
endogenous viruses must have mutations in the coding regions
for these proteins.
[image:2.612.97.257.68.199.2]These results indicated that the B and T cell lines were
capable of producing MMTV virions, since they produced both
viral RNA and processed proteins. To confirm this, the
super-natants of the infected cell cultures were collected and filtered
to remove cells and the virus fraction was obtained by PEG
precipitation. RNA or protein was isolated from the pellets
and subjected to RNase protection analysis with an MMTV
(C3H) U3-specific probe or to Western blot analysis,
respec-tively. Viral RNA could be detected in the supernatants of all
of the cell lines (Fig. 3A). Moreover, gp52 (Fig. 3B) and p27
(not shown) were also found in the supernatants. These results
showed that both cultured B- and T-lymphoma and
B-hybrid-oma cell lines could produce MMTV virions after infection.
FIG. 1. Southern blot analysis of clonal isolates of the infected B- and T-cell tumor lines. Genomic DNA was isolated from clonal isolates of infected popu-lations of B (LBB.A [LA], LBB.11 [L11], LK4.5 [LK], and A20) and T (BW5147 [BW]) tumor cell lines and digested with BglII and PstI, and the Southern blots were hybridized to an env-specific probe, as previously described (8, 19). The “1” denotes infected cells, and the “2” denotes the uninfected parental cell line. E, endogenous MMTV-specific band; V, exogenous MMTV-specific band.
on November 9, 2019 by guest
http://jvi.asm.org/
To show that the virus produced by the T and B cells was
infectious, we cocultured the infected BW5147 (T) and A20
(B) cell lines with an uninfected mouse mammary gland
(NMuMG) and a rat sarcoma (XC) cell line. After coculturing
the cells for 10 days in the presence of a 0.4-
m
m-pore-size
barrier filter, we removed the lymphoma cells. DNA was
pre-pared from the cultures, and PCR with primers specific for the
U3 region of the MMTV LTR was performed. The PCR
prod-ucts were digested with MunI; this site is unique to the HYB
PRO LTR derived from MMTV(C3H). Both the XC and
NMuMG cells were infected after being cocultured with either
the infected T or B cell line (Fig. 4). Thus, there is no general
block to the production of infectious MMTV virions in B- or
T-cell tumor lines, although some cell lines may not process
the viral proteins (16).
We also examined lymphocytes infected in vivo after
milk-borne transmission of MMTV. Four-month-old BALB/c and
C3H/HeN mice nursed on C3H/HeN MMTV(C3H)
1mothers
were sacrificed, and semiquantitative PCR analysis was
per-formed on genomic DNA isolated from their various lymphoid
cells and tissues to look for newly acquired MMTV(C3H)
proviral DNA, as described in Materials and Methods. In the
case of C3H/HeN mice infected by milk-borne MMTV(C3H),
we found that B cells were more infected than T cells (Fig. 5).
In contrast, in the MMTV(C3H)-infected BALB/c mice,
infec-tion of T cells and thymi was much greater than that of B cells
or spleens (Fig. 5). Infection of C3H/HeN T cells and thymi or
of BALB/c B cells and spleens could only be detected after
Southern blotting and hybridization of the gel shown in Fig. 5;
about 10 times more MMTV DNA was seen in C3H/HeN B
and BALB/c T cells than in C3H/HeN T or BALB/c B cells
(not shown). Similar results were obtained with 7-month-old
mice (not shown).
To determine whether both cell types were productively
infected in vivo by milk-borne virus, we assayed supernatants
from equal numbers of cultured B and T cells isolated from
MMTV(C3H)-infected BALB/c and C3H/HeN mice. We also
cultured primary lymphoid tissues from HYB PRO transgenic
mice to assay for virus production. HYB PRO mice express a
molecular clone of MMTV in both lymphoid tissues and
mam-mary glands and shed an infectious MMTV into milk (7). RNA
was isolated from the filtered supernatants of these cultures
and subjected to an RT-PCR assay that was specific for the
exogenous viral RNA. MMTV-specific RNA could be detected
in the supernatants of T and B cells from infected mice (Fig.
FIG. 2. Analysis of MMTV-specific cellular RNA and protein produced by MMTV-infected B- and T-cell tumor lines. (A) RNase T1protection analysis was carried out with RNA isolated from the various infected B and T cell lines, using a probe specific for the U3 region of HYB PRO. A protected fragment of 340 nt is present in cells expressing the HYB PRO RNA. (B) Western blot analysis of cellular extracts from the same cells with monospecific SU (gp52) or anti-GAG (p27) antisera. The arrows show the Env (gp78) (upper panel) and Gag (Pr77) (lower panel) precursor proteins. Abbreviations are the same as in Fig. 1.
[image:3.612.71.275.72.352.2]FIG. 3. Analysis of the virions shed by the MMTV-infected B- and T-cell tumor lines. (A) RNase T1protection analysis of virus pelleted from the super-natants of the various cell lines. Although the signal for the LK1cell line is weak, it was clearly visible upon longer exposure of the gel. (B) Western blot analysis of virus pelleted from the supernatants of the various cell lines. Monospecific polyclonal antibodies against the gp52 SU protein were used. Abbreviations are the same as in Fig. 1.
FIG. 4. T and B cell lines produce infectious virus. MMTV-infected B cell line A20 and T cell line BW5147 (BW) were cocultured with XC or NMuMG cells in the presence of a 0.4-mm-pore-size filter. DNA was isolated from the cocultured XC and NMuMG cells after an additional 10 days in culture and was subjected to PCR amplification followed by MunI digestion, as described in Materials and Methods. Mouse mammary tumor cell line Mm5MT was used as a positive control.
on November 9, 2019 by guest
http://jvi.asm.org/
6A) and in the thymi and spleens from the HYB PRO
trans-genic mice (Fig. 6B) but not in those from uninfected C3H/
HeN mice. B cells from C3H/HeN mice and T cells from
BALB/c mice appeared to shed the highest levels of virus,
approximately 2.2 and 4 times the amounts produced by C3H
T and BALB/c B cells, respectively. The ratio between the
numbers of proviruses in the C3H B and T cells or in the
BALB/c T and B cells was greater than was seen for virus
production. However, because we know nothing about the
kinetics of virus production from the various cell populations
either in vivo or in vitro, the assay for virus production is not
quantitative.
The results presented here show that both B and T cells
infected with MMTV can produce infectious virus particles
and thus are potential carriers of virus to the mammary gland.
It has been proposed that the role of the MMTV Sag is to
cause T-cell stimulation and cytokine production resulting in
B-cell proliferation and that infected B cells are ultimately
necessary for infection of the mammary gland (2, 10, 12). For
example, there is amplification of MMTV-infected B cells in
mice injected with MMTV (SW) even in the presence of
zidovudine, an inhibitor of retroviral replication (12). This
increase in the number of infected B cells has been suggested
to be an important step in the MMTV infection pathway.
However, it is also possible that this B-cell amplification only
reflects a strong humoral response to MMTV antigens and that
B cells are not the sole carriers of virus to the mammary gland.
We also found that the level of virus infection in different
lymphoid cell populations was dependent on the strain of
in-fected mice. In BALB/c mice inin-fected with milk-borne MMTV
(C3H), T cells were more highly infected than were B cells,
while B cells were more highly infected in C3H/HeN mice. We
have also found that the mammary glands of BALB/c mice are
more highly infected by MMTV(C3H) than those of C3H/HeN
mice (not shown). Moreover, the latency of mammary tumor
induction in BALB/c mice foster nursed on C3H/HeN
FIG. 5. Analysis of infected T and B cells from BALB/c and C3H/HeN mice nursed on C3H/HeN MMTV1mothers. (A) DNA isolated from the B cells, T
[image:4.612.86.266.76.322.2]cells, spleens, thymi, and tails (DNA) of MMTV(C3H)-infected BALB/c (lanes B) and C3H/HeN (lanes C) mice was used for PCR analysis, using primers that amplify MMTV(C3H) DNA. After amplification, the DNA was restricted with MunI, which cuts only in MMTV(C3H) and not in the endogenous proviral DNA. The Mtv-9 provirus present in BALB/c but not in C3H/HeN mice was also amplified with the primers (DNA, lanes B); however, this provirus lacks the MunI site (DNA, lane B1). The presence of the two lower-molecular-weight bands with MunI digestion in the different samples shows the presence of MMTV (C3H) DNA. These experiments were performed three times with cells and tissues isolated from different animals; a representative experiment is shown. (B) PCR analysis with primers that amplify endogenous as well as exogenous MMTVs was done as a control. The same DNAs used in A were amplified with these primers.
FIG. 6. Primary lymphoid cells from MMTV(C3H)-infected mice and HYB PRO transgenic mice shed virus. Lymphocytes isolated from HYB PRO mice and purified T and B cells from MMTV-infected C3H/HeN mice were cultured in vitro for 3 to 5 days, and their supernatants were harvested and filtered to remove cells. RNA isolated from the supernatants was subjected to RT-PCR, the amplified fragments were separated by gel electrophoresis, and Southern blotting with a LTR-specific probe was performed. (A) RT-PCR of RNA isolated from purified B and T cell supernatants was performed with or without reverse tran-scriptase. (B) RT-PCR of RNA isolated from cultured splenocytes (spleen) or thymocytes (thymus) from HYB PRO or nontransgenic (NT) mice was carried out. Mouse mammary tumor cell line Mm5MT RNA was used as a positive control.
FIG. 7. Tumor incidence in MMTV-infected BALB/c and C3H/HeN mice. The mice were foster nursed on C3H/HeN MMTV1mothers and were force bred.
on November 9, 2019 by guest
http://jvi.asm.org/
MMTV
1mothers was significantly decreased in comparison
with that of C3H/HeN mice (Fig. 7). This indicates that the
mammary gland virus load was indeed higher in BALB/c mice.
The fact that T cells were more infected than B cells in BALB/c
mice indicates that infected T cells may be an important source
of virus in vivo for the infection of mammary gland cells.
It is unlikely that the same virus shows different tropism in
the two inbred mouse strains, so these differences may reflect
other genes that influence host response to virus. BALB/c and
C3H/HeN mice differ at their major histocompatibility locus:
BALB/c mice are H-2
d, while C3H/HeN mice are H-2
k. It is
possible that Sag presentation or immune-cell recognition of
other virus proteins differs in these two strains. For example,
Sag presentation may be more effective in C3H/HeN mice and
bystander B cell activation more vigorous, resulting in greater
infection of this subset of cells. Another possibility is that
predominantly Sag-specific T cells are infected and that these
are more slowly deleted in BALB/c than C3H/HeN mice. This
seems unlikely, given that the experiments described here were
carried out on 4-month-old mice, in which the bulk of the
Sag-cognate T cells were already deleted (13). Moreover, this
would not explain why BALB/c mice had lower B-cell infection
levels.
Similarly, C3H/HeN mice may mount a stronger humoral
response to MMTV infection than BALB/c mice, resulting in
more extensive infection of B cells due to their increased
pro-liferation. Mice infected neonatally with MMTV do mount an
antivirus response (18). If virus clearance was also poorer in
BALB/c mice, they would be expected to succumb more
rap-idly to MMTV-induced mammary tumors, as we showed here.
In summary, both B and T cells are potential MMTV
carri-ers in vivo. Clarification of whether one or the other of these
lymphocyte subsets better transmits virus to the mammary
gland and of the mechanism by which this transfer occurs
awaits the results of future studies which address these
ques-tions.
ACKNOWLEDGMENTS
We thank Marta de Olano Vela and Bernadette van den Hoogen for
expert technical assistance and Jaquelin P. Dudley for helpful
com-ments on the manuscript. We thank J. P. Dudley for the A20 and
BW5147 cells, B. T. Huber for the LBB.11 and LBB.A cells, and Y.
Choi for the LK4.5 cells.
J. Dzuris is supported by PHS grant T32 CA09140, and T. V.
Golovkina was a Cancer Research Institute fellow. This work was
supported by PHS grant CA52646.
REFERENCES
1. Acha-Orbea, H., and H. R. MacDonald. 1995. Superantigens of mouse mam-mary tumor virus. Annu. Rev. Immunol. 13:459–486.
2. Beutner, U., E. Draus, D. Kitamura, K. Rajewsky, and B. T. Huber. 1994. B cells are essential for murine mammary tumor virus transmission, but not for presentation of endogenous superantigens. J. Exp. Med. 179:1457–1466.
3. Chirgwin, J. M., A. E. Prxybyla, R. J. MacDonald, and W. J. Rutter. 1979. Isolation of biologically active ribonucleic acid from sources enriched in ribonuclease. Biochemistry 18:5294–5299.
4. Desaymard, C., C. L. Tucek, B. Rocha, A. J. Korman, and M. Papiernik. 1993. Tissue distribution of Mtv-7-like exogenous retroviral transcripts and clonal deletion of Vb61T cells in Mls1b BALB/c mice. Int. Immunol.
5:217–222.
5. Golovkina, T. V., A. Chervonsky, J. P. Dudley, and S. R. Ross. 1992. Trans-genic mouse mammary tumor virus superantigen expression prevents viral infection. Cell 69:637–645.
6. Golovkina, T. V., A. Chervonsky, J. A. Prescott, C. A. Janeway, and S. R.
Ross.1994. The mouse mammary tumor virus envelope gene product is required for superantigen presentation to T cells. J. Exp. Med. 179:439–446. 7. Golovkina, T. V., J. P. Dudley, A. Jaffe, and S. R. Ross. 1995. Mouse mammary tumor viruses with functional superantigen genes are selected during in vivo infection. Proc. Natl. Acad. Sci. USA 92:4828–4832. 8. Golovkina, T. V., A. Jaffe, and S. R. Ross. 1994. Coexpression of exogenous
and endogenous mouse mammary tumor virus RNA in vivo results in viral recombination and broadens the virus host range. J. Virol. 68:5019–5026. 9. Golovkina, T. V., I. Piazzon, I. Nepomnaschy, V. Buggiano, M. de Olano
Vela, and S. R. Ross.1997. Generation of a tumorigenic milk-borne mouse mammary tumor virus by recombination between endogenous and exoge-nous viruses. J. Virol. 71:3895–3903.
10. Held, W., A. N. Shakhov, S. Izui, G. A. Waanders, L. Scarpellino, H. R.
MacDonald, and H. Acha-Orbea.1993. Superantigen-reactive CD41T cells are required to stimulate B cells after infection with mouse mammary tumor virus. J. Exp. Med. 177:359–366.
11. Held, W., G. Waanders, A. N. Shakhov, L. Scarpellino, H. Acha-Orbea, and
H. Robson-MacDonald.1993. Superantigen-induced immune stimulation amplifies mouse mammary tumor virus infection and allows virus transmis-sion. Cell 74:529–540.
12. Held, W., G. A. Waanders, H. Acha-Orbea, and H. R. MacDonald. 1994. Reverse transcriptase-dependent and -independent phases of infection with mouse mammary tumor virus: implications for superantigen function. J. Exp. Med. 180:2347–2351.
13. Ignatowicz, L., J. Kappler, and P. Marrack. 1992. The effects of chronic infection with a superantigen-producing virus. J. Exp. Med. 175:917–923. 14. Karapetian, O., A. N. Shakhov, J.-P. Kraehenbuhl, and H. Acha-Orbea.
1994. Retroviral infection of neonatal Peyer’s patch lymphocytes: the mouse mammary tumor virus model. J. Exp. Med. 180:1511–1516.
15. Nandi, S., and C. M. McGrath. 1973. Mammary neoplasia in mice. Adv. Cancer Res. 17:353–414.
16. Nusse, R., L. van der Ploeg, L. van Duijn, R. Michalides, and J. Hilgers. 1979. Impaired maturation of mouse mammary tumor virus precursor polypeptides in lymphoid leukemia cells, producing intracytoplasmic A par-ticles and no extracellular B-type virions. J. Virol. 32:251–258.
17. Scherer, M. T., L. Ignatowicz, G. M. Winslow, J. W. Kappler, and P.
Mar-rack.1993. Superantigens: bacterial and viral proteins that manipulate the immune system. Annu. Rev. Cell Biol. 9:101–128.
18. Schochetman, G., L. O. Arthur, C. W. Long, and R. J. Massey. 1979. Mice with spontaneous mammary tumors develop type-specific neutralizing and cytotoxic antibodies against the mouse mammary tumor virus envelope pro-tein gp52. J. Virol. 32:131–139.
19. Shackleford, G. M., and H. E. Varmus. 1988. Construction of a clonable, infectious, and tumorigenic mouse mammary tumor virus provirus and a derivative genetic vector. Proc. Natl. Acad. Sci. USA 85:9655–9659. 20. Tsubura, A., M. Inaba, S. Imai, A. Murakami, N. Oyaizu, R. Yasumizu, Y.
Ohnishi, H. Tanaka, S. Morii, and S. Ikehara.1988. Intervention of T-cells in transportation of mouse mammary tumor virus (milk factor) to mammary gland cells in vivo. Cancer Res. 48:6555–6559.
21. Waanders, G. A., A. N. Shakhov, W. Held, P. Karapetian, H. Acha-Orbea,
and H. R. MacDonald.1993. Peripheral T cell activation and deletion in-duced by transfer of lymphocyte subsets expressing endogenous or exoge-nous mouse mammary tumor virus. J. Exp. Med. 177:1359–1366.