0022-538X/81/030899-09$02.00/0 Vol.37, No.3
Avian
Reticuloendotheliosis
Virus:
Characterization of
Genome Structure by Heteroduplex Mappingt
SYLVIA S. F. HU,' MICHAEL M. C. LAI,2* TIMOTHY C.WONG,2ROBERT S. COHEN2AND
MARTINSEVOIAN3
Department of Pediatrics, City of Hope Medical Center, Duarte, California910101;Department of
Microbiology, University of SouthernCaliforniaSchool ofMedicine, Los Angeles, California900332; and
Department of Veterinary and Animal Sciences, University ofMassachusetts, Amherst, Massachusetts
01033
Thegenome structureof defective,oncogenicavianreticuloendotheliosisvirus
(REV) was studied byheteroduplex mapping between the full-length comple-mentary DNA ofthe helpervirusREV-Ti and the 30SREVRNA. The REV genome (5.5 kilobases) had adeletion of 3.69 kilobases in the gag-pol region,
confinming the genetic defectiveness of REV. In addition, REV lacked the sequences corresponding to the env gene but contained, instead, a contiguous
stretch (1.6to1.9kilobases) of the specificsequencespresumably related to viral
oncogenicity. Unlike those of other avian acute leukemia viruses, the
transfor-mation-specificsequencesof REVwere notcontiguouswith thegag-pol deletion.
Thus, REVhasa genome structuresimilarto that ofa defective mink cell
focus-inducing virus or a defective murine sarcoma virus. An additional class of
heteroduplex molecules containing the gag-pol deletionand two other smaller
deletionloopswasobserved. Thesemolecules probablyrepresentedrecombinants
between the oncogenic REV and itshelper virus.
Avian reticuloendotheliosis virus (REV)
causesvisceral reticuloendotheliosis and periph-eralnerve lesion invarious species of fowl (31,
32, 36, 38,42). It transforms fibroblasts aswell
ashematopoietic cellsderived from bone
mar-row and spleenin tissue culture (4, 10, 11, 15, 16). REV is unrelated to other avian leukosis
andsarcomaviruses(13,22,29).It isalso
defec-tive inreplication and requiresashelper virus other members of the reticuloendotheliosis
group of retroviruses for thesynthesis of viral
progeny (16). The defectiveness of the REV
genomeis consistent with the absenceof virion-related proteins in REV-transformed
nonpro-ducer cells (16). Recent studies further showed
that REV contained a 28S RNA genome, as comparedwith the358RNA for thehelpervirus
(4, 16). Oligonucleotide fingerprinting and
nu-cleic acidhybridizationstudies furthersuggested
thatatleast 30% of the REVgenomicsequences are specific for REV and are presumably the oncogenic sequences whichare responsible for
its acuteleukemogenicpotentialin vivo and its
transformingactivityinvitro.
To determine the genetic structure of REV
and to localize the transformation-specific se-quences in the REV genome,wecomparedthe genomes of REV and itshelpervirus REV-Aby
tContribution 2368 ofthe MassachusettsAgricultural
Ex-perimentStation.
an electron microscopic heteroduplex method. We found thatREVhadanextensive deletion inthe gap-polregion but containedastretchof transformation-specificsequencesin theenv
re-gion whichwas notcontiguouswith thegag-pol deletion. This genetic structure distinguishes REVfrom otheravian acuteleukemia viruses.
MATERIALS AND METHODS
Cells and viruses. The bonemarrow cell(BMC)
subline ofREV-transformedchicken BMCcame
orig-inally from H. Bose of the University ofTexas at
Austin(11) andwasobtainedthroughthecourtesy of
C.Moscovici Gainesville, Fla. The cellswere
main-tained in F-10 mediumsupplementedwith 10%
tryp-tosephosphatebroth,10% calf serum, and 5%chicken
serum. The REV-transformed lymphoblastoid cell
linesTV-1 and TV-2wereisolated from thespleensof
moribundSPAFAS and line6chicken, respectively,
infected with REV. These cells carry B-cell determi-nants ontheirsurfaces(23).TheTV-1 and TV-2cells weremaintainedinRPMI1640mediumsupplemented
with 10% calfserum. REV-Ti, anattenuated
prepa-ration ofREV, has been described previously (41).
Thisstrain does notcauseproliferationof reticuloen-dothelial cells but inducesnervelesionsinvivo (33,
42) and does not transform fibroblasts or BMC in
vitro.REV-Tiisgeneticallyvery similartothehelper
virus,REV-A, isolated fromtheviruses releasedfrom
theBMCline(4).
Preparation ofviral RNA. The RNA used for
heteroduplexmappingwaspreparedfrom theviruses
releasedover a12-hperiodfromtheinfected culture.
899
on November 10, 2019 by guest
http://jvi.asm.org/
Viruseswerepurifiedfrom the mediaaccordingtoa
modificationofpublishedprocedures(25). Briefly,
vi-rus waspelletedfrom the media inanSW27rotorat
25,000rpmfor 90minand thensedimentedon a25to
60% (wt/vol) sucrosegradient inanSW50.1rotorat
50,000rpmfor 150min.The70SRNAwasextracted
from the virionandpreparedaccordingtopublished
procedures (28).
Synthesisofgenome-lengthcDNAfrom
REV-Ti. Thegenome-length complementaryDNA(cDNA)
wassynthesizedfrom REV-Ti by endogenousreverse
transcription accordingtoapublishedprocedure(26).
This procedurewasestablishedfor severalstrains of avian and manmalianoncoviruses.Wefound thatit
wasalso applicabletothe REV-Ti strain. The
endog-enous reversetranscriptionwasperformedina
reac-tion mixturecontaining1 mMTris-hydrochloride (pH
8.0),30 mMdithiothreitol,3 mMmagnesiumacetate,
1mMeach dATP,dCTP,and dGTP,0.5 mM
[3H]-TTP (200mCi/mmol), 1mgofpurifiedvirusperml,
and 0.02 to 0.03% Triton X-100at37°Cfor 18 h.The
reaction productswereextracted with sodiumdodecyl
sulfate-phenol and then digestedwith RNase A (50
,ug/ml) at 37°C for 1 h. The full-length DNA was
separated by sedimentation through an alkaline
su-crosegradientinanSW40rotorat40,000rpmfor 11
h(26). Most of thefull-lengthDNAwas9kilobases
(kb) long. Theauthenticityof this invitro-synthesized
genome-length cDNA was confirmed by restriction
enzymemapping after itwasconverted intodouble-'
stranded DNA in vitro (37;K. Steele,J. M.Taylor, and M. M. C. Lai, unpublished data) and by
RNA-DNAhybridization (seeResults).
Heteroduplex mapping. REV-Ti cDNA(20,ig/
ml)washybridizedtoREV DNA (10,ug/ml)in20
pl
ofasolutioncontaining80%formamide,0.4 MNaCl, and 0.01 M piperazine-N,N'-bis-(2-ethanesulfonic
acid) (PIPES)buffer(pH 6.4)at46°Cfor 30min.The
hybridswerethendialyzed against1M glyoxalin0.01
M sodiumphosphatebuffer (pH 7.0)for 1 h at370C. Excessglyoxalwasremovedbydialysis against0.01 M
Tris-hydrochloride containing 0.001 M EDTA (pH
7.5) at 4°C overnight. Portions of thehybrids were
thenannealedtosimianvirus 40
(SV40)-polybromo-deoxyuridine[poly(BUdR)]markermoleculesatroom
temperature for2Xmin inmthe presenceof0.5 M N-tris(hydroxymethylXmethyl-2- aminoethanesulfonic
acid(TES), and0.05 MEDTA (pH7.5)(17).The final
product was then adjusted to 55% formamide and
spreadonto18%formamide.Spreadingwith66%urea
plusformamidewasperformedaccordingtothe
pro-ceduredescribedbyKungetal. (24).
TheSV40DNA(5.224kb)wasusedasthe internal
double-stranded nucleic acid-length standard. The
lengthofsingle-strandedDNAorRNAwastakento
be 0.714times the length ofacorresponding duplex
moleculeasdeterminedby spreading denatured SV40
DNA underthesameconditions (S. Hu, unpublished data).
Electrophoretic analy8is of RNA.
Polyacryl-amidegelelectrophoresiswascarriedoutaccordingto
a modification of the method of Duesberg (7). The
RNAwaselectrophoresedin 2%polyacrylamide gels
cross-linkedwithbisacrylamide(7 by0.6 cm)at50V for 5 h.Afterelectrophoresis, thegelswerefrozen and
sliced into 1-mm fractions which were incubated in
toluene-basedscintillation fluidcontaining 10%NCS
(Amersham/Searle) and 1% water at 50°C for 5 h
beforecounting.
RESULTS
Electrophoretic analysis of the RNA ge-nomeofREV. We have previously shown that the virus released from an REV-transformed BMC line (11) containsa355RNA, which
cor-respondstotheRNA of the helper virus REV-A, anda28SRNA, which represents thegenome
of the defectivetransforming component (4, 16). This cell linewasusedas oneof the virussources
forheteroduplex mapping in this study. In ad-dition, two different cell lines, TV-1 and TV-2,
established from the spleen cells of REV-in-fected chickenswerealso used (23). These two celllinescarryB-celldeterminants,and thevirus released from these cells is capable of causing reticuloendotheliosis (23). To identify the RNA
speciesof virusesreleasedbythese cell lines,we
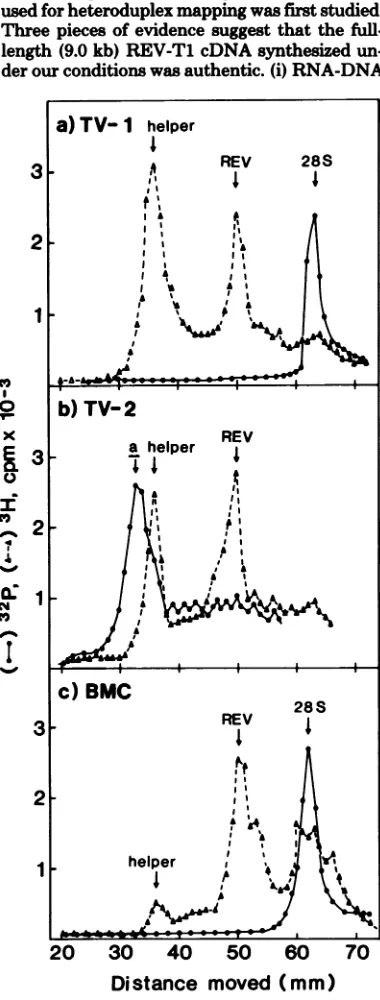
analyzed the 50 to 70S RNA genome of the viruses by polyacrylamide gel electrophoresis. The viruses from both TV-1 and TV-2 contained
a 35S RNA and asmaller RNA species which migrated slower than the 28S rRNA marker
(Fig. 1). Inanalogy to the REV released from the BMC line (Fig. lc), the 35S RNA probably represented helper REV-A and the smaller RNA species represented the REV genome which is responsible foroncogenicity in vivo and trans-forming capacity in vitro (4). We henceforth designate this small RNA as 30S REV RNA. However, incontrast totheBMC line, the virus released from TV-1 and TV-2 contained about equal ratios of the 35Shelper RNA and the 30S transforming component. The virus released from BMC occasionally contained, in addition to 35S and 30S RNA, a smaller RNA species which migrated almost together with the 28S rRNA in polyacrylamide gel electrophoresis
(Fig. lc). This RNA species has been noted before (4). From heteroduplex studies (see be-low), we suggest that this small RNA species might represent a defective form ofthe trans-forming RNA component.
Characterization ofthefull-length REV-TicDNA. Tocharacterize thesize and location of the genetic deletion as well as the
transfor-mation-specificsequences in the REV genome, weperformedelectronmicroscopic heteroduplex mapping between the full-length cDNA of the
helper virus REV-Ti andthe 30S REV RNA. Thefull-length REV-Ti cDNA was synthesized
by the endogenous reverse transcription from REV-Ti accordingtopublishedmethods (26).
Since it has been reported that the endoge-nousreversetranscriptionof
on November 10, 2019 by guest
http://jvi.asm.org/
osis viruses produces unfaithful cDNA (30), the authenticity of the full-length REV-Tl cDNA used forheteroduplexmapping wasfirst studied. Three pieces ofevidence suggestthat the
full-length(9.0kb) REV-Tl cDNA synthesized un-derourconditionswasauthentic. (i) RNA-DNA
20 30 40 50 60
Distance moved (mm)
FIG. 1. Polyacrylamidegel electrophoresis ofREV
RNA.The[3HJuridine-labeled50 to70Svirion RNAs
releasedfrom (a) TV-I, (b), TV-2, and(c)BMCcell
lines wereelectrophoresedin2%polyacrylamide gels
at50 Vfor5h. The markers usedwere32P-labeled 28SrRNA and35SclassaRNAofthePraguestrain
ofRoussarcomavirus(9).
REV GENOME 901
hybridization showed that this cDNA species
could annealtoREV-Tl 70SRNA to 85 to 100%,
dependingonthe hybridizationconditions
(Ta-ble 1). It did not hybridize to any other viral
RNA tested. These resultssuggest that this full-length cDNArepresents the entiregenomic se-quencesof REV-Tl and also that the cDNA is afaithfulcopyof REV-Tlgenome. (ii) Electron
microscopic homoduplex mapping between
REV-Tl full-length cDNA and 70S REV-Tl RNA showed that theywerecompletely homol-ogous (notshown). (iii) The full-length(9.0 kb)
REV-Tl cDNA was converted into
double-stranded DNA, using DNA polymerase I, and
thenanalyzed byrestrictionenzyme mapping as
described previously (37). The restriction pat-tems suggest that this cDNA species is a ho-mogeneous species of DNA and that it repre-sents an authentic copy of the REV-Tl RNA genome (Steeleetal., unpublished data).
Heteroduplex mapping betweenREV-Ti
cDNA and REV (REV-A) RNA from the transformed spleen cell lines 1 and
TV-2.Theheteroduplexmoleculeswerestudied by published procedures (18, 24). The 50 to 70S
RNA from TV-i and TV-2 cell lines wasfirst used. The 3'ends of the RNA molecules were
identified by binding apoly(BUdR)-tailed
cir-TABLE 1. Hybridization offull-length (9.0 kb)
REV-Ti cDNAwith REV-TI RNA'
% cDNA or
cDNA RNA RNA
hybrid-ized
3H-REV-T1 REV-Ti 100
PR-B 6
None 4
REV-Ti 3P-REV-T1 85
nP-PR-B
33P-RauscherMLV 2
None 1.5
a
Hybridization
was performed in a solutioncon-taining0.01MTris-hydrochloride, pH7.4,0.6 MNaCL
0.05% sodium dodecyl sulfate, 25pg oftRNA, and
appropriateamountsof cDNA and RNAat660C for
12h aspreviouslydescribed (4). In thefirst
experi-ment, 1,000 cpm (specific activity, 20,000cpm/Ag)of
full-length (9.0 kb) 3H-labeled REV-Ti cDNA was
hybridized with 0.2 pig of 70S RNA from different
viruses. Afterhybridization,thehybridizationmixture
wasdigested with nucleaseS1,andthepercentage of
[3H]DNA hybridizedwasdeterminedby precipitation
with trichloroacetic acid. In the second experiment,
1,000 cpmof3H-labeled REV-Ti cDNAwas
hybrid-ized with1,000 cpm of 32P-labeled 70S RNA(specific
activity, 106cpm/pg).Afterhybridization,the
hybrid-ization mixturewasdigested with RNase A (10 ug/
ml), andthepercentage of[32P]RNAhybridizedwas
determined by trichloroacetic acid precipitation.
MLV, Murine leukemia virus.
0
x
I 0
7-N
0
CO
I
A
on November 10, 2019 by guest
http://jvi.asm.org/
[image:3.488.50.240.93.593.2]cular SV40 DNA to the 3' polyadenylic acid
[poly(A)]
(1, 18, 27). Two types of moleculeswere observed. (i) The first type included the heteroduplex molecules formed between full-length REV-Ti cDNA and the helper virus,
REV-A RNA (not shown). These molecules
werecircularorlinear withcomplete homology.
Thisresult confirmedouroligonucleotide
map-pingshowing thatREV-Ti and REV-A are
al-most completely identical (4). (ii) The second
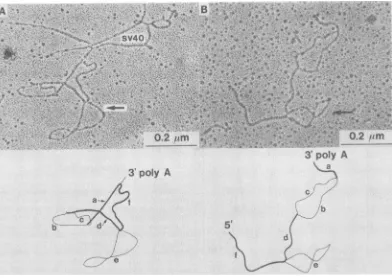
typeincluded theheteroduplexformed between REV-Ti cDNA and 30S REV RNA (Fig. 2).
This type of heteroduplex molecule had the following structural features. (i) Starting from the 3' end of thegenomeand immediately ad-jacenttothepoly(A)sequencewas astretch of homologous sequences of 0.73 ± 0.07 kb. This region ("a" in Fig. 2) corresponded to the C regionwhichisconstantin theRNA genomeof the avian leukosis-sarcoma virus complex (5).
Thisstretch ofhomologous sequencesmight also representthesequencesof similarnature in the
reticuloendotheliosis groupof viruses. (ii) Next
tothehomologoussequences atthe 3' endwas
a large substitution loop, with the two arms
measuring1.92±0.21("b" in Fig. 2) and1.64±
0.20kb ("c" inFig. 2),respectively. The
differ-enceinlength between thetwoarmsmightnot
besignificant, and theassignmentofthe strands
toeither virus isarbitrary.Thisnonhomologous
regionsuggeststhatREVlacks1.6to 1.9kb of the REV-Asequencesand containssome REV-specificsequencesofsimilar size whichare dis-tinct fromREV-Asequences.Thisregion (1.6to 1.9kb),therefore, represented theREV-specific
sequences whichmightberelatedtothe
onco-genicactivity ofREV. In itslocation, it
corre-spondedtothe envgeneof the avian
leukosis-sarcomaviruscomplex.(iii)Alargedeletionloop
("e" inFig. 2) of3.64 ± 0.25kbwaslocated at
1.64 ± 0.22 kb fromthe 5' end. This deletion looprepresented the secondstretch ofthe se-quences missing in REV and corresponded to
the3' half of thegag geneand 5' halfof thepol
FIG. 2. Electron micrographs ofREV-Ti -REV heteroduplexmolecules. (A) Circularheteroduplex; (B)
linearheteroduplex. About 15% of the heteroduplex moleculeswascircular because of thenoncolinearity
between the cDNA and the RNA(21). In the circularheteroduplex, the3'poly(A) was linked to an
SV40-poly(BUdR)marker. There were a few other linear RNA strands attached to the same marker molecule. In
the linearheteroduplexmoleculeshown (B), noSV40marker DNA was attached to the3'end.The orientation
of 3' and 5'endswasdeterminedfromother linearheteroduplexes (not shown). The larger gag-poldeletion
loop (shown by the arrows) wasfrequentlyseen with a figure eight structure, suggesting the presence of a
short stretch ofself-complementary sequences within the loop. See also Fig. 4.
on November 10, 2019 by guest
http://jvi.asm.org/
REV GENOME 903
gene, if we assume that REV-A has the same geneorderas dootheroncoviruses. This struc-ture confirms thegenetic defectivenessofREV (15, 16). Between the 3'-halfsubstitution loop and the 5'-half deletion loop was a stretch of
homologous sequencesof 1.04 t0.08 kb ("d" in
Fig. 2). This region should correspond to the partial sequence of the pol gene and possibly
alsopartof theenv gene. Thepresence of this
stretch of homologous sequences betweenthe
5'-end deletion loop and the 3'-half REV-specific
sequencesdistinguishes REV from other avian acuteleukemia viruses,such asMC29oravian
erythroblastosisvirus,in which thegag-poldele-tion is contiguouswith the leukemia virus-spe-cificsequences (19,28).
Depending on the strand assignment ofthe
substitution loop, thegenome size of REV-Awill be (1.64 + 3.64 + 1.04 + 1.64 + 0.73 + 0.20) = 8.89±0.82kbor9.17+0.83 kb,and the genome
size ofREVwill be (1.64 + 1.04 + 1.92 + 0.73 +
0.20) =5.53 ±0.58kb or 5.25 ±0.57kb.These
estimatesarein agreementwith thevalues ob-tained by electrophoresis in denaturing gels,
namely,8.7kbfor REV-A and 5.9 kb for REV
(16).
Heteroduplex mapping between
REV-Ti
cDNA and REV (REV-A) RNA released from the BMC line. To further confirm the geneticstructureofREV, weperformed heter-oduplex mappingbetweenREV-Tiand the
vir-ion released from anotherREV-transformedcell
line, the BMC line (11). This cell line releaseda
higher proportion of the transforming compo-nent, REV, than did the REV-transformed
spleen cell lines TV-1 andTV-2 (Fig. 1) (4). As
with the virusreleased from the TV celllines,
these heteroduplexes contained the same two
typesofheteroduplex molecules: (i)the hetero-duplexbetweenREV-TicDNAandRNAof the helperviruspresent inthe REV (REV-A) virus
population (these moleculeswerecompletely
ho-mologous [not shown]) and (ii) the heterodu-plexes between REV-Ti cDNA and 30S REV
RNA, which had a structure identical to that describedinFig.2.Thus, this RNAspecies likely
represented the REV RNA whichwas
respon-siblefor theoncogenicactivity ofthe virus. Inadditiontothese twokinds ofhybrid mol-ecules,athirdkindwasconsistentlyobserved in
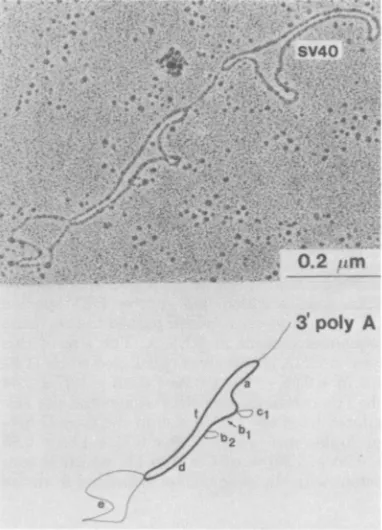
all of the viruspreparations released from the BMC line(Fig.3). Thisnewtype ofheteroduplex
ishenceforthtermed typeII,and the first type istermed type I. The type IIheteroduplex
mol-ecules (Fig. 3) had thefollowingstructural fea-tures.Startingat0.90+0.12kb from the 3'end,
there were threedeletionloops,with sizesof0.60 ±0.07 (cl), 1.09 +0.12 (b2), and3.69+ 0.25kb (e), in the middle of theheteroduplexmolecules.
-W2f.-0e|-pA,X
r7T i v<Sj
b21 A
FIG. 3. Electron micrograph ofatype II
hetero-duplex molecule betweenREV-TIandREV released
froma BMCline. AnSV4O-poly(BUdR)marker
mol-eculewasattachedtothe3'poly(A)sequencesofthe
hybrid. The molecule wascircular asexplained in
thelegendofFig.2.Theorientationof3'and5' ends
wasdeterminedfromthelinear heteroduplex
mole-cules with theattachedSV40 DNA(not shown).
The third deletion loop was located at 1.67 + 0.10 kb from the 5' end and was, therefore, similarinboth size and locationtothedeletion
loop inthetype Iheteroduplex. Thus,thisloop
represented the deletion in the gag-polgenes. Theothertwosmaller deletionloopsfellinthe
regionroughlycorrespondingtothe substitution loopinthetype Imolecules. Thissuggests that the type Iandtype IImoleculesmightdifferin
the extentof deletion and the size of the
REV-specificsequences in thisregion.
Todetermine the origin and nature of these
twosmall deletion loops, we further examined
the type II heteroduplexes under a spreading
condition (hyperphase, 50% formamide; hypo-phase, 15% formamide) under which
single-stranded DNA was well extended but
single-strandedRNA tendedtoform acollapsed
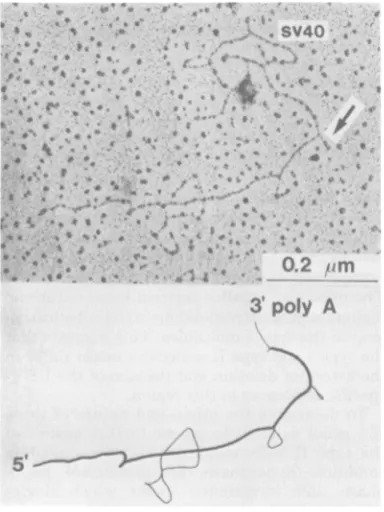
con-figuration. The two larger deletion
loops
(3.69and1.09kb)werealwayswellextended(Fig. 4),
suggestingthattheyrepresented DNAstrands
and, therefore, REV-A sequences which were
missing in REV. In contrast, the 0.60-kb
loop
37, 1981
on November 10, 2019 by guest
http://jvi.asm.org/
[image:5.488.250.441.76.341.2]904 HU ET AL.
(denoted byan arrowin Fig. 4)wascondensed
into a hairpin-like structure under this
condi-tion, suggesting that the 0.60-kb loop
repre-sented an RNA strand, and, therefore, REV-specific sequenceswhichweremissingin
REV-A. This stretch ofREV-specificsequences was much smaller than that in type I molecules. Likewise,the 1.09-kb deletionloop representing the REV-A-specific, and possibly env-related,
sequences which weremissingin REV wasalso smaller thanthe 1.6- to 1.9-kbdeletion observed
inthetypeImolecules.Furthermore, therewere
homologous sequences (0.55 kb) (b1) between thesetwodeletionloops. Thus, thetype II het-eroduplex could havebeenformed withanREV RNA species which had shorter REV-specific
sequencesandyetretainedpartof theenvgene sequences present in REV-A. The size ofthis
type IIRNA species wascalculated tobe (1.67
+0.75+0.55+0.60+ 0.90+0.20)=4.67+0.44
kb. ThecorrespondingREV-Agenomesize cal-culatedfromthecDNA sizein thetype II het-eroduplexwas (1.67 + 3.69 + 0.75 + 1.09 + 0.55 + 0.90 + 0.20) = 8.85 ± 0.74 kb,which is con-sistentwith thegenome size estimatedfrom the
type I heteroduplex and from electrophoresis (16). MeasurementsoftypeI andtypeII
heter-oduplexesaresummarized inFig.5.
The typeI andtypeII moleculeswerepresent in a 30:70 ratio in all of the RNA preparations obtained from the BMC line, regardless ofthe
passage history of the cells. To rule out the
possibility thatthe type IIheteroduplex might beanartifact of thespreading condition,
result-ingfrompartial homologyinthe two armsofthe
substitution loop observed in type I heterodu-plex molecules, we further studied the
hetero-duplex under various spreading conditions. We
compared the heteroduplexes after treatment with glyoxal followed by formamide spread or without glyoxal treatment and spread directly with 65% urea plus formamide. The latter method has been shown to cause lessrandom
denaturation and allow for detection of partially homologous sequences (18). If the type I and
type II molecules are derived from the same
RNAspecies, the ratio of these two heterodu-plexes will vary with the stringency of the spreading method (18). However,nodifference
in the ratio of typeI and typeII heteroduplex molecules was observed underdifferent condi-tions. We therefore conclude that these two structures are heteroduplexes formed with
dif-REV-TI f b
TYPE
IRV3
poly Ab2
TYEIREV-TI f db, a
REV (BMC)
_
|l|||l_
iu.=
MFIG. 4. Electronmicrograph ofa
type
HI
REV-TiREVheteroduplexwhenspreadunder50%'
formam-idewithoutpriorglyoxaltreatment.Thedeletionloop
closest tothe 3' end (arrow) is now seen collapsed,
whereas the other two deletion loops are still well
extended.
C,
LENGTHS OF SEGMENTS OF HETERODUPLEX (IN KILOBASES)
TYPE I TYPE A
a 0.73 ! 0.07 090 ! 0.12 b 1.92 ! 0.21 -
-0.55 t 0.05
1.09 ! 0.12
b2
1.64 * 0.20
C1 - - 0.60 0.07
d 1.04 ± 0.08 0.75 ± 0.10 e 3.64 0.25 3.69 0.25 f 164 ± 0.22 1.67 ± 0.10
FIG. 5. Schematic drawingandmeasurements of
type I andtypeIIheteroduplex moleculesformedwith
virionRNAreleasedfromaBMC line.A totalof78
type Iheteroduplex moleculesand144type II
heter-oduplex molecules were observed. Out ofthese, 21
type Imoleculesand 35 type IImoleculeswere used
for lengthmeasurements.
J. VIROL.
bi
on November 10, 2019 by guest
http://jvi.asm.org/
[image:6.488.53.245.338.594.2] [image:6.488.191.437.356.589.2]REV GENOME 905
ferent RNA species and that thetype II mole-cules are notthe products of partial denatura-tionoftype Imolecules. Thus,typeIImolecules
represent a newspecies of RNA with a smaller
size and smaller REV-specific andpossibly
on-cogenic sequences. These kinds of molecules
have so far only been found in virus released from theBMC cell line.
DISCUSSION
Biological datasuggested that REV is defec-tiveand requireshelper virus for itsreplication (11, 16).Electrophoresis andheteroduplex
map-ping showed that it has a5.5 to 5.9-kb or 30S
RNA genome (4, 16) and that 30% of its se-quences areREV specific andpresumably are
responsible for theoncogenic activityof the virus (4). Our heteroduplex studies further demon-strated that REV hasa3.64-kbdeletion in the
gag-polgenes,ifone assumesthatREVhas the
same gene order as other oncoviruses. Thus, REV is sinilar to other avian acute leukemia viruses in that it hasanextensive deletion in the
gag-polgenes (19, 28). The REV-specific, and presumably oncogenic, sequences in the REV
genomewhichareresponsibleforvisceral retic-uloendotheliosis in vivo and transformation of fibroblasts and bonemarrow cellsin vitro
con-stitute acontiguous stretch of1.6 to 1.9 kb in
length. These sequences account for30 to 35% of the genetic sequences of REV, which is in good agreement with the published data ob-tained by cDNA-RNA hybridization (4). Our study showed that theseREV-specificsequences werelocalized in theregioncorrespondingtothe
envgeneandwere notcontiguouswiththe
gag-pol deletion in the REVgenome. Thisgenetic
structure is in contrast to that of other avian
acute leukemia viruses (19, 28) and is rather similarto that ofthe defective Friend strain of
spleenfocus-formingvirus(39),defective murine
sarcomavirus (17), anddefective mink cell fo-cus-inducingviruses, whicharerecombinantsin
theenvgenebetween ecotropicandxenotropic
viruses (3, 40). Recently, we have also shown that the
transformation-specific
sequences ofREV are not related to those ofother known
avianormurine acuteleukemiavirusessuchas MC29, MH2,avian
erythroblastosis
virus,
avian myeloblastosis virus, orAbelson murine leuke-mia viruses (T. C. Wong and M. M. C. Lai,Virology,inpress).Thus,REV representsa new
classof avianacuteleukemia virus.
Allofthe avianacuteleukemiavirusesstudied
so far codefor apolyprotein which isafusion
proteinbetween part of the gag sequences and thetransformation-specificsequences(2,14,20).
Since the gag-pol gene and the
presumably
transformation-specific sequences in REV are not contiguous, we would not expect REV to
code for suchafusionpolyprotein.However, it is stillpossible thattheREV-specificsequences localized in the env region extend beyond the
termination point of thepol gene. If this is the case, apolyprotein containingpart of the
gag-pol and the transformation-specific sequences
might exist. It has beenreported that, in some
of the REV-transformed nonproducer cells, a
protein of 130,000 molecular weight could be
detected (16). However, this protein was not
observed in all of the REV-transformed cell lines and itwas notcertain whether itwas acellular protein or a virus-coded protein. Despite
re-peatedattempts, wehavenotbeenable to dem-onstrate such a polyprotein in various
REV-transformed cell lines. Alternatively, the
REV-specific sequenceslocated in the region
corre-spondingtotheenv geneof REV-A
might
codefor a transformation-related glycoprotein, in
analogytoFriendspleen focus-formingvirus (6, 34, 35). Further experiments are required to
resolve this issue.
Thetype IIRNA observed in thevirus
pop-ulationsreleased from theBMC line is particu-larly interesting.ThisRNAmight correspond to
the RNAspecies which
migrated
nearly together with the 28S rRNA marker in 2% polyacryl-amidegel electrophoresis (Fig. lc) (4). Gondaetal.have,also detected suchasmall RNA species (12).ThetypeII RNAmolecules containsmaller
REV-specific sequences andasmaller deletion intheenvregion than dotype IRNAmolecules, but it retainssome of the env gene sequences
homologoustothose of REV-A.As aresult,two
smallerdeletionloops,inplaceof thebig substi-tutionloop inthetype Imolecules,arepresent
in type II heteroduplexes. These two deletion loopsrepresentREV-specificsequencesand
de-letedsequences,respectively,intheenvgeneof REV-A.Sincewecould notdetect suchanRNA
species inREV-A populations,we suggestthat thetype IIRNAwasprobablyarecombination product of an unequal crossover between the
helpervirus, REV-A, and thetransforming com-ponent, REV.It isnotclearwhether thisRNA hasanybiological significance. Nevertheless,it
is interesting to note that an RNA species of similargenetic structurehas also been detected in avianmyelocytomatosisvirusMC29 released from a transformed quail cell line (Q10) (19). ThisRNAwastermed AMC29.Both the
REV-transforned BMC line and MC29-transformed
Q10line, inwhich this smallRNAspecieswas discovered, contain unusually low titers of the
helperviruses(4, 8).Thisfact suggests that this
smallRNAspecies might functionasdefective
on November 10, 2019 by guest
http://jvi.asm.org/
interfering particles andinterfere with the
rep-lication of the helper viruses. Further studiesare
requiredtodetermine thebiological significance of this small RNA species.
ACKNOWLEDGMENTS
Thisworkwassupported by PublicHealthService research grantCA16113 awarded by theNationalCancerInstitute and byAmericanCancerSocietyresearch grantMV-80. T.W.isa
postdoctoral fellowof the LeukemiaSocietyofAmerica. LITERATURE CITED
1. Bender, W., and N. Davidson. 1977.Mapping of poly (A)sequences intheelectronmicroscope reveals
un-usualstructure oftypeConcornaviruses RNA mole-cules. Cell7:595-607.
2.Bister, K., M. J. Hayman,and P.K.Vogt.1977. De-fectiveness of avian myelocytomatosis virus MC29: iso-lation oflong-term nonproducer culturesandanalysis ofvirus-specificpolypeptide synthesis. Virology 82:431-448.
3. Bosselman, R. A., L. J. L.D.vanGriensven,M.Vogt, and L.M.Verma. 1980. Genomeorganizationof
Ret-roviruses.IX.Analysisof thegenomesofFriend spleen focusforming (F-SFFV) andhelpermurine leukemia viruses byheteroduplex formation.Virology 102:234-239.
4. Breitman,M.L.,M. M. C.Lai,and P.K.Vogt.1980. Thegenomic RNA ofavianreticuloendotheliosisvirus REV.Virology100:450-461.
5. Coffin, J. M., M. Champion, and F. Chabot. 1978. Nucleotide sequence relationships between the ge-nomesofanendogenousandanexogenousaviantumor virus.J.Virol.28:972-991.
6. Dresler, S.,M. Ruta,M. J.Murray,and D. Kabat. 1979.Glycoprotein encoded bytheFriendspleen focus-formingvirus. J. Virol.30:564-575.
7.Duesberg,P. H. 1968.Physical propertiesofRous sar-comavirusRNA. Proc. Natl. Acad. Sci. U.S.A. 60:1511-1518.
8. Duesberg,P.H.,K.Bister, andP.K. Vogt.1977.The RNAofavianacuteleukemia virus MC29.Proc.Natl. Acad.Sci. U.S.A.74:4320-4324.
9. Duesberg,P.H.,and P. K.Vogt. 1973. RNAspecies obtained fromclonal lines of aviansarcomaand from avianleukosisviruses.Virology54:207-219.
10.Franklin, R. B., C. Y.Kang, M. K. Wan, and H. R. Bose.1977.Transformationofchickembryofibroblasts byreticuloendotheliosisvirus.Virology83:313-321.
11. Franklin,R.B., R. L.Maldonado, andH. R.Bose. 1974.Isolation and characterizationof reticuloendothe-liosisvirustransformed bonemarrowcells.
Intervirol-ogy3:342-352.
12. Gonda, M. A., N.R.Rice, andR.V. Gilden.1980.Avian reticuloendotheliosisvirus:characterization of the high-molecular-weightviralRNA intransforming and helper viruspopulation.J.Virol. 34:743-751.
13. Halpern,M. S., E. Wade, E. Rucker, K. L. Baxter-Gabbard, A.S.Levine,and R. E. Friis. 1973. A study oftherelationshipof reticuloendotheliosis virustothe
avianleukosis-sarcomacomplex of virus. Virology53:
287-299.
14. Hayman,M. J., B. Royer-Pokora, and T. Graf.1979. Defectivenessof avian erythroblastosisvirus: synthesis
ofa75Kgag-related protein. Virology 92:31-45. 15. Hoelzer,J. D., R. B. Franklin, and H.R.Bose. 1979.
Transformationbyreticuloendotheliosisviruses:
devel-opmentofafocusassayandisolation ofa nontransform-ingvirus. Virology 93:20-30.
16. Hoelzer,J. D., R. B. Lewis, C. R. Wasmuth,and H. R.
Bose. 1980. Hematoieticcelltransformation byavian
reticuloendotheliosis: characterization of the genetic de-fect.Virology 100:462-474.
17.Hu, S., N. Davidson, and L.M. Verma. 1977. A
hetero-duplex study of thesequencerelationships between the RNAsofM-MSV and M-MLV.Cell 10:469-477. 18.Hu, S. S. F., M. M. C. Lai, and P. K. Vogt. 1978.
Characterization of theenv genein avian oncoviruses by heteroduplex mapping.J. Virol.27:667-676. 19.Hu,S. S.F.,M. M. C.Lai,and P. K.Vogt.1979.Genome
of avian myelocytomatosis virus MC29: analysis by heteroduplex mapping. Proc.Natl. Acad. Sci. U.S.A. 76:1265-1268.
20. Hu,S. S.F., C.Moscovici,and P. K.Vogt. 1978.The defectivenessofMillHill2,acarcinoma-inducingavian oncovirus.Virology 89:162-178.
21. Junghans, R. P.,S. Hu, C. A. Knight, and N.
David-son.1977.Heteroduplex analysis of avian RNA tumor viruses. Proc.Natl.Acad. Sci. U.S.A. 74:477-481. 22. Kang,C.Y.,and H.M. Temin. 1973. Lack ofsequence
homologyamongRNAs of avian leukosis-sarcoma vi-ruses,reticuloendotheliosisviruses,and chicken
endog-enousRNA-directed DNApolymeraseactivity.J.Virol.
12:1314-1324.
23. Keller,L.H.,R.Rufner,and M. Sevoian. 1979.
Isola-tion anddevelopment ofareticuloendotheliosis virus-transformed lymphoblastoid cell line from chicken spleencells.Infect.Ixnmun. 25:694-701.
24. Kung, H. J., J.M.Bailey,N.Davidson,M. D. Nicol-son,and R.M.McAllister.1975.Structure,subunit compositionand molecularweightofRD-114 RNA.J. Virol. 16:397-411.
25. Lai, M. M. C. 1976.Phosphoproteins of Roussarcoma
viruses.Virology74:287-301.
26. Lai,M. M.C.,and S. S.F.Hu. 1978. Invitrosynthesis and characterization of full- and half-genome-length complementaryDNA from avian oncoviruses. Nature (London) 271:481-483.
27. Lai, M. M. C., S. S. F.Hu, and P. K. Vogt. 1977. Occurrence ofpartialdeletionand substitution of the
srcgenein the RNAgenomeofaviansarcoma virus. Proc.Natl. Acad.Sci.U.S.A.74:4781-4785.
28. Lai,M. M. C., S. S. F. Hu, and P. K.Vogt.1979. Avian erythroblastosis virus: transformation-specific
se-quencesformacontiguoussegment of 3.25 kb located
inthe middle of the 6-kbgenome.Virology97:366-377. 29. Maldonado,R.L.,and H.R.Bose. 1973.Relationship of the reticuloendotheliosis virus tothe aviantumor
viruses: nucleic acid and polypeptide composition. J. Virol. 11:741-747.
30. Mizutani, M.,and H. M. Temin. 1975. Purificationand propertiesofspleennecrosis virus DNApolymerase.J. Virol. 16:797-806.
31. Olson, C. D. 1967. Histopathological and hematologic changesinmoribund chicks infected with T-virus.Am. J. Vet. Res. 28:1501-1507.
32. Purchase,H.G.,C.Ludford,N.Nazerian,and H.W. Cox.1973. Anewgroupofoncogenicviruses: reticulo-endotheliosis, chicksyncytial,duckinfectiousanemia, andspleennecrosisvirus.J. Natl. Cancer Inst. 51:489-499.
33. Purchase,H.G.,and R. L.Witter. 1975. The reticulo-endotheliosis viruses. Curr.Top.Microbiol.Immunol. 71:103-124.
34. Racevskis, J.,and G. Koch. 1978.Viralprotein synthesis inFrienderythroleukemiacelllines.Virology
87:345-365.
35. Ruscetti, S.K.,D.Linemeyer,J.Field,D.Troxler, andE. M.Scolnick. 1979. Characterization ofaprotein
found in cells infected with thespleenfocus-forming virus that sharesimmunological cross-reactivitywith thegp7Ofound in minkcellfocus-inducingvirus parti-cles.J. Virol. 30:787-798.
36. Sevoian,M.R.,N.Larose,and D. M.Chamberlain.
on November 10, 2019 by guest
http://jvi.asm.org/
1964. Avian lymphomatosis. VI. A virus of unusual potencyandpathogenicity.Avian Dis.8:336-347. 37.Taylor, J. K,T. W. Hsu, and M. M. C. Lai. 1978.
RestrictionenzymesitesontheavianRNAtumorvirus genome.J.Virol.26:479-484.
38. Theilen, G. H, R. F. Zeigel, and KL J. Twiehaus.1966. Biological studies with RE virus (strain T) that induces reticuloendotheliosis in turkeys, chickens, and Japanese quail. J. Natl. Cancer Inst.37:731-743.
39. Troxler,D.H.,andE. M.Scolnick.1978.Rapid
leuke-mia inducedbycloned Friendstrain ofreplicating
mu-rine type-C virus. Virology 85:17-27.
40. Van Griensven, L. J. L. D., and M. Vogt. 1980. Rauscher "minkcellfocus-inducing"(MCF) viruscause
erythroleukemia in mice: its isolationandproperties. Virology 101:376-388.
41. Vogt, P. K., J.L.Spencer,W.Okazaki,R.L.Witter,
andLB. Crittendon. 1977. Phenotypic mixing
be-tweenreticuloendotheliosis virus andsarcomaviruses. Virology80:127-135.
42. Witter, R. L,H. G.Purchase,and G. H.Burgoyne. 1970.Peripheralnervelesions similartothose of Ma-rek's diseaseinchickens inoculated with reticuloendo-theliosisvirus.J.Natl. Cancer Inst. 45:567-577.
on November 10, 2019 by guest
http://jvi.asm.org/