The Human Cytomegalovirus IE1 Protein
Antagonizes PML Nuclear
Body-Mediated Intrinsic Immunity via the
Inhibition of PML
De Novo
SUMOylation
Eva-Maria Schilling,
aMyriam Scherer,
aNina Reuter,
aJohannes Schweininger,
bYves A. Muller,
bThomas Stamminger
aInstitute of Clinical and Molecular Virology, Friedrich-Alexander-Universität Erlangen-Nürnberg, Erlangen,
Germanya; Division of Biotechnology, Friedrich-Alexander-Universität Erlangen-Nürnberg, Erlangen, Germanyb
ABSTRACT
PML nuclear bodies (NBs) are accumulations of cellular proteins embedded
in a scaffold-like structure built by SUMO-modified PML/TRIM19. PML and other NB
pro-teins act as cellular restriction factors against human cytomegalovirus (HCMV); however,
this intrinsic defense is counteracted by the immediate early protein 1 (IE1) of HCMV. IE1
directly interacts with the PML coiled-coil domain via its globular core region and
dis-rupts NB foci by inducing a loss of PML SUMOylation. Here, we demonstrate that IE1
acts via abrogating the
de novo
SUMOylation of PML. In order to overcome reversible
SUMOylation dynamics, we made use of a cell-based assay that combines inducible IE1
expression with a SUMO mutant resistant to SUMO proteases. Interestingly, we observed
that IE1 expression did not affect preSUMOylated PML; however, it clearly prevented
de
novo
SUMO conjugation. Consistent results were obtained by
in vitro
SUMOylation
as-says, demonstrating that IE1 alone is sufficient for this effect. Furthermore, IE1 acts in a
selective manner, since K160 was identified as the main target lysine. This is
strength-ened by the fact that IE1 also prevents As
2O
3-mediated hyperSUMOylation of K160,
thereby blocking PML degradation. Since IE1 did not interfere with coiled-coil-mediated
PML dimerization, we propose that IE1 affects PML autoSUMOylation either by directly
abrogating PML E3 ligase function or by preventing access to SUMO sites. Thus, our
data suggest a novel mechanism for how a viral protein counteracts a cellular restriction
factor by selectively preventing the
de novo
SUMOylation at specific lysine residues
with-out affecting global protein SUMOylation.
IMPORTANCE
The human cytomegalovirus IE1 protein acts as an important antagonist
of a cellular restriction mechanism that is mediated by subnuclear structures termed
PML nuclear bodies. This function of IE1 is required for efficient viral replication and thus
constitutes a potential target for antiviral strategies. In this paper, we further elucidate
the molecular mechanism for how IE1 antagonizes PML NBs. We show that tight
bind-ing of IE1 to PML interferes with the
de novo
SUMOylation of a distinct lysine residue
that is also the target of stress-mediated hyperSUMOylation of PML. This is of
impor-tance since it represents a novel mechanism used by a viral antagonist of intrinsic
im-munity. Furthermore, it highlights the possibility of developing small molecules that
spe-cifically abrogate this PML-antagonistic activity of IE1 and thus inhibit viral replication.
KEYWORDS
PML nuclear bodies, human cytomegalovirus, immediate early 1,
intrinsic immunity, nuclear domain 10, sumoylation
P
romyelocytic leukemia protein (PML) is known to act as organizer of subnuclear,
matrix-associated structures which can be detected as spherical dots of
approxi-mately 0.1 to 1.0
m (1). These structures, termed PML nuclear bodies (NBs) or nuclear
domain 10 (ND10), are multiprotein complexes consisting of more than 160 proteins.
Received15 October 2016Accepted28 November 2016
Accepted manuscript posted online30 November 2016
CitationSchilling E-M, Scherer M, Reuter N, Schweininger J, Muller YA, Stamminger T. 2017. The human cytomegalovirus IE1 protein antagonizes PML nuclear body-mediated intrinsic immunity via the inhibition of PMLde novoSUMOylation. J Virol 91:e02049-16. https://doi.org/10.1128/JVI.02049-16.
EditorRozanne M. Sandri-Goldin, University of California, Irvine
Copyright© 2017 American Society for Microbiology. All Rights Reserved.
Address correspondence to Thomas Stamminger,
thomas.stamminger@viro.med.uni-erlangen.de.
VIRUS-CELL INTERACTIONS
crossm
on November 7, 2019 by guest
http://jvi.asm.org/
PML NBs are implicated in several cellular processes: transcription, apoptosis,
senes-cence, response to DNA damage, and antiviral defense (2). The last is mediated by PML
itself and further NB-associated proteins, which act as cellular restriction factors,
thereby forming the first line of antiviral defense, the so-called intrinsic immunity.
Several viruses from different families, including herpes-, adeno-, polyoma-, rhabdo-,
and retroviruses, have been shown to be repressed by the NB-mediated intrinsic
immunity (3, 4).
In humans, there are 7 isoforms (I to VII) of PML (TRIM19), which are characterized
by one common feature at their N terminus: the TRIM/RBCC motif, consisting of a RING
domain, B boxes, and a coiled-coil region (5). It has been suggested that PML initiates
NB
de novo
formation by dimerization and subsequent oligomerization via this RBCC
motif. However, the maintenance and maturation of this structure, as well as the
recruitment of further proteins such as hDaxx and Sp100, highly depend on
posttrans-lational modification of PML by small ubiquitin-like modifiers (SUMOs) (2, 6–8). This
modification occurs on three major lysine residues (K65, K160, and K490) by covalent
attachment of SUMO molecules which exist in paralogs, termed SUMO1 to -4 and the
recently identified SUMO5 (9–12). While SUMO2 and SUMO3 share a high similarity and
therefore are commonly referred to as SUMO2/3, SUMO1 displays only 50% sequence
identity with SUMO2/3. SUMO2/3 efficiently forms polySUMO chains due to the internal
residue K11, which enables conjugation to itself and other SUMO paralogs. SUMO1, in
contrast, which is devoid of K11, is conjugated to its substrates only as single SUMO1
protein, or it can serve as a SUMO chain terminator at the end of a polySUMO chain (13).
Covalent attachment of the 11-kDa SUMO moiety to its substrate requires a four-step
enzymatic process (14). First, immature SUMO is processed by cellular SUMO proteases
(SENPs) to expose a diglycine motif required for conjugation. Second, mature SUMO is
activated in an ATP-dependent manner by the E1-activating enzyme, a heterodimer of
SAE1 and SAE2 in mammals. Third, SUMO is passed to the E2-conjugating enzyme UBC9
and finally conjugated to its substrate at the SUMO consensus sites (
-K-X-E/D, where
is a large hydrophobic amino acid and X is any amino acid). The last step is
accomplished by substrate-specific SUMO E3 ligases which play an important role for
conferring selectivity and efficiency of the SUMOylation reaction. Interestingly, PML
itself has been suggested to possess SUMO E3 ligase activity because of its ability to
interact with SUMO and Ubc9 as well as to stimulate SUMOylation in yeast and
mammalian cells (15–17). Therefore, the SUMOylated species of PML may arise at least
partially via autoSUMOylation, as also suggested by Shen and colleagues (18). The
dynamics between SUMOylation and deSUMOylation of NB-associated PML are
pro-posed to be indispensable for NB architecture and function (19). These dynamics are
fostered by the presence of all components of the SUMOylation machinery and also
SENPs, which are required both for processing of immature SUMO and for
deconjuga-tion of SUMO proteins (20). In addideconjuga-tion to physiological changes, there are several
stimuli that affect PML SUMOylation (2). The most extensively studied substance,
arsenic trioxide (As
2O
3), provokes the hyperSUMOylation of PML, which is followed by
recruitment of the SUMO-targeted ubiquitin ligase (STUbL) RNF4 and the subsequent
ubiquitin-dependent proteasomal degradation of PML (21).
Viruses have evolved several mechanisms to overcome PML NB-mediated intrinsic
immunity by exploiting the host cell SUMOylation system (22). For example, the herpes
simplex virus 1 (HSV-1) immediate early protein ICP0 acts as a STUbL which targets
SUMOylated PML and Sp100 as well as other SUMOylated factors for proteasomal
degradation (23, 24). The immediate early protein 1 (IE1) of human cytomegalovirus
(HCMV) also induces the loss of SUMOylated forms of PML, which is followed by the
disruption of NBs, and it has been shown by several studies that this correlates with the
antagonization of PML NB-mediated intrinsic immunity (25–31). For IE1, no such STUbL
activity has been reported (32). Since IE1 is SUMOylated itself, the hypothesis has been
raised that it acts as a decoy substrate by competing for SUMO (32, 33). However, this
idea was disproved in earlier studies, as a SUMOylation-negative mutant of IE1 is still
able to induce a loss of PML SUMOylation (32). Importantly, the direct binding of IE1 to
on November 7, 2019 by guest
http://jvi.asm.org/
PML seems to be a prerequisite for mediating the loss of SUMOylated PML (29, 34).
Previous studies identified a central large core region of IE1 as being sufficient for
interaction with PML; however, further attempts to narrow down the binding region by
using mutations and deletions failed (29, 30, 34–38). Indeed, the crystal structure of the
rhesus cytomegalovirus IE1
CORE, which was recently solved by our group, provided
strong evidence that IE1
COREconsists of a single contiguous domain which is very
sensitive to deletion mutagenesis (30). The crystal structure further revealed a similarity
between the crystal structure of IE1
COREand that of TRIM coiled-coil domains.
Inter-estingly, experiments from our group and others demonstrated that IE1 targets the
coiled-coil domain of PML and also interacts with other members of this protein family,
such as TRIM5
␣
and TRIM33 (30, 39). This led the authors to propose a model whereby
IE1 targets PML via an extended binding surface which involves coiled-coil interactions,
thereby interfering with PML SUMOylation (30).
In the present study, we set out to further clarify the mechanism that leads to a loss
of PML SUMOylation by IE1. By using
in vivo
as well as
in vitro
studies, we found that
IE1 alone is sufficient to inhibit the
de novo
SUMOylation of PML without affecting
preSUMOylated PML. This clearly demonstrates that IE1 exclusively inhibits the on-rate
of SUMO modification. Interestingly, IE1 seems to selectively modulate the
SUMOyla-tion of lysine 160, which is also known to be the main SUMO target site during
As
2O
3-mediated PML degradation. Indeed, we could observe that the effect of IE1 on
PML SUMOylation is evident not only under physiological conditions but also under
stress conditions. Taken together, our data suggest a novel mechanism for how a viral
antagonistic protein counteracts a cellular restriction factor by selectively preventing
the
de novo
SUMOylation at specific lysine residues without affecting global protein
SUMOylation.
RESULTS
IE1 induces the loss of PML polySUMOylation.
The antiviral activity of PML NBs is
known to be disturbed by HCMV in an IE1-dependent manner (27, 34). It is supposed
that the ability of IE1 to inhibit the accumulation of SUMO-conjugated forms of PML is
crucial for this effect, since SUMOylated PML is essential for assembling genuine NB
structures (1, 6, 7, 18, 28, 29). To get further insights into the exact mechanism of
IE1-mediated loss of PML SUMOylation, we performed infection experiments using
primary human foreskin fibroblast (HFF) cells. For this, we conducted time course
analyses with the HCMV strain AD169 at a multiplicity of infection (MOI) of 3. In
accordance with previous studies, PML was found to be partially deSUMOylated upon
infection (Fig. 1 A) (29, 40). While most high-molecular-weight bands of PML
disap-peared, one presumably modified species of PML seemed to be resistant against IE1
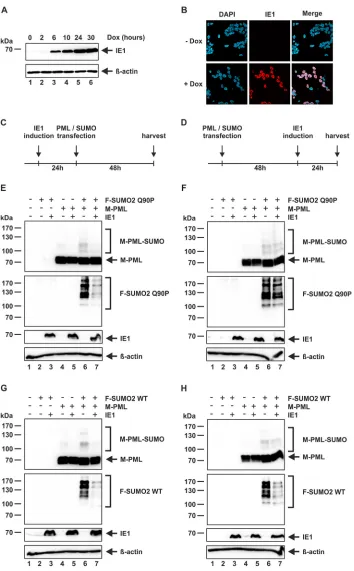
action (Fig. 1A, asterisk). The same was true in HFF cells with inducible expression of IE1,
thereby demonstrating a direct effect of IE1 on PML polySUMOylation (Fig. 1 B). Since
we speculated that the unaffected band may correspond to a SUMO-modified form of
PML, we reintroduced a FLAG-tagged version of PML (F-PML) I into PML knockdown
(PML-kd) HFFs by lentiviral transduction. Indeed, consistent results were obtained with
this PML isoform upon infection: while most SUMOylated PML species disappeared, one
isoform remained unaffected (Fig. 1C). To confirm that this resistant isoform
corre-sponds to a SUMO-modified form of PML, we conducted immunoprecipitation studies
with the F-PML I-expressing PML-kd HFFs. Cells were either mock infected or infected
with AD169, lysed at 24 h postinfection, and subjected to immunoprecipitation under
denaturing conditions. Modification of F-PML I with endogenous SUMO proteins was
subsequently detected with a SUMO2/3-specific antibody. Strikingly, while several
SUMOylated species of F-PML I were observed in mock-infected cells, the infected cells
displayed only one distinct band (Fig. 1D, right panel, compare lanes 2 and 4). Since this
band exhibited an approximately 15-kDa shift in molecular mass compared to
unmod-ified F-PML I, we assume that it corresponds to a monoSUMOylated F-PML I. Hence, our
data provide evidence that IE1 selectively affects PML polySUMOylation, while a
monoSUMOylated isoform of PML appears to be resistant.
HCMV IE1 Inhibits PMLDe NovoSUMOylation Journal of Virology
on November 7, 2019 by guest
http://jvi.asm.org/
IE1 specifically affects the SUMOylation of PML K160.
As depicted in Fig. 2A, PML
is modified mainly at three lysine residues, K65, K160, and K490. Each of these residues
can be either monoSUMOylated or polySUMOylated (10). To analyze whether the
IE1-resistant, monoSUMOylated PML species corresponds to a defined lysine residue,
we generated SUMOylation site mutants by changing the respective lysine to arginine.
Next, we characterized the SUMOylation patterns of single, double, and triple SUMO
site mutants of PML in HEK293T cells. As expected, due to the absence of all major
SUMOylation sites, expression of the triple mutant (3KR) did not result in any
slower-migrating, SUMOylated forms of PML (Fig. 2A, lane 8). Consistent results were obtained
with the K160R/K490R mutant (Fig. 2A, lane 7), as SUMOylation of K65 is tightly linked
to SUMOylation of K160 (21). The K65R/K490R mutant displayed one SUMO-modified
band (Fig. 2A, lane 6), corresponding to the SUMOylation of K160 (Fig. 2A, marked by
a circle), which obviously is the main SUMO conjugation site in wild-type PML (PML WT)
(Fig. 2A, lane 1). The K65R/K160R (Fig. 2A, lane 5) and the K160R (Fig. 2A, lane 3)
mutants revealed one distinct band presumably corresponding to the SUMOylation of
K490 (marked by an asterisk), which is in turn lacking in all K490R mutants (Fig. 2A, lanes
4, 6, 7, and 8). The K490R mutant (Fig. 2A, lane 4) displayed one additional
SUMO-modified species besides K160-SUMO, corresponding to the SUMOylation of K65/160
FIG 1IE1 induces the loss of polySUMOylated PML. (A and C) HFF cells (A) or PML-kd HFFs stably expressing F-PML I (C) were either mock infected or infected with AD169 at an MOI of 3. At the indicated time points postinfection, cells were harvested and analyzed by Western blotting for IE1,-actin, and PML (A) or F-PML (C). (B) HFF F-IE1 cells with doxycycline-inducible expression of F-IE1 were either not induced or induced with 0.5g/ml doxycycline. Cells were harvested at the indicated time points and analyzed by Western blotting for PML, F-IE1 and-actin. Asterisks indicate the residual monoSUMOylated PML species. (D) PML-kd HFFs stably overexpressing F-PML I were either mock infected or infected with AD169 at an MOI of 3. At 24 h postinfection the cells were lysed under denaturing conditions in the presence of 20 mM NEM and subjected to immunoprecipitation using anti-FLAG antibody or mouse IgG Fc fragment as a control. Lysates (left panel) and immunoprecipitates (IP) (right panel) were analyzed by Western blotting to detect SUMO2/3 and F-PML I. Lysates were additionally analyzed to detect IE1 as an infection control and-actin as a loading control.on November 7, 2019 by guest
http://jvi.asm.org/
[image:4.585.42.397.74.389.2](marked by a diamond). Interestingly, PML K490-SUMO migrates approximately 15 kDa
higher than the unmodified PML and seems to be exclusively monoSUMOylated. Thus,
we speculated that PML K490-SUMO represents the monoSUMOylated species that is
unaffected by IE1. To prove this hypothesis, we cotransfected an IE1 expression plasmid
FIG 2IE1 specifically affects the SUMOylation of PML K160. (A and B) HEK293T cells were transfected with expression plasmids encoding myc-tagged PML (M-PML) variants (A) or cotransfected with expression plasmids encoding M-PML variants and IE1 (B) as indicated. At 48 h posttransfection, the cells were harvested and analyzed by Western blotting for the detection of M-PML, IE1, and-actin. Asterisks indicate the residual monoSUMOylated PML species referring to K490 SUMOylation. Circles and diamonds indicate the disappearing SUMOylated PML species referring to K160 and K65/160 SUMOylation, respectively. (C to F) HFFs stably expressing F-PML VI WT (C), F-PML VI K160R (D), F-PML VI K490R (E), or F-PML VI 3KR (F) were either mock infected or infected with AD169 at an MOI of 3. At the indicated time points postinfection, cells were harvested and analyzed by Western blotting for F-PML, IE1, and-actin.
HCMV IE1 Inhibits PMLDe NovoSUMOylation Journal of Virology
on November 7, 2019 by guest
http://jvi.asm.org/
[image:5.585.44.420.71.604.2]together with plasmids for PML WT, K160R, K490R, and 3KR. Strikingly, the expression
of IE1 led to a complete loss of PML K160-SUMO as well as PML K65/160-SUMO, while
the SUMOylation of PML K490 seemed to be resistant against IE1 action (Fig. 2B). In
order to confirm these data in the context of infection, we generated HFF cells stably
overexpressing FLAG-tagged versions of PML VI WT, K160R, K490R, and 3KR. Under
these conditions, we surprisingly observed a residual modified band (arrowhead) in
HFFs expressing PML 3KR (Fig. 2F, lane 1). This band probably arises from one further
diSUMOylated or two monoSUMOylated SUMO sites, as it migrates approximately 30
kDa higher than unmodified PML. Interestingly, this SUMOylation was also unaffected
by IE1, since the modified band did not disappear during AD169 infection (Fig. 2C, D,
E, and F, lanes 6). In addition to this uncharacterized band, only K490 SUMOylation
remained during AD169 infection, thereby demonstrating that this SUMO modification
represents the resistant monoSUMOylation during infection (Fig. 2C and D, lanes 6).
Taken together, we provide evidence that IE1 selectively affects SUMOylation at
K65/K160 of PML in order to disrupt NB integrity, whereas SUMOylation of K490, and
probably SUMOylation of one or two additional sites, remains unaffected.
IE1 prevents arsenic trioxide-mediated PML degradation.
Lysine 160 is thought
to be the main conjugation site for polySUMO chains during the process of arsenic
trioxide (As
2O
3)-induced degradation of PML (6). These polySUMO chains are targeted
by the cellular STUbL RNF4, which mediates ubiquitination and subsequent
protea-somal degradation (21, 41). Having shown that lysine 160 is selectively affected by
HCMV IE1, the question arose whether IE1 can disturb this As
2O
3-mediated degradation
of PML. To clarify this issue, we utilized HFF cells with doxycycline-inducible expression
of wild-type IE1 or truncated IE1 1–382 (IE1
CORE), which was described to be sufficient
for loss of PML SUMOylation (30, 42). As illustrated in Fig. 3A and B, lanes 1 to 6, As
2O
3provokes the hyperSUMOylation of endogenous nuclear PML isoforms, starting at 0.5 h
after treatment and finally resulting in the degradation of PML at 24 h. As previously
described, this degradation of PML could be reversed by addition of the proteasome
inhibitor MG132 (Fig. 3A and B, lanes 7) (6). In agreement with the data shown in Fig.
1, PML was found to be partially deSUMOylated after doxycycline-induced expression
of wild-type IE1 (Fig. 3A, lane 8) as well as IE1
CORE(Fig. 3B, lane 8). Interestingly, IE1 and
IE1
COREnot only caused the loss of physiological PML SUMOylation but also prevented
As
2O
3-mediated hyperSUMOylation (Fig. 3A and B, lanes 9, 10, 11, and 12). Strikingly,
this activity of IE1 blocked the degradation of PML (Fig. 3A and B, compare lanes 13 and
6). These findings further strengthen the assumption that IE1 selectively affects the
SUMOylation, and also the hyperSUMOylation, of PML lysine 160, which is a crucial
residue during As
2O
3-mediated degradation of PML.
FIG 3IE1 prevents arsenic trioxide-mediated PML degradation. (A and B) HFF cells with doxycycline-inducible expression of F-IE1 (A) or F-IE1CORE(B) were either not induced or induced with 0.5g/ml doxycycline for 16 h, followed by As2O3(1M) and simultaneous MG132 (10M) treatment when indicated. Cells were harvested at the indicated time points and analyzed by Western blotting for PML, F-IE1, and-actin.
on November 7, 2019 by guest
http://jvi.asm.org/
[image:6.585.45.544.72.244.2]A SUMO mutant resistant to SUMO proteases is stably conjugated to PML.
The
SUMOylation and deSUMOylation of PML is a very dynamic and reversible process due
to the presence of SUMO E3 ligases on the one hand and SUMO proteases on the other
hand (20). Unfortunately, these dynamics impede the ability to discriminate whether
IE1 mediates proteolytic PML deSUMOylation or inhibition of PML
de novo
SUMOyla-tion. To overcome these dynamics, we generated a nondeconjugatable mutant of
SUMO2 by mutagenizing SUMO residue 90 from glutamine to proline (43). Since SUMO
proteases are necessary not only for deconjugation of SUMO but also for processing
and thereby conjugation, we utilized an already processed version of SUMO2, termed
SUMO2 Q90P. First, we tested whether SUMO2 Q90P is conjugated to PML as efficiently
as SUMO2 WT. Therefore, we transiently transfected HEK293T cells with expression
vectors for PML VI and SUMO2 WT or SUMO2 Q90P and harvested the cells at 48 h
posttransfection by immediately lysing the cells under strictly denaturing conditions. As
shown in Fig. 4A, both SUMO2 WT and SUMO2 Q90P were conjugated to PML to
approximately the same extent. Next, we set out to examine whether SUMO2 Q90P is
more stably conjugated to PML than SUMO2 WT due to its resistance to SUMO
proteases. For this purpose, we again transfected HEK293T cells with aforementioned
constructs and lysed the cells using a nondenaturing NP-40 lysis buffer with or without
the cysteine protease inhibitor
N
-ethylmaleimide (NEM). It is known that rapid
de-SUMOylation of proteins occurs under nondenaturing lysis conditions, which is a
consequence of the constitutive activity of endogenous SUMO proteases (44). By
addition of NEM, SUMOylation can be retained to a certain degree. As expected, no
SUMO2 WT conjugates of PML could be detected in the absence of NEM (Fig. 4B, lane
5). However, the addition of NEM to the nondenaturing lysis buffer was sufficient to
stabilize the SUMOylation of PML mediated by SUMO2 WT (Fig. 4C, lane 5). In contrast,
the SUMO2 Q90P mutant was found to be conjugated to PML in both the absence and
presence of NEM, thereby confirming a stable conjugation despite the presence of
SUMO proteases (Fig. 4B and C, lanes 6). Taking these findings together, due to its
resistance to SUMO protease cleavage, SUMO2 Q90P could be identified as a tool to
overcome the SUMOylation dynamics and thus to discriminate between proteolytic
PML deSUMOylation and inhibition of PML
de novo
SUMOylation.
FIG 4A SUMO mutant resistant to SUMO proteases is stably conjugated to PML. HEK293T cells were cotransfected with expression plasmids encoding M-PML and F-SUMO2 WT or F-SUMO2 Q90P as indicated. At 48 h posttransfection, the cells were lysed by immediately boiling the cell pellet in SDS loading buffer (A) or by incubating the cells with NP-40 buffer without NEM (B) or with 30 mM NEM (C). The lysates were analyzed by Western blotting for the detection of M-PML, -actin, F-SUMO2 WT, and F-SUMO2 Q90P. Densitometric analysis of the F-SUMO2 signals in panel A, lanes 5 and 6, revealed relative intensities of 100% and 121%, respectively.
HCMV IE1 Inhibits PMLDe NovoSUMOylation Journal of Virology
on November 7, 2019 by guest
http://jvi.asm.org/
[image:7.585.43.545.70.294.2]IE1 prevents PML
de novo
SUMOylation
in vivo
.
Next, we set out to develop a
cell-based system that, by using the SUMO protease-resistant SUMO2 Q90P mutant,
allows the discrimination between IE1-mediated proteolytic deSUMOylation of PML
and inhibition of PML
de novo
SUMOylation. For this, we generated HEK293 cells with
doxycycline-inducible expression of IE1 by lentiviral transduction. Examination of the
IE1 expression kinetics revealed increasing protein levels until 24 h after doxycycline
stimulation but no further accumulation at 30 h (Fig. 5A). As our own observations and
earlier studies indicate that a high expression level of IE1 is crucial for its effect on PML
SUMOylation, we selected an induction time of 24 h for further experiments (29).
Immunofluorescence analysis at this time point disclosed a high percentage of positive
cells (
⬎
85%) (Fig. 5B). Next, we combined this inducible system and the
nondeconju-gatable SUMO mutant in an assay that should have the following outcomes depending
on the mode of IE1 action. If IE1 inhibits PML
de novo
SUMOylation, expression of IE1
after transfection of a plasmid for SUMO2 Q90P should not affect the SUMOylation
pattern of PML, whereas expression of IE1 before SUMO2 Q90P transfection should
result in a loss of PML SUMOylation. Alternatively, if IE1 acts in a proteolytic way either
by acting as or recruiting a SUMO protease, the time point of IE1 expression should not
have major effects on the PML SUMOylation pattern. Intriguingly, a clear difference in
the SUMOylation state of PML was observed in the two experimental settings: IE1
expression before transfection of plasmids for PML and SUMO2 Q90P induced an
almost complete loss of PML SUMOylation (Fig. 5C and E, lanes 6 and 7), while PML was
still heavily SUMOylated when the expression of IE1 was started after transfection of
plasmids for PML and SUMO2 Q90P (Fig. 5D and F, lanes 6 and 7). In a control
experiment with SUMO2 WT, we observed a depletion of SUMOylated PML irrespective
of whether IE1 was expressed before or after transfection (Fig. 5G and H). These results
demonstrate that the reversible, dynamic nature of the SUMOylation system strongly
influences the experimental outcome and underline the importance of utilizing a
nondeconjugatable SUMO mutant. Furthermore, the observation that IE1 is able to
prevent the modification of PML with this SUMO2 Q90P mutant strongly suggests that
IE1 disrupts NBs via the inhibition of PML
de novo
SUMOylation.
IE1 prevents PML
de novo
SUMOylation
in vitro
.
In order to confirm the data that
we obtained in HEK293 cells using an alternative experimental system, we produced
prokaryotically expressed and purified proteins for subsequent
in vitro
SUMOylation
assays, in which the mixtures contain all components necessary and sufficient for
protein SUMOylation (45). As these systems are devoid of SUMO proteases as well as
SUMO E3 ligases present in eukaryotic extracts, they provided a further possibility to
overcome the rapid SUMOylation dynamics and, in addition, to determine whether IE1
alone is sufficient for the loss of PML SUMOylation. In order to achieve a stable complex
of purified PML (GST-PML VI) and IE1 (His-IE1
CORE), we utilized a system that contains
two plasmids with different antibiotic resistance genes and origins of replication (ori),
thereby allowing simultaneous expression of the two proteins in one prokaryotic cell.
The IE1/PML protein complex was purified via binding of His-IE1
COREto an Ni-Sepharose
matrix to ensure that each PML molecule was occupied by IE1
CORE. The specific
interaction between IE1
COREand PML VI was confirmed by glutathione
S
-transferase
(GST) pulldown analysis (Fig. 6A). Initially, we tested the ability of GST-PML VI, which
was purified using glutathione Sepharose, to become
in vitro
SUMOylated. Notably,
while a monoSUMOylated form of PML was observed after 1 h of incubation with the
SUMOylation kit (Fig. 6B, lane 2), polySUMOylation of PML occurred only after a longer
incubation time of 4 h (Fig. 6B, lane 3). Comparative analysis of the SUMOylation of PML
alone with SUMOylation of the PML/IE1
COREcomplex after a 4-h
in vitro
reaction
revealed a drastic reduction of SUMO-modified PML variants in the presence of IE1
CORE:
polySUMOylation was completely abolished, while monoSUMOylation remained to a
minor extent (Fig. 6C, compare lanes 2 and 4). In contrast, the addition of IE1
COREto
preSUMOylated PML did not result in any reduction of PML SUMOylation, thereby
confirming the
in vivo
results (Fig. 6D, lanes 2 and 3). Altogether, our data provide direct
on November 7, 2019 by guest
http://jvi.asm.org/
FIG 5IE1 prevents PMLde novoSUMOylationin vivo. (A and B) HEK293-IE1 cells were not induced or induced with doxycycline (0.5g/ml). (A) At the indicated time points, the cells were harvested and analyzed by Western blotting for IE1 and-actin. (B) At 24 h post induction, immunodetection of IE1 was conducted by using an anti-IE1 antibody. (C) Timeline for the experiments for panels E and G. (D) Timeline for the experiments for panels F and H. (E to H) HEK293-IE1 cells were cotransfected with expression plasmids encoding M-PML and F-SUMO2 WT or F-SUMO2 Q90P as indicated. IE1 expression was induced with doxycycline (0.5g/ml). Cells were harvested after a total time of 72 h and analyzed by Western blotting for the detection of M-PML, IE1, -actin, F-SUMO2 WT, and F-SUMO2 Q90P. Densitometric analysis of the F-SUMO2-Q90P signals in panel E, lanes 6 and 7, revealed relative intensities of 100% and 31%, while the signal intensities in panel F, lanes 6 and 7, were 100% and 92%, respectively. The signal intensities for F-SUMO2 WT in panel G, lanes 6 and 7, were 100% and 22%, respectively. For panel H, lanes 6 and 7, the signal intensities were 100% and 31%, respectively.
HCMV IE1 Inhibits PMLDe NovoSUMOylation Journal of Virology
on November 7, 2019 by guest
http://jvi.asm.org/
[image:9.585.41.395.67.638.2]evidence that IE1 alone is able to prevent
de novo
SUMOylation of PML but does not
abrogate the SUMOylation of preSUMOylated PML.
IE1 interacts with the dimeric PML coiled coil without interfering with PML
dimerization.
The coiled-coil domain of TRIM family proteins has been described to be
involved in homointeractions which are necessary for the biochemical activity of TRIM
proteins (46, 47). For PML, the coiled-coil-mediated dimerization is thought to be a
prerequisite for its SUMO E3 ligase activity and for PML autoSUMOylation. Since IE1
COREwas shown to interact with the coiled-coil domain of PML, we asked whether IE1 affects
PML SUMOylation by disrupting the coiled-coil-mediated self-interaction (30). For this
purpose, we performed competitive coimmunoprecipitation experiments using two
differentially tagged versions of PML and increasing amounts of IE1
CORE. As shown in
Fig. 7A, we could not detect any decrease in PML homodimerization upon coexpression
of IE1
CORE. The same results were obtained when utilizing the isolated PML coiled-coil
domains in combination with IE1
COREor with full-length IE1 (Fig. 7B). These data
indicate that IE1 directly interacts with the dimeric coiled-coil domain of PML to prevent
PML
de novo
SUMOylation, without interfering with PML dimerization.
DISCUSSION
It is well known that HCMV IE1 induces the loss of PML SUMOylation, thereby
antagonizing NB-mediated intrinsic immunity (26–30, 32, 34–36, 48). However, the
underlying mechanism remained elusive for the last decades. IE1-mediated loss of PML
SUMOylation could be achieved either by proteolytic deSUMOylation or by inhibition of
de novo
SUMOylation. In the present study, we report for the first time that IE1 is able
to prevent the
de novo
SUMOylation of PML. By using a nondeconjugatable SUMO
mutant, described by Békés and colleagues, we were able to overcome the rapid and
reversible dynamics of PML SUMOylation (43). These dynamics are caused by the
FIG 6IE1 prevents PMLde novoSUMOylationin vitro. (A) GST pulldown analysis of prokaryotically expressed GST, GST-PML, and His-IE1CORE. Bound proteins were eluted by boiling the glutathione-Sepharose beads in SDS loading buffer. Eluted proteins and input were analyzed by Western blotting for the detection of GST fusions and His-IE1CORE. (B to D) Bacterially expressed and purified proteins were incubated together with anin vitroSUMOylation kit containing E1 (SAE1/SAE2), Ubc9, ATP, and SUMO3 for the indicated times (B) or for 4 h (C and D) as described in Materials and Methods. (B and C) The reaction mixes were analyzed by Western blotting using anti-PML antibody (B) or both anti-PML and anti-His antibodies (C). (D) After terminating the reaction by adding 10 mM each EDTA and DTT, the mix was again incubated for 1 h in the presence or absence of bacterially expressed and purified His-IE1CORE.The reaction mixes were analyzed by Western blotting using anti-PML and anti-His antibodies.
on November 7, 2019 by guest
http://jvi.asm.org/
[image:10.585.72.338.70.310.2]presence of both SUMO conjugating enzymes and deconjugating SUMO proteases
inside PML NBs, which are considered to represent nuclear “SUMO hot spots” (19). An
experimental approach which combined this SUMO mutant with inducible expression
of IE1 revealed that IE1 affects the PML SUMOylation status only when it is present
before SUMOylation takes place (Fig. 5C and E). In contrast, when IE1 is expressed after
PML SUMOylation, no loss of PML SUMOylation is detectable (Fig. 5D and F).
Further-more, this effect seems to be independent of additional cellular proteins, as we
observed the same outcome
in vitro
by using prokaryotically expressed and purified
proteins (Fig. 6). Notably, we obtained further evidence that formation of a tight
complex between IE1 and PML is critical for IE1 action, since the inhibitory effect on
PML SUMOylation as detected by
in vitro
SUMOylation was evident only when we
utilized a purified IE1/PML complex and not when using separately purified proteins
(data not shown). This may explain the contradicting results reported by Kang and
colleagues, who pursued the latter strategy and did not observe effects of IE1 on PML
SUMOylation by
in vitro
SUMOylation and therefore proposed that IE1 neither interferes
with PML SUMOylation nor directly causes PML deSUMOylation by acting as a SUMO
protease itself (48). The latter was actually confirmed by our experiments: the addition
of purified IE1 to preSUMOylated PML did not affect the SUMOylation status of PML
(Fig. 6D). This finding is additionally supported by structural analysis of IE1
CORE, which
provides no evidence for the presence of a potential active site with hydrolase activity
(30).
The observation that IE1 exclusively affects PML polySUMOylation has already been
made in earlier studies (Fig. 1) (29, 40). However, by utilizing a variety of PML
SUMOylation site mutants, we now provide evidence that the residual
monoSUMO-ylated form can be assigned to a PML, which is SUMO modified on lysine 490 (Fig. 2C
to F). Three major SUMOylation sites have been reported for PML: K65, K160, and K490
(10). When we mutated these three lysine residues to arginine, followed by expression
of the triple mutant (3KR) in HEK293T cells, SUMOylation of PML was completely
abolished (Fig. 2A, lane 8). Surprisingly, the same mutant, stably expressed in HFF cells,
revealed one residual slower-migrating band which might correspond either to a PML
that is monoSUMOylated on two additional SUMO sites or to a diSUMOylated PML
FIG 7IE1 does not act by interfering with PML dimerization. (A) HEK293T cells were cotransfected with expression plasmids encoding F-PML VI and M-PML VI with increasing amounts of M-IE1COREconstruct. (B) HEK293T cells were cotransfected with expression plasmids encoding the F-PML coiled-coil domain (CC) and GFP-PML CC with increasing amounts of M-IE1COREor M-IE1 FL constructs. Upper panels, Western blot detection of M-PML VI (A) or GFP-PML CC (B) after immunoprecipitation with an anti-FLAG antibody (IP). Lower three panels, detection of M-PML VI (A) or GFP-PML CC (B) and F-PML VI (A) or F-PML CC (B) and M-IE1 variants (as indicated) in cell lysates before precipitation (input).
HCMV IE1 Inhibits PMLDe NovoSUMOylation Journal of Virology
on November 7, 2019 by guest
http://jvi.asm.org/
[image:11.585.44.463.71.287.2](Fig. 2F). The same modified band was detected in PML WT, K160R, and K490R cells after
mock as well as HCMV infection, thereby suggesting the existence of at least one
additional SUMO site that is not affected by IE1 (Fig. 2C to E). A similar phenomenon
was observed by Cuchet-Lourenço and colleagues when expressing the PML I 3KR
mutant in HFFs. However, the mutation of one further minor SUMOylation site (K616)
eliminated all modified bands (49). PML VI lacks the K616 SUMO site; however, four
additional minor SUMOylation sites (K226, K380, K497, and K400) were reported for this
isoform, which might be SUMOylated under specific conditions in a cell type-specific
manner (50, 51). In summary, when taking into account previous results demonstrating
that K65 is modified only when K160 is SUMOylated, IE1 appears to selectively inhibit
K160 SUMOylation in order to induce PML NB disruption (21). This assumption is further
strengthened by the finding that IE1 and also its central domain IE1
COREis able to
prevent As
2O
3-mediated hyperSUMOylation of PML (Fig. 3), which takes place on K160
(6). Additionally, this finding highlights the capability of IE1 to prevent not only
physiological polySUMOylation but also hyperSUMOylation under cellular stress
con-ditions.
Having demonstrated that IE1 exclusively affects the on-rate of PML SUMO
modi-fication without requiring additional proteins, we can now narrow down the possible
mechanistic scenarios. IE1 is able to prevent PML SUMOylation
in vitro
, which limits the
number of IE1 target proteins to the E1-activating enzyme, a heterodimer of SAE1 and
SAE2, the E2-conjugating enzyme Ubc9, the SUMO protein, and PML itself. Surprisingly,
despite the fact that IE1 itself is a SUMO substrate, interactions between IE1 and Ubc9
or SUMO could not be demonstrated, making it rather unlikely that IE1 targets these
factors (32, 33). To our knowledge, an interaction with the E1-activating enzyme has
also not been observed for IE1, as is true for the avian adenoviral protein Gam1, which
is able to induce a global loss of SUMOylation (52). In contrast to Gam1, IE1 acts not by
reducing global SUMO conjugation but by specific interference with the SUMOylation
of PML and Sp100 (53). Consequently, the hypothesis arose that there must be a more
targeted action of IE1, presumably by acting on PML itself. This is strengthened by the
fact that a direct binding of IE1 to PML is a prerequisite for IE1 action (29, 34). Due to
the following reasons, we would further suggest that the integrity of the central
globular domain of IE1, termed IE1
CORE, is needed for the loss of PML SUMOylation.
First, IE1
COREis described as an elongated molecule consisting of a single contiguous
domain with conserved residues distributed evenly over the entire surface (30). Second,
IE1
COREis hypersensitive to mutational interventions (36–38). Third, further attempts to
narrow down the binding region by mutations and deletions failed. We strongly favor
a model whereby the IE1 globular core domain targets the PML coiled-coil domain via
coiled-coil interactions, thereby preventing the
de novo
SUMOylation of PML.
An interesting mechanism was described for influenza virus protein NS1, which
prevents TRIM25 oligomerization, thereby abrogating its ubiquitin E3 ligase activity
(54). However, the idea that IE1 interferes with SUMO E3 ligase activity by preventing
PML dimerization was rejected, as IE1 does not decrease homointeractions of PML (Fig.
7). Thus, one possible scenario is that IE1 blocks SUMO E3 ligase activity of PML by a
mechanism independent of PML dimerization. By mutating key zinc-chelating residues,
Chu and Yang demonstrated that RING and B-box domains are likely required for the
SUMO E3 ligase activity of PML (16). Interestingly, when expressing these RING and
B-box mutants, PML was found only monoSUMOylated, which led us to speculate that
the SUMO E3 ligase activity of PML is necessary for its own polySUMOylation (16). The
assumption that IE1 selectively affects the SUMOylation of PML K160, which is nearby
the IE1 binding site on PML, raised another idea. IE1 may also act by simply covering
the K160 SUMO site or by blocking the accessibility of PML for the components of the
cellular SUMOylation machinery, thereby preventing SUMOylation by a steric hindrance
mechanism. This idea is supported by the fact that IE1 does oligomerize, thereby
potentially providing an enlarged surface (30). In addition to this, the assumption that
the interaction with PML is a prerequisite for IE1 action bears similarity to the case for
on November 7, 2019 by guest
http://jvi.asm.org/
the adenoviral E4-ORF3 protein, which is suggested to inactivate PML via tight binding
using a multivalent matrix built by extensive multimerization (55).
Taken together, the results presented in this study clearly demonstrate that IE1
selectively abrogates the
de novo
SUMOylation of PML at specific lysine residues, which
may be due either to interference with the E3 ligase function of PML or to steric
hindrance which blocks access to specific SUMOylation sites. Further attempts to
understand the molecular architecture of the IE1-PML protein complex by
cocrystalli-zation will help to distinguish between these two possibilities.
MATERIALS AND METHODS
[image:13.585.41.376.82.281.2]Oligonucleotides and plasmid constructs.The oligonucleotide primers used for this study were purchased from Biomers GmbH (Ulm, Germany) and are listed in Table 1. Lentivirus vectors expressing FLAG-tagged versions of PML isoforms I to VI were kindly provided by Roger Everett (Glasgow, UK) (56). All cDNAs encoding the PML isoforms have been made resistant to the anti-PML short hairpin RNA (shRNA) siPML2; the altered sequence is 5=-AGATGCTGCAGTTAGCAAG-3=(31). For the generation of Myc-PML VI as well as FLAG-PML VI and FLAG-PML 228 –399 (CC), PML isoform VI was amplified from pAS-PML (a gift from Gerd Maul) and inserted into pHM1580 Myc) or pHM971 (pcDNA3.1-FLAG) as described elsewhere (30, 31, 57, 58). The construct for transient expression of green fluorescent protein (GFP)-PML 228 –399 (CC) was generated by amplification using the above-mentioned pcDNA3.1-Myc PML VI and primers 5=PMLaa228_SalI and 3=PMLaa399_BamHI, followed by insertion via SalI and BamHI into pEGFP C1 (Clontech, Mountain View, CA, USA). The constructs for transient expression of IE1 (pHM494), Myc-IE1, and Myc-IE1COREwere generated as described elsewhere (30, 57). Derivatives of Myc-PML isoform VI with point mutations in the SUMO modification sites (SUMOmut; K160R, K490R, and 3KR) were constructed by site-directed mutagenesis using a QuikChange site-directed mutagenesis kit as instructed by the manufacturer (Stratagene, La Jolla, CA, USA). The constructs for the stable overexpres-sion of PML VI WT, PML VI K160R, PML VI K490R, and PML VI 3KR were generated by amplification using the above-mentioned PML VI constructs and primers 5=PML_SalI and 3=EcoRI_PML6, followed by insertion via SalI and EcoRI into a modified pLKO-based lentiviral vector. The modified pLKO-based lentiviral vector was generated by exchanging the CMV major immediate early promoter (MIEP) via NdeI/PacI for an EF1␣ promoter amplified with primers 5=EF1alpha-NdeI and 3=EF1alpha-PacI. For inducible expression of IE1, the entry vector pDONR221-IE1 was introduced into the pINDUCER20 destination vector using the Invitrogen Gateway recombination technology (Thermo Fisher Scientific, Waltham, MA, USA) (53, 59). The FLAG-SUMO2 Q90P mutant was generated by using the GeneArt site-directed mutagenesis kit as instructed by the manufacturer (Thermo Fisher Scientific, Waltham, MA, USA) with pcDNA3-FLAG-SUMO2 as the template (57). The processed FLAG-SUMO2 Q90P was generated by amplification using the mutagenized construct and primers 5=SUMO2_BamHI and 3=Xba_SUMO2_GG, followed by insertion via BamHI and XbaI into pHM971 (pcDNA3.1-FLAG) (57). For bacterial expression of GST-tagged full-length PML, the synthetic, codon-optimized cDNA of PML VI was synthesized by the GeneArt gene synthesis service (Thermo Fisher Scientific, Waltham, MA, USA) and was cloned into the prokaryotic expression vector pGEX-6P-1 (GE Healthcare, Munich, Germany) via BamHI/XhoI. His-IE1 14 –382 was obtained by subcloning a synthetic codon-optimized version of the IE1 cDNA sequence purchased from the GeneArt gene synthesis service into pCOLADuet-1 (Novagen, Merck Millipore, Billerica. MA, USA) via BamHI and NotI. The integrity of all newly generated plasmids was confirmed by automated DNA sequence analysis.
TABLE 1Oligonucleotides used for plasmid construction
Purpose and oligonucleotide Sequence
Mutagenesis
5=PML-SUMO65-mut CCAGGCGGAAGCCAGGTGCCCGAAGCTGC
3=PML-SUMO65-mut GCAGCTTCGGGCACCTGGCTTCCGCCTGG
5=PML-SUMO160-mut CCAGTGGTTCCTCAGGCACGAGGCCCGGC
3=PML-SUMO160-mut GCCGGGCCTCGTGCCTGAGGAACCACTGG
5=PML-SUMO490-mut CCAGGAAGGTCATCAGGATGGAGTCTGAGGAGG
3=PML-SUMO490-mut CCTCCTCAGACTCCATCCTGATGACCTTCCTGG
5=SUMO2_Q90Pmut GATGTGTTCCAACAGCCAACGGGAGGTGTCTAC
3=SUMO2_Q90Pmut GTAGACACCTCCCGTTGGCTGTTGGAACACATC
Amplification
5=PMLaa228_SalI CATAGTCGACGACATCAGCGCAGAGATCCA
3=PMLaa399_BamHI CATAGGATCCTCATGGATACAGCTGCATCTTTC
5=PML-SalI CATAGTCGACATGGAGCCTGCACCCGCCCG
3=EcoRI_PML6 CATAGAATTCTCACCACAACGCGTTCCTCTC
5=EF1alpha-NdeI CATACATATGCCCGTCAGTGGGCAGAG
3=EF1alpha-PacI CATATTAATTAATATTAGTACCAAGCTAATTC
5=SUMO2_BamHI CATAGGATCCGCCGACGAAAAGCCCAAGGA
3=Xba_SUMO2_GG CATATCTAGATCAACCTCCCGTTGGCTGTTG
HCMV IE1 Inhibits PMLDe NovoSUMOylation Journal of Virology
on November 7, 2019 by guest
http://jvi.asm.org/
Cells and viruses. HEK293 and HEK293T cells were cultivated in Dulbecco’s minimal essential medium (DMEM) (Gibco, Thermo Fisher Scientific, Waltham, MA, USA) containing 10% fetal calf serum (FCS). HEK293 cells with inducible overexpression of IE1 (HEK293-IE1) were cultured in DMEM supple-mented with 10% tetracycline-free fetal bovine serum (Clontech, Palo Alto, CA, USA) and 500g/ml Geneticin. Primary human foreskin fibroblasts (HFFs) were prepared from human foreskin tissue and were maintained in Eagle’s minimal essential medium (MEM) (Gibco, Thermo Fisher Scientific, Waltham, MA, USA) supplemented with 7.5% fetal calf serum. HFFs stably overexpressing F-PML VI constructs (WT, K160R, K490R, and 3KR) were maintained in MEM supplemented with 7.5% fetal calf serum and 500 g/ml Geneticin. HFFs with a small interfering RNA-mediated knockdown of PML (siPML2 cells) and stable overexpression of F-PML I were maintained in MEM supplemented with 10% fetal calf serum and 5g/ml puromycin as well as 500g/ml Geneticin. HFFs with inducible expression of IE1 or IE1 1–382 were cultured in MEM supplemented with 10% tetracycline-free fetal bovine serum (Clontech, Palo Alto, CA, USA), 5g/ml puromycin, and 500g/ml Geneticin. Infection experiments were performed with the HCMV laboratory strain AD169 at specified multiplicities of infection (MOIs). Viral stocks were titrated via IE1p72 fluorescence (60). For this purpose, HFFs were infected with various dilutions of virus stocks. After 24 h of incubation, cells were fixed and stained with monoclonal antibody (MAb) p63-27, directed against IE1p72 (61). Subsequently, the number of IE1-positive cells was determined and used to calculate viral titers, expressed as IE protein-forming units (IEU).
Lentivirus transduction and selection of stably transduced cells.HFF PML-kd cells stably ex-pressing FLAG-PML I were generated via lentiviral transduction using the ViraPower lentiviral system according to the manufacturer’s protocol (Invitrogen, Karlsruhe, Germany) and were selected in Dulbec-co’s minimal essential medium supplemented with 10% fetal calf serum plus 500g/ml Geneticin (31). For the generation of HFF cells stably expressing FLAG-tagged PML VI WT, K160R, K490R, and 3KR or HEK293 cells stably expressing inducible IE1, replication-deficient lentiviruses were generated using pLKO-based or pINDUCER20-based expression vectors. For this purpose, HEK293T cells seeded in 10-cm dishes (5⫻106cells) were cotransfected with a pLKO vector encoding FLAG-tagged PML versions or pINDUCER20 vector encoding IE1 together with packaging plasmids pLP1, pLP2, and pLP/VSV-G using the Lipofectamine 2000 reagent (Invitrogen, Karlsruhe, Germany). Viral supernatants were harvested 48 h after transfection, cleared by centrifugation, filtered, and stored at⫺80°C. HFFs and HEK293 cells were incubated for 24 h with lentiviral supernatants in the presence of 7.5g/ml Polybrene (Sigma-Aldrich, St. Louis, MO, USA). Stably transduced HFF and HEK293 cell populations were selected by adding 500 g/ml Geneticin to the cell culture medium. HFFs with inducible expression of IE1 or IE1 1–382 were generated as described elsewhere (42).
Antibodies. Endogenous PML was detected by rabbit polyclonal antibodies A301-167A (Bethyl Laboratories, Montgomery, TX, USA) and A301-168A (Bethyl Laboratories, Montgomery, TX, USA) in combination. Mouse monoclonal antibody p63-27, which recognizes IE1, has been described elsewhere (61). Myc-tagged IE1 and Myc-tagged IE1COREwere detected by MAb-Myc (1-9E10.2; ATCC). MAb-FLAG 1804 (Sigma-Aldrich, St. Louis, MO, USA) was utilized for the detection of FLAG-IE1 and FLAG-IE1CORE. For the detection of Myc-, FLAG-, and GFP-tagged versions of PML the mouse monoclonal antibodies MAb-Myc (1-9E10.2; ATCC), MAb-FLAG 1804 (Sigma-Aldrich, St. Louis, MO, USA), and MAb GFP (Roche, Penzberg, Germany) were utilized. A mouse IgG Fc fragment from Dianova (Hamburg, Germany) served as control antibody. Rabbit monoclonal antibody EPR4602 (Abcam, Cambridge, UK) was utilized for the detection of endogenous SUMO2/3. Mouse monoclonal antibody AC-15, which recognizes beta-actin, was purchased from Sigma-Aldrich (St. Louis, MO, USA). Prokaryotically expressed GST-PML was detected by using rabbit polyclonal antibody H-238 (Santa Cruz Biotechnology, Santa Cruz, CA, USA) or mouse anti-GST monoclonal antibody BZL01642 (Biozol, Eching, Germany). Mouse monoclonal antibody HIS.H8 (Thermo Fisher Scientific, Waltham, MA, USA) against 6⫻His tag was utilized for the detection of prokaryotically expressed His-IE1CORE. Horseradish peroxidase-conjugated anti-mouse and anti-rabbit secondary antibodies for Western blot analysis were obtained from Dianova (Hamburg, Germany), while Alexa Fluor 555-conjugated secondary antibody for indirect immunofluorescence experiments was purchased from Molecular Probes (Karlsruhe, Germany).
Transfection and doxycycline induction. HEK293T and HEK293-IE1 cells were transfected by applying the standard calcium phosphate coprecipitation method. For this, 7 ⫻ 105 HEK293T or HEK293-IE1 cells were seeded into six-well dishes 1 day before transfection; 1.5 to 2g of plasmid DNA was used for each transfection reaction. At about 16 h later, the cells were washed two times with phosphate-buffered saline without calcium and magnesium (PBSo) and provided with fresh medium. At 48 h after transfection, cells were harvested for further analyses. The HEK293-IE1 cell line was transfected using the Lipofectamine 2000 reagent (Invitrogen, Karlsruhe, Germany), according to the instructions of the manufacturer, when transfection and doxycycline induction were combined. Cells (7⫻105) were seeded into six-well dishes and were grown until the next day. IE1 expression was induced either 24 h prior to transfection or 24 h posttransfection by the addition of 0.5 g/ml doxycycline. At 48 h posttransfection, cells were harvested or IE1 expression was induced for 24 h, depending on the experimental setting.
Immunoblotting.Lysates from infected HFF, transfected HEK293T, or transfected/induced HEK293-IE1 cells were prepared in sodium dodecyl sulfate (SDS)-Laemmli buffer and boiled for 10 min at 95°C (57). To analyze the stability of SUMO2 WT in comparison to SUMO2 Q90P conjugation, transfected HEK293T cells were lysed for 20 min at 4°C by addition of 800l NP-40 buffer (50 mM Tris-HCl [pH 8.0], 150 mM NaCl, 5 mM EDTA, 0.5% NP-40, 1 mM phenylmethylsulfonyl fluoride [PMSF], 2g each of aprotinin, leupeptin, and pepstatin per ml buffer) in the presence or absence of a 30 mM concentration of the cysteine modifierN-ethylmaleimide (NEM) (Sigma-Aldrich, St. Louis, MO, USA). After centrifugation,
on November 7, 2019 by guest
http://jvi.asm.org/
the supernatant was diluted in SDS-Laemmli buffer and boiled at 95°C for 10 min. Proteins were separated by SDS-polyacrylamide gel electrophoresis (SDS-PAGE) on 8% to 12.5% polyacrylamide gels and transferred onto nitrocellulose membranes (GE Healthcare, Munich, Germany), followed by chemi-luminescence detection according to the manufacturer’s protocol (ECL Western blotting detection kit; Amersham Pharmacia Europe, Freiburg, Germany).
Immunoprecipitation. HFF cells stably overexpressing FLAG-PML were infected with HCMV AD169 or mock infected. At specified time points postinfection, the cells were harvested and lysed for 5 min at 4°C using denaturing radioimmunoprecipitation assay (RIPA) buffer (50 mM Tris-HCl [pH 7.4], 150 mM NaCl, 1% NP-40, 0.1% SDS, 0.5% sodium deoxycholate, 1 mM PMSF, 2g each of aprotinin, leupeptin, and pepstatin per ml buffer) in the presence of 20 to 30 mM NEM. After centrifugation, the supernatant was incubated with the appropriate antibody coupled to protein A-Sepharose beads for 1.5 h at 4°C. The Sepharose beads were collected and washed three times in NP-40 buffer. Antigen-antibody complexes were recovered by boiling in SDS sample buffer and analyzed via Western blotting. Coimmunoprecipitation analysis using HEK293T cells was performed as described elsewhere (30).
Indirect immunofluorescence.For indirect immunofluorescence analysis, 3.5⫻105HEK293-IE1 cells were seeded on coverslips. One day after seeding, the expression of IE1 was induced by addition of 0.5 g/ml doxycycline. At 24 h postinduction, the cells were fixed with a 4% paraformaldehyde solution for 10 min at room temperature (RT) and then washed twice. Permeabilization of the cells was achieved by incubation with 0.2% Triton X-100 in PBSo on ice for 20 min. Cells were washed again with PBSo over a time period of 5 min and incubated with the monoclonal antibody p63-27 detecting IE1 diluted in PBSo–1% FCS for 30 min at 37°C. Excessive antibodies were removed by washing four times with PBSo, followed by incubation with the fluorescence-coupled secondary antibody rabbit anti-mouse 555 diluted in PBSo–1% FCS for 30 min at 37°C. The cells were mounted using the DAPI (4=,6= -diamidino-2-phenylindole)-containing Vectashield mounting medium (Alexis, Grünberg, Germany) and analyzed using a Leica TCS SP5 confocal microscope with the 543-nm laser line. The images were exported, processed with Adobe Photoshop CS5, and assembled using CorelDraw⫻5. To quantify the amount of IE1-expressing cells after induction, IE1-expressing cells were counted in three fields of view, and the mean value as well as the standard deviation was calculated.
Protein generation and purification.GST, GST-PML FL, His-IE1CORE, and the complexes consisting of GST-PML FL and His-IE1COREwere produced inEscherichia colistrain BL21(DE3). For this, LB media with appropriate antibiotics were inoculated with transformed BL21(DE3). GST fusion proteins and the complex were induced by treatment with 0.1 mM isopropyl--D-thiogalactoside (IPTG) for 16 h at 25°C. His-IE1COREwas induced by using 1 mM IPTG for 5 h at 30°C. Cell pellets were resuspended in the appropriate lysis buffer for either GST fusion proteins (1⫻PBSo, 2 mM PMSF, 1g/ml aprotinin, 1g/ml leupeptin, 1g/ml pepstatin, 2g/ml DNase, and 10 mM dithiothreitol [DTT]) or His fusion proteins (2⫻ PBSo, 250 mM NaCl, 2 mM PMSF, 1g/ml aprotinin, 1g/ml leupeptin, 1g/ml pepstatin, 2g/ml DNase, and 5 mM DTT), followed by sonication and centrifugation to separate the lysate from the cell debris. The fusion proteins were purified from the lysate by affinity chromatography using glutathione-Sepharose or Ni-glutathione-Sepharose according to the specifications of the manufacturer (GE Healthcare, Freiburg, Germany). Eluted proteins were analyzed by SDS-PAGE and Coomassie blue staining.
In vitroSUMOylation assays.In vitroSUMOylation reactions were performed at 37°C for 1 to 4 h in a 20-l volume containing prokaryotically expressed proteins along with a 50 nM concentration of the heterodimeric human E1-activating enzyme (SAE1/SAE2), a 62.5M concentration of the E2-conjugating enzyme UbcH9, 1 mM Mg-ATP, 25M SUMO3, and reaction buffer (500 mM HEPES [pH 8], 1,000 mM NaCl, 10 mM DTT) according to the manufacturer’s protocol (BostonBiochem, Cambridge, MA, USA). After termination of the reaction using SDS sample buffer containing beta-mercaptoethanol, the reaction products were fractionated by 8% SDS-PAGE for immunoblot detection of IE1COREand PML.
ACKNOWLEDGMENTS
We thank Roger Everett (Glasgow, UK) for providing expression vectors for PML
isoforms and Regina Müller (Erlangen, Germany) for excellent technical assistance.
REFERENCES
1. Ishov AM, Sotnikov AG, Negorev D, Vladimirova OV, Neff N, Kamitani T, Yeh ET, Strauss JF, III, Maul GG. 1999. PML is critical for ND10 formation and recruits the PML-interacting protein daxx to this nuclear structure when modified by SUMO-1. J Cell Biol 147:221–234. https://doi.org/ 10.1083/jcb.147.2.221.
2. Lallemand-Breitenbach V, de The H. 2010. PML nuclear bodies. Cold Spring Harb Perspect Biol 2:a000661. https://doi.org/10.1101/ cshperspect.a000661.
3. Tavalai N, Stamminger T. 2008. New insights into the role of the sub-nuclear structure ND10 for viral infection. Biochim Biophys Acta 1783: 2207–2221. https://doi.org/10.1016/j.bbamcr.2008.08.004.
4. Everett RD, Chelbi-Alix MK. 2007. PML and PML nuclear bodies: implica-tions in antiviral defence. Biochimie 89:819 – 830. https://doi.org/ 10.1016/j.biochi.2007.01.004.
5. Jensen K, Shiels C, Freemont PS. 2001. PML protein isoforms and the RBCC/TRIM motif. Oncogene 20:7223–7233. https://doi.org/10.1038/ sj.onc.1204765.
6. Lallemand-Breitenbach V, Zhu J, Puvion F, Koken M, Honore N, Doubeik-ovsky A, Duprez E, Pandolfi PP, Puvion E, Freemont P, de The H. 2001. Role of promyelocytic leukemia (PML) sumolation in nuclear body for-mation, 11S proteasome recruitment, and As2O3-induced PML or PML/ retinoic acid receptor alpha degradation. J Exp Med 193:1361–1371. https://doi.org/10.1084/jem.193.12.1361.
7. Zhong S, Muller S, Ronchetti S, Freemont PS, Dejean A, Pandolfi PP. 2000. Role of SUMO-1-modified PML in nuclear body formation. Blood 95: 2748 –2753.
8. Borden KL. 2008. Pondering the puzzle of PML (promyelocytic leukemia) nuclear bodies: can we fit the pieces together using an RNA regulon?
HCMV IE1 Inhibits PMLDe NovoSUMOylation Journal of Virology
on November 7, 2019 by guest
http://jvi.asm.org/
Biochim Biophys Acta 1783:2145–2154. https://doi.org/10.1016/ j.bbamcr.2008.06.005.
9. Liang YC, Lee CC, Yao YL, Lai CC, Schmitz ML, Yang WM. 2016. SUMO5, a novel poly-SUMO isoform, regulates PML nuclear bodies. Sci Rep 6:26509. https://doi.org/10.1038/srep26509.
10. Kamitani T, Kito K, Nguyen HP, Wada H, Fukuda-Kamitani T, Yeh ET. 1998. Identification of three major sentrinization sites in PML. J Biol Chem 273:26675–26682. https://doi.org/10.1074/jbc.273.41.26675.
11. Kamitani T, Nguyen HP, Kito K, Fukuda-Kamitani T, Yeh ET. 1998. Cova-lent modification of PML by the sentrin family of ubiquitin-like proteins. J Biol Chem 273:3117–3120. https://doi.org/10.1074/jbc.273.6.3117. 12. Duprez E, Saurin AJ, Desterro JM, Lallemand-Breitenbach V, Howe K,
Boddy MN, Solomon E, de The H, Hay RT, Freemont PS. 1999. SUMO-1 modification of the acute promyelocytic leukaemia protein PML: impli-cations for nuclear localisation. J Cell Sci 112:381–393.
13. Tatham MH, Jaffray E, Vaughan OA, Desterro JM, Botting CH, Naismith JH, Hay RT. 2001. Polymeric chains of SUMO-2 and SUMO-3 are conju-gated to protein substrates by SAE1/SAE2 and Ubc9. J Biol Chem 276:35368 –35374. https://doi.org/10.1074/jbc.M104214200.
14. Hay RT. 2005. SUMO: a history of modification. Mol Cell 18:1–12. https:// doi.org/10.1016/j.molcel.2005.03.012.
15. Quimby BB, Yong-Gonzalez V, Anan T, Strunnikov AV, Dasso M. 2006. The promyelocytic leukemia protein stimulates SUMO conjugation in yeast. Oncogene 25:2999 –3005. https://doi.org/10.1038/sj.onc.1209335. 16. Chu Y, Yang X. 2011. SUMO E3 ligase activity of TRIM proteins. Oncogene
30:1108 –1116. https://doi.org/10.1038/onc.2010.462.
17. Guo L, Giasson BI, Glavis-Bloom A, Brewer MD, Shorter J, Gitler AD, Yang X. 2014. A cellular system that degrades misfolded proteins and protects against neurodegeneration. Mol Cell 55:15–30. https://doi.org/10.1016/ j.molcel.2014.04.030.
18. Shen TH, Lin HK, Scaglioni PP, Yung TM, Pandolfi PP. 2006. The mech-anisms of PML-nuclear body formation. Mol Cell 24:331–339. https:// doi.org/10.1016/j.molcel.2006.09.013.
19. Van Damme E, Laukens K, Dang TH, Van OX. 2010. A manually curated network of the PML nuclear body interactome reveals an important role for PML-NBs in SUMOylation dynamics. Int J Biol Sci 6:51– 67. 20. Wilkinson KA, Henley JM. 2010. Mechanisms, regulation and
conse-quences of protein SUMOylation. Biochem J 428:133–145. https:// doi.org/10.1042/BJ20100158.
21. Lallemand-Breitenbach V, Jeanne M, Benhenda S, Nasr R, Lei M, Peres L, Zhou J, Zhu J, Raught B, de The H. 2008. Arsenic degrades PML or PML-RARalpha through a SUMO-triggered RNF4/ubiquitin-mediated pathway. Nat Cell Biol 10:547–555. https://doi.org/10.1038/ncb1717. 22. Wimmer P, Schreiner S, Dobner T. 2012. Human pathogens and the host
cell SUMOylation system. J Virol 86:642– 654. https://doi.org/10.1128/ JVI.06227-11.
23. Boutell C, Cuchet-Lourenco D, Vanni E, Orr A, Glass M, McFarlane S, Everett RD. 2011. A viral ubiquitin ligase has substrate preferential SUMO targeted ubiquitin ligase activity that counteracts intrinsic antiviral de-f e n c e . P L o S P a t h o g 7 : e 1 0 0 2 2 4 5 . h t t p s : / / d o i . o r g / 1 0 . 1 3 7 1 / journal.ppat.1002245.
24. Everett RD, Freemont P, Saitoh H, Dasso M, Orr A, Kathoria M, Parkinson J. 1998. The disruption of ND10 during herpes simplex virus infection correlates with the Vmw110- and proteasome-dependent loss of several PML isoforms. J Virol 72:6581– 6591.
25. Kelly C, van DR, Wilkinson GW. 1995. Disruption of PML-associated nuclear bodies during human cytomegalovirus infection. J Gen Virol 76:2887–2893. https://doi.org/10.1099/0022-1317-76-11-2887. 26. Korioth F, Maul GG, Plachter B, Stamminger T, Frey J. 1996. The nuclear
domain 10 (ND10) is disrupted by the human cytomegalovirus gene product IE1. Exp Cell Res 229:155–158. https://doi.org/10.1006/ excr.1996.0353.
27. Ahn JH, Hayward GS. 1997. The major immediate-early proteins IE1 and IE2 of human cytomegalovirus colocalize with and disrupt PML-associated nuclear bodies at very early times in infected permissive cells. J Virol 71:4599 – 4613.
28. Muller S, Dejean A. 1999. Viral immediate-early proteins abrogate the modification by SUMO-1 of PML and Sp100 proteins, correlating with nuclear body disruption. J Virol 73:5137–5143.
29. Lee HR, Kim DJ, Lee JM, Choi CY, Ahn BY, Hayward GS, Ahn JH. 2004. Ability of the human cytomegalovirus IE1 protein to modulate sumoy-lation of PML correlates with its functional activities in transcriptional regulation and infectivity in cultured fibroblast cells. J Virol 78: 6527– 6542. https://doi.org/10.1128/JVI.78.12.6527-6542.2004.
30. Scherer M, Klingl S, Sevvana M, Otto V, Schilling EM, Stump JD, Muller R, Reuter N, Sticht H, Muller YA, Stamminger T. 2014. Crystal structure of cytomegalovirus IE1 protein reveals targeting of TRIM family member PML via coiled-coil interactions. PLoS Pathog 10:e1004512. https:// doi.org/10.1371/journal.ppat.1004512.
31. Tavalai N, Papior P, Rechter S, Leis M, Stamminger T. 2006. Evidence for a role of the cellular ND10 protein PML in mediating intrinsic immunity against human cytomegalovirus infections. J Virol 80:8006 – 8018. https://doi.org/10.1128/JVI.00743-06.
32. Xu Y, Ahn JH, Cheng M, ap Rhys CM, Chiou CJ, Zong J, Matunis MJ, Hayward GS. 2001. Proteasome-independent disruption of PML onco-genic domains (PODs), but not covalent modification by SUMO-1, is required for human cytomegalovirus immediate-early protein IE1 to inhibit PML-mediated transcriptional repression. J Virol 75:10683–10695. https://doi.org/10.1128/JVI.75.22.10683-10695.2001.
33. Spengler ML, Kurapatwinski K, Black AR, Azizkhan-Clifford J. 2002. SUMO-1 modification of human cytomegalovirus IE1/IE72. J Virol 76: 2990 –2996. https://doi.org/10.1128/JVI.76.6.2990-2996.2002.
34. Ahn JH, Brignole EJ, III, Hayward GS. 1998. Disruption of PML subnuclear domains by the acidic IE1 protein of human cytomegalovirus is medi-ated through interaction with PML and may modulate a RING finger-dependent cryptic transactivator function of PML. Mol Cell Biol 18: 4899 – 4913. https://doi.org/10.1128/MCB.18.8.4899.
35. Lee HR, Huh YH, Kim YE, Lee K, Kim S, Ahn JH. 2007. N-terminal determinants of human cytomegalovirus IE1 protein in nuclear targeting and disrupting PML-associated subnuclear structures. Biochem Biophys Res Commun 356:499 –504. https://doi.org/10.1016/j.bbrc.2007.03.007. 36. Wilkinson GW, Kelly C, Sinclair JH, Rickards C. 1998. Disruption of
PML-associated nuclear bodies mediated by the human cytomegalovirus major immediate early gene product. J Gen Virol 79:1233–1245. https:// doi.org/10.1099/0022-1317-79-5-1233.
37. Hayhurst GP, Bryant LA, Caswell RC, Walker SM, Sinclair JH. 1995. CCAAT box-dependent activation of the TATA-less human DNA polymerase alpha promoter by the human cytomegalovirus 72-kilodalton major immediate-early protein. J Virol 69:182–188.
38. Lafemina RL, Pizzorno MC, Mosca JD, Hayward GS. 1989. Expression of the acidic nuclear immediate-early protein (IE1) of human cytomegalo-virus in stable cell lines and its preferential association with metaphase chromosomes. Virology 172:584 – 600. https://doi.org/10.1016/0042 -6822(89)90201-8.
39. Martinez FP, Tang Q. 2013. Identification of cellular proteins that interact with human cytomegalovirus immediate-early protein 1 by protein array assay. Viruses 6:89 –105.
40. Tavalai N, Adler M, Scherer M, Riedl Y, Stamminger T. 2011. Evidence for a dual antiviral role of the major nuclear domain 10 component Sp100 during the immediate-early and late phases of the human cytomegalo-virus replication cycle. J Virol 85:9447–9458. https://doi.org/10.1128/ JVI.00870-11.
41. Tatham MH, Geoffroy MC, Shen L, Plechanovova A, Hattersley N, Jaffray EG, Palvimo JJ, Hay RT. 2008. RNF4 is a poly-SUMO-specific E3 ubiquitin ligase required for arsenic-induced PML degradation. Nat Cell Biol 10: 538 –546. https://doi.org/10.1038/ncb1716.
42. Scherer M, Otto V, Stump JD, Klingl S, Muller R, Reuter N, Muller YA, Sticht H, Stamminger T. 2016. Characterization of recombinant human cytomegaloviruses encoding IE1 mutants L174P and 1-382 reveals that viral targeting of PML bodies perturbs both intrinsic and innate immune responses. J Virol 90:1190 –1205. https://doi.org/10.1128/JVI.01973-15. 43. Bekes M, Prudden J, Srikumar T, Raught B, Boddy MN, Salvesen GS. 2011.
The dynamics and mechanism of SUMO chain deconjugation by SUMO-specific proteases. J Biol Chem 286:10238 –10247. https://doi.org/ 10.1074/jbc.M110.205153.
44. Golebiowski F, Tatham MH, Nakamura A, Hay RT. 2010. High-stringency tandem affinity purification of proteins conjugated to ubiquitin-like moieties. Nat Protoc 5:873– 882. https://doi.org/10.1038/nprot.2010.40. 45. Werner A, Moutty MC, Moller U, Melchior F. 2009. Performing in vitro sumoylation reactions using recombinant enzymes. Methods Mol Biol 497:187–199. https://doi.org/10.1007/978-1-59745-566-4_12.
46. Meroni G, Diez-Roux G. 2005. TRIM/RBCC, a novel class of ‘single protein RING finger’ E3 ubiquitin ligases. Bioessays 27:1147–1157. https:// doi.org/10.1002/bies.20304.
47. Li Y, Wu H, Wu W, Zhuo W, Liu W, Zhang Y, Cheng M, Chen YG, Gao N, Yu H, Wang L, Li W, Yang M. 2014. Structural insights into the TRIM family of ubiquitin E3 ligases. Cell Res 24:762–765. https://doi.org/ 10.1038/cr.2014.46.
on November 7, 2019 by guest
http://jvi.asm.org/
48. Kang H, Kim ET, Lee HR, Park JJ, Go YY, Choi CY, Ahn JH. 2006. Inhibition of SUMO-independent PML oligomerization by the human cytomegalo-virus IE1 protein. J Gen Virol 87:2181–2190. https://doi.org/10.1099/ vir.0.81787-0.
49. Cuchet-Lourenco D, Boutell C, Lukashchuk V, Grant K, Sykes A, Murray J, Orr A, Everett RD. 2011. SUMO pathway dependent recruitment of cellular repressors to herpes simplex virus type 1 genomes. PLoS Pathog 7:e1002123. https://doi.org/10.1371/journal.ppat.1002123.
50. Galisson F, Mahrouche L, Courcelles M, Bonneil E, Meloche S, Chelbi-Alix MK, Thibault P. 2011. A novel proteomics approach to identify SUMO-ylated proteins and their modification sites in human cells. Mol Cell Proteomics 10:M110.004796.
51. Vertegaal AC, Andersen JS, Ogg SC, Hay RT, Mann M, Lamond AI. 2006. Distinct and overlapping sets of SUMO-1 and SUMO-2 target proteins revealed by quantitative proteomics. Mol Cell Proteomics 5:2298 –2310. https://doi.org/10.1074/mcp.M600212-MCP200.
52. Boggio R, Colombo R, Hay RT, Draetta GF, Chiocca S. 2004. A mechanism for inhibiting the SUMO pathway. Mol Cell 16:549 –561. https://doi.org/ 10.1016/j.molcel.2004.11.007.
53. Scherer M, Reuter N, Wagenknecht N, Otto V, Sticht H, Stamminger T. 2013. Small ubiquitin-related modifier (SUMO) pathway-mediated en-hancement of human cytomegalovirus replication correlates with a recruitment of SUMO-1/3 proteins to viral replication compartments. J Gen Virol 94:1373–1384. https://doi.org/10.1099/vir.0.051078-0. 54. Gack MU, Albrecht RA, Urano T, Inn KS, Huang IC, Carnero E, Farzan M,
Inoue S, Jung JU, Garcia-Sastre A. 2009. Influenza A virus NS1 targets the ubiquitin ligase TRIM25 to evade recognition by the host viral RNA sensor RIG-I. Cell Host Microbe 5:439 – 449. https://doi.org/10.1016/ j.chom.2009.04.006.
55. Ou HD, Kwiatkowski W, Deerinck TJ, Noske A, Blain KY, Land HS, Soria C, Powers CJ, May AP, Shu X, Tsien RY, Fitzpatrick JA, Long JA, Ellisman MH,
Choe S, O’Shea CC. 2012. A structural basis for the assembly and functions of a viral polymer that inactivates multiple tumor suppressors. Cell 151:304 –319. https://doi.org/10.1016/j.cell.2012.08.035.
56. Cuchet D, Sykes A, Nicolas A, Orr A, Murray J, Sirma H, Heeren J, Bartelt A, Everett RD. 2011. PML isoforms I and II participate in PML-dependent restriction of HSV-1 replication. J Cell Sci 124:280 –291. https://doi.org/ 10.1242/jcs.075390.
57. Hofmann H, Floss S, Stamminger T. 2000. Covalent modification of the transactivator protein IE2-p86 of human cytomegalovirus by conjuga-tion to the ubiquitin-homologous proteins SUMO-1 and hSMT3b. J Virol 74:2510 –2524. https://doi.org/10.1128/JVI.74.6.2510-2524.2000. 58. Hofmann H, Sindre H, Stamminger T. 2002. Functional interaction
be-tween the pp71 protein of human cytomegalovirus and the PML-interacting protein human Daxx. J Virol 76:5769 –5783.