Vol.63,No. 1 JOURNALOFVIROLOGY, Jan. 1989, p.64-75
0022-538X/89/010064-12$02.00/0
CopyrightC) 1989, AmericanSociety forMicrobiology
Detection
of
Multiple, Novel Reverse Transcriptase
Coding
Sequences in Human Nucleic Acids:
Relation
to
Primate
Retroviruses
ANDY SHIH, RAVIMISRA, AND MARKG. RUSH*
DepartmentofBiochemistry, New York University SchoolofMedicine, New York,New York10016
Received 30 June 1988/Accepted 8 September 1988
Avariety ofchemically synthesized oligonucleotides designed onthe basis of amino acid and/or nucleotide sequencedatawereusedtodetectalargenumberofnovelreversetranscriptasecodingsequencesin humanand mouseDNAs. Procedures involving Southern blotting, library screening, and thepolymerasechainreaction were all used to detect such sequences; the polymerase chain reaction was the most rapid and productive
approach. In thepolymerasechainreaction, oligonucleotide mixturesbasedonconsensussequencehomologies
to reverse transcriptase coding sequences and unique oligonucleotides containing perfect homology to the coding sequences of human T-cell leukemia virus types I andII were both effective in amplifying reverse
transcriptase-related DNA.Itisshown that human DNA containsawide spectrum of retrovirus-relatedreverse
transcriptase codingsequences,includingsomethatareclearlyrelatedtohuman T-cell leukemia virustypes
I andII,somethatarerelatedtothe L-1family of long interspersednucleotidesequences,andothers thatare
relatedtopreviously describedhuman endogenous proviralDNAs. Inaddition, human T-cell leukemia virus
typeI-relatedsequences appeartobetranscribed in both normal human T cells and inacell line derived from ahuman teratocarcinoma.
Two majorclasses ofeucaryotic dispersed repetitive
se-quences, retrotransposons (retroviruslike transposable ele-ments) and retroposons (short andlonginterspersed nucle-otide sequences as well as processed pseudogenes), appear
totransposeby amechanisminvolvingthe reverse transcrip-tionofanRNAintermediate(4,47, 65). In retrotransposons (such as vertebrate retrovirus proviral DNAs, mouse intra-cisternalAparticle [IAP] proviralDNAs, Drosophila copia and copialike DNAs, and yeast Ty DNAs), the reverse transcriptase required fortransposition is clearlya transpo-son-coded
function,
whereas in most retroposons (such as the primate Alu family of short interspersed nucleotide sequences and processed pseudogenes in general) there-versetranscriptase required for transposition is assumed to besupplied from retrovirusesand/or retroviruslike transpos-ableelements. However, oneclass ofretroposons, the L-1 family of mammalian long interspersed nucleotide se-quences, almostcertainly contains some members that
pos-sessreverse transcriptase coding capacity (13, 23, 54). The potential importanceof retrotransposons and retroposons in regard to the evolution of the mammalian genome and to mutagenesis andoncogenesis (64) has been most thoroughly studied in mice, where the total mass of these repetitive sequences represents atleast 10%of the genome (4, 47, 60, 65).Although short interspersed nucleotide sequences (hun-dreds of thousands of copies per cell) and long interspersed nucleotide sequences (tens of thousands of copies per cell) accountfor most of this material, endogeneous C, B, IAP, and viruslike 30S-type proviral DNAs (here considered as retrotransposons) are also present at significant levels (thou-sands ofcopies percell). Indeed, it is these proviral DNAs and the viruses that some of them generate that have been the most thoroughly characterized eucaryotic transposable elements at the genetic and biochemical levels, and it is
*Correspondingauthor.
possible that similar proviral DNAs may be amenable to
serious functionalinvestigation in humans.
A few investigators have recently begun to search suc-cessfully for retroviruslike and/or proviral DNAs in humans. In essence, these searches have been based primarily on
threeapproaches: (i) thelow-stringencyscreening of human genomic libraries with proviral DNA probes of murine leukemia virus(MLV) (27, 44, 45), murine mammarytumor
virus(MMTV) (1, 2), baboonorchimpanzee virus (33, 34), and IAPs (35); (ii) thescreeningof humangenomic libraries with oligonucleotide probes homologous to the primer-binding sites of known mammalian retroviruses (12,18);and (iii) the fortuitous identification ofpreviously uncharacter-ized repetitive elements in genomic clones followed by subcloning and sequencing (24, 25, 39). All of these ap-proaches have been successful, and the results may be briefly summarized bystating thefollowing:(i) the elements identifiedto datearesignificantly different from each other; (ii) six of the elements that have been extensively or com-pletely sequenced each use a different tRNA for putative priming of reverse transcription; (iii) some of the elements clearlycontain all of theinformation associated with proviral DNA(longterminal repeats and gag, pol, andenvregions), whereas otherscontain extensive deletions; (iv) copy
num-bers of different elements range fromasinglecopyto tensof thousands;(v) manyof the elementsaretranscribed (7, 9, 14, 36, 39, 42); and (vi) thelong terminal repeats ofmostof the sequenced elements contain all oftheregulatory sequences generally associated with theirfunction as both promoters andterminatorsoftranscription, althoughgagand pol genes have acquired deletions oraccumulated additional termina-tion codons within their predicted functional open reading frames. (We have adopted the terminology of Kroger and Horak [18], in which human retrovirus-related sequences [HuRRS] are classified on the basis of putative tRNA primer-binding sites. Thus HuRRS-E [45] and HuRRS-K 64
on November 10, 2019 by guest
http://jvi.asm.org/
HTLV-1 TGGAAAGTACTACCCCAAGGG TTTAAAAATAGTCCCACCCTGTTCGAAATGCA\GCTGGCCC\TA\TCCTGCAGCCCATTCGGCAAGCTTTCCCCCAATGCA\CTA\TTCTTCAG TA\CATGGATGACATTCTCCTAGCAAGC
U K V L P Q G P K N SPYT L F EM Q L A HRI L Q P1I R Q A F P Q C T I L Q Y MND DI LL AlS
HTLV-II TGGACTGTCCTTCCACAGGGG TTTAAAAACAGCCCCACCCTCTTCGAACAACAA\TTAGCAGCCGTCCTCAACCCCA\TGAGGAAAATGTTTCCCACATCGACCATTGTCCAA TACATGGATGACATACTTTTAGCCAGC V T V L P Q G P K HISP T LYrE Q Q L A A V L N P M RKKmNr P T IT I V Q TNM D D I LL AlS
HIV-1 TACAATGTGCTTCCACAGGGA TGGAAAGGATCACCAGCAATATTCCAAAGTAGCATGACAAAAATCTTIAGAGCCTTTTA)AAAACA\AAATCCAGACATAGTTATCTATCAA TACATGGATGATTTGTATGTAGGATCT TRY L P Q G U K G SPA IFP QII RTM K I L E P r K K Q N P D I V ITY Q T M D DL T V G S
SRV-11 TGGAAAGTGTTG;CCTCAAGGC ATGGCCAATAGTCCTACCTTGTGTCAAAAATATGTAGCTGCTGCTATAGAGCCAGTCAGAAA\ATCTTGGGCACAAATGTACATTATACAC TATATGGATGACA\TTCTAATAGCAGGA
U K V L P Q G MARNS PT L C Q K T V A A Al IE P VRKKS WVAQ N T I I H T M D D I L I A G
NPMV TGGAAGoGTTTTACCACAAGGT ATGGCCJ
U K V L P Q G M A
BaEV TGGACTCGACTTCCCCAGGGG TTCAAAj
VT K L PQ G F K
MoMLV
PTV
Mtu-IA?
Pr-RSV
SLY
VLV
EIAY
CAkYv
RHY
CaNY
HRuRS-E HRRMS-K
RISP T L C Q K T V A T A IRHK V R H AUW KG MTH I I H
RISP T L P D E AL H R D L TD FRYT Q H P E VT L L Q
TGGACCAGACTCCCACAGGGT TTCAAAAACAGTCCCACCCTGTTTGATGAGGCACTGCACAGAGACCTAGCAGACTTCCGGATCCAGCACCCAGACTTGATCCTGCTACAG V T R L P Q G F KRN SP T L FPD E AL H RD0 L AD F R I Q H P DL I LL Q
TGGAAAGTTTTGCCCCAGGGT ATGAAAAATAGCCCTACTTTATGTCAAAAATTTGTGGACAAAGCTATATTGACTGTAAGGGATAAATACCAAGACTCATATATTGTGCAT WKYV L PQG N K H S PT LCQ K F V D K A I L TV R D KY Q D S T I V H
TGGAAGGTCTTACCACAGGGA ATGTCCAATAGTCCTACAATGTGCCAACTTTATGTGCAAGAAGCTCTTTGCCAGTGAGGGAACAATTCCCCTCTTTAATTTTGCTCCTT
WKV L PQ G NIH S P TMNC Q L Y VQ E A LL P V REQ F P S L I L L L
TGGAAGGTCTTGCCCCAAGGG ATGACCTGTTCTCCCACTATCTGTCAGTTGGTAGTGGGTCAGGTACTTGAGCCCTTGCGACTCAAGCACCCATCTCTGTGCATGTTGCAT V K V L P Q G M T CS?P T IC Q L VV G Q V L E P L R L KR? ISL CMNL H TGGCGGGTCCTACCTCAAGGCTTCATTAACAGCCCAGCTCTTTTCGAACGGGCACTACAGGAACCCCTTCGCCAAGTTTCCGCCGCCTTCTCCCAGTCTCTTCTGGTGTCC
WRV L P QG FIRNS P ALFr E RA LQ E P L RQV s A A rS Q I L LVI
TGGAAAGTGTTACCACAAGGA TGGAAATTAAGTCCTGCAGTGTATCAATTTACAATGCAAATTAGGAGATAGA.AGAACACCCTATGATACAATTTGGAATA
V K V L P Q G V K LISP A V T Q FT MQ K I L R G V IE ER? MIr Q P G I
TGGAATTGTTTACCACAAGGA TTCGTGTTGAGCCCATATATATATCAGAAAACATTACAGGAAATTTTACAACCTTTTAGGGAAAGATATCCTGAAGTACAATTGTATCAA
V H CL P Q G P V LISP T I T Q K T LQ ElI LQ PPF RE RKY?P E VQ L T Q
TGGAAG-GTTCTACCACAAG4GT TGGAAACTGAGTCCATCTGTATATCAATTTACCATGCAGGAGATCTTAGGGGAATGGATACAGGAGCATCCTGAAATTCAATTTAGAATA
U K V L P Q G WUK LISP S VTY Q FT MQ ElI L G E V I Q ER? ElI Q FR I TTTCGCAAGATTCCTATGGGA GTGGGCCTCAGTCCGTTTCTCCTGGCTCAGTTTACTAGTGCCATTTGTTCAGTGGTTCGCAGGGCTTTCCCCCACTGTTTGGCTTTCAGT
F R K I1PM G V G LISP F L L A Q FYI A I C I V V R K A F PRH CLAPFS
TGGAATGTGG;TCCCTTTCGGC CTAAAGCAGGCACCATCCATATTCCAGAGACACATGGACGAAGCATTTCGTGTGTTCAGAAAATTCTGTTGCGTG
V KY vvP P G L K Q A PSI P Q R H M D E APfR VP K K F C C V
TGGACCCAGCTTCCCCAAAGGTTCAAGAACTCCCCCACCATCTTTGGGGAGGCGTTGGCTCGAGACCTCCAGAAGTTTCCCACCAGAGACCTAGGCTGCGTGTTGCTCCAG
WYT Q L P QR KP K R P TIPF GE A L A RD L Q K P P Y R D L G C V L L Q
TGGAAAGTGTTACCTCAGGGAATGCTTAATAGTCCAACTATrTGTCAGACTTTTGTAGGTCGAGCTCTTCAACCAGTGAGAGAAAAGTTTTCAGACTGTrATATTATFCAT
V K V L P Q G ML RISP T IC Q T F V G R A LQ PYVRE KY ID C TI I H
UPSTREAM PRIMER TG A A
MOP 5. TGGAAAGTG/T/CC/CA/GG 3'
C A C G
HTLV-I 5' TGGA.AAGTACTACCCCAAGG 3' HTLV-11 5' TGG3ACTGTCCTTCCACAGGG 3'
TMN D D I L LC H
TMN D DL L L A A TATATGGACGATATCCTTATCGCTTCG
TMN D DI L I A S
TMH D D I TI RIS
TATATGGATGATGTGGTATTGGGGGCC
YTM D D V V L G A
TV D DI L V P S
TACGTTGATGACCTTTTGCTGGGACAC
TV D D L L L G H
DOWNSTREAM PRIMER
T TG A
3' ATG/ACCTACTG/A/GACGACCGG/GG 5'
C GC C
3' ATGTACCTACTGTAAGAGGA S.
3' ATGTACCTACTGTATGAAAA 5'
FIG. 1. Nucleotide and amino acid comparisons ofconserved sequences identified in the amino-terminal coding regions of reverse
transcriptasesandnucleotide sequencesof six synthetic oligonucleotidesusedforpriming polymerase chainreactions. Thesource ofeach
reversetranscriptase coding regionisindicatedontheleft,asis thenatureof theoligonucleotide primer.Standardsingle-letterabbreviations
are usedto designate aminoacids, and all nucleotide sequences, except for the downstream primers, are oriented in the 5'-to-3' (sense)
direction. A block ofsevenamino acidsattheamino-terminal ends of theprotein sequencesshownandablock of nine amino acidsattheir carboxyterminirepresentexceptionally highlyconservedregions.Thesequencespresentedhereareasfollows: HTLV-I(50);HTLV-II(51); HIV-1(43);
SRV-1I,
simian retrovirustype11(61); MPMV,Mason-Pfizermonkeyvirus(57); BaEV,baboonendogenousvirus(15); MoMLV, MoloneyMLV(52);MMTV(31); Mu-IAP,murinelAP(29); Pr-RSV,Roussarcomavirus(PragueC strain) (49); BLV,bovine leukemia virus(46); VLV,visna lentivirus(56); EIAV,equineinfectious anemia virus(3); CAEV,caprinearthritisencephalitisvirus(3); HBV, hepatitisB virus(ADR strain) (17); CaMV,cauliflower mosaic virus(6);HuRRS-E (45);and HuRRS-K(37).
[37] refer to such sequences
containing
glutamic
acid- andlysine-specific tRNA-binding
sites,respectively.)
The detection of new retrovirus-related sequences in humans hasbeen based
primarily
upontheirhybridization
to known clonedfamily
members, apropertyrequiring
exten-sive
homology
overrelatively long
(kilobase [kb]) stretches of DNA and thusbiasing
the results in favor ofspecific
family
types. Inthepresentstudy
wedescribetheefficacy
ofa new detection
procedure,
thepolymerase
chain reaction (PCR), based upon the use ofsynthetic oligonucleotides
homologous
tohighly
conserved regions of reversetran-scriptase coding
genes. Thepolymerase
chain reaction is atechnique
in which small(picogram)
amounts ofa definedregion
of DNAareamplified (chemically
cloned)byrepeated
rounds of DNA
synthesis primed by
an excess of twospecific oligonucleotides,
one homologous to an upstream target in one strand of thetemplate
DNA and the other homologoustoadownstreamtarget intheopposite
strand of thetemplate
DNA.Theprimers
aredesigned
sothatthetwochain extensions occur in convergent directions;
repeated
(-30) rounds ofdenaturation,
primer annealing,
and DNAsynthesis
result in thegeneration
ofmicrogram
quantities
of double-stranded, blunt-endedfragments
(bounded by theprimers)
that are suitable for dir'ectcloning
into M13se-quencing
vectors. The reversetranscriptase-homologous
primers
used inourstudy (Fig.
1)resulted in the amplifica-tion of a wide spectrum of retrovirus-related reversetran-scriptase coding
sequences, bothexpected
andunexpected.
Among
theunexpected
werethose withconsiderablehomol-ogytohuman T-cellleukemiavirustypeI(HTL
V-I),
HTLV-II, and some monkey retroviruses as well as those with asimilarity
to the L-1 family, whereas among theexpected
were those with
homologies
topreviously
definedendoge-nous human
proviral
DNAs. The advantages of the PCRprocedure
over conventionaloligonucleotide probing
foridentifying
members ofamultigene family arediscussed,asisthe
possible significance
ofourfindingsto thenature and evolution of human retroviruses.on November 10, 2019 by guest
http://jvi.asm.org/
66 SHIH ET AL.
MATERIALS AND METHODS
Standardprocedures.Restriction endonucleasedigestions, gelelectrophoresis, Southern and Northern (RNA) blotting, hybridization of blots to oligonucleotide-primed labeled probes, labeling ofsynthetic oligonucleotides with polynu-cleotide kinase, and preparation of high-molecular-weight chromosomal and bacteriophage lambda DNAs were per-formed as previously described (30) and/or according to standard procedures (26). Sequencing of single-stranded M13 cloned DNAs was performed by the dideoxy-chain terminator method with amodifiedT7 polymerase enzyme (Sequenase)and[at-355]dATPwith thereagents andprotocol oftheUnited States Biochemical Corp. All sequences were obtainedfrom multiple gelreadings. Experimentsinvolving theprobingof blotted gels containing PCRreactionmixtures (or total genomic DNA) always included stringency and copy number controls. Specifically,random oligonucleotide-primedprobes synthesized from single-stranded M13 clones were hybridized to both blotted gels and control dots of filter-bound complementary M13 RF, and both blots and dots werewashed in 0.03 M NaCl-3mMtrisodium citrate-0.1% sodium dodecyl sulfate at a series of temperatures rangingfrom50to70°C at 5°C intervals. For theexperiments shown in Fig. 6 and7, blots and controls exhibited compa-rable losses in signal intensity, indicatingthatprobes were extensively homologous to their complementary blotted DNAs and, inthe case of PCR mixtures, not due solely to the 30% expected primerhomology. The filters autoradio-graphed for Fig. 6 were washed at 60°C, and thoseforFig. 7 were washed at 55°C.
High-molecular-weightDNAs.High-molecular-weightDNAs isolatedfrom livers ofC57L/JandAKR/Jmice were obtained from P. D'Eustachio (New York University, Department of Biochemistry), as was DNAisolatedfromthe A9 mousecell line(derived from strain C3H); DNAisolatedfrom spleens of RF/Jmice wasfromS. Sloan (New YorkUniversity, Depart-mentofPathology);DNAisolated fromthe 293 humancell line (11) was fromR. Schneider(New York University, Depart-ment ofBiochemistry); DNA isolated from the CEMhuman T-cell lymphoblastoid cell line wasfrom J. Krolewski (New YorkUniversity,Department ofPathology);and DNAisolated from human scalp skin and the human melanoma cell lines BSFMand HM20werefromD. Zouzias (New York Univer-sity, DepartmentofDermatology).
Genomic and cDNA libraries. Ahumangenomic placental library was obtained from J. Hall (New York University, Department of Biochemistry), a human teratocarcinoma (TC)-derived cell line cDNA library (54, 55) was from J. Skowronski (ColdSpringHarborLaboratory), and a human T-cell (PTL)-derived cell line cDNA library was from J. Krolewski. All ofthesebacteriophagelambdalibrarieswere plated on Escherichia coliLE392and screened with oligo-nucleotideprobes by the hybridization procedure described below.
Total and poly(A)+ RNAs. Total and poly(A)+ RNAs isolated from the Burkitt lymphoma cell line ST486, were obtained fromJ. Krolewski. HeLa total poly(A)+ RNA was isolated by the guanidinium isothiocyanate procedure of Maniatis etal. (26).
Hybridization of oligonucleotides to filter-immobilized DNAs. Alow-stringencyhybridization (37°C in 0.9 M NaCl, 0.09 M trisodium citrate, 10% dextran sulfate) followed by high-stringency washes in the presence of 3 M tetramethyl ammonium chloride (a compound thateliminates the effect of base composition on duplexDNA meltingtemperatures)
wasused in all studiesexactlyas describedbyWoodetal. (66).Inaddition,controls ofMoloneymurine leukemia virus proviralDNAwereincluded in mostexperiments(oneof the mixed probe sequences is about 95% homologous to the known conserved Moloney 27-base-pair region) as a
rough
measure of probe homology, at least as determined by comparative reductions in signal intensity as a function of washing temperatures (60 to 75°C). Hybridizations were performed in 10-ml solutions containing 200 ng of 5' 32p_ labeledoligonucleotideswithspecificactivitiesrangingfrom 2 x 108 to 1 x
109
cpm/,ug. Forthe experiments shown in Fig. 2 and 3 the washing temperature was 600C.PCR. PCRs were performed essentially as described
by
Saiki et al. (48)with Thermus aquaticus DNApolymerase
(Perkin ElmerCetus) to increase reaction specificity (since chain extension occurs at 72°C) as well as to reduce the overall time of amplification(sincethis enzyme isstable ata94°C denaturation temperature). In short, reactions were performedin aDNA Thermal Cycler (Perkin Elmer Cetus) in 100-,ul solutionscontaining1to2,ugofDNA,50 mMKCl,10 mMTris (pH8.4), 2.5 mM MgCl2, 1 ,uM each uniqueprimer or10,uM each mixed oligonucleotide primer (MOP), 200 ,uM eachdeoxyribonucleoside triphosphate,200
Rg
ofgelatinper ml, and 2 U of polymerase. Samples overlaid with anequal volume of mineral oil were subjected to 30 cycles of ampli-fication by (i) heating from 72 to 94°C (over 30 s) and incubatingat94°Cfor 2min(denaturation), (ii) cooling(over 1 min) from 94°C to a primer annealing temperature of between 37 and 60°C (depending on the experiment) and incubating for 3 min, and (iii) heating from the annealing temperature(over 1min)to achain extension temperatureof 72°C and incubating for 4 min. The final extension wasincubated for 7min, and 15-,ul samples were then resolved oncomposite gels of 3% NuSieve-1%electrophoresisgrade agarose (Schwartz/Mann) containing 0.5 ,g of ethidium bromideper ml.Fragments to be subcloned(20-,I samples) werefirstphosphorylated by adding 20 U (2
RI)
T4 polynu-cleotide kinase and 1RI
of 50 mM ATP, and incubating for 45 min at 37°C. After ethanolprecipitationandsuspensionin 20 p1 ofH20, thefragments were clonedbyblunt-end ligation (10 U of T4 DNA ligase) into the SmaI site of alkaline phosphatase-treated M13mpl9.Comparisonofamino acidsequences. Computer analysisof sequence data was performed on a Digital VAX 11/750 microcomputer by using the protein information resource (40, 41) and/or Staden (58, 59) nucleic acid sequence pro-grams.
Synthetic primers and probes. All oligonucleotides were synthesizedinthelaboratory ofB.Goldschmidt (NewYork
University,
Department of EnvironmentalMedicine) with an Applied Biosystems380A Synthesizer.RESULTS
Synthetic oligonucleotide primers and probes. In the inter-est of detecting new human endogenous retroviruses and retroviral related sequences that might lack extensive
(kb)
homologies with known retroviruses, two procedures with reverse transcriptase-specific synthetic oligonucleotides were employed. The first was based on conventional ge-nomic library and Southern blot screenings, whereas the second was based on the use ofPCR. As shownbelow, the PCRprocedure represented a far moreproductive approach to characterizingthemultigene reversetranscriptase family than did conventionalscreeningandresulted in the identifi-cation of sequences that are
apparently
related to known J. VIROL.on November 10, 2019 by guest
http://jvi.asm.org/
primate retroviruses. Figure 1 presents published nucleotide and amino acid sequences of 18 retroviral or retroviruslike reverse transcriptase genes in a region that contains a variety of particularly well-conserved, so-called diagnostic amino acids (38, 62, 63). Also included in this figure are the sequences of six synthetic oligonucleotides used as primers for PCR analyses of human (and in some cases, mouse) DNAs. One of these oligonucleotides (see below) was also used as a conventional probe for library and blot screening. The primers represent putative (MOP) or actual(HTLV-I andHTLV-II)coding sequences for exceptionally well
con-served amino acid blocks. The rationale for using unique HTLVprimers wasthat human DNA might contain endog-enous proviral DNAs related to human retroviruses, whereas the rationale for using MOP was that humanDNA is already known to contain a variety of murine and other retrovirus-related sequences. The biases used for determin-ing the sequence of the mixed primers, in which both the
upstream and downstream mixtures are composed of 16 distinct oligonucleotides, were based both on theoretical considerations (20) and on the amino acid composition of the conserved reverse transcriptase regions in birds, mice, and humans. The downstream MOP contains 27 nucleotides rather than the 20 present in the other primers, since this mixed oligonucleotide (as well as its complement) was also designed for use as a probe for conventional screening of genomic and cDNA libraries.
The theories of probe selectivity and sensitivity upon which the latter screening was based indicated that rather long unique syntheticoligonucleotides (>25 residues), con-taining sequences based on primate codon usage and pos-sessing about 85% homology to a target sequence, should be more than adequate for selecting truepositivenonrepetitive clones from cDNA libraries. Slightly better homologies and/ or longer probes are required for selecting true positive unique clones from genomic libraries. Our mixed 27-nucle-otide oligomer represents a compromise between the useof a single long probe with about 85% target site homology (based on codon usage) and the use of a mixture of many smaller probes representing allpossiblecodingpossibilities. This compromise was adopted since the amino acid sequence(s) upon which the probe was based is itself a consensus. The expectation that nearly all of our target sequences should be present in repetitive elements,andthat the selection of positive clones with at least90%homology to our probe (estimated by stringency comparisons with a Moloney MLV proviral DNA control), were twoadditional factors that favored the selection of true positive clones.
Of the two complementary probes, one, 5' TAC
A A C T
-TGGATGAC-T-CTGCTGGCC-GC 3', represents the
G C G G
sequence expected in an mRNA coding for reverse
tran-scriptase and is known as the sense probe, whereas the
A GT T
other, 5' GG-GGCCAGCAG-A-GTCATCCA-GTA 3'
C C G C
represents the
complement
of this and is known as theantisense probe.
Detection of conserved reverse transcriptase coding
se-quences by conventional screening. The detection of
con-served reverse transcriptase coding sequences in Southern
and Northern blots of human DNA and RNA is shown in
Fig. 2 and 3. Also shown inFig. 2 isthe detection of such
sequences in mouse DNAs. The resultsoftheSouthern blot
analysis shown in Fig. 2 and ofotherSouthernblotanalyses,
including additional mouse and human DNAs (data not
A Y BgP X K B9P X K BgP X K BnH K X
< 0 A 9 C57L/J AKR/J 293
8Kb- _
2Kb- o
a
wV
S
0.
W.V.
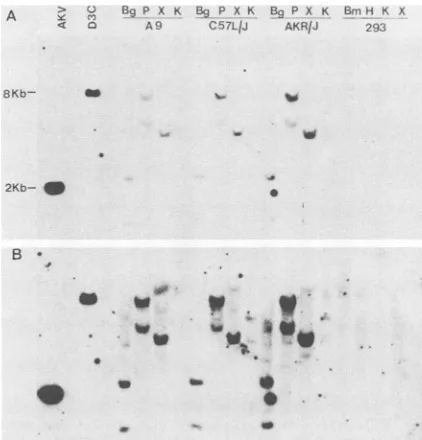
FIG. 2. Southem blot analysis of mouse and human DNAs
probed witharadioactively labeledsynthetic oligonucleotide
mix-ture homologous to a conserved region of mammalian reverse
transcriptase codingsequences. Restriction endonuclease-digested DNAsampleswereelectrophoresedfor11 h in1%agaroseat2V/
cm, transferred to nitrocellulose, and hybridized to the sense
oligonucleotide mixture(200ng, 109cpm/,ug) asdescribedin Mate-rials and Methods. Panels A and B represent, respectively, short
(16-h) andlong(60-h)exposuresof thesameprobedfilter.Thelanes contain thefollowing (from lefttoright). AKV, 1 ,ugofan EcoRI-cleavedmouse cDNA clone ofAKV viralRNA in phage lambda. D3C, 1 jig of an EcoRI-cleaved human genomic clone (D3C) in phagelambda. (Thisclone was selectedby usingthe 27nucleotide mixed probe; although it contains a reasonably homologous target
[see below], its reverse transcriptase context is not clear.) A-9,
C57L/J and
AKRIJ,
10 ,ug ofhigh-molecular-weight DNA isolated from mouse cell line A-9 and mouse strains C57L/J and AKR/Jdigestedwith thefollowingrestriction endonucleases: Bg,BglII; P, PstI;X,XbaI;K, KpnI. 293, 10 ,ugofhigh-molecular-weight DNA isolatedfromthe humancellline293 anddigestedwith thefollowing endonucleases: Bm,BamHI; H, HindIll. Theapproximate sizesof
the AKVand D3C insertshomologous totheoligonucleotide probe are indicated on the left. The signal from the 2-kb AKV insert corresponds to about 10,000 copies ofthe oligonucleotide target
sequence per cell. (ThetargetinAKVis about 95%homologous to
onemember ofthemixed27nucleotideprobe,whereas thatinD3C is about85% homologous toanother member.) A 2-,ug
sample
ofHindIll-digestedphage lambdaDNAincluded for visible(ethidium bromide-stained) size markers was electrophoresed in a lane (not shown)adjacent toD3C.
shown), were as follows:
(i)
distinct,
moderately
repetitive
bandswerereadilydetected inmouseDNAsaftera16-h blot
exposure, whereas
lower-copy-number
species
became vis-ible after 60 h;(ii)
the patterns ofmoderately
repetitive
mouseDNAs were
essentially
identicalfor all threestrains,
whereasthoseof the
lower-copy-number
bandsdiffered;
and (iii) human DNA revealed a diffuse pattern ofhomologous
material,withperhapsafew distinct visible bands in the 60-h exposure.
At present itappears thatthepolgenerepresentsthemost highly conserved
region
among theendogenous
proviral
DNAs of mice, and the results shown in
Fig.
2strongly
suggest that our
probe
detected many members ofdifferent proviral(retrotransposon)
families.Unfortunately,
it ison November 10, 2019 by guest
http://jvi.asm.org/
[image:4.612.331.542.74.294.2]68 SHIH ET AL.
'Sense Probe
XR
c >1-a
(0x
0'Q
O~ O o
Antisense Probe
2_~ C3 >~
_ X
._ .
0 I
m CD,
2 UD
o
i..
5Kb
-2Kb
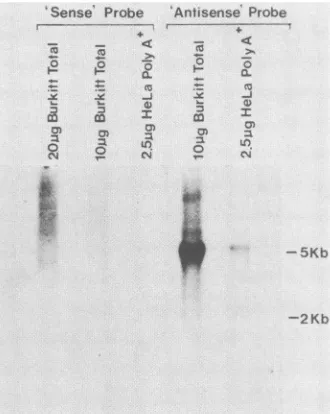
FIG. 3. Northern blot analyses of human total and poly(A)+
RNAs probed with reverse transcriptase-related DNA-coding
se-quences. The indicatedamounts of totalorpoly(A)+ RNAs were
electrophoresedfor 13 h in 1% agarosecontaining6.6%
formalde-hydeat 2.5 V/cm, transferredto nitrocellulose, andhybridized to
either a sense or antisense synthetic oligonucleotide mixture as
described inMaterials and Methods. Thepositionsof 5-kb(28S)and
2-kb(18S)rRNAsweredetermined from ethidiumbromide-stained
gels beforetransferandareindicatedontheright.Thesenseprobe
representsanoligonucleotide mixturecomposed ofsequences
ex-pected in a reverse transcriptase coding mRNA, whereas the
antisenseproberepresentsamixtureofcomplementarysequences. Thus, it is the antisense probe that is expected to hybridize to mRNAscontaininganhomologousconservedreversetranscriptase
codingsequence. Filterswereautoradiographedfor 16 h.
ficulttocorrelate thebands shown in thisfigurewithknown endogenous viral loci due to the fact that gag- and
env-specific probes have been routinely used to characterize suchloci andbythe fact thatmousecellscontain thousands ofIAPs, hundreds of viruslike30S,tensofC-type xenotro-pic and polytropic, a few C-type ecotropic, a few B-type, andvarious quantities of other endogenous proviral DNAs (60). Since the IAP family seems to be the most abundant andnon-strain-specific mouseendogenous proviral DNA,it
has beententativelycorrelated with the moderately repeti-tive bands shown in Fig. 2 (29). The less intense bands (in
some cases strainspecific) have been tentativelycorrelated with the remaining classesofproviral DNAs.
The failuretodetect discrete bandshomologoustothe 27 nucleotide mixed probe in Southernblots ofhuman DNAs
probablyreflectsthepresenceof thousands ofdivergentand
dispersed reverse transcriptase sequences rather than their absence (see below).
Theresults oftheNorthern blot analysisshown inFig. 3 andthatof otherNorthernblotanalyseswere asfollows: (i) HeLapoly(A)+ RNAandtoalesserextent, poly(A)+RNA isolated from a Burkitt's lymphoma cell line (data not shown) contained an abundant 5- to 6-kb species
homolo-goustotheantisense (butnotsense)oligonucleotide probe; (ii)Burkitt'slymphomacelltotal RNAcontainedavariety of abundantspecieshomologoustotheantisense probe [includ-ing the 5- to 6-kb RNA detected in poly(A)+ samples] but contained much lower amountsof material (withan
appar-ently different size distribution) homologous to the sense
probe; and (iii) 28S (5-kb) rRNA failed to hybridize to the antisenseprobeunder the conditions used forourscreenings (datanotshown). Surprisingly, 28S rRNAcontains atarget
sequence with about 70% homology to one member of the probe mixture (10).
The extensive hybridization of total RNA [incontrast to poly(A)+ RNA] withthe oligonucleotide probeistypical of
many highly conserved, repetitive dispersed sequences,
whereas the detectionofa discrete band ofpoly(A)+ RNA homologous to a highly conserved region of endogenous retroviruses is consistent withtherecent identification ofa variety ofsuch RNAsinmanyother celltypes(7, 9, 14, 36, 42).
Despite thepositiveresultsobtainedin theblotting
exper-iments, attempts to isolate reverse transcriptase homologs fromhuman genomicand cDNA librariesby screeningwith the MOP downstream probe failed. We therefore investi-gated the use of PCR; the requirement that a template contain sequences homologous to two distinct probes was expected to increase the specificity ofthe screen, whereas theamplificationinherent in the PCRwasexpectedtoallow detectionofrare sequences.
Detection of conserved reverse transcriptase coding se-quencesby the PCR.PCRamplificationhas been showntobe
an extremely powerful tool for the analysis of unique or
extremely low-copy-number genes (19, 21, 48); it mayalso
represent one of themost rapid and productive approaches for the characterization of multigene families. Specifically, thereversetranscriptase-specific primers presented in Fig. 1
were used to amplify avariety of human (and one mouse) DNAs at annealing temperatures ranging from 37 to 60°C, and the reactionproducts were analyzed by gel electropho-resis. In successful cases of amplification, the fragment mixtures werecloned by blunt-end ligation into M13mpl9, sequenced, and then compared attheamino acid level both with each other andwith known reverse transcriptase genes. Theresultsofsomegelelectrophoresis analysesand of all of the amino acid sequence comparisons are shown in Fig. 4 andTable 1,respectively. The results of the electrophoresis analysis shown in Fig. 4 indicate that (i) the brightest band in
anyPCR generally correspondedtothe sizeexpected(-130 base pairs) for amplification of retroviruslike reverse
tran-scriptases; (ii) the MOP functioned inaspecific PCR ampli-ficationeven though it was relatively ineffective as aprobe for screening libraries and Southern genomic blots; (iii) higher annealing temperatures (higher PCR stringencies) generatedfewer and more clearly defined amplification prod-ucts; and (iv) HTLV-I- andHTLV-II-specific primers func-tioned at annealing temperatures as high as 60°C, even though it isgenerally assumed that amplification at annealing temperatures of 55°C requires nearly perfect primer-tem-plate homology (48). In addition the results ofmany other electrophoresis analyses (data not shown) indicate the fol-lowing: (i) all of our primers, including those corresponding tohumanimmunodeficiency virus(HIV),generate aproduct of theexpected size when humangenomic DNAs areusedas templates at an annealing temperature of 60°C; (ii) only HTLV-I primers generate a product of the expected size when TC cDNA and T-cell (PTL) cDNA are used as templates at an annealing temperature above 55°C; and (iii) theupperlimitforeffective annealingofour primersis in the range of 65 to 70°C. On the whole, gel analysis of PCR-amplified fragments strongly suggested that conditions of bothmoderate and very high stringency allowed the synthe-sisof reverse transcriptase-related sequences. The results of the amino acid sequence comparisons (Table 1) confirmed J. VIROL.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:5.612.99.264.72.279.2]MOP HTLV-I
0, ~ U)<
-<r
LLAJ fQ2 wJ UJ uj2 WUJI.-WLU wUw
L)CIz)0 (O QLm0 X
246bp-123
bp- 246bp-
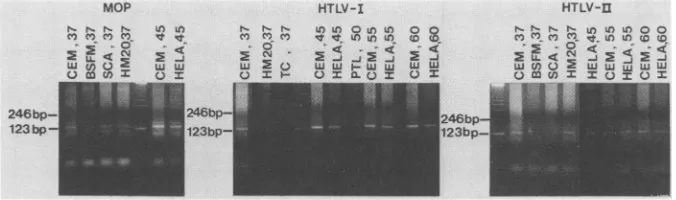
123bp-FIG. 4. Electrophoretic analysis of products from25 polymerase chain reactions withseven human DNAs and three distinctreverse
transcriptase-relatedprimermixtures. Gelswere runfor2.5hat5V/cmasdescribed in Materials andMethods. The composite photographs
shownherearegroupedonthe basis of primers (MOP,HTLV-I,and HTLV-II,seeFig. 1), and the templateDNAandannealingtemperature
areindicatedabove each lane. A marker lane (unlabeled) containing 1.5,ugofa123-base-pair ladder (Bethesda Research Laboratories, Inc.)
ispresentin eachgroup, and thepositions of the123- and246-base-pair fragmentsareindicatedontheleft. Template abbreviations: CEM, human T-cell lymphoblastoidcell line DNA; BSFM and HM20, human melanoma cell line DNAs; SCA, human scalp DNA; HELA, human HeLa cell lineDNA;TC,humanteratocarcinomacell line cDNA (in phagelambda);PTL,human T-cell cDNA (in phage lambda). The lowest broad band in allof the lanesrepresentsunincorporated primers.
this suggestion. Although some of the group designations
anddiagnostic amino acid assignments in Table1are
admit-tedlyabitarbitrary, thereversetranscriptasecontextsof all of thesequences outside ofgroupsC and Fareclear. Some
ofthemorerelevant technical observations related toTable 1 as well as to the nucleotide sequences from which it is
derived are as follows. (i) Blunt-end ligation appeared to
resultin the cloning of full-length fragments. (ii) All primer
sequencesinthe cloned fragmentswereexactlyasexpected,
and in thecaseof MOP primerusageappearedtobe random. (iii) Mixed HTLV-I-HTLV-II priming yielded two clones (A.9 and F.6) with only HTLV-I primer ends and two(B.4 andB.5) witha5'HTLV-I anda3' HTLV-II primer end. (iv)
Fewer than 20% (groups C and F) of the total sequences were not in a reverse transcriptase context, and of these about halfwerederived from mixedoligonucleotide priming at 37°C (F.1 through F.4). (v) Of the 40 sequences in a reverse transcriptase context, only 2 (A.1 and A.11)
con-tained terminationcodons. Onlyoneamino acidsequencein
the entiretable, B.8,wasnotderived by direct translation of its cloned fragment. The nucleotide sequence of B.8
con-tained an A after the sixth codon which, when removed,
allowedaperfectnucleotidesequencematchwith clone B.7.
(vi) All examples of identical amino acid sequences (except
for B.7 and B.8; see above) in cloned fragments, whether
thesefragments werederived fromasingle reaction mixture
suchasthe three clonesofgroupCorfrom different reaction
mixtures such as clones A.3/A.4, A.5/A.16, A.9/A.12, and B.3/B6,wereduetoidentical nucleotidesequences.(vii) The
reported mutation frequency of the T. aquaticus polymerase (48) suggests that one out of three clones generated by amplification of the same template should contain a single
nucleotide substitution. Thus, clones differing by a single
nucleotide (datanotshown) (suchas A.2andA.3, A.6 and
A.8, A.12 and A.14, B.4 and B.5, and D.1 and D.2) and also differing by a single amino acid may have been generated from the samesite.
In addition to these technical points and the overall efficacy of the PCR procedure,afewadditional observations
concerning the homologies of some sequences shown in
Table 1 should benoted. (i) None ofthe human sequences
was identical tothat ofany published retrovirus orhuman
endogenous provirus, although many were similar to HuRRS-E and K. B.1differedby onlyoneaminoacid from
HuRRS-E. Onemousesequence(D.1) differed byoneamino
acid from a known IAP, and two other mouse amino acid
sequences(D.2 and D.3)wereidenticaltoIAPandMMTV,
respectively. The nucleotidesequencecomparisons between
B.1 and HuRRS-E, D.2 and IAP, and D.3 and MMTV are
shown in Fig. 5. Since clones B.1, D.2, and D.3 differ from theirknown chromosomal homologsattwoorfour internal
(nonprimer) positions, it is likely that these particular homo-logs did not initiate amplification. (HuRRS-E is part ofa
rather large human endogenous provirus family, as is the
IAP family of mice. Mice may also contain a variety of
sequences that are highly homologous to MMTV.) The clustering of mismatches in primer regions suggests that mixedoligonucleotide priming at37°Ccan amplify relevant
regions evenwhenprimers and targetsarerelatively poorly
matched. (ii) Some human sequences, especially those
gen-eratedby high-stringency PCR with HTLV-I primers, bore limited butstriking resemblancestoprimate-specificreverse
transcriptases. Note, for example, groupE andmembers of
groupAcontaining the HTLV-I diagnostic FPQCgroupfour
aminoacids in from theircarboxy termini. (iii) Some identi-cal human sequences were present in both genomic and cDNAs (A.5/A.16, A.9/A.12), and many cDNA-related se-quences also showed the resemblance to primate-specific
reverse transcriptases described above. (iv) The final
inter-esting homology relationship shown in Table I is that of
group C, in which both the length and amino terminal
residuesof theamplified clones suggestedasimilaritytothe L-1 family. (The hepatitis B virus homolog is included to show that these amino-terminal residues, or functionally
conservedones,arepresentin yet anotherreverse
transcrip-tase gene.) Although the relationships in this group are
unclear, the MOP was not expected to detect L-1 family members; it is thereforepossible that PCRhas amplifiedan
L-1-like elementinwhich thesequencesof theMOP-binding
sitesresemble those ofretroviruses.
Distribution of some reverse transcriptase-related, cloned DNA fragments in total PCR products. To investigate the
distribution ofspecific amplified PCRfragments in various
reaction mixtures, four reverse transcriptase-related M13
cloneswerelabeled and usedtoprobeblots ofgels
contain-ing electrophoretically resolved PCR products. An experi-mentinvolvingblots of thegelsshown in Fig.4 ispresented in Fig. 6. The four reverse transcriptase-related cloned
probes are as follows (see Table 1): (i) A.12, an HTLV-I
high-stringency-primed fragment containing both monkey
andHTLV-I-like characteristics; (ii) B.3,an HTLV-II high-stringency-primed fragment containing some monkey char-acteristics but quite different from any reported reverse
transcriptase; (iii) C.1,anMOPmoderate-stringency-primed
HTLV-E3
on November 10, 2019 by guest
http://jvi.asm.org/
[image:6.612.143.481.72.172.2]TABLE 1. Predicted amino acid sequences encodedby cloned PCR-amplified DNAsa
Anneal- No. of
Template ing indepen- Primer Internal amino acid sequence
DNA temp dent
(OC) clones
A.1CEM 37 A.2CEM 55 A.3CEM 55 A.4HeLa 55 A.5 HeLa 55 A.6CEM 55 A.7CEM 60 A.8CEM 60 A.9 HeLa 60 A.10 HeLa 60 A.11 HeLa 60 A.12TC 60 A.13 PTL 60 A. 14 PTL 60 A.15 PTL 60 A.16 PTL 60 A. 17 PTL 60
0 00000 00 000 00 0
1 MOP MLNSPTICQTYVGKVIKPVREQF*KCYSIH
1 HTLV-I MLNSPTLCQYFVGRVLQPVRDQFPRCYIVH
1 HTLV-I MLNSPTLCQYFVGRVLQPVRDQFPRCYIVY
3 HTLV-I MLNSPTLCQYFVGRVLQPVRDQFPRCYIVY
1 HTLV-I MLNNSTICQYYVGTILKPVRDQFPQCYVIH
2 HTLV-I MLNSIISACCRIGIKG--ASEYVSTAYIRH
1 HTLV-I MLNSPTIYQYFVGHGLQPVRDQFPRCYIVH
1 HTLV-I MLNSIISACCRIGIKG--ASEYLSTAYIRH
1 HTLV-1-HTLV-Il MLNSPTICQYFVGRVLQPVRDQFPRCYIVH 1 HTLV-I MLNSPTICQYFVGRVLQPVRISFPRCYIVH
1 HTLV-1 MLNSLTICQTYVRKAIK*VRE*FKKCYIIH
1 HTLV-I MLNSPTICQYFVGRVLQPVRDQFPRCYIVH
1 HTLV-I MLNSPTICQYFVGRVLQPVRDQFPRCYIVY 1 HTLV-I MLNSPTICQYFVGHVLQPVRDQFPRCYIVH
1 HTLV-I RLNSIISACCRIGIKG--ASDMFPTAYIRJH
1 HTLV-I MLNNSTICQYYVGTILKPVRDQFPQCYVIH 1 HTLV-I MLNSPTICQYFVGRVLQPCQGHVFRCYILH
FKNSPTLFEMQLAHILQPIRQAFPQCTILQ FKNSPTLFEQQLAAVLNPMRKMFPTSTIVQ MANSPTLCQKYVAAAIEPVRKSWAQMYIIH
MANSPTLCQKYVATAIHKVRHAWKQMYIIH
MKNSPTLCQKFVDKAILTVRDKYQDSYIVH MLNSPTICQTFVGRALQPVREKFSDCYIIH
HTLV-IHomolog HTLV-IIHomolog
SRV-IIHomolog MPMVHomolog
MMTVHomolog HuRRS-KHomolog
B.1CEM 37 B.2CEM 37 B.3CEM 55 B.4CEM 60 B.5 HeLa 60 B.6 HeLa 60
B.7TC 60
B.8 PTL 60
0 00 0000 00000o00 000
1 MOP FKNSPTIFGEALARDLQKFPTRDLGCVLLK
1 MOP FRDSSHLFGEALTRALSQFSYLD--TLVLW
2 HTLV-I1 FRDSPHYFGQALQLDLSQLHLQ-PS-ILLQ 1 HTLV-I-HTLV-I1 FRDSPHYFGQALQLDLSQLHLQ-PS-IILQ 1 HTLV-I-HTLV-I1 FRDSPHYFGQALQLDLSQIHIQ-PS-ILLQ 1 HTLV-lI FRDSPHYFGQALQLDLSQLHLQ-PS-ILLQ 1 HTLV-I FKDSPLYLASISPR-LEPILI--PDTFVLQ
1 HTLV-I FKDSPLYLASISPR-LEPILI--PDTFVLQ
WKGSPAIFQSSMTKILEPFKKQNPDIVIYQ FKNSPTLFDEALHRDLTDFRTQHPEVTLLQ FKNSPTLFDEALHRDLADFRIQHPDLILLQ FKNSPTIFGEALARDLQKFPTRDLGCVLLQ
CA CEM 37 3 MOP
HIVHomolog
BaEVHomolog MoMLV Homolog
HuRRS-EHomolog
000 00
RGLSCL-FII*LCCGKTCFFHAQAICYADTLGNWSFGSGK
VGLSP--FLLAQFTSAICSVVRRAFPHCLAFS
CPLSPLLFNIVLEVLARAIRHELEIKKIQLGKEEVKLSL
HBVHomolog
Li Homolog
D.1MU-S 37
D.2 MU-S 37
D.3 MU-S 37 D.4MU-S 37
3 MOP
1 MOP
1 MOP
1 MOP
000000000 0 000 0
MSNSPTMCQLYMQEALLPVREQFPSLILLL MSNSPTMCQLYVQEALLPVREQFPSLILLL
4-MKNSPTLCQKFVDKAILTVRDKYQDSYIVH
*-MANSPTMCQLYVGKAVEPIRKEYPKLRCVH
MKNSPTLCQKFVDKAILTVRDKYQDSYIVH MSNSPTMCQLYVQEALLPVREQFPSLILLL
MMTVHomolog MU-IAP Homolog E.1 CEM 37
E.2 TC 60
3 MOP
2 HTLV-I
0 0000 00 00 00 000 0000
MSNSPTICQTYVGQAIEPTRKKFSQCY-IH MLNSPTVCQIYVRKAILPVREQFKKCYIIH
MANSPTLCQKYVAAAIEPVRKSWAQMYIIH
F.1CEM 37
F.2CEM 37
F.3CEM 37
F.4CEM 37
F.5CEM 60 F.6HeLa 60
F.7HeLa 60
SRV-IIHomolog
1 MOP RSRG*RTVNHAFYLCLPAAGGLWCSLEVDF*WGSL
1 MOP KKDAPSPPSAAIFLYCKQHDSDANQPQIV
1 MOP FSSPR*VYR*VLALR*
1 MOP SQGEAGEVGQRSHSY
1 HTLV-I RLNSLFVKLMSGKLLSQLENSSKNGILS
1 HTLV-I-HTLV-II FINSLLCVIILFGENLIAFHIRQIHHWS
1 HTLV-II YQCSSPLLHFIHQVN*RILIAFPCQKMPHWS
70
A3 = A4 A9 = A12 A5 = A16
B3 = B6
on November 10, 2019 by guest
http://jvi.asm.org/
(B.1)
---__---__
**** ** * * *
CEM/ 37/MOP TGGAAAGTGTTGCCACAAGGG TTCAAGAACTCCCCCACCATCTTCGGGGAGGCATTGGC 1111 -111 111 11 11111111111111111111111111111 11111 HuRRS-E HOMOLOGUE TGGACCCAGCTTCCCCAAAGG
TTCAAGAACTCCCCCACCATCTTTGGGGAGGCGTTGGC
* *
TCGAGACCTCCAGAAGTTCCCACCAGAGACCTAGGCTGCGTGTTGCTCAAG TACGTGG
1111111I*I 1111111111111111 11111111111111111
IITi
TCGAGACCTCCAGAAGTTTCCCACCAGAGACCTAGGCTGCGTGTTGCTCCAG TACGTTG
** ****
ATGACCTGCTGCTGGCCGCC 1111111
H11IM1
ATGACCTTTTGCTGGGACAC
---__---__
(D.2)
---__---__
*** ~~~**
MU-S/37/MOP TGGAAAGTGCTGCCACAGGG AATGTCCAATAGTCCTACTATGTGTCAACTTTATGTGCA
11111111
1Illi
11111 111111MU-IAP HOMOLOGUE TGGAAGGTCTTACCACAGGG AATGTCCAATAGTCCTACAATGTGCCAACTTTATGTGCA
AGAAGCTCTTTTGCCAGTGAGGGAACAATTCCCCTCTTTAATTTTGCTCCTT TACGTGG
1111111111111111111111111111111111111111111111111111
11AGAAGCTCTTTTGCCAOTGAGGGAACAATTCCCCTCTTTAATTTTGCTCCTT
TACATGG* * **
ATGACCTCCTGCTGGCC ___11111 III
ATGACATCCTCCTGTGC
(D.3)
---__---__
* * *
MU-S/37/MOP TGGAAAGTGTTGCCACAGG GTATGAAAAATAGCCCTACTTTATGTCAAAAATTTGTAGA 11111 1111
IIIIIIIIIIIIIIIIIIIIIIIIIIIIIIIIIIIII11111 I1
MMTV HOMOLOGUE TGGAAAGTTTTGCCCCAGG GTATGAAAAATAGCCCTACTTTATGTCAAAAATTTGTGGA
* * * *
TAAAGCTATATTGACTGTAAGGGATAAATATCAAGACTCATATATTGTGCAC TACGTGG
11111111111111111111111111II11111111111111111
IllIlI111l
CAAAGCTATATTGACTGTAAGGGATAAATACCAAGACTCATATATTGTGCAT TACATGG
* * ** ***
ATGACCTGCTGCTGGCCGCC 11111 1 11
1III
ATGACATTCTTTTGGCACAC
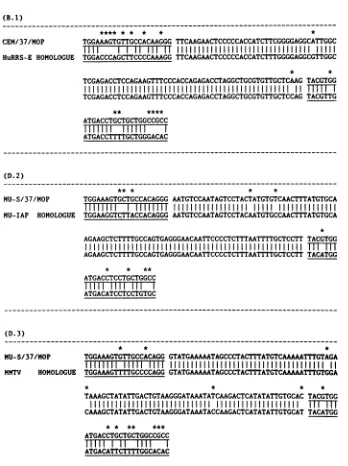
FIG. 5. Comparison of the nucleotide sequences of three PCR-amplified DNA fragments and three previously cloned chromosomal sequences to which theyare highly homologous. The sequenced fragments correspond to those indicated by arrows in Table 1, but the nucleotide sequences include the upstream and downstream primers in addition to theinternalregion translated in Table 1.(The chromosomal homologsareshown in Fig. 1 and described in the legend to Fig. 1). Primer regions are underlined, sequence mismatches are indicated by
anasterisk, and sequence identities are indicated by vertical lines. fragment containing possible L-1-like characteristics; and (iv) E.2, an HTLV-I high-stringency-primed fragment con-taining monkey characteristics. Assuggested by the data of Table 1, the results oftheexperiments shown in Fig. 6 (as
well as those ofother experiments with these same filters afterstripping[60%formamideat70°C])indicateahighlevel ofprimer selectivity.Forexample,
referring
tothe terminol-ogy ofFig.
6, the L-1-like sequence (C.1) was present inaSequencesareorganizedin groups(Athrough F)based onsimilaritiestoknown reversetranscriptase homologs,and the sequencesofthe latter areincluded forreferenceatthe endof eachgroup.Gaps (dashes)havebeen introduced into thesequences tomaximizehomologies;starsindicate terminationcodons. The opencirclesatthetopof eachgroup representpositionsthat areoftenhelpfulindiagnosing this region ofthe reversetranscriptasegene. Note that group F represents sequenceswithnoclearreversetranscriptasecontext.Forabbreviations ofprimersandhomolog,seethelegendtoFig.1.The L-1homologrepresents thecompilation sequence of SingerandSkowronski (53). TemplateDNAs wereisolated from thefollowing: CEM,ahuman T-celllymphoma-derivedcellline (5); HeLa,ahumancervicalcarcinoma-derived cellline; TC,abacteriophagelambda cDNAlibrary prepared fromahumanteratocarcinoma-derivedcell line(54); PTL,abacteriophage lambdacDNAlibrary prepared from normalhumanrestingTcells (22); andMU-S, total RF/J strainmousespleen.Thearrowsindicate cloneswhoseentire nucleotidesequences arepresentedinFig.5.In groups A andB,identicalnucleotide sequencesthatweregeneratedfromdifferenttemplates
and/orannealing temperaturesarealsoindicated(for example,A.3 = A.4).
on November 10, 2019 by guest
http://jvi.asm.org/
[image:8.612.140.488.61.527.2]72 SHIH ET AL.
A B C E
rie .-I
<> , , -1 1 E ". :;se
)-3tE
3..'-~ Woslsm
B R K BR K B R K B RK
23Kb- e
964Kb-
*6.5Kb-~.
4t4Kb- * .
-23Kb-
I
2QKb-its.2
C,l.v- 2)
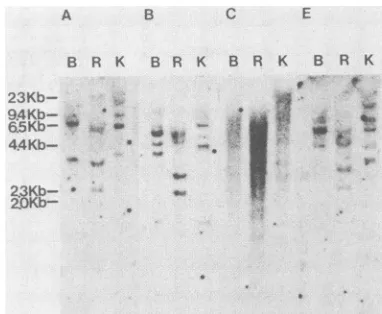
FIG. 7. Southernblotanalysisofhuman CEMDNAprobedwith 2'l6t,;:-J* e ,,>t;t,1,
~~~~four
cloned PCR amplified, reversetranscriptase-related,
DNA1?:1i;-
Upit9,,-
' - 1:fragments (see legend toFig. 6 and the text). Restrictionendonu-clease-digestedDNAsamples (10
,ug
perlane)wereelectrophoresed for 11 h in 1% agarose at2V/cm,transferred tonitrocellulose, and hybridized as previously described (see Materials and Methods).Eachpanelshowstheautoradiographofafiltercontainingthesame
K'F A DNA (CEM), digested with the same enzymes (B,
BamHI;
R,FIG. 6. Southern blot analysis of PCR-amplified DNAs. The gels EcoRI;K, KpnI) buthybridized todifferent probes as follows: A, photographed for Fig. 4 were blot transferred to nylon filters A.12;B,B.3;C,C.1; E,E.2. Autoradiographs wereexposedfor15 (Biotrans; ICN Pharmaceuticals Inc.), and cut sections of these h.
filters were hybridized separately to one of four cloned
PCR-amplified, reverse transcriptase-related DNA fragments. Lane des- . .
ignations and groupings shown on the top of each autoradiograph
containing
electrophoretically
resolved
restriction
endonu-are as inFig. 4; labeled lines within each autoradiograph span the clease-digested DNAs. All of the reverse transcriptase-regions hybridizedto probe A.12, B.3, C.1, or E.2 (Table 1, see the related cloned
probes
detectedrepetitive
sequencesinchro-text). The autoradiographs correspond to the following hybridiza- mosomal DNA (Fig. 7). In the case of the monkey tions. MOP was probedwith A.12; MOP (after stripping of the filter retroviruslike probes (A.12, B.3, and E.2), bands with signal
probed withA.12) probed with C.1 (note that the material hybrid- intensities corresponding to 10 to 100 copies per
cell
wereizing to C.1migrated more slowly than did that hybridizing to A.12, evident, and there were definite differences in the
patterns
as expected for the larger size of group C clones). HTLV-I lane obtained with different probes. In the case of the L-1-like CEM,37 was probed with B.3, lanes HM20,37 to CEM,55 was
probed with A.12, and lanes CEM,60 and HELA,60 were probed probe
(C.1),
bands and smears withcombinedsignal
inten-with E.2.HTLV-1I
lanes CEM,37 to HM20,37 were probed with sities ofmorethan 100copies
percellwereevident,
andtheA.12, and lanes HELA,45 to HELA,60 (right) were probed with pattern is
suggestive
of along
interspersed
nucleotidese-B.3.Autoradiographswere exposed for 1 h. quence
family.
Blotted humanscalp
orHeLa DNAsyielded
similar if not identical patterns when hybridized to each of these probes (data not shown). Interestingly, C.1exhibited MOP-primed reaction mixtures but not inHTLV-I-primed
no obvious homology toany region
ofan L-1compilation
mixtures (data not shown), whereas the HTLV-I-like se- sequence (53) or to any published primate L-1 sequences. quence (A.12) was present in both. Also, the monkeylike The results of the genomic Southern blotting experiments sequence (E.2) was present in HTLV-I- but not HTLV-11- presented here suggest that many of the reverse transcrip-primed reaction mixtures (data not shown), whereas the tase-related sequences amplified by PCR are members of rather novelsequence (B.3) was present inHTLV-II-
but not multigene subfamilies. In addition, since most of the bands HTLV-I-primed mixtures. (The nature of the lower band of in Fig. 7 are significantly larger than the reverse transcrip-the doublet in the 123-base-pair range, observed by both tase coding region, it is likely that each reverse transcriptase blotting and UV absorption in many PCR products, is is part of alargerrepetitive elementprobably due to the amplification of slightly shorter
tem-plates asreflected by products such as A.6 or B.7, shown in DISCUSSION Table 1.) Due to theefficiency of amplification after the first
successful priming reaction and the reasonable selectivity of The human genome apparently contains a wide spectrum mixed oligonucleotide priming at 37°C (MOP clones), it is of sequences that are closely related to the reverse transcrip-probable that additional blotting studies will reveal isolated tase genes of retroviruses, and it is assumed that most of cases ofunexpected priming. In any event, the data shown in these sequences are part of defective proviral DNAs. In this Table 1 and Fig. 6 indicate that high-stringency-primed communication we show that synthetic oligonucleotides amplificationsgenerally yield specific and distinct families of homologous to highly conserved regions of reverse tran-related sequences. scriptase genes are particularly effective in detecting such
DistributiopD
of some reverse transcriptase-related cloned sequences when used as primers in PCR amplification. DNAfragmentsingenomic DNA. To investigate the distribu- These primers may be either unique(HTLV-I
andHTLV-II;
tion of some specific amplified PCR fragments in human Fig. 1), or mixed (MOP; Fig. 1), and the nature of the DNA, the four reverse transcriptase-related clones de- amplified sequences appears to depend on both the primer scribedabove were labeled and used to probe blots of gels sequence and annealing temperature (although the relativeJ. VIROL.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:9.612.339.530.76.233.2] [image:9.612.60.298.77.311.2]importance of each has not been
investigated
systemati-cally).
Surprisingly, MOP amplification of
genomic
DNA orcDNA (21) is certainly a more productive approach to identifying reverse transcriptase sequences than is conven-tionalmixed-oligonucleotidescreening of
genomic
orcDNA libraries.Amplification
ofa true sequenceis lessdependent ontheextentofprimer-target homologythanisthedetection ofatruepositivesignal in oligonucleotideprobing.
Published data on MOP-amplified cDNA (21) and the results shown here (Fig. 5) on MOP-amplified genomic DNA suggest that primer-target homologies of less than 70% can amplify expected sequences, atleastwith 37°C annealingand either 37 or72°C extension. It is assumed that thisfidelity
ofPCR is due primarily to the requirement for two independent primer-target interactions in specific orientation within a defined distance, and that the use of increasedannealing
temperaturesand/or uniqueprimers would limitthe compo-sition ofthe amplified products.Indeed,
uniqueoligonucle-otide primers of20residues and
annealing
temperatures of550C
supposedly require perfect or nearlyperfect
primer-target matching for successful amplification (48).
(The
an-nealing limit for our HTLV-I and HTLV-IIprimers
was between 65 and70°C;
thelimitofour mixedprimers
wasnot tested.)The nature of the human reverse
transcriptase-related
amino acidsequencesdetected byPCR inboth
genomic
and cDNAs was quite varied (Table 1).Neglecting
putative
L-1-likesequences(groupC) andthose sequences innoclear context(groupF), one might distributethe
remaining
clones between two major mouse-related classes, onebroadly
ho-mologous to MMTV (including groups A and E) and the otherbroadly homologoustoMoloney
MLV(group
B).
This classification is reasonably consistent withprevious
studies of human endogenousretroviruses,
whichsuggested
that MMTV- andMoloney
MLV-relatedspecies
account for thousands of humanproviral
DNAs.However,
it does not take into account otherstriking
amino acid sequence rela-tionships,nordoesit considerthemorelikely
origin
ofmanyor most human endogenous retroviruses or retrovirus-re-lated genes, which would be
primate
rather than murine.Many of the amino acid sequences shown in Table 1 corresponding to
HTLV-I-primed,
high-stringency (55
and 60°C)PCR
amplifications
revealedstriking
similarities toreverse transcriptases of
primate
retroviruses.Specifically,
thesearetheFPQC-likeoridenticalblocksnoted
previously
formost
genomic
and cDNA membersofgroupA(related
to the HTLV-I homolog), 18 out of 30 amino acid identities noted for group E(related
to the simian retrovirus type II [SRV-II] homolog), andpossibly
the LEP block noted for the two cDNA-derived(TC,
PTL)
members of group B (relatedtothe HIVhomolog).
Thesehomologies
are admit-tedly limited, as is the entireregion
here understudy,
but whenconsidered inconjunction
with theiramplification by
veryhigh-stringency
priming
withprimate-specific
oligonu-cleotides and present views
regarding
the evolution of ret-roviruses(28),manyorall of thesereversetranscriptase-like regions may be primate related.Evolutionarily,
MMTV appearstobecloselyrelatedtoSRV-I,
SRV-II,
andMason-Pfizer monkey virus, whereas
Moloney MLV,
itselfquite
distantlyrelatedtoMMTV,is very
closely
relatedtobaboon endogenous virus. At least in terms of the amino acid sequencespresentedin Table 1, groupsA and B canjust
aseasilyberelatedtotheir
monkey
homologs
(SRV-II/Mason-Pfizer
monkey
virus and baboonendogenous
virus,
respec-tively) as to their murinehomologs;
as notedabove,
theinclusion ofHTLV-I and HIV
homologs
oftenstrengthens
the
primate
relationship.
Asmight
beexpected
fromgenetic
drift and recombination(including
geneconversion)
amongpresumed
predominantly
defective humanproviruses,
thisrelationship
is notstraightforward
and may bequite
com-plex.
Thepeculiar
Mason-Pfizermonkey
virus/HTLV-I
hy-bridlikecharacterofmany group A sequences
(such
asA.16)
could reflect this
complexity,
as could the MMTV/SRVhybridlike
character of HuRRS-K notedby
Onoet al.(37).
Eventually,
additional PCRamplifications
with differentprimers
andstringencies,
characterizations ofgenomic
li-brary
clones,
andcross-probed
Southerngenomic
blots of manyprimate
DNAs should reveal theevolutionary
history
ofhuman
endogenous
retrovirusesand/orrelatedsequences. If human andmonkey
retrovirusesdiverged
and evolvedrelatively
independently,
it ispossible
that many humanendogenous
retrovirus-related sequences are derived fromhuman viruses.
Irrespective
of theorigin
of humanretroviruses,
their presence leads to bothpractical
and theoretical concerns.Presently,
themajor
practical
concern is that the effectiveuseof PCRas a
screening procedure
forHTLV-I,
HTLV-Il,
and HIVinfectionsmust
always
includeappropriate
controls to ensurethat noendogenous
sequencescontribute to posi-tivesignals.
Aspreviously noted,
HIVunique
primers
corresponding
tothehighly
conservedreversetranscriptase
region
showninFig.
1function well in the PCRamplification
of HeLa DNA even atannealing
temperatures
of60°C.
Although
it isunlikely
that the internal nucleotidesequence(s)
would be detected afterstringent
hybridization
to anHIV-specific probe,
other sites in HIV areprobably
more
appropriate
foritsunambiguous
detectionby
PCR. Anotherpractical
concern is that the use of PCR fordetermining
thepossible
retroviraletiology
ofavariety
ofhumandiseases may be
complicated by endogenous
retrovi-ruses. Even if cDNAs are used for PCR
templates,
thetranscriptional
activities ofendogenous
sequences must be considered.The
major
theoretical concern related to thepresence ofendogeneous
human retroviruses is theirpotential
expres-sion.Inanalogy
with murinesystems,
thefollowing might
beexpected: (i)
that such sequences in humans will be classedas
low-,
high-,
or, mostoften,
non-virus-producing
andthat this classification willdepend
on the nature of both theprovirus
and thehost;
(ii)
that some humanendogenous
proviral
DNAs that aredefective nonvirusproducers might
yield
infectious and/oroncogenic
recombinants withexoge-nous viruses or with otherdefective ornondefective
provi-ruses;
(iii)
that some human sequencesmight
bepartially
transcribed and translated intofunctionalgeneproducts;
and(iv)
thatsomeendogenous
elementsmight
beresponsible
forknown or new mutations as well as for the insertional activation of oncogenes, as are some human L-1 elements
(16, 32).
In
addition,
it is alsopossible
that the human genome containsreversetranscriptases
that are not associated withendogeneous
viruses. The use of MOPamplification
ofcDNA
[either
from cloned cDNAsorfromreversetranscrip-tase-copied poly(A)+
RNAs]
coupled
with cDNAlibrary
characterizations,
shouldprovide
areasonably
rapid
ap-proach
toinvestigating
reversetranscriptase
geneexpres-sion.
Investigations
with ourhumnan
teratocarcinoma andT-cell cDNA PCR
amplification products
are inprogress. Inconclusion,
theimportance
ofreversetranscription
inregard
to thedevelopment
and evolution of mammalian genes and itspresent
importance
as a mechanism of DNAon November 10, 2019 by guest
http://jvi.asm.org/
74 SHIH ET AL.
transposition suggest that a variety of reverse transcriptase genes, including those in unexpected contexts,arepresent in humans and mice. Moreover, the homology relationships of human putative reverse transcriptases identified by PCR amplification suggest that many or most of themare related to primate retroviruses. Such a relationship, if meaningful, impliesan ancient
(>105
years)associationbetween retrovi-ruses and humans. It is possible that the proposed transfer of anHTLV-I-like monkey virus to humans in the 16th century and of an HIV-like monkey virus to humans in the 20th century merely represent the most recentdetectableexam-ples of monkey-human cross-infections. It is also possible that both human and monkey exogenous retroviruses have beenevolving relatively independently (8),and thatmany of the human endogenoussequences were derived from human retroviruses.
ACKNOWLEDGMENTS
Wethank Horace Lozina and HeinzAnnusfor excellent technical help,RogerKaress, PeterD'Eustachio, HannahKlein,and Robert Schneider for helpful discussions and support, Steve Sloane for assistance with the polymerase chainreaction,LaraSchulman and Ross Smith for help withcomputeranalysis, and Aileen Siegel for typing the manuscript.
Thisworkwassupported by National Science Foundationgrant
PCM8203006 andby grantBRSGS07RR5399-25 of the Division of ResearchResources, National Institutes of Health. A.S. andR.M. arepredoctoral traineessupported by PublicHealth Servicetraining grants 1GM07827 and
5T32GM07238208,
respectively, from the National Institutes of Health.LITERATURECITED
1. Callahan,R., I. M.Chiu, J.H.F.Wong,S.Tronick,B. A.Roe, S.A.Aaronson,andJ. Schlom. 1985.Anewclass ofendogenous
human retroviralgenomes. Science 228:1208-1211.
2. Callahan, R., W. Drohan, S. Tronick, and J. Schlom. 1982. Detection andcloning of humanDNA sequencesrelatedtothe
mouse mammary tumorvirusgenome. Proc. Natl. Acad. Sci. USA 79:5503-5507.
3. Chiu, I.-M.,A.Yaniv, J. E.Dahlberg, A.Gazit,S. F.Skuntz,
S. R. Tronick,andS. A. Aaronson. 1985. Nucleotidesequence
evidence for relationship of AIDS retrovirus to lentivirus.
Nature(London) 317:366-368.
4. Finnegan, D. J.1985.Transposable elements in eukaryotes.Int. Rev.Cytol. 93:281-326.
5. Foley, G. E., H. Lazarus, S. Farber, B. G. Uzman, B. A. Boone, and R. E.McCarthy. 1965. Continuous cultureofhuman lym-phoblasts from peripheral blood ofachild withacuteleukemia. Cancer 18:522-529.
6. Franck,A., H. Guilley, G.Jonaro, K.Richards, and L. Hirth. 1980. Nucleotide sequence of cauliflower mosaic virus DNA.
Cell 21:285-294.
7. Franklin, G. C., S. Chretien, I. M. Hanson, H. Rochefort,
F.E. B. May, and B. R. Westley. 1988. Expression of human
sequencesrelatedtothose ofmouse mammarytumorvirus.J. Virol. 62:1203-1210.
8. Fukasawa, M., T.Miura,A.Hasegawa,S.Morikawa,H. Tsuji-moto, K.Miki, T. Kitamura, and M. Hayami. 1988.Sequence of simian immunodeficiency virus from Africangreenmonkey,a newmember of theHIV/SIVgroup. Nature(London) 333:457-461.
9. Gattoni-Celli, S., K. Kirsch, S. Kalied, and K.J. Isselbacher.
1986. Expression oftype C-related endogenous retroviral se-quences in human colon tumors and colon cancer cell lines.
Proc. Natl.Acad. Sci. USA83:6127-6131.
10. Gonzalez,I.L., J. L. Gurski, T. J. Campen, D. J. Dorney, J. M.
Erickson, J.E.Sylvester,and R. D. Schmickel. 1985. Variation
amonghuman28S ribosomalRNAgenes. Proc. Natl.Acad.Sci.
USA 82:7666-7670.
11. Graham, F. L.,J. Smiley, W.C. Russell, andR.Nairn. 1977.
Characteristics ofahuman cell line transformedbyDNA from humanadenovirus types. J. Gen. Virol. 36:59-72.
12. Harada, F.,N.Tsukada,and N. Kato. 1987. Isolation of three
kinds of human endogenous retrovirus-like sequences using tRNAPROas aprobe.Nucleic Acids Res. 15:9153-9162. 13. Hattori,M., S.Kuhara,0.Takenaka,and Y. Sakaki. 1986. Li
familyofrepetitivesequences inprimatesmay be derived from
a sequence encoding a reverse transcriptase-related protein. Nature(London)321:625-628.
14; Kato, N.,S.Pfeifer-Ohlsson,M. Kato,E. Larsson,J. Rydnert, R.Ohlsson, and M.Cohen. 1987.Tissue-specific expressionof human provirusERV3 mRNA in human placenta: two of the
threeERV3 mRNA contain human cellular sequences. J. Virol. 61:2182-2191.
15. Kato, S., K. Matsuo, N. Nishimura, N. Takahashi, and T. Takano.1987. The entire nucleotidesequenceof baboon
endog-enousvirus DNA: achimericgenomestructureof murinetype C andsimiantype Dretroviruses. Jpn.J. Genet. 62:127-137. 16. Kazazian, H. H., Jr., C. Wong, H. Youssoufian, A. F. Scott,
D.G. Phillips, and S. E. Antonarakis. 1988. Haemophilia A resultingfrom denovo insertionof Li sequencesrepresentsa
novel mechanism for mutationinman. Nature (London) 332: 164-166.
17. Kobayashi, M., and K. Koike. 1984. Complete nucleotide
se-quenceofHBV DNAofsubtypeADRand its conserved gene organization.Gene 30:227-234.
18. Kroger, B., and I. Horak. 1987. Isolation of novel human
retrovirus-related sequences by hybridization to synthetic oli-gonucleotides complementary to the tRNAPro primer-binding
site.J. Virol. 61:2071-2075.
19. Kwok,S.,D. H. Mack,K. B. Mullis,B.Poiesz,G. Ehrlich,D.
Blair,A.Friedman-Kien,andJ. J.Sninsky.1987. Identification ofhumanimmunodeficiencyvirussequencesbyusingin vitro
enzymatic amplification and oligomer cleavage detection. J.
Virol. 61:1690-1694.
20. Lathe,R.1985.Synthetic oligonucleotide probesdeduced from amino acid sequence data: theoretical andpractical consider-ations.J. Mol.Biol. 183:1-12.
21. Lee,C.C.,X.Wu,R. A.Gibbs,R.G. Cook,D. M.Muzny,and C. T. Caskey. 1988. Generation of cDNA probes directed by
amino acid sequence: cloning ofurate oxidase. Science 239:
1288-1291.
22. Littman, D. R., Y. Thomas, P. J. Maddon, L. Chess, and R. Axel. 1985.Theisolation andsequenceofgeneencodingT8:a
moleculedefiningfunctional classes ofTlymphocytes.Cell40:
237-246.
23. Loeb,D.D.,R. W.Padgett,S. C.Hardies,W.R.Shehee,M. B. Comer,M. H.Edgell,andC. A. Hutchison.1986. Thesequence
ofalargeLlMd element revealsatandemlyrepeated5'end and several features found inretrotransposons. Mol. Cell. Biol.6:
168-182.
24. Mager, D. L., and J. D. Freeman. 1987. Human endogenous retrovirus-like genome with type C pol sequences and gag sequences related to human T-cell lymphotropic viruses. J.
Virol.61:4060-4066.
25. Mager, D. L., and P. S. Henthorn. 1984. Identification ofa
retrovirus-likerepetitiveelement in human DNA. Proc. Natl. Acad. Sci. USA 81:7510-7514.
26. Maniatis, T.,E. F. Fritsch,andJ. Sambrook. 1982. Molecular cloning:alaboratorymanual. ColdSpringHarborLaboratory, ColdSpring Harbor,N.Y.
27. Martin, M. A., T.
Bryan,
S. Rasheed, and A. S. Khan. 1981.Identification andcloning ofendogenous retroviral sequences present in human DNA. Proc.Natl. Acad. Sci. USA 78:4892-4896.
28. McClure,M.A.,M.S.Johnson,D.-F.Feng,and R. F.Doolittle. 1988. Sequence comparisons of retroviral proteins: relative
ratesofchange andgeneral phylogeny. Proc. Natl. Acad. Sci. USA 85:2469-2473.
29. Mietz, J.,Z. Grossman,K. K.Lueders, and E. L. Kuff. 1987.
Nucleotide sequence of a complete mouse intracisternal A-particle genome: relationship to known aspects of particle assemblyandfunction.J.Virol.61:3020-3029.
J. VIROL.