0022-538X/94/$04.00+0
Copyright© 1994, American Society for Microbiology
DNA
Strand Exchange and Selective
DNA
Annealing
Promoted
by the Human
Immunodeficiency Virus Type
1
Nucleocapsid Protein
ZENTATSUCHIHASHIt ANDPATRICK0. BROWN*
HowardHughes MedicalInstitute andDepartment ofBiochemistry, Stanford University, Stanford, Califomia 94305
Received31 March 1994/Accepted 7 June 1994
Nucleocapsidprotein (NC) ofhumanimmunodeficiencyvirustype 1 (HIV-1) wasexpressed inEscherichia
coli and purified. Theprotein displayed a variety ofactivities on DNAstructure, all reflectingan ability to promotetransitionbetweendouble-helicalandsingle-stranded conformations.We found that, in additiontoits previouslydescribedabilitytoacceleraterenaturationofcomplementaryDNA strands, the HIV-1 NC protein couldsubstantiallylowerthemeltingtemperatureofduplexDNA and couldpromotestrand exchange between double-stranded andsingle-strandedDNAmolecules.Moreover,inthepresenceof HIV-1 NC, annealingofa single-strandedDNAmoleculetoacomplementaryDNA strand that would yielda morestabledouble-stranded
productwas favoredoverannealing toalternative complementary DNA strands that wouldform less stable
duplexproducts (selective annealing). NC thusappearstolowerthekinetic barriersothat double-strand<-> single-strand equilibrium is rapidly reached tofavor the lowestfree-energynucleic acid conformation. This
activityofNCmaybeimportant forcorrectfolding of viral genomic RNA andmayhavepractical applications.
Nucleocapsid (NC) proteins are small structural proteins present in the cores ofall retroviruses (51). In a virion, the
dimeric viral RNAgenome is thoughttobe coveredwith NC proteins. NC proteins are basic, typically include one or two zincfinger-like motifs, andcan bind toeither DNA orRNA,
with apreference forsingle strands (13, 24, 25, 28, 29, 31, 37, 46, 48, 52).NC protein is synthesized as apart of the Gagor Gag-Pol precursor polyprotein. These polyproteins assemble into virus particles andpackage the viral RNA. The
polypro-teins are subsequently cleaved by the viralprotease to form
mature proteins, including NC protein (51). Viruses with mutations intheregion encoding NC showdisrupted packag-ing of viral RNA, without other apparent defects in viral particle formation (4, 14, 16-20, 32, 33, 38).Inthecaseofavian
leukosisvirus,it isnotyetclear whethermutations inCys-His motifs directly block packaging of its genome or reduce the
stability ofpackaged RNA(5). These results have led to the
proposal that the NC regionoftheGag polyprotein is
respon-sible forrecognitionof thepackagingsignalsonviralRNAs. In
support of thisnotion, purified Gag polyproteinaswellasthe
NCprotein ofhumanimmunodeficiencyvirustype 1(HIV-1)
boundHIV RNAwithsomespecificity (6, 30).
In a test tube, NC protein has been shown to promote
annealing ofprimertRNA and DNAoligonucleotidestoviral RNA and to promote dimerization of model viral genomic
RNAs containing a dimer linkage sequence (7, 12, 42, 43).
Recently, a DNA renaturation activity of HIV-1 NC was described, and its kinetic properties were characterized (13).
NC has also been shown to stimulate both formation of strong-stopcDNAand strandtransferbyreversetranscriptase (28).
Inthisstudy,wetested thepossibilitythat HIV-1NCcould promote transfer of DNA strands between alternative DNA
*Corresponding author. Phone: (415) 723-0039. Fax: (415)
723-1399.
tPresent address: Somatix Therapy Corporation, Alameda, CA
94501.
duplexes by virtue of its DNA-annealing and DNA-melting
activities. We found that NC could overcome the kinetic
barrier to equilibrium between alternative pairs of
comple-mentarystrands. Thisactivity mightplay importantroles inthe viral life cycle.
MATERIALS AND METHODS
DNAoligonucleotides.DNAoligonucleotideswereobtained
from the PAN facility (Stanford University) and Operon (Alameda,Calif.).Thesequencesof theseoligonucleotidesare as follows: 24A-, ACT GCT AGA GAT TTlT CCA CAA
GTC; 24A+, GACTTG TGG AAA ATC TCT AGCAGT;
28X-, TCGAAC TGC TAG AGA lFlTl TCCACA GACT; 21A-, ACTGCT AGAGAT TTTCCA CAT;21A+, ATG
TGG AAA ATC TCT AGCAGT; 28A-,ATG CAC TGC
TAG AGA lTTTTCC ACA AGTC;28A+, GACTTGTGG AAAATCTCT AGC AGT GCA T; and32A-, ACTGCT AGA GATTl?TCCA CAT AGT ATC GAATT.
Preparation of HIV-1 NC protein. Thegene encoding the 71-amino-acid form of HIV-1 NCwasamplified byPCR(19)
from pNL4-3 plasmid DNA (1). At the same time, codons encoding the sequence Met-(His)6 were added to the N-terminal end of the native NCgeneforthepurposeof one-step
purification using Ni-nitrilotriacetricacid resin(Qiagen,
Chats-worth,Calif.).Thishybridgenewascloned into the T7-7vector
(49) to make plasmid pHIVp7hisl. This plasmid was intro-duced intoEscherichia coliBL21(DE3),which has an isopro-pyl-l-thio-13-D-galactopyranoside (IPTG)-inducible T7 RNA
polymerasegene(47).Transformed cellsweregrownin 2 liters of LB medium with 70 ,ugof carbenicillinpermlat37°Cuntil
they reached an optical density at 595 nm of 0.5. IPTGwas
then added at 1 mM, and incubation was continued for 2 h before harvest. Harvested cellsweresuspendedin 10 ml of 50 mM Tris-Cl (pH 7.5)-10% sucrose (TS) and frozen in liquid nitrogen.Frozen cellswerethawed, and 4 ml ofTS, 1.5 ml of 50 mM Tris-Cl(pH7.5)-(0.3 Mspermidine-10% sucrose-O.1
MNaCl, 14 ,ulof ,B-mercaptoethanol-2MTrisbase, 20 ,ulof 5863
on November 9, 2019 by guest
http://jvi.asm.org/
0.25 M
phenylmethylsulfonyl
fluoride(PMSF),
0.35 ml of10-mg/ml lysozyme,
and 0.2 ml of 10% Triton X-100 wereadded. Aftersuccessive incubations at
0°C
for 50 min and at37°C
for 3min,the cellsuspension
wascentrifuged
inaSorvall SS34rotor at14,000
rpmfor 30 min.NCprotein
wasextracted from thepellet
with 10 mlof extraction buffer(50
mMTris-Cl[pH 7.5],
100mMNaCl,
15%glycerol,
0.1%TritonX-100,
10mM
3-mercaptoethanol,
0.25 mMPMSF,
6 Mguanidium-HCl),
and insoluble material was removed by centrifugation(Beckman
SW41 rotor, 35,000rpmfor 20min).
The superna-tant was loadeddirectly
onto a 1-ml column of Ni-nitrilotri-acetic acid agarose resin ina3-mlsyringe.
The resinwasthen washed with 40 ml of extractionbuffer with 1 mM imidazole and30 ml of bufferA(50
mMTris-Cl[pH 7.5],
100mMNaCl,15%
glycerol,
0.1%TritonX-100, 10 mM ,B-mercaptoethanol,0.25 mM
PMSF) plus
20 mM imidazole. Recombinant NCprotein
wastheneluted with 25 ml of bufferAplus
100 mM imidazole followedby
25 ml of buffer Aplus
200 mMimidazole.
Approximately
15 mg of NCwasrecovered,
andthepeak
fractions used in this workwerejudged
tobe >95%pureby
Coomassie brilliant bluestaining
of sodiumdodecyl
sulfate(SDS)-polyacrylamide gel.
Protein concentration was deter-minedby using
a Bradfordprotein
determination kit(Bio-Rad)
with bovineserumalbumin as astandard.Labeling of DNA substrates. The 93-mer DNA fragment
wasderived from
pT7-5
DNA(49) digested
withHindlll and PvuII. The 5' ends of this DNAfragment
werelabeled with[,y-32P]ATP
(ICN)
and T4polynucleotide
kinase(New
En-gland Biolabs) (44),
and the labeled DNAfragment
waspurified
by electrophoresis through
a10%polyacrylamide-7
Murea
gel
in Tris-borate-EDTA(TBE)
buffer(44).
Specificactivity
of the radiolabeled DNAwas estimatedby thin-layer
chromatography.
5'labeling
ofoligonucleotides
wasdone with[y-32P]ATP
(ICN)
and T4polynucleotide kinase,
and labeledfragments
werepurified through
aspin
column ofSephadex
G-15
(Pharmacia) (44).
DNA
annealing
assay.The standard reaction mixture(10
RI)
contained 50 mM Tris-CI
(pH 7.9),
1 mMEDTA,
1 mMdithiothreitol,
0.1%TritonX-100,
and 3 mMMgCl2
(standard
buffer for
annealing
assay).
32P-labeled DNA substrates andHIV-1 NCwere added as described for each assay. Double-stranded DNA
(dsDNA)
substrateswereinitially
heat dena-turedby
incubation at94°C
for 5 min inTE(10
mM Tris-Cl[pH
7.9],
1 mMEDTA)
and thenrapidly
chilledonice before addition tothe reaction mixture.Complementary
oligonucle-otideDNAswereadded
separately
atthestartof the reaction. After thereaction,
5,lI
ofstopping
solution(0.25%
bromo-phenol blue,
20%glycerol,
20 mMEDTA,
0.2%SDS,0.4 mg of yeast tRNA[Sigma]
perml)
wasadded,
and thesamples
were
electrophoresed
on a 10 or 15%polyacrylamide
gel inTBEat
4°C.
Thegel
wasthendried,
andanautoradiogramwasexposed
at-40°C. Quantitation
of theradioactivity
wasdone witha MolecularDynamics Phosphorlmager.
RESULTS
Stimulation of DNAannealing byHIVNC.Figure 1A shows the timecoursefor
annealing
of93-mercomplementaryDNAs withandwithoutpurified
recombinantHis6-tagged
HIV-1NCprotein.
Inthe absenceof NCprotein, only10% ofthe DNAhad renatured after 30 min, while in the presence of the recombinant
His6-tagged
NC protein preparation,renatur-ationwas almost
complete
atthefirst timepoint(10
s). Thus,thisNC
protein preparation
acceleratedannealing by atleast2,000-fold
under these conditions. Overexpressioninbacterial cells withahistidine tag, and theguanidium
chloridetreatmentA
75
0
.)
50
-25
oc
-0-- -NC
.--*- +NC
time(sec)
B
C 0
S.
a
0 0.1 1 10 100 1000
NCconc.(nM)
FIG. 1. (A) Time courseofannealingof93-mercomplementary
DNAs. The 12.5-,ulreaction mixture contained 50 mM Tris-Cl (pH 7.9),1 mMEDTA,and0.1% TritonX-100,with 1 nM heat-denatured 32P-labeled 93-mer DNAwithorwithout 1 puMHIV-1 NCprotein.
Afterincubation at 37°C for the indicated length oftime, reaction
mixtures were treated asdescribed in Materials and Methods, and
ssDNA andrenatured dsDNAwereresolvedby10%
polyacrylamide-TBE gel electrophoresis. (B)Titration of HIV-NC in an annealing
reaction.The indicatedamountof NCproteinwasaddedto12.5 pulof standard reactionmixture with 1 nM32P-labeled 93-mer DNA. After incubation at 37°Cfor 2min, sampleswere resolvedby gel electro-phoresisand theamountsof ssDNA anddsDNAweredetermined.
andrefolding protocolused forpurification, therefore didnot appeartocompromisetheprotein's activity. MgCl2stimulated thisreaction,withanoptimalconcentration of between 2and 4mM. Noeffect of ATPwasobserved(datanotshown).For 1 nM (as double-stranded molecules) of 93-mer DNA, full
activity required 11 nMNC under these conditions(Fig. 1B), oroneNC moleculeper17nucleotides.
Stimulation ofannealing byNCwasalso observed withshort
DNA oligonucleotides (Fig. 2A),but at high concentrations,
NC prevented annealing of these short DNAs (data not shown).Apossible explanationfor this inhibitionwasthatby bindingmorestronglyto single-strandedDNA(ssDNA) than
dsDNA, NC could shift the equilibrium toward the
single-r
,r-- I
8 8
%n § %n
on November 9, 2019 by guest
http://jvi.asm.org/
[image:2.612.317.556.64.459.2]A
NC (pmol) 0 30
time (min) 0 1 3 10
3 0 C 370C 30WC 37C
..I.1~ 7s F
-*--ds
4 s s
* -* * 24mer
.. 21mer
30
temperature (°C)
ds
ss
-35 45 55 25 35 45 55 30 40 50
40 50 60 30 40 50 25 35 45 55
;.
[image:3.612.137.485.70.505.2]. ~~~~~~~~W
FIG. 2. (A2 Effectof NConannealing of shortDNAoligonucleotides. A 10-pul reaction mixture contained standard reaction buffer with2fmol (0.2 nM)of3P-labeled24A+ oligonucleotideand10 fmol(1 nM) of unlabeled 21A- oligonucleotide withorwithout 30 pmol (3 ,uM) of NC.
24A+ and 21A- canform 20contiguous base pairs (see Materials and Methods). 21A- wasaddedat thestart of the incubation. After the reaction,thesamplesweretreatedasdescribedinMaterials andMethods and analyzedon a 15% polyacrylamide-TBE gel. (B) Effect of NCon
melting of short duplex DNA.A5-,ul reaction mixture containedastandard buffer and32P-labeled24A+ (1fmol,0.2nM)and unlabeled
24A-(5 fmol, 1nM). Oligonucleotides 24A+ and 24A- have the potentialtoformacomplete duplex (see Materials and Methods). ThesetwoDNAs wereincubatedtogetherat30°C for 10min before addition of NCtoallow all labeled 24A+toformaduplex. Indicatedamountsof NCwerethen
addedtothereactionmixture,andincubationwascarriedoutsuccessivelyat25°C for5min,30°C for 5min,35°C for5min,40°C for5min,45°C
for5min,50°Cfor5min,55°C for 5min, and 60°C for 5 min. After eachstepof incubation,onetesttubewastransferredtoice, and 5 ,ul of stopping
solutionwasaddedimmediately. Sampleswereanalyzedon a 15%polyacrylamide-TBE gel.
stranded form. Totestthishypothesis,excessNCwasaddedto apair ofpreannealed complementary 24-mer DNAs (0.2 nM 32P-24A+and 1 nM of24A-), and thetemperaturewasraised
in steps of5°C (Fig. 2B). In the absence of NC, the single-stranded form was detected only above 55°C, but in the
presence of 30 pmol (6 ,uM) of NC, substantial ssDNAwas
observed at 40°C, and at 150 pmol (30 ,uM) of NC, the single-stranded form was detected at 25°C. Thus, NC can
promotedenaturation ofadsDNAoligonucleotide.
Strandexchange reaction promoted by NC protein. Sincewe
found that NC could stimulate both single-strand--*double-strand (ss->ds) andds->ss transitions under different experi-mentalconditions,wetested whether both of these transitions could be promoted by NC under identical conditions, in the
sametest tube, by monitoringstrandexchangebetween
com-plementary double-stranded and single-stranded DNA oligo-nucleotides. In theexperimentshown inFig. 3A, a32P-labeled
21-mer double-stranded oligonucleotide (32P-21A+unlabeled 21A-) was mixed with a 50-fold excess of a single-stranded
32-mer (32A-) and incubated at 37°C in the presence or
1 3 10 30 1 3 10 30 30 1 3 10 30 0
B
NC (pmol)
0
-I
150
;;J
on November 9, 2019 by guest
http://jvi.asm.org/
A
*
21
(20fmol)
21
32
(lpmol)
2 : h ot
* 21 o
32
4-21
a U c a
B
[image:4.612.141.474.71.268.2] [image:4.612.350.517.485.625.2]28
|lI __--
(20fmol)
4 28< 21 \ (1pmol)
a b c d
\t~Il-.
2828
28
* -*-2
-21
FIG. 3. Strand exchange of short DNAspromoted byNC.(A)32P-labeled21A+/21A- duplex(20 fmol,2nM)and 32A-(1 pmol,100nM)
ssDNAsweremixed ina10-,ulstandardbuffer(seeMaterials andMethods)andincubatedasdescribed below. 21A+ and 32A- have thepotential
toform 21contiguousbasepairs.After thereaction,1,ulof10%SDSwasadded,and DNAwasextracted withphenol-chloroform(1:1)and then subjectedtoelectrophoresis througha15%polyacrylamide-TBE gelat4°C.Incubationconditionswere asfollows: lane a,noincubation;laneb,
37°Cfor 10 minwithoutNC;lane c,37°Cfor 10 min with 12pmol(1.2F.M)ofNC;laned, 94°Cfor 5 min andthen50°Cfor 5 minwithout NC. (B)32P-labeled28A+/28A- duplex (20 fmol,2nM)and 21A- (1 pmol,100nM)ssDNAsweremixed ina10-p.1standard buffer(seeMaterials andMethods),and reaction mixtureswereincubated andanalyzedasdescribed for thecorrespondinglanes inpanelA. 28A+ and 21A- havea
potentialtoforma20-bp duplex region. Asterisksin this andsubsequentfiguresindicate radioactivelabeling.
absence of NC protein. The 21 nucleotides at the 5' end of
32A- are identical in sequence to the corresponding nucle-otide of21A-. Strand exchange between these
oligonucleo-tideswasobserved in the presence of NCprotein (lane c),but
nostrandexchangewasdetectable in the absence of NC(lane
b).
With a second combination of double- andsingle-stranded
DNAs, a strikingly different result was obtained (Fig. 3B).
While heat denaturation followed by reannealing promoted
strand exchange between a double-stranded 28-mer (28A+/ 28A-) andashortersingle-stranded oligonucleotide (21A-)
that could form 20bpwith 28A+ (Fig. 3B, laned), nostrand exchangewasobserved between these substratesat37°Ceven
in the presence of NC protein (lane c). This excludes the
possibilitythat NCprotein's onlyrole in the strand exchange
process was to promote denaturation of the dsDNA, with
annealing taking place after removal of NC. When the
sub-strates were first heat denatured and then NC was added,
labeled28A+againhybridizedexclusivelyto28A- instead of 21A-, althoughthe latterwaspresentin50-foldexcess (data
notshown). Thus, NCprotein alsoplays a role in thestrand
exchange processafter thestrandseparation step.
Selective annealingofcomplementary oligonucleotides. To
further examine theeffectof NCprotein onthespecificityof strand exchange, we tested oligonucleotides of different lengths in competition for annealing with complementary
DNAstrands. Figure4showsadiagram of the substrates used in this experiment. The actual DNA sequences are given in
Materials and Methods. 28A- can form a 28-bp duplex with 28A+, while24A- canform24bp and 21A- can form only 20
bpwith 28A+. 28X- isa28-mer, but itcan formonly 20 bp with28A+.
Several combinations of theseoligonucleotides were tested in competition for annealing with 28A+ (Fig. 5) in the presence ofNCprotein. When28A- waspresent at a concen-tration of4nM, themajorityof28A+washybridizedto
28A-even when 24A- or21A- waspresent in 25-foldexcess (Fig.
5, lanes1 to3).
Onthe otherhand,40fmol of 24A- hybridized to28A+ in
the presence ofexcess21A-butnotinthe presence ofexcess
28A- (Fig. 5, lanes 4 to 6). 21A- was unable to compete against eitherexcess28A-or24A- (lanes7 to9). Thus,in the presence of NC, hybridization to 28A- was favored over
hybridizationto24A- and21A-, andhybridizationto
24A-was favored over hybridization to 21A-. To determine
* 4 20
4 20
-4
20 4
20 1
20 K 4
28A+
28A-
2421
A-
28X-FIG. 4. Substrates for the selectiveannealing assay.Sequences of these DNAoligonucleotidesaredescribedinMaterials and Methods. 28A+, 24A+,and 21A have the same polarity,while 28A-,24A-, 21A- and 28X- have theopposite polarity. The numbers in their
namesindicate thelengthsof theseoligonucleotides.28A+ and28A-, 24A+ and24A-, and 21A+ and 21A- can formcomplete duplex
structures.24A+ isidenticaltothe 24-base sequenceatthe 3'end of 28A+. Twentynucleotides of 21A+ are identical to themiddle 20 nucleotides of 28A+. Twenty nucleotides in the middle of 28X- are identical to the 20 nucleotides in thecorresponding region of 28A-.
000,
4 4
on November 9, 2019 by guest
http://jvi.asm.org/
DNA
(40fmol)
28A- 24A- 21 A- 28X- noDNA-T I
DNA 11 (1 pmol)
S,°°
c ', °~<°lc")'~
Ny"vl
(5%O1
'N~
)-'1 z228~~~pI
l,(-r
28
* >.-qI' ~SSP
28
* -*.
21 2
1 2 3 4 5 6 7 8 9 10 1 1 12 13
*
A;r-
282-.0 *
-p-
W28
== * -*.
28
1. 421
[image:5.612.138.473.79.222.2]\28
FIG. 5. Effect of the length of the base-paired region on the annealing of competing DNAs in the presence of HIV-1 NC protein. Reaction mixtures (10
[lI)
contained32P-labeled 28A+ (18fmol) with different combinations of two competing complementary oligonucleotides (40 fmol and 1 pmol, respectively) in standard annealing buffer with 25 pmol of HIV-1 NC protein. Incubation was at 30°C for 15 min and was followed by addition of 5plofstopping solution (see Materials and Methods). The samples were analyzed byelectrophoresis through a15%polyacrylamide gel inTBEat4°C.whether this discrimination reflected an effect of the length of
the base-paired region rather than simply an effect of the
overall length of the DNA, we used 28X- DNA, which,
although 28 nucleotides long, could only form 20contiguous base pairs with 28A+. Both 24A- and 21A- effectively competed with 28X- for annealing to 28A+, indicating that
the selective annealing was determined by the length of the
potential base-paired region and not the total length of the
oligonucleotides (lanes 10 to 12).
From these results, we proposethat NC, by catalyzing the
ss*->ds transition and thereby accelerating the approach to
equilibrium, differentiallyenhancesannealingbetweenpairs of
DNA strands that can form the most stable duplex (in this
no NC
example, by forming a longer base-paired region) among
competing alternatives.
The effect oftemperature onselective annealing by NCwas
tested as shown in Fig. 6. In the absence of NC, selective
annealing of 28A-to28A+ inthepresenceofa25-foldexcess
of 21A- was observed only at 65°C, which is between the predicted melting temperature of 28A+/28A- and that of 28A+/21A- under these conditions. In thepresence of NC, this selectivity was observed at all the temperatures tested, ranging from 0 to700C. At thehighesttemperatures (65 and 700C), selectivitywasreduced,presumablyas aresult of either partial denaturation of NC orinefficientDNAbinding by NC
atthese temperatures.
+ NC (2Spmol)
cE E EE E E c o o o o o E E o N N N N N N 0 0
o NI)n LI) CD - 0 LoI
C r C fS C~. = c =
E E E E E E E E E
0 0 LIn 0 O 0 0 0
N N m N N N N N N\1
0 (D CD Ln O N LI) 0 r _ - C\)m r LO CD
N-28
28
\ 28
*1
-*FIG. 6. Effectof NCproteinonannealing ofanssDNAoligonucleotide (28A+)withtwocompeting complementaryDNAs(28A-and21A-).
The10-,ulreaction mixturescontained 32P-5'-end-labeled 28A+ (20 fmol), unlabeled28A- (40 fmol),and unlabeled 21A- (1 pmol)instandard
annealingbuffer, withorwithout 25pmolof HIV-1 NCprotein. Incubationwasperformedatthe indicatedtemperatureand forthe indicated lengthof time. After eachreaction,thesampleswerechilledonice,1 p.lof10%SDSwasadded,and DNAwasextracted withphenol-chloroform (1:1)andanalyzedon a 15%polyacrylamide-lX TBEgel.
28A±
* - ..
(I
8fmol)
28A-(40fmol)
+
21
A-(1
pmol)-iA.,4 .. Ah,.-A I.A.&&A
%W
.,",t bw Wd40wmm#90tw:I& .. Iso 0,
6i
".g, II
I
"?
on November 9, 2019 by guest
http://jvi.asm.org/
[image:5.612.127.494.463.673.2]Coaggregation of DNA and NC protein.Oneof thepossible mechanisms forstimulation of DNA renaturation involves the formation of a multimolecular DNA-NC complexin whichthe overallconcentration of complementaryDNAs inthereaction
is irrelevant. We indeed found that NC and DNA could form a coaggregate even when the DNA molecules were asshortas
24 nucleotides long. NC protein, which is soluble in the absenceof DNA, could be inducedtoprecipitate bya24-mer
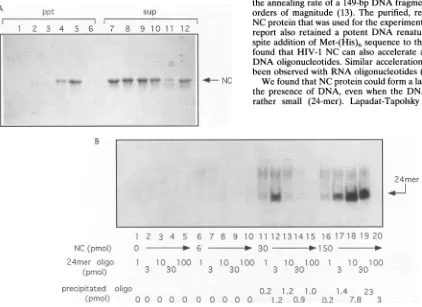
oligonucleotide (Fig. 7A). When 150pmol of NC waspresent in 10 ,ul of solution, addition of 30 pmolof theoligonucleotide resulted in efficient precipitation of NC (lanes 5 and 11), but precipitationwas less efficient when the amount of DNA was
greater or less.
Precipitation of the labeled 24-mer oligonucleotide in the presence ofNC proteinwas monitored under similar condi-tions(Fig. 7B). Tenor30pmol of the oligonucleotides in 10
RI
of solutionwasefficientlyprecipitated by150pmolofNC(78and77%, respectively) (lanes 19 and 20), but addition ofmore DNA inhibited precipitation (lane 20). In thepresence of30 pmol of NC, 1.2 pmoloutofatotalof3pmolofDNAcould beprecipitated, but addition ofmore NC didnot cause more
A ppt
B
sup
1 2 3 4 5 NC (pmol) 0
--4- NC
precipitation. From these results, we conclude that NC and DNA canformalarge coaggregate. The ratio ofDNA andNC was critical for formation of this precipitable complex. In
contrast,catalysis ofannealingorstrandexchange couldoccur
under awider range ofconditions, including conditions that
didnotresult in appreciable precipitation. DISCUSSION
There have been many reports of enhanced nucleic acid renaturation mediated by DNA- or RNA-binding proteins
(45). These proteins include T4 gene 32 protein (3), E. coli
single-strand-binding protein (9, 10), RecA protein (8), his-tones (11), and heterogeneous nuclear ribonucleoprotein
(hnRNP) Al (26, 35,39),aswellasotherproteins.hnRNPAl is able to accelerateannealing ofa200-bpDNAmolecule by
several thousand-fold (39),while the rate enhancements pro-moted by the other proteins cited above are significantly
smaller. TheextentofstimulationDNAannealing byhnRNP Al appearstobetoogreattobe explained solelyby inhibition of electrostatic repulsion or intrastrand secondary structure
formation (39).
A recentreporthas shown that HIV-1NCproteinincreases theannealingrateofa149-bpDNAfragmentbymorethan4
orders of magnitude (13). The purified, recombinant HIV-1 NCprotein that was used for the experiments described in this report also retained a potent DNA renaturation activity de-spite addition ofMet-(His)6 sequencetothe Nterminus. We
found that HIV-1 NC can also accelerate annealing of short
DNA oligonucleotides. Similar acceleration hassubsequently
beenobserved withRNAoligonucleotides (22).
We foundthat NCproteincould formalargecoaggregatein the presence ofDNA, evenwhen the DNA moleculeswere
rather small (24-mer). Lapadat-Tapolsky et al. also have
24mer
<_21
6 7 8 9 101112131415 1617181920
6 -* - 30 - 150 -* 0
1 10 100 1 10 100 1 10 100 1 10 100
3 30 3 30 3 30 3 30
precipitated oligo
(pmol) 0 0 0 C 0 0 0 0 0 0
0.2 1.2 1.0 1.4 23
[image:6.612.64.486.302.609.2]1.2 0.9 0.2 7.8 3
FIG. 7. (A) NC proteinaggregatesin thepresenceofashortoligonucleotides. The10-RIreaction mixtures contained 150 pmol (15 FM) ofHIV
NCprotein in the standardannealing buffer with variousamountsof 24A+ DNAoligonucleotides ina1.5-ml Eppendorftesttube (lanes 1 and
7,noDNA; lanes2and8, 1 pmol of DNA; lanes 3and 9, 3 pmol of DNA; lanes 4 and 10, 10 pmol of DNA; lanes 5 and 11, 30 pmol of DNA;
lanes 6 and12, 100pmol of DNA). Aftera20-min incubationat0°C, tubeswere spunat14,000rpmfor 10min.Supernatant(sup) and pellet (ppt) wereseparated, boiled in the sample buffer, and analyzedon anSDS-18% polyacrylamide gel (27). Protein bandswerevisualized by staining with Coomassiebrilliant blue.(B) Precipitation ofa24-meroligonucleotide by NC protein. The10-plreaction mixtures containedstandard annealing buffer, variousamountsof 24-meroligonucleotide (oligo; 24A+), and NC protein,asindicated. Theamountof32P-labeled 24A+ in the reaction waskeptconstant,although the totalamountof 24A+wasvariedby adding unlabeled oligonucleotide. After 20 min of incubationat0°C inan Eppendorf Microfuge tube, the sampleswerecentrifugedat14,000rpmfor20min. The pelletsweredissolved in 5 pulof 10 mMTris-Cl (pH7.9)-l
mMEDTA, 5
RIl
ofstopping solution (see Materials and Methods)wasadded, and the sampleswereanalyzedon a15%polyacrylamide-TBE gel. Thegelwasthendried andexposed for autoradiography. Quantitation of the radioactivitywasdone withaMolecular Dynamics Phosphorlmager. Theamountofprecipitated oligonucleotidewascalculatedby comparing radioactivities in the pellet andsupernatantfractions.1 2 3 4 5 6 7 8 9 10 11 12
24mer oligo
(pmol)
Us
on November 9, 2019 by guest
http://jvi.asm.org/
observed coaggregates of NC withlongerDNA(plasmidDNA containing a 6-kb region from the HIV-1 genome) (28). Since the DNA molecules in our experiment were short, it is not plausible thatacomplex ofasingleDNAmolecule and several boundNC molecules would be large enough to beprecipitated.
This result therefore suggests the possibility of an interaction between NC molecules bound to nucleic acids. Khan and Giedroc have previously observed cooperative binding of NC
to nucleic acids, which mayreflect the same protein-protein
interactions (25). Alternatively, simultaneous binding of single NCproteins to two DNAmolecules could provide an expla-nationfor the coaggregation that weobserved.
Sikorav and Church haveproposedthataggregationofDNA
provides a general mechanism for acceleration ofDNA rena-turation by a variety of methods(45).DNAaggregation in the presenceofNC is consistent with suchamechanismplayinga
role here. There is no direct evidence, however, that this aggregate is an intermediate in DNA renaturation, and the conditions favoring annealing were not identical to those favoring formation of precipitable aggregates.
Some of the activities previously shown for NC, such as
promoting dimerization of the retroviral genome or annealing ofa tRNAprimerto theprimer bindingsite, may largely be due to NC's ability to destabilize secondary structure. Inthe absence of proteins, these reactions proceed much more
rapidly after the nucleic acids are heat denatured (42, 43),
suggesting that removal of intramolecular secondarystructure
is rate limiting. Melting of tRNA by HIV NC protein was
demonstrated directly by circular dichroism spectroscopy(25).
We were able to show complete melting of short dsDNA at
highNCconcentrations, althoughat lowerprotein
concentra-tions, NC stimulated renaturation. These results suggest that NC is able to stimulate both the
ds-'ss
and the ss->dstransition,at least under differentprotein concentrations.
Results from ourDNAannealing experiments with compet-ing DNA strands(Fig.5and6)indicate that in the presence of NCprotein, these DNAs areinequilibrium, witharapidrate
of ss<->ds transition. hnRNPAlproteinalso is abletofacilitate
asimilar approach toequilibrium, albeitat ahigher
tempera-ture (40). Recent results suggest that NCcan playthe same
rolewith RNAsubstrates (22, 50).
What is the significanceof these activities of NCprotein in vivo? The propensityof nucleic acid moleculestoform
meta-stable base-paired structures that dissociate slowly can limit the rate and specificity of reactions that involve specific
base-pairing (21). One prediction from the results described aboveis that NC could facilitatecorrectbase-pairingorfolding
of nucleicacids inafashionsimilartothatofprotein
chaper-ones(2).When DNA or RNAismisfolded, thetypically high
energybarrier toabandoning themisfolded state under ordi-nary conditions at physiological temperature means that the time required to reach the most thermodynamically stable folded conformationcanbeextremely long.Inthe presence of
NC, this process could be greatly accelerated, although it remains to be determined whether NC can indeed promote
folding of a large RNA molecule. Such an activity on RNA conformation hasrecentlybeenproposedforhnRNPs,
includ-ing hnRNP Al (41). This activity need not be limited to
correction of misfolded nucleic acids. Since RNAfoldingcould
start before completion of transcription, the initial folded conformation of a nascent transcript might be different from that of the most thermodynamically stable structure for the
complete RNA.Thus, theabilityof NCto reduce thekinetic barrierstostructural transformations of nucleic acids could be essential in allowing the transcripts to achieve a mature
conformation. Dimerization of the retroviral genome can be
considered an example of such a folding problem. The NC
domain of the Gag polyprotein is required for thisprocessin vivo(34),andpurifiedNCproteincanpromotethis reaction in vitro(42, 43).Theseconformationalchangesin viral RNA may becritical for efficientpackagingand forsubsequentactivities ofgenomicRNA.Involvement of the NC domain of theGag polyprotein in genomicRNApackaginghas been established
by genetic experiments (4, 5, 14, 16-20, 32, 33, 38). After dimerization and
packaging,
a further step in maturation ofmurineleukemia virusgenomicRNArequiresthe presence of
matureNCprotein andturnsthegenomic RNAinto a more
compactconformation
(16).
Genetic evidence suggests that NC
protein
is also involved inreverse transcription (32). The ability of NC to promote the ds->ss transition could play a role in facilitating strand
dis-placementin the final stage of viral DNAsynthesis.NCprotein mightalso assistreverse
transcription
bydestabilizing
second-arystructuresin thetemplateDNA.NC has beenreportedtoincrease the initial product of reverse transcription, but the basisfor this effect remainstobedefined
(32).
The selective annealing activity of NC may have
practical
applications. One might be to enhancespecificity
in PCR.Annealingofprimerstoincorrect sitescanleadtoproduction
of spurious PCR products. The ability of NC protein to
promoteselectiveannealing mightbeemployedtoincrease the ratioofcorrect to incorrectproducts.This activity mightalso prove useful in promoting accurate renaturation of
complex
DNAs in solution, which is required for genomic mismatch
scanning
(36),
or DNAannealing
to an immobilized DNA target on achip
(15)
or onothermaterials(e.g.,
in Southernhybridization
orinsituhybridization).
Whether NCwill prove useful in theseapplications
remainstobeexplored.
ACKNOWLEDGMENTS
WethankDanielHerschlagforcomments onthemanuscript.
This workwas supported by a grant from the NIAID-NCDDG-AIDS program andbythe HowardHughesMedical Institute(HHMI). P.O.B. is an assistant investigator of the HHMI, and Z.T. was an HHMIresearch associate.
REFERENCES
1. Adachi, A.,H. E.Gendelman, S. Koenig,T. Folks,R.Willey, A. Rabson,and M. A. Martin. 1986. Production ofacquired
immu-nodeficiency syndrome-associated retrovirus in human and
non-human cells transfected with an infectious molecular clone. J. Virol. 59:284-291.
2. Agard,D. A.1993. Tofoldor nottofold. Science260:1903-1904. 3. Alberts, B. M.,and L. Frey. 1970. T4bacteriophagegene 32;a
structuralprotein inthereplicationandrecombinationof DNA. Nature(London)227:1313-1318.
4. Aldovini,A.,andR. A.Young.1990.Mutations of RNA andprotein
sequencesinvolved in humanimmunodeficiencyvirus type 1
packag-ingresult inproductionofnoninfectious virus. J. Virol.64:1920-1926.
5. Aronof,R.,A.Hajjar,and M.L.Linial.1993. Avianretroviral RNA
encapsidation: reexamination of functional 5' RNA sequences and the role ofnucleocapsid Cys-Hismotifs.J. Virol.67:178-188.
6. Berkowitz,R.D., J. Luban,and S. P.Goff.1993. Specific binding
of human immunodeficiencyvirus type 1 Gag polyprotein and nucleocapsid protein to viral RNAs detected by RNA mobility shiftassays. J.Virol. 67:7190-7200.
7. Bieth,E.,C.Gabus,andJ.-L.Darlix. 1990. Astudyofthedimer formation of Rous sarcomavirus RNA and ofitseffectonviral protein synthesisin vitro.Nucleic Acids Res. 18:119-127. 8. Bryant, F. R., and I. R. Lehman. 1985. On the mechanism of
renaturation of complementary strandsby the recA protein of
Escherichiacoli.Proc. Natl. Acad. Sci. USA82:297-301. 9. Chase, J. W., and K. R. Williams. 1986. Single-stranded DNA
bindingproteins requiredfor DNAreplication.Annu. Rev. Bio-chem. 55:103-136.
on November 9, 2019 by guest
http://jvi.asm.org/
10. Christiansen, C., and R. L. Baldwin. 1977. Catalysis of DNA
reassociation by the EscherichiacoliDNA binding protein. J. Mol. Biol. 115:441-454.
11. Cox, M. M., and I. R. Lehman. 1981. Renaturation of DNA: a novel activity of histones. Nucleic Acids Res. 9:389-400. 12. De Rocquigny, H., C. Gabus, A. Vincent, M.-C. Fournie-Zaluski,
B.Roques, and J.-L. Darlix. 1992. Viral RNA annealing activities of human immunodeficiency virus type 1 nucleocapsid protein require only peptide domainsoutside the zinc fingers. Proc. Natl. Acad. Sci. USA 89:6472-6476.
13. Dib-Hajj, F., R. Kahn, and D. P. Giedroc. 1993. Retroviral nucleocapsid proteins possess potent nucleic acid strand renatur-ation activity. Protein Sci. 2:231-243.
14. Dupraz, P., and P.-F. Spahr. 1992. Specificity of Rous sarcoma virus nucleocapsid protein in genomic RNA packaging. J. Virol. 66:4662-4670.
15. Fodor, S. P. A., P. R. Rava, X. C. Huang, A. C.Pease,C.P. Holmes,
and C. L. Adams. 1993. Multiplexed biochemical assays with biological chips. Nature (London) 364:555-556.
16. Fu, W., and A. Rein. 1993. Maturation of dimeric viral RNA of Moloney murine leukemia virus. J. Virol. 67:5443-5449. 17. Fu, X., R. A. Katz, A. M. Skalka, and J. Leis. 1988. Site-directed
mutagenesis of the avian retrovirus nucleocapsid protein, pp12. J. Biol. Chem. 263:2140-2145.
18. Gorelick,R.J., D. J.Chabot, A. Rein, L. E. Henderson,and L.0. Arthur. 1993. The two zincfingers in the human immunodeficiency virus type 1 nucleocapsid protein are not functionally equivalent. J. Virol. 67:4027-4036.
19. Gorelick, R. J., L. E. Henderson, J. P.Hanser, and A. Rein. 1988. Point mutants of Moloney murine leukemia virus that fails to package viral RNA: evidence for specific RNA recognition by a "zinc finger-like" protein sequence. Proc. Natl. Acad. Sci. USA 85:8420-8424.
20. Gorelick, R. J., S. M. Nigida, Jr., J. W. Bess, Jr., L. 0. Arthur, L.E.
Henderson, and A. Rein. 1990. Noninfectious human immunode-ficiencyvirus type 1 mutantsdeficient ingenomic RNA. J. Virol. 64:3207-3211.
21. Herschlag, D. 1991.Implication ofribozyme kinetics for targeting thecleavage ofspecificRNAmoleculesin vivo: more isn't always better. Proc. Natl. Acad. Sci. USA 88:6921-6925.
22. Herschlag, D., M.Khosla, Z. Tsuchihashi, and R. L. Karpel. A RNAchaperone activity ofnonspecific RNAbinding proteins in hammerhead ribozyme catalysis. EMBO J., in press.
23. Innis, M. A., andD.H.Gelfand. 1990. Optimizationof PCRs, p. 3-12. In M. A. Innis, D. H. Gelfand, J. J. Sninsky, and T. J. White (ed.),PCRprotocols. Academic Press, SanDiego,Calif. 24. Karpel, R. L., L.E. Henderson, and S. Oroszlan. 1987.
Interac-tions ofretroviral structuralproteinswith single-strandednucleic acids. J. Biol. Chem. 262:4961-4967.
25. Khan, R., and D. P. Giedroc. 1992. Recombinant human immu-nodeficiency virus type 1 nucleocapsid (NCP7) protein unwinds tRNA.J. Biol. Chem. 267:6689-6695.
26. Kumar,A., and S. H. Wilson. 1990.Studies of the strand-annealing activities of mammalian hnRNP complexprotein Al. Biochemis-try 29:10717-10722.
27. Laemmli, U. K. 1970. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature (London) 227:680-685.
28. Lapadat-Tapolsky, M., H. De Rocquigny, D. van Gent, B. Roques,
R. Plasterk, andJ.-L. Darlix. 1993. Interaction between HIV-1 nucleocapsid protein and viral DNA may have important functions in the viral lifecycle. Nucleic Acids Res. 21:831-839.
29. Leis, J.,andJ.JentofL1983.Characteristics and regulation of interaction ofavian retrovirus ppl2 with viral RNA. J. Virol. 48:361-369. 30. Luban, J., and S. P. Goff. 1991. Binding of human
immunodefi-ciency virus type 1 (HIV-1) RNA to recombinant HIV-1 Gag polyprotein. J. Virol. 65:3203-3212.
31. Meric, C., J.-L. Darlix, and P.-F. Spahr. 1984. It is Rous sarcoma virusproteinp12and notp19that binds tightly to Rous sarcoma
virus RNA. J. Mol. Biol. 173:531-538.
32. Meric, C., and S. P. Goff. 1989. Characterization of Moloney murine leukemia virus mutants with single-amino-acid substitu-tions in the Cys-His box of the nucleocapsid protein. J. Virol. 63:1558-1568.
33. Meric,C., E.Gouilloud,and P.-F. Spahr.1988.Mutations in Rous
sarcoma virus nucleocapsid proteinp12 (NC): deletions of Cys-His boxes. J. Virol. 62:3328-3333.
34. Meric, C., and P.-F. Spahr. 1986. Rous sarcoma virus nucleic acid-binding protein p12 is necessary for viral 70S RNA dimer formation andpackaging. J. Virol. 60:450-459.
35. Munroe, S. H., and X. Dong. 1992. Heterogenous nuclear ribonu-cleoprotein Al catalyzes RNA-RNA annealing. Proc. Natl. Acad. Sci. USA 89:895-899.
36. Nelson, S. F., J. H. McCusker, M. A. Sander,Y. Kee, P. Modrich, and P. 0. Brown. 1993. Genetic mismatch scanning: a new approach to genetic linkage mapping. Nat. Genet. 4:11-18. 37. Nissen-Meyer, J., and A. K. Abraham. 1980. Specificity of RNA
binding by thestructural protein (plO) of Friend murine leukemia virus. J. Mol. Biol. 142:19-28.
38. Oertle, S., and P.-F. Spahr. 1990. Role of the Gag polyprotein precursor in packaging and maturation of Rous sarcoma virus genomic RNA. J. Virol.64:5757-5763.
39. Pontius, B. W., and P. Berg. 1990. Rapid renaturation of comple-mentary DNAstrands mediated by purified mammalian hetero-genous nuclear ribonucleoprotein Al protein: implication for a mechanism for rapid molecular assembly processes. Proc. Natl. Acad.Sci. USA87:8403-8407.
40. Pontius, B. W., and P. Berg. 1992. Rapid assembly and disassembly of complementary DNA strands through an equilibrium interme-diate state mediated by Al hnRNP protein. J. Biol. Chem. 267:13815-13818.
41. Portman, D. S., and G. Dreyfuss. 1994. RNAannealingactivities
in HeLa nuclei.EMBO J. 13:213-221.
42. Prats, A.-C., V. Housset, G. de Billy, F. Cornille, H. Prats, B.
Roques, and J.-L. Darlix.1991. Viral RNA annealing activities of the nucleocapsid protein of Moloney murine leukemia virus are zincindependent. Nucleic Acids Res. 19:3533-3541.
43. Prats, A.C., L. Sarih, C. Gabus, S. Litvak, G. Keith, and J.-L. Darlix. 1988.Small finger protein of avian and murine retrovirus has nucleic acid annealing activity and positions the replication primer tRNA onto genomic RNA. EMBO J. 7:1777-1783. 44. Sambrook, J., E. F. Fritsch, and T. Maniatis. 1989. Molecular
cloning: alaboratory manual, 2nd ed. Cold Spring Harbor Labo-ratory Press, Cold SpringHarbor, N.Y.
45. Sikorav, J.-L., and G. M.Church. 1991. Complementary recogni-tion in condensed DNA: accelerated DNA renaturation. J. Mol. Biol.222:1085-1108.
46. Smith, B. J., and J. M. Bailey. 1979. The binding of an avian myeloblastosis virus basic 12,000 dalton protein to nucleic acids. Nucleic AcidsRes. 7:2055-2072.
47. Studier, W. F., A. H.Rosenberg, J. J. Dunn, and J. W.Dubendorff. 1990.Use of T7 RNA polymerase to direct expression of cloned genes. MethodsEnzymol. 185:60-89.
48. Surovoy, A., J.Dannull, K. Moelling, and G. Jung. 1993. Confor-mational and nucleic acid binding studies on the synthetic nucleo-capsid protein of HIV-1. J. Mol. Biol. 229:94-104.
49. Tabor, S., and C. C. Richardson. 1989. Selective inactivation of the exonuclease activity of bacteriophage T7 DNA polymerase by in vitro mutagenesis. J. Biol. Chem. 264:6447-6458.
50. Tsuchihashi, Z., M. Khosla, and D. Herschlag. 1993. Protein enhancement of hammerhead ribozyme catalysis. Science 262:99-102.
51. Varmus, H., and P. 0. Brown. 1989. Retroviruses, p. 53-108. In D. E. Berg and M. M. Howe (ed.), Mobile DNA. American Society for Microbiology, Washington, D.C.
52. You, J. C., and C. S. McHenry. 1993. HIV nucleocapsid protein: expression in Escherichiacoli, purification, and characterization. J. Biol.Chem.268:16519-16527.