0022-538X/01/$04.00
⫹
0
DOI: 10.1128/JVI.75.23.11811–11820.2001
Copyright © 2001, American Society for Microbiology. All Rights Reserved.
Complete Genome Sequence of the Shrimp White Spot
Bacilliform Virus
FENG YANG, JUN HE, XIONGHUI LIN, QIN LI, DENG PAN, XIAOBO ZHANG,
ANDXUN XU*
The Third Institute of Oceanography, Xiamen 361005, People’s Republic of China
Received 11 June 2001/Accepted 1 August 2001
We report the first complete genome sequence of a marine invertebrate virus. White spot bacilliform virus
(WSBV; or white spot syndrome virus) is a major shrimp pathogen with a high mortality rate and a wide host
range. Its double-stranded circular DNA genome of 305,107 bp contains 181 open reading frames (ORFs). Nine
homologous regions containing 47 repeated minifragments that include direct repeats, atypical inverted repeat
sequences, and imperfect palindromes were identified. This is the largest animal virus that has been completely
sequenced. Although WSBV is morphologically similar to insect baculovirus, the two viruses are not detectably
related at the amino acid level. Rather, some WSBV genes are more homologous to eukaryotic genes than viral
genes. In fact, sequence analysis indicates that WSBV differs from all known viruses, although a few genes
display a weak homology to herpesvirus genes. Most of the ORFs encode proteins that bear no homology to any
known proteins, either suggesting that WSBV represents a novel class of viruses or perhaps implying a
significant evolutionary distance between marine and terrestrial viruses. The most unique feature of WSBV is
the presence of an intact collagen gene, a gene encoding an extracellular matrix protein of animal cells that has
never been found in any viruses. Determination of the genome of WSBV will facilitate a better understanding of
the molecular mechanism underlying the pathogenesis of the WSBV virus and will also provide useful
infor-mation concerning the evolution and divergence of marine and terrestrial animal viruses at the molecular level.
White spot bacilliform virus (WSBV) or white spot
syn-drome virus (WSSV) is a major shrimp pathogen that is highly
virulent in penaeid shrimp, the most important species used in
aquaculture, and can also infect most species of crustacean (15,
32). Infection of penaeid shrimp by WSBV can result in
mor-tality of up to 90 to 100% within 3 to 7 days (57). A major
outbreak of WSBV infection in 1993 resulted in a 70%
reduc-tion in shrimp producreduc-tion in China (14, 57) and has raised
major concerns in aquaculture around the world. Prevention
and inhibition of infection by this virus can be difficult due
largely to the ability of WSBV to survive for a long time in the
environment (2 years in a shrimp pond) and also due to a poor
understanding of this virus at the molecular level.
WSBV was originally classified as an unassigned member of
the
Baculoviridae
because of its rod-shaped, enveloped
mor-phology (20). However, it was recently excluded from the
bac-ulovirus family and is temporarily unclassified due to the lack
of molecular information (53). The virus is known generally as
white spot syndrome virus (WSSV) (31), and a new genus
name,
Whispovirus
, was proposed by Vlak et al. (48). Sequence
analysis of individual genes and proteins later showed that
most WSBV proteins bear poor sequence homology to
bacu-lovirus proteins but have repeated regions similar to those of
some baculoviruses. To understand the molecular basis of viral
replication and infection, we decided to sequence the whole
genome of WSBV.
MATERIALS AND METHODS
Isolation and sequencing of WSBV genomic DNA.Intact WSBV genomic
DNA was isolated from dead and moribund WSBV-infectedPenaeus japonicus
shrimp which were collected from shrimp ponds in Tongan, Xiamen, east China, in October 1996 as previously described (56). A whole-genome random sequenc-ing method (19) was used to obtain the complete genome sequence for WSBV. Genomic DNA was cloned by the shotgun method intoSmalI-linearized pUC18 vector, amplified, and sequenced using ABI BigDye Terminator chemistry on ABI 377 and ABI 3700 capillary sequencers. Large DNA fragments of 5 to 10 kb were also obtained by partial digestion withSau3A1 and cloned into the pBlue-script vectors (41). This was used to form a genome scaffold and to verify the orientation and integrity of the contigs formed from the shotgun library. A total of 5,770 sequences for sevenfold coverage were assembled using the InnerPeace software by Charles Lawrence based on the Phred, Phrap, and Consed program originally developed at the University of Washington.
The WSBV genome sequence was confirmed by comparison of the observed restriction fragments from seven restriction enzymes (BamHI,EcoRI,HindIII,
KpnI,PstI,SalI, andXbaI) to those predicted from the sequence data and was also confirmed by the genome scaffold produced by sequence pairs from 1,495 large-insert clones, which covered 90% of the main genome.
Gaps were closed by a combination of sequence-walking of shotgun and PCR large-fragment libraries.
DNA sequence analysis.Genome DNA composition, structure, repeats,
re-striction enzyme patterns, and translation were analyzed with the DNAMAN software (Lynnon BioSoft, Vaudreuil, Canada). Open reading frames (ORFs) consisted of more than 60 codons that are initiated with a methionine codon. For detection of potential protein-coding regions, the codon usage bias and posi-tional base preference were evaluated by determining the codon frequency of known WSBV genes or cDNA cloned from the WSBV cDNA library. Homology searches were performed with the FASTA (38) and BLAST programs (3). Pro-tein motifs were analyzed by using the PROSITE database, release 16 (25). Trans-membrane domains and signal peptides were predicted with ANTHEPROT (23).
Preparation and screening of a WSBV cDNA library.Poly(A) mRNA was
isolated from WSBV-infected shrimp tissue using the PolyATtract System 1000 kit (Promega). Double-stranded cDNAs were synthesized using the SUPER-SCRIPT plasmid system for cDNA synthesis and plasmid cloning (GIBCO BRL). WSBV cDNA clones were selected by hybridization with the digoxigenin (DIG)-labeled WSBV genomic DNA probe (DIG labeling kit; Boehringer Mannheim) and sequenced. The transcription of some ORFs was also verified by PCR on a cDNA cocktail using ORF-specific primers.
Nucleotide sequence accession number.The complete WSBV sequence can be
obtained from the GenBank database (accession no. AF332093).
* Corresponding author. Mailing address: The Third Institute of
Oceanography, Xiamen 361005, People’s Republic of China. Phone:
86-592-2195296. Fax: 86-592-2085376. E-mail: xxu@public.xm.fj.cn.
11811
on November 8, 2019 by guest
http://jvi.asm.org/
RESULTS AND DISCUSSION
General features of the WSBV genome.
We have previously
developed a unique method that enables us to highly purify the
WSBV virus from infected shrimp tissue (56). A random
shot-gun method was employed to sequence the entire genome of
WSBV; the sequence was subsequently confirmed by the
ge-nome scaffold formed by sequencing a large-fragment DNA
library. The complete WSBV genome is a double-stranded
circular DNA of 305,107 bp, similar to a previous estimate of
290 kb (56). Since the origin of replication was unknown, the
start of the largest
Bam
HI fragment was chosen to be base 1
(Fig. 1). Three percent of the WSBV genome is made up of
nine homologous regions (
hr
s), while the remaining 97% of the
sequences are unique (see description below). The genome has
a total G
⫹
C content of 41%.
A total of 531 putative ORFs were identified by sequence
analysis, among which 181 ORFs are likely to encode
func-tional proteins (Table 1). This corresponds to an average gene
density of one gene per 1.7 kb. Thirty-six of the 181 ORFs
annotated here either have been identified by screening and
sequencing a WSBV cDNA library (Table 1) or have been
reported previously to encode functional proteins (45, 46, 48,
49, 50). Transcription of another 52 ORFs was confirmed by
reverse transcription-PCR (RT-PCR; see Material and
Meth-ods) (Table 1). The relative positions of the ORFs and
hr
s in
the genome are shown in Fig. 1. For 80% of the putative 181
ORFs there is a potential polyadenylation site (AATAAA)
downstream of the ORF (Table 1).
WSBV ORFs encode gene products homologous to known
proteins.
Table 1 contains a list of the 181 predicted WSBV
ORFs. Among 181 ORFs, the proteins encoded by 18 ORFs
show 40 to 68% identity to known proteins from other viruses
or organisms or contain an identifiable functional domain.
These proteins include enzymes involved in nucleic acid
me-tabolism and DNA replication, a collagen-like protein, and
three viral structure proteins (for details, see below). Thirty
FIG. 1. Circular representation of the WSBV genome. Arrows, positions (outer ring) of the 181 ORFs (red and blue indicate the different
directions of transcription); green rectangles, 9
hr
s. B, sites of
Bam
HI restriction enzymes (inner ring; their positions are in parentheses).
on November 8, 2019 by guest
http://jvi.asm.org/
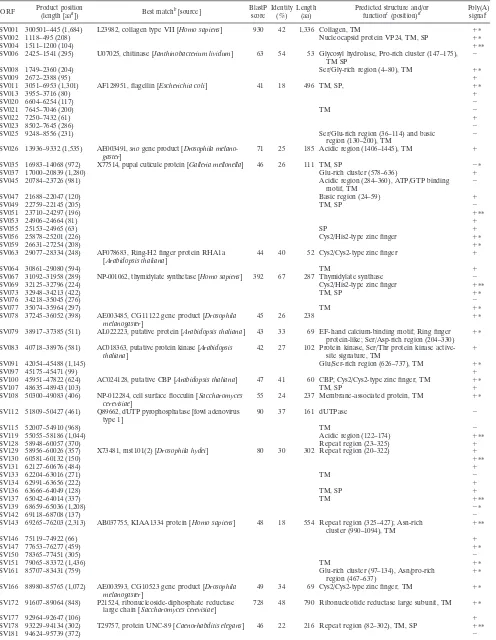
TABLE 1. Listing of potentially expressed ORFs in WSBV
ORF Product position
(length [aaa]) Best matchb[source]
BlastP score
Identity (%)
Length (aa)
Predicted structure and/or functionc(position)d
Poly(A) signalc
WSV001 300501–445 (1,684) L23982, collagen type VII [Homo sapiens] 930 42 1,336 Collagen, TM ⫹ⴱ
WSV002 1118–495 (208) Nucleocapsid protein VP24, TM, SP ⫹ⴱ
WSV004 1511–1200 (104) ⫹ⴱⴱ
WSV006 2425–1541 (295) U07025, chitinase [Janthinobacterium lividum] 63 54 53 Glycosyl hydrolase, Pro-rich cluster (147–175), TM SP
⫺
WSV008 1749–2360 (204) Ser/Gly-rich region (4–80), TM ⫹ⴱ
WSV009 2672–2388 (95) ⫹
WSV011 3051–6953 (1,301) AF128951, flagellin [Escherichia coli] 41 18 496 TM, SP, ⫹ⴱ
WSV013 3955–3716 (80) ⫹
WSV020 6604–6254 (117) ⫺
WSV021 7645–7046 (200) TM ⫺
WSV022 7250–7432 (61) ⫹
WSV023 8502–7645 (286) ⫺
WSV025 9248–8556 (231) Ser/Glu-rich region (36–114) and basic
region (130–200), TM
⫺
WSV026 13936–9332 (1,535) AE003491,snogene product [Drosophila
melano-gaster]
71 25 185 Acidic region (1406–1445), TM ⫹
WSV035 16983–14068 (972) X77514, pupal cuticule protein [Galleria mellonella] 46 26 111 TM, SP ⫺ⴱ
WSV037 17000–20839 (1,280) Glu-rich cluster (578–636) ⫹
WSV045 20784–23726 (981) Acidic region (284–360), ATP/GTP binding
motif, TM
⫺
WSV047 21688–22047 (120) Basic region (24–59) ⫹
WSV049 22759–22145 (205) TM, SP ⫺
WSV051 23710–24297 (196) ⫹ⴱⴱ
WSV053 24906–24664 (81) ⫹
WSV055 25153–24965 (63) SP ⫹
WSV056 25878–25201 (226) Cys2/His2-type zinc finger ⫹ⴱ
WSV059 26631–27254 (208) ⫹ⴱ
WSV063 29077–28334 (248) AF078683, Ring-H2 finger protein RHA1a
[Arabidopsis thaliana]
44 40 52 Cys2/Cys2-type zinc finger ⫹
WSV064 30861–29080 (594) TM ⫹
WSV067 31092–31958 (289) NP-001062, thymidylate synthetase [Homo sapiens] 392 67 287 Thymidylate synthase ⫺
WSV069 32125–32796 (224) Cys2/His2-type zinc finger ⫹ⴱⴱ
WSV073 32948–34213 (422) TM, SP ⫹ⴱ
WSV076 34218–35045 (276) ⫺
WSV077 35074–35964 (297) TM ⫹ⴱ
WSV078 37245–36052 (398) AE003485, CG11122 gene product [Drosophila
melanogaster]
45 26 238 ⫹ⴱ
WSV079 38917–37385 (511) AL022223, putative protein [Arabidopsis thaliana] 43 33 69 EF-hand calcium-binding motif; Ring finger protein-like; Ser/Asp-rich region (204–330)
⫹ⴱ
WSV083 40718–38976 (581) AC018363, putative protein kinase [Arabidopsis
thaliana]
42 27 102 Protein kinase, Ser/Thr protein kinase active-site signature, TM
⫹
WSV091 42054–45488 (1,145) Glu/Ser-rich region (626–737), TM ⫹ⴱ
WSV097 45175–45471 (99) ⫹
WSV100 45951–47822 (624) AC024128, putative CBP [Arabidopsis thaliana] 47 41 60 CBP; Cys2/Cys2-type zinc finger, TM ⫹ⴱ
WSV107 48635–48943 (103) TM, SP ⫹
WSV108 50300–49083 (406) NP-012284, cell surface flocculin [Saccharomyces
cerevisiae]
55 24 237 Membrane-associated protein, TM ⫹ⴱ
WSV112 51809–50427 (461) Q89662, dUTP pyrophosphatase [fowl adenovirus type 1]
90 37 161 dUTPase ⫺
WSV115 52007–54910 (968) TM ⫺
WSV119 55055–58186 (1,044) Acidic region (122–174) ⫹ⴱⴱ
WSV128 58948–60057 (370) Repeat region (23–325) ⫹
WSV129 58956–60026 (357) X73481, mst101(2) [Drosophila hydei] 80 30 302 Repeat region (20–322) ⫹
WSV130 60581–60132 (150) ⫹ⴱⴱ
WSV131 62127–60676 (484) ⫹
WSV133 62204–63016 (271) TM ⫺
WSV134 62991–63656 (222) ⫹
WSV136 63666–64049 (128) TM, SP ⫹
WSV137 65042–64014 (337) TM ⫹ⴱⴱ
WSV139 68659–65036 (1,208) ⫺ⴱ
WSV142 69118–68708 (137) ⫺
WSV143 69265–76203 (2,313) AB037755, KIAA1334 protein [Homo sapiens] 48 18 554 Repeat region (325–427); Asn-rich cluster (990–1094), TM
⫹ⴱⴱ
WSV146 75119–74922 (66) ⫹
WSV147 77653–76277 (459) ⫹ⴱ
WSV150 78365–77451 (305) ⫺
WSV151 79065–83372 (1,436) TM ⫹ⴱ
WSV161 85707–83431 (759) Glu-rich cluster (97–134), Asn/pro-rich
region (467–637)
⫹ⴱ
WSV166 88980–85765 (1,072) AE003593, CG10523 gene product [Drosophila
melanogaster]
49 34 69 Cys2/Cys2-type zinc finger, TM ⫹ⴱ
WSV172 91607–89064 (848) P21524, ribonucleoside-diphosphate reductase large chain [Saccharomyces cerevisiae]
728 48 790 Ribonucleotide reductase large subunit, TM ⫹ⴱ
WSV177 92964–92647 (106) ⫹
WSV178 93229–94134 (302) T29757, protein UNC-89 [Caenorhabditis elegans] 46 22 216 Repeat region (82–302), TM, SP ⫹ⴱⴱ
WSV181 94624–95739 (372) ⫺
Continued on following page
on November 8, 2019 by guest
http://jvi.asm.org/
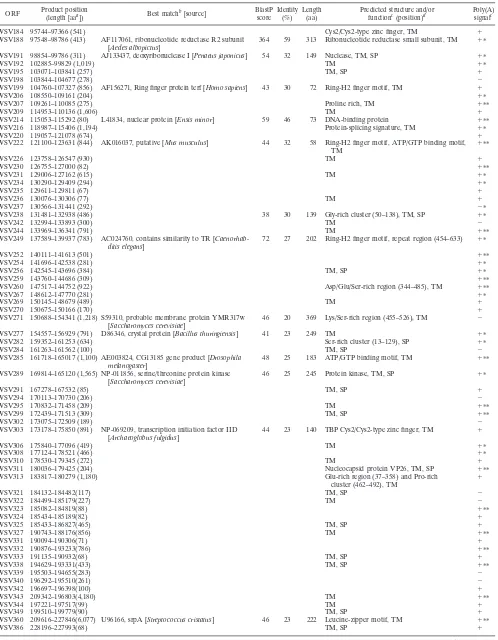
TABLE 1—
Continued
ORF Product position
(length [aaa]) Best matchb[source]
BlastP score
Identity (%)
Length (aa)
Predicted structure and/or functionc(position)d
Poly(A) signalc
WSV184 95744–97366 (541) Cys2/Cys2-type zinc finger, TM ⫹
WSV188 97548–98786 (413) AF117061, ribonucleotide reductase R2 subunit
[Aedes albopictus]
364 59 313 Ribonucleotide reductase small subunit, TM ⫹ⴱ
WSV191 98854–99786 (311) AJ133437, deoxyribonuclease I [Penaeus japonicus] 54 32 149 Nuclease, TM, SP ⫹ⴱ
WSV192 102885–99829 (1,019) TM ⫹ⴱ
WSV195 103071–103841 (257) TM, SP ⫹
WSV198 103844–104677 (278) ⫺
WSV199 104760–107327 (856) AF156271, Ring finger protein terf [Homo sapiens] 43 30 72 Ring-H2 finger motif, TM ⫹
WSV206 108550–109161 (204) ⫹ⴱ
WSV207 109261–110085 (275) Proline rich, TM ⫹ⴱⴱ
WSV209 114953–110136 (1,606) TM ⫹
WSV214 115053–115292 (80) L41834, nuclear protein [Ensis minor] 59 46 73 DNA-binding protein ⫹ⴱⴱ
WSV216 118987–115406 (1,194) Protein-splicing signature, TM ⫹ⴱ
WSV220 119057–121078 (674) ⫹
WSV222 121100–123631 (844) AK016037, putative [Mus musculus] 44 32 58 Ring-H2 finger motif, ATP/GTP binding motif, TM
⫹ⴱⴱ
WSV226 123758–126547 (930) TM ⫹
WSV230 126755–127000 (82) ⫹ⴱⴱ
WSV231 129006–127162 (615) TM ⫹ⴱ
WSV234 130290–129409 (294) ⫹ⴱ
WSV235 129611–129811 (67) ⫹
WSV236 130076–130306 (77) TM ⫹
WSV237 130566–131441 (292) ⫺ⴱ
WSV238 131481–132938 (486) 38 30 139 Gly-rich cluster (50–138), TM, SP ⫹ⴱ
WSV242 132994–133893 (300) TM ⫺
WSV244 133969–136341 (791) TM ⫹ⴱⴱ
WSV249 137589–139937 (783) AC024760, contains similarity to TR [
Caenorhab-ditis elegans]
72 27 202 Ring-H2 finger motif, repeat region (454–633) ⫹ⴱ
WSV252 140111–141613 (501) ⫹ⴱⴱ
WSV254 141696–142538 (281) ⫹ⴱ
WSV256 142545–143696 (384) TM, SP ⫹ⴱ
WSV259 143760–144686 (309) ⫹ⴱⴱ
WSV260 147517–144752 (922) Asp/Glu/Ser-rich region (344–485), TM ⫹ⴱⴱ
WSV267 148612–147770 (281) ⫹ⴱ
WSV269 150145–148679 (489) TM ⫹
WSV270 150675–150166 (170) ⫹
WSV271 150688–154341 (1,218) S59310, probable membrane protein YMR317w
[Saccharomyces cerevisiae]
46 20 369 Lys/Ser-rich region (455–526), TM ⫺
WSV277 154557–156929 (791) D86346, crystal protein [Bacillus thuringiensis] 41 23 249 TM ⫹ⴱ
WSV282 159352–161253 (634) Ser-rich cluster (13–129), SP ⫹ⴱ
WSV284 161263–161562 (100) TM, SP ⫺
WSV285 161718–165017 (1,100) AE003824, CG13185 gene product [Drosophila
melanogaster]
48 25 183 ATP/GTP binding motif, TM ⫹ⴱⴱ
WSV289 169814–165120 (1,565) NP-011856, serine/threonine protein kinase
[Saccharomyces cerevisiae]
46 25 245 Protein kinase, TM, SP ⫹ⴱ
WSV291 167278–167532 (85) TM, SP ⫹
WSV294 170113–170730 (206) ⫺
WSV295 170832–171458 (209) TM ⫹ⴱⴱ
WSV299 172439–171513 (309) TM, SP ⫹ⴱⴱ
WSV302 173075–172509 (189) ⫺
WSV303 173178–175850 (891) NP-069209, transcription initiation factor IID
[Archaeoglobus fulgidus]
44 23 140 TBP Cys2/Cys2-type zinc finger, TM ⫹
WSV306 175840–177096 (419) TM ⫹ⴱ
WSV308 177124–178521 (466) ⫹ⴱ
WSV310 178530–179345 (272) TM ⫹
WSV311 180036–179425 (204) Nucleocapsid protein VP26, TM, SP ⫹ⴱⴱ
WSV313 183817–180279 (1,180) Glu-rich region (37–358) and Pro-rich
cluster (462–492), TM
⫹
WSV321 184132–184482(117) TM, SP ⫺
WSV322 184499–185179(227) TM ⫺
WSV323 185082–184819(88) ⫹ⴱⴱ
WSV324 185434–185189(82) ⫹
WSV325 185433–186827(465) TM, SP ⫹
WSV327 190743–188176(856) TM ⫹ⴱⴱ
WSV331 190094–190306(71) ⫹
WSV332 190876–193233(786) ⫹ⴱⴱ
WSV333 191135–190932(68) TM, SP ⫹
WSV338 194629–193331(433) TM, SP ⫹ⴱⴱ
WSV339 195503–194655(283) ⫺
WSV340 196292–195510(261) ⫺
WSV342 196697–196398(100) ⫹
WSV343 209342–196803(4,180) TM ⫹ⴱⴱ
WSV344 197221–197517(99) TM ⫹
WSV349 199510–199779(90) TM, SP ⫹
WSV360 209616–227846(6,077) U96166, srpA [Streptococcus cristatus] 46 23 222 Leucine-zipper motif, TM ⫹ⴱⴱ
WSV386 228196–227993(68) TM, SP ⫹
Continued on following page
on November 8, 2019 by guest
http://jvi.asm.org/
ORFs predicted proteins that show a partial homology (20 to
39% identity) to known proteins or contain one or two
se-quence motifs (versus a real functional domain). The
remain-ing 133 ORFs encode proteins with no homology to any known
proteins or motifs.
[image:5.587.47.542.77.601.2]Enzymes involved in nucleotide metabolism.
Among the 18
ORFs encoding proteins that show extensive homologies with
previously identified proteins, WSV067, WSV112, WSV172,
WSV188, and WSV395 may encode the WSBV homologues of
enzymes involved in nucleic acid metabolism (Table 1). The
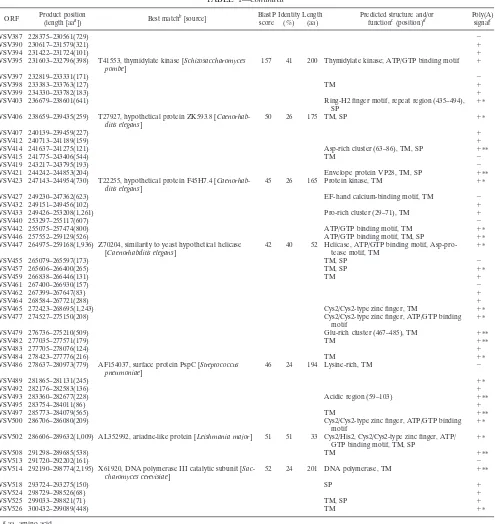
TABLE 1—
Continued
ORF Product position
(length [aaa]) Best matchb[source]
BlastP score
Identity (%)
Length (aa)
Predicted structure and/or functionc(position)d
Poly(A) signalc
WSV387 228375–230561(729) ⫺
WSV390 230617–231579(321) ⫹
WSV394 231422–231724(101) ⫹
WSV395 231603–232796(398) T41553, thymidylate kinase [Schizosaccharomyces
pombe]
157 41 200 Thymidylate kinase; ATP/GTP binding motif ⫹
WSV397 232819–233331(171) ⫺
WSV398 233383–233763(127) TM ⫹
WSV399 234330–233782(183) ⫹
WSV403 236679–238601(641) Ring-H2 finger motif, repeat region (435–494),
SP
⫹ⴱ
WSV406 238659–239435(259) T27927, hypothetical protein ZK593.8 [
Caenorhab-ditis elegans]
50 26 175 TM, SP ⫹ⴱ
WSV407 240139–239459(227) ⫹
WSV412 240713–241189(159) ⫹
WSV414 241637–241275(121) Asp-rich cluster (63–86), TM, SP ⫹ⴱⴱ
WSV415 241775–243406(544) TM ⫺
WSV419 243217–243795(193) ⫺
WSV421 244242–244853(204) Envelope protein VP28, TM, SP ⫹ⴱⴱ
WSV423 247143–244954(730) T22255, hypothetical protein F45H7.4 [
Caenorhab-ditis elegans]
45 26 165 Protein kinase, TM ⫹ⴱ
WSV427 249230–247362(623) EF-hand calcium-binding motif, TM ⫺
WSV432 249151–249456(102) ⫹
WSV433 249426–253208(1,261) Pro-rich cluster (29–71), TM ⫹
WSV440 253297–255117(607) ⫺
WSV442 255075–257474(800) ATP/GTP binding motif, TM ⫹ⴱ
WSV446 257552–259129(526) ATP/GTP binding motif, TM, SP ⫹ⴱ
WSV447 264975–259168(1,936) Z70204, similarity to yeast hypothetical helicase
[Caenorhabditis elegans]
42 40 52 Helicase, ATP/GTP binding motif, Asp-pro-tease motif, TM
⫹ⴱ
WSV455 265079–265597(173) TM, SP ⫺
WSV457 265606–266400(265) TM, SP ⫹ⴱ
WSV459 266838–266446(131) TM ⫹
WSV461 267400–266930(157) ⫺
WSV462 267399–267647(83) ⫹
WSV464 268584–267721(288) ⫹
WSV465 272423–268695(1,243) Cys2/Cys2-type zinc finger, TM ⫹ⴱ
WSV477 274527–275150(208) Cys2/Cys2-type zinc finger, ATP/GTP binding
motif
⫹ⴱ
WSV479 276736–275210(509) Glu-rich cluster (467–485), TM ⫹ⴱⴱ
WSV482 277035–277571(179) TM ⫹ⴱⴱ
WSV483 277705–278076(124) ⫹
WSV484 278423–277776(216) TM ⫹ⴱ
WSV486 278637–280973(779) AF154037, surface protein PspC [Streptococcus
pneumoniae]
46 24 194 Lysine-rich, TM ⫺
WSV489 281865–281131(245) ⫹ⴱ
WSV492 282176–282583(136) ⫹
WSV493 283360–282677(228) Acidic region (59–103) ⫹ⴱⴱ
WSV495 283754–284011(86) ⫹
WSV497 285773–284079(565) TM ⫹ⴱⴱ
WSV500 286706–286080(209) Cys2/Cys2-type zinc finger, ATP/GTP binding
motif
⫹ⴱ
WSV502 286606–289632(1,009) AL352992, ariadne-like protein [Leishmania major] 51 51 33 Cys2/His2, Cys2/Cys2-type zinc finger, ATP/ GTP binding motif, TM, SP
⫹ⴱ
WSV508 291298–289685(538) TM ⫹ⴱⴱ
WSV513 291720–292202(161) ⫺
WSV514 292190–298774(2,195) X61920, DNA polymerase III catalytic subunit [
Sac-charomyces cerevisiae]
52 24 201 DNA polymerase, TM ⫹ⴱⴱ
WSV518 293724–293275(150) SP ⫹
WSV524 298729–298526(68) ⫹
WSV525 299033–298821(71) TM, SP ⫹
WSV526 300432–299089(448) TM ⫹ⴱ
a
aa, amino acid.
b
Accession numbers are from the GenBank or SwissProt database.
c
Function was deduced from the degree of amino acid similarity to either products of known genes or Prosite signatures. TM, transmembrane domains; SP, signal peptides.
d
Positions of amino acid residues.
e⫹, polyadenylation signal present;⫺, signal absent.ⴱ, the transcription of the ORF was also verified by RT-PCR;ⴱⴱ, the ORF was confirmed by cDNA sequencing.
on November 8, 2019 by guest
http://jvi.asm.org/
highest degree of homology (67% identity over 287 amino
acids) was detected between the product of WSV067 and the
human thymidylate synthase. The 29-amino-acid thymidylate
synthase prosite motif (PS00091), which contains the catalytic
cysteine residue, is 100% conserved in the product of WSV067.
In addition, WSV112 may encode a WSBV homologue of
dUTPase (37% identity over 161 amino acids) since the five
conserved regions of dUTPase, especially the highly conserved
substrate-binding residues, were identified in the product of
WSV112 (13, 35). dUTPase has been shown to be essential for
the replication of DNA viruses (5). Consistent with the
previ-ous reports by van Hulten et al. (48) and Tsai et al. (45, 46),
WSBV contains ribonucleotide reductases (products of
WSV172 and WSV188) and also thymidylate/thymidine kinase
(product of WSV395). Among these enzymes, thymidylate
syn-thase catalyzes the methylation of dUTP to yield the
nucleo-tide precursor dTMP. This is an important step in the de novo
pathway of biosynthesis of pyrimidine (12). Despite its
ubiqui-tous distribution in nature, a viral thymidylate synthase was
found only in a few herpesviruses (2, 10, 26, 39),
Melanoplus
sanguinipes
entomopoxvirus (MsEPV) (1),
Chilo
iridescent
vi-rus (CIV) (36), and bacteriophages (9). Most vivi-ruses do not
contain thymidylate synthase, as they depend mostly on the
host enzymatic machinery for the replication of their genomes
so as to keep the viral genome small (36). WSBV and other
thymidylate synthase-containing viruses may therefore exhibit
a considerable independence from the host
deoxyribonucleo-tide synthesis. This may represent a significant advantage for
viral genome replication that may ultimately lead to
persis-tence of infection and a broad host range for viral infection
(36). It is possible that WSBV acquires these
replication-re-lated genes from its host and/or from a coinfecting virus that
might occur at an earlier period in evolution. However, since
the shrimp homologues of these genes have not been cloned,
we are not able to test this hypothesis.
Proteins involved in DNA replication and transcription.
WSBV contains genes encoding proteins involved in DNA
replication such as DNA polymerase (product of WSV514).
The WSBV DNA polymerase was putatively identified by the
presence of three highly conserved motifs, YGDTDSVFC
(DNA polymerase family B signature PS00116), KLG
MNSMYG, and DMTSLYP (conserved amino acid residues
are underlined), that are found in most eukaryotic DNA
poly-merases (4) as well as in some viral polypoly-merases (18, 29, 43).
However, since the degree of amino acid similarity between the
product of WSV514 and known DNA polymerases is low (24%
identity over 201 amino acids), its putative activity as a DNA
polymerase still awaits future experimental verification.
Inter-estingly, the size of this putative WSBV polymerase (2,195
amino acids) is much larger than those of the regular
poly-merases found in other organisms.
Products of ORFs that show weak similarity (BlastP score,
⬍
100; identity,
⬍
20 to 39%) to known proteins include
puta-tive TATA-box binding protein (TBP) (product of WSV303,
containing partial conservation with transcription initiation
factor IID repeat signature PS00351) (Fig. 2A), the putative
CREB-binding protein (CBP) (product of WSV100) (Fig. 2B),
nuclease (product of WSV191, containing most residues of
DNA/RNA nonspecific endonuclease active site PS01070), the
putative helicase (product of WSV447), and protein kinases
(products of WSV083, WSV289, and WSV423). Most of them
play important roles in the regulation of gene transcription.
TBP and CBP, which have never been reported in a virus
genome, deserve special attention since they are critical basal
transcription regulators in eukaryotic cells (21, 51). However,
their functions in virus are yet to be determined.
Structure proteins.
A unique feature of WSBV is that it
contains a collagen-like gene, WSV001, which encodes a
pre-dicted 1,684-amino-acid protein and whose transcription has
been confirmed by RT-PCR. The product of this ORF displays
FIG. 2. Multiple amino acid sequence alignment of products of WSV303 and WSV100. The homology regions are shaded (black, 100%; pink,
⬎
75%; blue,
⬎
50%). The positions of the amino acid sequence are indicated on both ends. (A) Alignment of product of WSV303 with a known
TBP. Human,
Homo sapiens
, accession no. XP_004534; yeast,
Saccharomyces cerevisiae
, accession no. M26403; fly,
Drosophila melanogaster
,
accession no. A35615; At,
Arabidopsis thaliana
, accession no. AC005223; Metha,
Methanothermobacter thermautotrophicus
, accession no.
AE000921; Archa,
Archaeoglobus fulgidus
, accession no. AE001078; Halob,
Halobacterium
sp. strain NRC-1, accession no. AE005110. (B)
Alignment of product of WSV100 with the CBP. Human,
Homo sapiens
, accession no. U47741; mouse,
Mus musculus
, accession no. S39161; fly,
D. melanogaster
, accession no. U88570; At,
A. thaliana
, accession no. AC024128.
on November 8, 2019 by guest
http://jvi.asm.org/
the highest degree of homology to human collagen type VII
(42% identity over 1,336 amino acids) (Fig. 3). This is the first
time that an intact collagen gene has been reported in a virus
genome. The collagen-like protein of WSBV contains a typical
repeat of Gly-X-Y (X is mostly proline, and Y can be any
amino acid) that can form the triple-helical structure
charac-teristics of animal collagen fiber. The presence of this
collagen-like protein may help to protect the WSBV from
environmen-tal factors and may contribute to its ability to survive for a long
time in a shrimp pond.
Previously only a short segment of collagen-homologous
se-quence was found in the structural proteins of ectocarpus
sil-iculosus virus 1 (EsV-1) (16), hepersvirus saimiri (HVS) (2,
22), and bacteriophage PRD1 (6, 7) (Fig. 3). In EsV-1, the
collagen-like sequence was found in the N-terminal half of
both vp55 and vp74 (16), which were encoded by the EsV-1
genome and which are likely to be the components of the viral
core structure. In HVS, the Gly-X-Y motif is repeated 18 times
and is located in the central region of saimiri
transformation-associated protein (STP). These collagen-like repeats may
serve as a hinge to extend the active domain of STP to its site
of action (2). Finally, in bacteriophage PRD1, a minor capsid
protein was found to contain a short collagen-like region
(Gly-X-Y)
6(7). All of the collagen-like segments present in these
proteins are short. These segments may play only a
supple-mentary role in protein functions.
In addition, WSV002 and WSV311 encode a nucleocapsid
protein, and the product of WSV421 shows characteristics of
an envelope protein. These proteins have recently been
puri-fied from the nucleocapsid and envelope of WSBV (49, 50).
WSV214 encodes a polypeptide with 44.2% basic amino acid
residues (Arg/Lys) and 24.6% Ser residues. This amino acid
composition is similar to that of the DNA-binding protein of
insect baculoviruses (34, 40, 55). Homologs of these
DNA-binding proteins have also been found in granulosis virus (47).
The basic residues of these DNA-binding proteins have a high
affinity for the phosphate backbone of DNA, enabling the
generation of a highly compact form of viral genomic DNA.
Upon entry into a host cell, the DNA-binding protein may
become phosphorylated by a protein kinase, resulting in the
unpacking of the viral DNA (54).
Protein motifs.
ORFs containing zinc finger and leucine
zipper motifs have been found in WSBV (Table 1). These
motifs have been shown to be involved in DNA-protein
inter-action and in regulation of transcriptional activation. Ring-H2
finger motifs, a variation of the Ring finger motif (30, 44)
found in proteins critical for virus survival and replication (11,
42), are also detected. Products of WSV079 and WSV427
contain an EF-hand calcium-binding motif (PS00018). Proteins
with these motifs are found in some prokaryotic and all
eu-karyotic organisms and play important roles in the regulation
and control of normal cellular functions. The detection of
these motifs in proteins of a marine virus suggests that some of
these basic regulatory activities are well conserved throughout
evolution.
The remaining 133 ORFs encode novel proteins of unknown
function. These novel genes obviously will provide ample
op-portunities for future research and for exploration of
molecu-lar mechanisms by which a virus and its host interact to survive
in the marine environment.
Among the 181 ORFs examined, the products of 96 have
potential transmembrane domains and 32 proteins contain
both signal peptide sequences and substantial hydrophobic
do-mains, suggesting that they may be membrane-associated
pro-teins and that they may play an important role in the
WSBV-host cell interaction and WSBV-host range determination. Other than
the putative signal sequences and hydrophobic domains, these
proteins are not obviously related to other known proteins.
[image:7.587.93.502.73.146.2]Repetitive regions.
Three percent of the WSBV genome is
composed of highly repetitive sequences, and the repeats are
distributed throughout the genome. We found nine
hr
s with a
total of 47 repeated minifragments encompassing direct
re-peats, atypical inverted repeat sequences, and imperfect
pal-indromic sequences. The nine
hr
s vary in size from 0.76 to 3.62
FIG. 3. Multiple amino acid sequence alignment of the product of WSV001 with human (
Homo sapiens
) type VII collagen, accession no.
L23982; fruit fly (
Drosophila melanogaster
) collagen, accession no. P08120; sea urchin (
Strongylocentrotus purpuratus
) collagen, accession no.
A43426; brown alga virus (BAV; ectocarpus siliculosus virus) collagen-like protein, accession no. NP_077542; HVS strain 484-77) collagen-like
protein, accession no. P25050; and bacteriophage PRD1 (PRD1) coat protein, which contains a short collagen-like region, accession no. P22536.
The homology regions are shaded (black, 100%; pink,
⬎
75%; blue,
⬎
50%). Repeat sequence density is shown as a ratio of
a
/
b
, in which
a
indicates
the length of the typical repeat sequence and
b
indicates the full length of the protein.
on November 8, 2019 by guest
http://jvi.asm.org/
kb, and
hr
1 to
hr
9 are separated in the WSBV genome by about
49, 13, 15, 28, 20, 28, 46, 36, and 55 kb of DNA, respectively.
Each
hr
contains several repeated minifragments, each with a
size around 300 bp. These minifragments are referred here as
a, b, c, d, e, f, etc. (Table 2). The percentage of homology
among the consensus sequences within the same homologous
region is over 73%, while the identity among the
hr
s is 61.6%
(Table 2). A few sequence motifs were found to be present at
very high copy numbers. For example, sequences CCAGAAA
or TTTCTGG, AGNGGTCCACC, and AACTTGACAT are
repeated 219, 88, and 47 times, respectively.
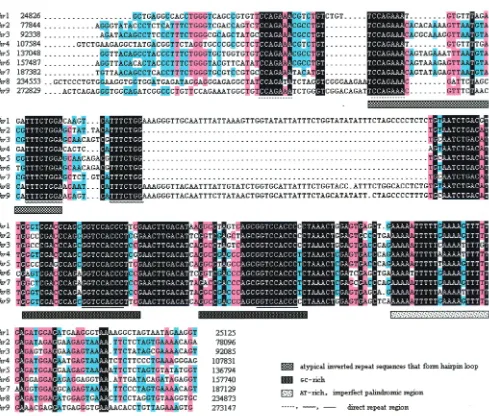
[image:8.587.43.284.85.195.2]As an example of such repetitive region, the homology
among the b minifragments of the nine
hr
s is shown in Fig. 4.
Both GC-rich sequences and AT-rich sequences are found in
the repeats. In the imperfect palindromic sequences, there are
2- or 3-bp mismatches that always exist in the same location
within every palindrome (Fig. 4), suggesting a functional
sig-nificance for the mismatch. Atypical inverted repeat sequences
that can form one or two hairpin loops are also found within
the repeat segments. The AT-rich elements, inverted repeat
sequences, and loop structures are reminiscent of the origin of
TABLE 2. Positions and identities of hrs in WSBV genome
hr Position Minifragment Identity (%)
Identity between
hrs (%)
hr
1
24528–28184
a, b, c, d, e, f, g
73.87
61.62
hr
2
77591–78859
a, b, c, d, e
87.98
hr
3
91832–92592
a, b, c,
88.85
hr
4
107335–108339
a, b, c, d
87.26
hr
5
136540–137301
a, b, c,
91.41
hr
6
157231–159211
a, b, c, d, e, f, g
74.35
hr
7
186876–188141
a, b, c, d, e,
89.65
hr
8
234231–236419
a, b, c, d, e, f, g
79.77
hr
9
272510–274432
a, b, c, d, e, f
80.86
FIG. 4. Alignment of partial consensus sequences within each
hr
. The consensus minifragments b are shown in order:
hr
1 to
hr
9. The
hr
s are
shaded (black, 100%; pink,
⬎
75%; blue,
⬎
50%), and the numbers on both ends refer to the positions of consensus sequences in the WSBV
genome. The direct repeat region, the atypical inverted repeat sequence that may contribute to the hairpin loop, the imperfect palindrome, and
GC-rich and AT-rich regions are shown.
on November 8, 2019 by guest
http://jvi.asm.org/
[image:8.587.49.538.275.691.2]replication in eukaryotic cells and also in some of the viruses
(17, 37). The presence of
hr
s is a feature of many baculovirus
genomes. The
hr
s may serve as transcription enhancers and
origins of DNA replication and play a fundamental role in the
viral life cycle (24, 27, 28, 33). The presence of nine
hr
s
sug-gests that WSBV may contain multiple replication origins. This
may account for the fast replication and the growth rate of
WSBV. Furthermore, although the organization of WSBV
hr
s
is similar to that of baculovirus, no homology among most of
their ORFs is detected. Thus, future investigations are needed
to determine whether WSBV is a seawater baculovirus and
whether the ancestors of WSBV and insect baculoviruses
evolved by separate routes, acquiring genes independently in
different environments.
In summary, we have obtained the complete genome
se-quence of WSBV. This is the first complete genome sese-quence
from a marine invertebrate virus. It is also the largest animal
virus genome sequenced (8, 52). As the genomic data
demon-strated, more than 80% of WSBV proteins bear no homology
to previously identified proteins. This leads us to consider a
separate evolutionary origin for this virus. Among the proteins
that show homology with known proteins, most seem to be
related to eukaryotic proteins and relatively few seem to be
related to viral proteins (Table 1). Although a few genes show
weak similarities to genes of herpesvirus (data not shown), the
morphology and the double-stranded circular WSBV genome
differ significantly from those of herpesvirus, which contains an
icosahedral capsid and a linear double-stranded DNA
mole-cule. On the other hand, WSBV shares some complex
mor-phological traits with the insect baculovirus, and a pattern of
interspersed repetitive regions in WSBV is similar to that
found in some of the insect baculoviruses, but sequence
com-parison indicates that they are not detectably related at the
amino acid level. Unfortunately, until now there were no
ge-nome sequence data available for the nonoccluded
baculovi-rus. Based on genetic analysis, WSBV clearly should not be
included in any of the currently recognizable baculovirus
sub-families and perhaps should be classified in a new virus family.
It is possible that other WSBV-like viruses that can infect other
organisms may exist. As the sequence of a representative of a
marine DNA virus, the complete WSBV genome sequence
should provide valuable information to serve as the genetic
basis for future studies. Future work may shed more light on
the evolution of these viruses.
ACKNOWLEDGMENTS
We thank Mei He and Yun Ye for their assistance, and we
acknowl-edge the support of Mingwei Wang, Lin Zao, and Yan Shen. We thank
Mark Yandell, Jennifer R. Wortman, Chinnappa Kodira, P. W. Li, and
Z. Deng of Celera Genomics for coordinating the project at Celera.
We thank Kunxin Luo of Lawrence Berkeley National Laboratory and
UC Berkeley for data analysis and critical reading of the manuscript.
This work is funded by the Chinese High Tech “863” Program
(Z19-02-05-01), Fujian Science Fund (C97053), and Science
Founda-tion of the State Oceanic AdministraFounda-tion.
REFERENCES
1.Afonso, C. L., E. R. Tulman, Z. Lu, E. Oma, G. F. Kutish, and D. L. Rock.
1999. The genome ofMelanoplus sanguinipesEntomopoxvirus. J. Virol.73:
533–552.
2.Albrecht, J. C., J. Nicholas, D. Biller, K. R. Cameron, B. Biesinger, C.
Newman, S. Wittmann, M. A. Craxton, H. Coleman, B. Fleckenstein, and
R. W. Honess.1992. Primary structure of the herpesvirus saimiri genome.
J. Virol.66:5047–5058.
3.Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman.1990.
Basic local alignment search tool. J. Mol. Biol.215:403–410.
4.Arif, P.1988. A sequence motif in many polymerases. Nucleic Acids Res.
16:9909–9916.
5.Baldo, A. M., and M. A. McClure.1999. Evolution and horizontal transfer of
dUTPase-encoding genes in viruses and their hosts. J. Virol.73:7710–7721.
6.Bamford, D. H., and J. K. Bamford.1990. Collagenous proteins multiply.
Nature344:497.
7.Bamford, J. K., and D. H. Bamford.1990. Capsomer proteins of
bacterio-phage PRD1, a bacterial virus with a membrane. Virology177:445–451.
8.Bankier, A. T., S. Beck, R. Bohni, C. M. Brown, R. Cerny, M. S. Chee, C. A.
Hutchinson, T. Kouzarides, J. A. Martignetti, E. Preddie, S. C. Satchwell, P.
Tomlinson, K. M. Weston, and B. G. Barrell.1991. The DNA sequence of
the human cytomegalovirus genome. DNA Seq.2:1–12.
9.Belfort, M., A. Moelleken, G. F. Maley, and F. Maley.1983. Purification and
properties of T4 phage thymidylate synthase produced by the cloned gene in an amplification vector. J. Biol. Chem.258:2045–2051.
10.Bodemer, W., H. H. Niller, N. Nitsche, B. Scholz, and B. Fleckenstein.1986.
Organization of the thymidylate synthase gene of herpesvirus saimiri. J. Vi-rol.60:114–123.
11.Borden, K. L.2000. RING domains: master builders of molecular scaffolds?
J. Mol. Biol.295:1103–1112.
12.Carreras, C. W., and D. V. Santi.1995. The catalytic mechanism and
struc-ture of thymidylate synthase. Annu. Rev. Biochem.64:721–762.
13.Cedergren-Zeppezauer, E. S., G. Larsson, P. O. Nyman, Z. Dauter, and K. S.
Wilson.1992. Crystal structure of a dUTPase. Nature355:740–743.
14.Cen, F.1998. The existing condition and development strategy of shrimp
culture industry in China, p. 32–38.InY. Q. Su (ed.), The health culture of shrimps. China Ocean Press, Beijing, People’s Republic of China.
15.Chen, X. F., C. Chen, D. H. Wu, H. Huai, and X. C. Chi.1997. A new
baculovirus of cultured shrimp. Sci. China Ser. C40:630–635.
16.Delaroque, N., S. Wolf, D. G. Muller, and R. Knippers.2000.
Characteriza-tion and immunolocalizaCharacteriza-tion of major structural proteins in the brown algal virus EsV-1. Virology269:148–155.
17.DePamphilis, M. L.1993. Origins of DNA replication that function in
eu-karyotic cells. Curr. Opin. Cell Biol.5:434–441.
18.Earl, P. L., E. V. Jones, and B. Moss. 1986. Homology between DNA
polymerases of poxviruses, herpesviruses, and adenoviruses: nucleotide se-quence of the vaccinia virus DNA polymerase gene. Proc. Natl. Acad. Sci. USA83:3659–3663.
19.Fleischmann, R. D., M. D. Adams, O. White, R. A. Clayton, E. F. Kirkness,
A. R. Kerlavage, C. J. Bult, J. F. Tomb, B. A. Dougherty, J. M. Merrick, et al.1995. Whole-genome random sequencing and assembly ofhaemophilus
influenzaeRd. Science269:496–512.
20.Francki, R. I. B., C. M. Fauquet, D. L. Knudson, and F. Brown.1991.
Classification and nomenclature of viruses. Fifth report of the International Committee on Taxonomy of Viruses. Arch. Virol.1991(Suppl. 2):1–450.
21.Furukawa, T., and N. Tanese.2000. Assembly of partial TFIID complexes in
mammalian cells reveals distinct activities associated with individual TATA box-binding protein-associated factors. J. Biol. Chem.275:29847–29856
22.Geck, P., S. A. Whitaker, M. M. Medveczky, and P. G. Medveczky.1990.
Expression of collagenlike sequences by a tumor virus, herpesvirus saimiri. J. Virol.64:3509–3515.
23.Geourjon, C., and G. Deleage.1995. ANTHEPROT 2.0: a three-dimensional
module fully coupled with protein sequence analysis methods. J. Mol. Graph.
13:209–212.
24.Guarino, L. A., and W. Dong.1991. Expression of an enhancer-binding
protein in insect cells transfected with theAutographa californicanuclear polyhedrosis virus IE1 gene. J. Virol.65:3676–3680.
25.Hofmann, K., P. Bucher, L. Falquet, and A. Bairoch.1999. The PROSITE
database, its status in 1999. Nucleic Acids Res.27:215–219.
26.Honess, R. W., W. Bodemer, K. R. Cameron, H. H. Niller, B. Fleckenstein,
and R. E. Randall. 1986. The A⫹T-rich genome ofHerpesvirus saimiri
contains a highly conserved gene for thymidylate synthase. Proc. Natl. Acad. Sci. USA83:3604–3608.
27.Kool, M., P. M. Van Den Berg, J. Tramper, R. W. Goldbach, and J. M. Vlak.
1993. Location of two putative origins of DNA replication ofAutographa
californicanuclear polyhedrosis virus. Virology192:94–101.
28.Kool, M., J. T. Voeten, R. W. Goldbach, J. Tramper, and J. M. Vlak.1993.
Identification of seven putative origins ofAutographa californicamultiple nucleocapsid nuclear polyhedrosis virus DNA replication. J. Gen. Virol.
74:2661–2668.
29.Larder, B. A., S. D. Kemp, and G. Darby.1987. Related functional domains
in virus DNA polymerases. EMBO J.6:169–175.
30.Leverson, J. D., C. A. Joazeiro, A. M. Page, H. K. Huang, P. Hieter, and T.
Hunter.2000. The APC11 RING-H2 finger mediates E2-dependent
ubiq-uitination. Mol. Biol. Cell11:2315–2325.
31.Lightner, D. V.1996. A handbook of pathology and diagnostic procedures
for diseases of penaeid shrimp. World Aquaculture Society, Baton Rouge, La.
32.Lo, C. F., C. H. Ho, S. E. Peng, C. H. Chen, H. C. Hsu, Y. L. Chiu, C. F.
Chang, K. F. Liu, M. S. Su, C. H. Wang, and G. H. Kou.1996. White spot
on November 8, 2019 by guest
http://jvi.asm.org/
syndrome baculovirus (WSBV) detected in cultured and captured shrimp, crabs and other arthropods. Dis. Aquat. Org.27:215–226.
33.Lu, A., P. Krell, J. M. Vlak, and G. F. Rohrmann.1997. Baculovirus DNA
replication.InL. K. Miller (ed.), The baculoviruses. Plenum Press, New York, N.Y.
34.Maeda, S., S. G. Kamita, and H. Kataska.1991. The basic DNA-binding
protein of Bombyx morinuclear polyhedrosis virus: the existence of an additional arginine repeat. Virology180:807–810.
35.McGeoch, D. J.1990. Protein sequence comparisons show that the
’pseudo-proteases’ encoded by poxviruses and certain retroviruses belong to the deoxyuridine triphosphatase family. Nucleic Acids Res.18:4105–4110.
36.Muller, K., C. A. Tidona, U. Bahr, and G. Darai.1998. Identification of a
thymidylate synthase gene within the genome of Chilo iridescent virus. Virus Genes17:243–258.
37.Pearson, M., R. Bjornson, G. Pearson, and G. Rohrmann.1992. The
Au-tographa californica baculovirus genome: evidence for multiple replication origins. Science257:1382–1384.
38.Pearson, W. R.1990. Rapid and sensitive sequence comparison with FASTP
and FASTA. Methods Enzymol.183:2444–2448.
39.Richter, J., I. Puchtler, and B. Fleckenstein.1988. Thymidylate synthase
gene of herpesvirus ateles. J. Virol.62:3530–3535.
40.Russell, R. L., and G. F. Rohrmann.1990. The p6.5 gene region of a nuclear
polyhedrosis virus ofOrgyia pseudotsugata: DNA sequence and transcrip-tional analysis of four late genes. J. Gen Virol.71:551–560.
41.Sambrook, J., E. F. Fritsch, and T. Maniatis.1989. Molecular cloning: a
laboratory manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
42.Saurin, A. J., K. L. Borden, M. N. Boddy, and P. S. Freemont.1996. Does
this have a familiar RING? Trends Biochem. Sci.21:208–214.
43.Tomalski, M. D., J. G. Wu, and L. K. Miller.1988. the location, sequence,
transcription, and regulation of a baculovirus DNA polymerase gene. Virol-ogy167:591–600.
44.Torii, K. U., C. D. Stoop-Myer, H. Okamoto, J. E. Coleman, M. Matsui, and
X. W. Deng.1999. The ring finger of photomorphogenic repressor COP1
specifically interacts with the RING-H2 motif of a novelArabidopsisprotein. J. Biol. Chem.274:27674–27681.
45.Tsai, M. F., C. F. Lo, M. C. van Hulten, H. F. Tzeng, C. M. Chou, C. J.
Huang, C. H. Wang, J. Y. Lin, J. M. Vlak, and G. H. Kou.2000.
Transcrip-tional analysis of the ribonucleotide reductase genes of shrimp white spot syndrome virus. Virology277:92–99.
46.Tsai, M. F., H. T. Yu, H. F. Tzeng, J. H. Leu, C. M. Chou, C. J. Huang, C. H.
Wang, J. Y. Lin, G. H. Kou, and C. F. Lo.2000. Identification and
charac-terization of a shrimp white spot syndrome virus (WSSV) gene that encodes a novel chimeric polypeptide of cellular-type thymidine kinase and thymidy-late kinase. Virology277:100–110.
47.Tween, K. A., L. A. Bulla, and R. A. Consigli.1980. Characterization of an
extremely basic protein derived from granulosis virus nucleocapsid. J. Virol.
33:866–876.
48.van Hulten, M. C., M. F. Tsai, C. A. Schipper, C. F. Lo, G. H. Kou, and J. M.
Vlak.2000. Analysis of a genomic segment of white spot syndrome virus of
shrimp containing ribonucleotide reductase genes, and repeat regions. J. Gen. Virol.81:307–316.
49.van Hulten, M. C., M. Westenberg, S. D. Goodall, and J. M. Vlak.2000.
Identification of two major virion protein genes of white spot syndrome virus of shrimp. Virology266:227–236.
50.van Hulten, M. C., R. W. Goldbach, and J. M. Vlak.2000. There functionally
diverged major structural proteins of white spot syndrome virus evolved by gene duplication. J. Gen. Virol.81:2525–2529.
51.Van Orden, K., and J. K. Nyborg.2000. Insight into the tumor suppressor
function of CBP through the viral oncoprotein tax. Gene Expr.9:29–36.
52.Vink, C., E. Beuken, and C. A. Bruggeman.2000. Complete DNA sequence
of the rat cytomegalovirus genome. J. Virol.74:7656–7665.
53.Volkman, L. E.1995. Baculoviridae, p. 104–113.InF. A. Murphy and C. M.
Fauquet (ed.), Virus taxonomy. Sixth report of the International Committee on Taxonomy of Viruses. Springer-Verlag, New York, N.Y.
54.Wilson, M. E., and R. A. Consigli.1985. Functions of a protein kinase activity
associated with purified capsids of the granulosis virus infectingplodia
inter-punctella. Virology143:526–535.
55.Wilson, M. E., T. H. Mainprize, P. D. Friesen, and L. K. Miller.1987.
Location, transcription, and sequence of a baculovirus gene encoding a small arginine-rich polypeptide. J. Virol.61:661–666.
56.Yang, F., W. Wang, and X. Xu.1997. A simple and efficient methods for
purification of prawn baculovirus DNA. J. Virol. Methods67:1–4.
57.Zhan, W. B., and Y. H. Wang.1998. White spot syndrome virus infection of
cultured shrimp in China. J. Aquat. Anim. Health10:405–410.
on November 8, 2019 by guest
http://jvi.asm.org/