JOURNAL OF VIROLOGY,Sept.1983,p.631-636
0022-538X/83/090631-06$02.00/0
Copyright©1983,AmericanSocietyfor Microbiology
Vol. 47, No. 3
Loss
of Viral Genomes from
Hamster Tumor Cells and
Nonrandom Alterations in
Patterns
of Methylation of
Integrated
Adenovirus
Type 12 DNA
INGRID KUHLMANNt ANDWALTERDOERFLER*
Instituteof Genetics, University of Cologne, Cologne, Germany Received 11 March1983/Accepted 31 May 1983
The
insertion
stability and DNA methylation patterns of integratedadenovirustype 12(Adl2)genomes wereinvestigatedin Adl2-inducedtumors and in tumor
cell lines established from them as afunction of time ofpassage under culture
conditions. Upon subcultivation of cells from some of the tumors, the viral
genomes wereeliminated, apparently inastepwise process with segments of the
left termini ofAdl2DNAs persisting the longest. Morphological variants of these
tumorcells lost all viral DNA and yet retained the oncogenic phenotype. All 13
independently isolated clones fromone revertantlinewere devoid ofAdl2DNA.
Itcouldnotbe ruledoutthat very short sequence elements of viral DNA, such as
promoters orenhancing sequences, could have persisted in these variants. The
extentof viral DNA methylation was minimal inAdl2-induced tumors, although
theviralgenome was notextensively expressed, ifatall.Upon passage in culture,
the levels of viral DNA methylation increased. It was interesting that establish-ment
of
thefinalmethylation pattern of integrated Adl2DNAs required many cellgenerations after the fixation of foreign DNA in the host genome. The shift in
methylationwasnonrandom. The latepartsoftheinserted viralgenomes became
methylated moreextensively than did the early gene segments.
The results ofa detailed study of the integra-tion andmethylation patterns of adenovirustype 12(Adl2) DNA in 39Ad12-inducedtumors and
cell lines established from these tumors have been
reported previously
(9, 10). In over 70 different adenovirus-transformed cell lines and Adl2-inducedtumorsor tumorlines,
thesiteof viral DNA insertion differs from cell lineto cellline (3). In general, the
patterns
of viral DNAinsertion have proven to be quite stable for a
given
cell line (16). However, morphological variants of Ad12-transformed hamster cells (6) orofAdl2-induced hamstertumorcells (9) have been observed whichappearedmorefibroblastic than theoriginal
tumorcells. Althoughsome of these variants have been characterized by thecomplete
elimination of viral DNA fromthehostgenome(5, 9), these cells retain their oncogenic
phenotype upon injection into hamsters (9).
From thesefindings, it has been concluded that
theoncogenic phenotype is compatible withthe
complete loss of Adl2DNAfrom cells that had
originally
been transformed by Adl2. Presum-ably, in hamster cells, Adl2 infection triggers hereto unknown mechanisms that entailonco-genic transformation,
and thistransformation istPresent address:Institute ofVirology,Universityof
Co-logne, CoCo-logne,Germany.
accompanied by
fixation ofmultiple copies
of viralDNA. Atleast inthecells in whichtheviralgenome was
completely lost,
stability
of theoncogenic phenotype
is notpremised
on the persistence of viral genes(3, 9).
Conceivably,
the decisivetransforming
events areelicitedby
thefixation
of viral genes,expression of
viralgenes, or
both,
but thecontinuedrealization
oftransformation
once initiated may be indepen-dentof viralfunctions.
There is nowextensive evidence for the
no-tion that DNA
methylation
inhighly
specific
sequences can play a decisive role in the
long-term shutoff ofgenes in eucaryotes
(for
recentreviews, seereference4and W.
Doerfler,
Curr. Top. Microbiol.Immunol.,
inpress). It appearsthat theabsence ofDNA
methylation
isaneces-sary but not sufficient
precondition
for geneexpression to proceed (9, 17). This latter con-cepthadbeensupported
by
thestriking
observa-tion thatthelevels ofAdl2DNAmethylation
inAdl2-induced tumors were very
low,
if DNAmethylation
could be detected atall,
at 5'-CCGG-3' and 5'-GCGC-3' sites (10).However,
viral gene expression not at all detectable inthese tumors or at an
exceedingly
low level isnot commensurate with the low level ofDNA
methylation (10). Upon
explantation
andcontin-ued cultivation ofthesetumor
cells,
the extent631
on November 10, 2019 by guest
http://jvi.asm.org/
632
NOTESJ.Vo.
0
I
:---EcoRIl C D B
FIG. 1. Integration p~
Ad12-induced hamstert and in cell lines Hllll from these tumors, respi fromtumorsorfromcell
explantationfrom thetul
arations (10 p.g) were cl
ments were separated
agarose gels and blotted and Ad12-specific fragnf bridizationto 32P-labele4
diography. Adl2 DNA genome equivalents [20.
10 p.gofcellular DNA
cells)wasmixed with 10 treated in thesame manr
fragments (A to F) and
(Kbp), as well as theEK also shown.
of DNA methylationi
sequences increases g In the present cor
strate that Adl2 DNA cells inastepwisefast,
the viral DNAs vanis the levels of Adl2 D' lished hamster tumor
nonrandom. The 5'-C termini of Adl2 DNA
ylated. Inthisway, tI
patterns approached I
Ad12-transformed cel
co periods oftime
(8, 15).
Viral geneexpression
in~~ ~ -~~ Ad12-inducedtumors wasminimal.
~~~L'Ci
~The
induction oftumorsby
Adl2
in newbornE
~~~~~~hamsters
and theestablishmentof cell lines fromX
~~~~Kbp
these tumors, as well as the methods used todeterminepatternsofviral DNA
integration
andmethylation,
have beendescribedearlier(9,
10).A1387
In theseexperiments,
the Southernblotting
B7
technique (13)
and a DNA-DNAhybridization
procedure (20)
wereused,
employing
nick-trans-*-C5.4
lated(11)
3P-labeled Adl2 DNAorcloned viral DNAfragments (18).
DNAmethylation
ofinte-4"4
-0)4.0
grated viral DNA sequences was detectedby
hybfidization
of _Plabeled Adl2 DNA to blot-ted tumor cell DNA that hadpreviously
been-E24 cleaved with the isoschizomeric restriction
en--E24donuclease
pair Hpall
orMspI (19)
or withHhaIL
Inthe latterinstance, Hhal-cleaved Adl2
virion
DNA,
which isunmethylated (7),
wasusedas areference.
From 39
Ad12-induced
tumors, 13 cell lineswere established, and in 8 cases, the cell lines
Fo6 were carried
continuously
over 10or morepas-sages. In three lines, the patterns of
integration
and copy numbers of viral DNA per cell didnot
change.
In three otherlines,
viral DNA was no- FA
longer
detectable even after the first passage.One cell line was not
analyzed
further. In cell)atterns of viral DNA in the ln 11() nymnrcagswr b
,umors Tl111(1) and Tl11112
linvedHFig.12) only, sminor chaneofsiwee
ob-nd
(1) and H1111(2) established
dsaperved(i.
1);e.g. some
otHell()
offsiz bandsectively.
DNAwasextracteddisappgearmed.I
eluvlet
lierHe
)allt
10tosag
11Isatstagesof passage(p)after virlgnm qiaet eels npsae2
mnors
asindicated. DNA prep-(Fig.
1). In passage 12, 3 to 4 genome equiva-leaved with EcoRI, the frag- lentsremained,
exhibiting
the sameintegration
byelectrophoresis
on 0.5% pattern as inprevious
passages. The numberof1 (13) tonitrocellulose filters,
Adl2
genomeequivalents
persisting
in passageients were visualized by hy- 12 of line
Hllll(1)
was estimated from thed1(11)
Adl2DNAandautora-intensity
of the internal viral DNA bands as (0.5 ng is equivalent to 20 shown inFig.
1by
aspectrophotometric
scan-x] percell, correspondingto cdr sdsrbdpeiul 1)
and theequivalent numberof ning procdrasecibdpvoul(1)
I
±g
ofB3(BHK-21)DNAand Both the internalAdl2
fragments
and the Adl-rier.Theauthentic virion DNAspecific
off-size bandsweredecreasedinintensi-Itheir sizes in kilobase pairs ty. Thus, it was most
likely
that total Adl2IcoRI mapofAdl2 DNA, are genomes had been lost
during
passage of the tumorcells. In passage 13 of cell lineHllll(l),
one to two
copies
of the left terminus ofAdl2DNA
persisted,
the EcoRI Dfragment
and aniin
theintegrated
Adl2 off-sizefragment
hybridizing
to bothtermini. In;,radually
(10). thisexperiment,
themolecularly
cloned terminimmunication, we demon- of Adl2 DNA, the EcoRI C and A*
fragments
A can be lost from tumor (18), were used as
hybridization probes
(refer-iionwith theleft termini of ence9,cf. maps in
Fig.
1 and2b).
In DNAfrom;hing
last. The increase in passage 20 onward, viral DNA could not be INAmethylation
in estab-detected,
eventhough
these cells continued tocell lines wasfound tobe be
oncogenic
in hamsters(9). Thesensitivity
of.,CGG-3'
sites at theright
the blothybridization
experiments
wassuch that Ls wereincreasingly
meth- onecopyof the left 10% of the genome per cell ieAdl2
DNAmethylation
would have been detected. In passage 12 of cellthose described earlierfor line Hllll(l), 13 cellular clones were
estab-LI lines cultured over
long
lished. In the DNA of these clones, viral DNAJ. VIROL.
0
'r :I? C114 Cl. a a
C--;=. -,;=. -;:x ;.
x T m I
-- - -- ----Cl
on November 10, 2019 by guest
http://jvi.asm.org/
[image:2.492.51.245.63.349.2]NOTES 633
a Approximatesize
of DNAfragment
inbase pairs [bp]
4150
2464
Band pattern 0kd12DNA MspI
A-B
1700 1650
1250 1150 1050 940 850
760 710 580 550 504
E.F-
GE,
H,-J
K.L
M
NO0 P Q-R,S
Ad12DNA,leftterminus
b
Ela ElbMspl R
od.
Hpall
bpI
Locatedin
fragment
EcoRIC
EcoRIC
EcoRIX FIG. 2. Restriction pattern and partial map ofAdl2 DNAfor the isoschizomeric restriction enzyme pair
EcoRIB
MspI
andHpaII.
(a)Adl2
DNA was cleaved withEco
RIA*
MspI,
and thefragments
wereseparated by
gel
elec-EcoRI A trophoresis on a 1% agarose gel and blotted to nitrocel-lulose paper. Individualfragments were located on the Adl2 map by hybridizing independent blots with 32p-EcoRI B labeled clonedfragments of Adl2 DNA as indicated.
EcoRIB Theestimated sizes of
MspI
fragments
AtoSarealsoindicated in base pairs (bp). The results have been EcoRI A summarized schematically. (b) Restriction maps of the EcoRIB left(26%) and theright
(40%)
termini of Adl2 DNA Eco RIC aftercleavage with HpaII or MspI. These maps have EcoRIC beenpublished previously(8).punits
MspI
HpaII bp
90
iMap
unitscouldnotbedetectedby Southernblotting,even
whenupto 300pgof cellular DNA wasblotted
on nitrocellulose filters. In these hybridization
experiments, the32P-labeled totalAdl2genome
aswellasclonedfragments,inparticularthe left
terminal EcoRI Cfragment (comprising the El
region of Adl2 DNA) and EcoRI D fragment,
wereusedashybridization probes (formaps, see
Fig. 2).
We conclude that viral DNA can be deleted
apparently in a stepwise manner from
Adl2-induced hamstertumorcells.The morphological
variants that lost all viral DNAcan retain their
oncogenic phenotypes, although they do not
carrythe Elregion of Adl2 DNA. It will bevery
difficulttorigorouslyruleoutthepossibility that
smallelements of viralDNA,e.g., apromoteror
enhancer sequence (2, 21), do persist in the
variant cell line. It shouldbe pointedoutthat the
celllinesestablished fromtumorswereprobably
closeto being clonal, as most tumorsexhibited
rather homogeneous integration patterns of
S B Q A
5soi22 2464 | AQ 4150 1
EcoC Eco D Eco B
10 20 3)0 40
E2b (notepPressed)
A1DE2a E3 E4
Adl12 DNAright terminus ,N
D I C G0
li 1
JU
1Ro I1s
|X|11700
l>dT EcoA ] A
60 70 80
VOL. 47,1983
on November 10, 2019 by guest
http://jvi.asm.org/
[image:3.492.53.448.51.551.2]634 NOTES
rwtl - rx at l
U- C]-e: 3
*~~~~~~~~~
.m
-N O* t. SI
FIG. 3. Nonrandom changes of methylation
pat-ternsofAdl2DNAinanAdl2-induced hamstertumor
line. DNAwasisolated from theAdl2-inducedtumor
T1111(2) andfrom cell line H1111(2)establishedfrom this tumor. DNA waspurifiedin passage 3(p3)or16
(p16) of this cell line. All procedures used in DNA
isolation have beenreportedelsewhere(10, 15).These DNApreparations (10 ,ug perslot)or0.5 ngof Adl2
virionDNA(equivalentto20genomeequivalentsper
cell), mixed with 10,ugof B3 hamster DNAascarrier, were cleaved withMspIorHpaIIasindicated. Frag-ments were separated by electrophoresis on a 1% agarosegel, blotted (13), andhybridizedtoAdl2 DNA that was32P-labeledby nick translation (11).Letters A
toSdesignate marker DNAfragments.
Adl2 DNA. Moreover, the cell lines described
herewere derived fromclonedcells.
We alsoinvestigatedin detailthe shifts in viral
DNA methylation that occurred upon
cultiva-tion of Ad12-inducedhamstertumor cells. These
shifts fromlow-level DNA methylation to more
extensive levels did not occur randomly but
followed certain patterns. Theincrease in Adl2
DNAmethylation was surveyed by cleaving the
DNA fromtumor cells at various passages with
HpaIIorMspI, by gel electrophoresis and
blot-ting,
and by hybridization to32P-labeled
Adl2 DNA. The MspI (recognition sequence5'-CCGG-3')
mapof theleftterminal 26%and therightterminal40% ofthe Adl2genome has been
J. VIROL.
establishedpreviously (8). AlthoughMspI sites
in the remaining 34% ofthe viralgenome have notyetbeenmapped,the MspIfragments E, F, K, L, M, and Pwere localized in the EcoRI B
fragment by blotting MspI-cleaved Adl2 DNA
and by hybridization to the cloned EcoRI B
fragment(18). Inaddition, fragmentB harbored
a large number ofvery small MspI fragments.
The schemes inFig. 2aassigned map locations
to MspI fragments A to S (4,150 to about 504
base pairs in length). Figure 2b presents
previ-ously published partialMspI maps (8).
The result presented inFig. 3 actually
demon-strated the increase in DNA methylation in the
integrated Adl2 genomes by comparing the
HpaIlandMspI cleavage patterns of viral DNA
from the tumor T1111(2) with those from cell
lineH1111(2)inpassage3 andpassage16. Itwas apparentthat HpaII bands predominantly from
the (late) EcoRI B fragment and from the right
terminal EcoRI A fragmentstarted disappearing
atlowpassagelevels,whereas theHpaIIbands
7-.
C - I
7n- cnc)
I
-M'sp1.
[image:4.492.87.212.73.361.2]FK
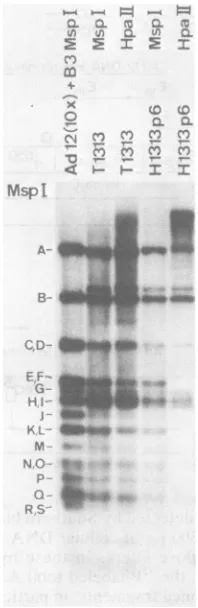
FIG. 4. Changes in methylation
patterns
in Ad12-induced hamstertumorline H1313.Experimental con-ditions were similar to those described for Fig. 3. T1313 and H1313 p6 refer to DNA from the Ad12-induced tumorT1313 and the cell line H1313 inpas-sage6, respectively.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:4.492.300.399.309.613.2]VOL. 47, 1983 NOTES 635
o r - t:
+ + + + + ++++ ++ ++++ ++ + ++ + + + + + + *+ + +++++ ++ +++++ ++ +++++ + ++++ ++ ++++ ++ ++ + + + ++ + +++++ + + + + ++ + + ++++ + ++ + + + + + + ++ + ++ + +++ + +++ + + t0J X W _ I.
-gkJ
- 0-_ _ -6- b_-t-J 0 -o _ _ I._ x C) " <n PZ 0e Pz aF a n (L) v) C., (A 9 (A 0z
0 n- I.-0. 0 CL co t= 0. 0I0
Cs m CDO0
Ds 0 3 Cs rJ C0. 0. CD Cs 0. U) Cs 0z 0q Cs Osin theearly regions of the genome, particularly
in region El, remained unaltered. The data
presentedin Fig.3and4demonstratedthat with
increasing passage ofcells inculture the HpaII
bands C, D, E, F,H, I, K, L, M,andprobably
smaller fragments as well started to disappear
frompassage 3 onward.Disappearance of HpaII
bands obviously signified methylation at the corresponding 5'-CCGG-3' sites. At higher pas-sage levels, mainly the HpaII sites in early regions oftheintegrated viralgenomeremained unmethylated. Thus, the pattern of DNA meth-ylation approached thatobserved in Adl2-trans-formed cell lines (15). Similar resultswere
pre-sented for DNA fromtumorT1313andcellline H1313 (Fig. 4). It was interesting to note that establishment of the finalpatternofmethylation required many cellgenerations after fixation of the viral DNA in the host genome. Similar observationswere made with othertumors and
tumorlines and also after cleavage of the DNA
withHhaI (10). In severalof the Ad12-induced
tumorsandtumorlinesa
1,400-base-pair off-size
fragmentwasapparentthatcorrespondedtothe junction between the left terminus of viral DNA and cellular DNA. In some cell lines, the 5'-CCGG-3' sites
flanking
thisfragment
became methylated with continued passage; in other lines, thesame sites remainedunmethylated.In Table 1, the results ofmany experiments
aresummarized.Ingeneral, the conclusion held thatwith
increasing
passage the lateregions
oftheAdl2genome wereprogressively methylated
at5'-CCGG-3' sites.
A total of 17 Adl2-induced tumors and 3
tumor cell lines were screened for the
occur-rence of Adl2-specific cytoplasmic RNA. By
Northernblotting (1, 12), viralRNA was
detect-edinonlytwo tumorsin smallamounts(datanot
shown). Thus, the low level of viralDNA
meth-ylation
did not correspond tofull-fledged
expression of the inserted viralgenome. On thecontrary, the viral genomesseemed to be
com-pletelyshutoff; not eventhe El region seemed
tobeexpressed. Itwill be
interesting
toinvesti-gatewhat factors influence the levels of methyl-ation of inserted Adl2DNA andby what mecha-nisms the
expression
of the viral genome is controlled atearly
and late times after the fixa-tion offoreignDNAin thetumorcells. Obvious-ly, the stringency ofthese control mechanismscould
change
with timeofpassage under cultureconditions.
I.K.wassupported byafellowship oftheFriedrichEbert
Foundation. This researchwasmadepossible byagrantfrom
the Deutsche Forschungsgemeinschaft through
Sonderfors-chungsbereich74C1.
We thank Hanna Mansi-Wothke for the preparation of media.
on November 10, 2019 by guest
http://jvi.asm.org/
636 NOTES
LITERATURECITED
1. Alwine, J.C.,D.J. Kemp,andG.R.Stark. 1977.Method for detection ofspecific RNAs inagarosegels by transfer
to diazobenzyloxymethyl-paper and hybridization with DNA probes. Proc. NatI. Acad. Sci. U.S.A. 74:5350-5354.
2. Banerji,J., S. Rusconi,andW.Schaffner. 1981. Expres-sion ofa 3-globingeneisenhancedbyremoteSV40 DNA
sequences.Cell 27:299-308.
3. Doerfler, W. 1982. Uptake, fixation, and expression of foreign DNA in mammalian cells: the organization of integrated adenovirus DNAsequences.Curr. Top. Micro-biol. Immunol. 101:127-194.
4. Doerfier, W. 1983. DNAmethylation andgene activity. Annu. Rev. Biochem. 52:93-124.
5. Eick, D.,S. Stabel,and W.Doerfler. 1980.Revertants of adenovirustype12-transformed hamster cell line T637as
tools in the analysis of integration patterns. J. Virol. 36:41-49.
6. Groneberg, J.,D.Sutter, H. Soboll,and W.Doerfler.1978. Morphological revertants of adenovirus type 12-trans-formed hamster cells.J. Gen. Virol. 40:635-645.
7. Gunthert, U.,M. Schweiger,M.Stupp,and W.Doerfler. 1976.DNAmethylation in adenovirus, adenovirus-trans-formed cells, and host cells. Proc. Natl. Acad. Sci. U.S.A. 73:3923-3927.
8. Kruczek, I., and W. Doerfier. 1982. The unmethylated
stateof thepromoter/leader and 5'-regions of integrated adenovirusgenescorrelateswithgeneexpression. EMBO J. 1:409-414.
9, Kuhlmann, I.,S.Achten, R.Rudolph,andW.Doerfler. 1982. Tumor inductionby human adenovirustype12 in hamsters:loss of the viralgenomefromadenovirustype
12-induced tumorcells is compatible withtumor forma-tion. EMBOJ. 1:79-86.
10. Kuhlmann, I.,and W.Doerfler. 1982. Shiftsin theextent
andpatternsof DNAmethylationuponexplantation and subcultivation of adenovirus type 12-induced hamster
tumor cells. Virology 118:169-180.
11. Rigby, P. J. W., M.Dieckmann, C. Rhodes, and P. Berg. 1977. Labeling deoxyribonucleic acid to high specific
activity in vitro by nick translation with DNA polymerase I. J. Mol.Biol. 113:237-251.
12. Schirm, S., and W. Doerfler. 1981. Expressionof viral DNA inadenovirus type 12-transformedcells,in tumor cells,and in revertants. J. Virol. 39:694-702.
13. Southern, E. M. 1975. Detection ofspecific sequences among DNAfragments separated by gel electrophoresis. J. Mol. Biol. 98:503-517.
14. Stabel, S., W.Doerfier, and R. R. Friis. 1980. Integration sites of adenovirus type 12 DNA intransformedhamster cells and hamster tumor cells. J.Virol. 36:22-40. 15. Sutter, D., and W.Doerfier. 1980. Methylationof
integrat-ed adenovirus type 12 DNA sequences intransformed cells isinversely correlated with viral gene expression.
Proc.Natl. Acad. Sci. U.S.A. 77:253-256.
16. Sutter, D., M. Westphal, and W. Doerfler. 1978.Patterns ofintegration of viral DNA sequences in the genomes of adenovirus type 12-transformed hamster cells. Cell 14:569-585.
17. van der Ploegh, L. H. T., and R. A.Flavell.1980. DNA methylation in the human ybI-globinlocus in erythroid andnonerythroidtissues. Cell 19:947-958.
18. Vogel, S., M.Brotz,I.Kruczek, R. Neumann, D. Eick, U. Winterhoff, and W. Doerfler. 1981.Cloned fragments of humanadenovirus type 12-DNA. Gene 15:273-278. 19. Waalwik, C., and R. A. Flavell. 1978.MspI, an
isoschi-zomerof HpaII which cleaves bothunmethylated and methylatedHpaIIsites. Nucleic Acids Res. 5:3231-3236. 20. Wahl, G. M., M. Stern, and G. R. Stark. 1979. Efficient
transfer of large DNA fragments from agarose gels to
diazobenzyloxymethyl-paper and rapid hybridization by using dextran sulfate. Proc. Natl. Acad. Sci. U.S.A. 76:3683-3687.
21. Weiher, H., M.Konig,and P. Gruss.1983.Multiple point
mutationsaffecting the simian virus 40 enhancer. Science 219:626-631.
J. VIROL.
![FIG.(0.5 ng is equivalent to 20x] per cell, corresponding toand the equivalent number ofI±g of B3 (BHK-21) DNA andrier](https://thumb-us.123doks.com/thumbv2/123dok_us/1444055.96819/2.492.51.245.63.349/fig-equivalent-cell-corresponding-toand-equivalent-number-andrier.webp)