JOURNALOF VIROLOGY,Jan. 1986, 343-348 0022-538X/86/010343-06$02.00/0
Copyright © 1986, American Societyfor Microbiology
Functional cDNA Library
For
Efficient
Expression of Measles
Virus-Specific
Gene Products in Primate Cells
TIMOTHY C. WONG*AND AKIKO HIRANO
DepartmentofMicrobiology and Immunology, Universityof Washington Schoolof Medicine, Seattle, Washington 98195
Received 1July 1985/Accepted 10 September 1985
AcDNA library designed for high-levelexpressionof measlesvirus-specificgeneproductsin mammalian cells was generated. From this library, functional clones which contained the entireprotein-codingsequences of the nucleocapsid (N) and the phosphoprotein (P) genes were isolated. By DNA-mediated gene transfer into a line of simian virus 40-transformed monkey kidney cells, the N-specific cDNA was expressed into a single
polypeptide of about 60,000Mr, which wasimmunoprecipitated by monoclonal antibodiesagainstthemeasles
virus N protein. In contrast, the P-specific cDNA could be expressed into either one or two species of polypeptides of 75,000 or 70,000
Mr,
bothof which wereimmunoprecipitated bymonoclonal antibodiesagainstthe measles virus P protein.
Measles virus possesses a negative-strand RNA genome which encodes six virion proteins: a 60,000-dalton
nucleo-capsid protein
(N), a 70,000-dalton phosphoprotein (P), a 38,000-dalton matrixprotein (M), a60,000-dalton precursorof the fusionprotein (Fo),a80,000-daltonhemagglutinin (H), and a high-molecular-weight protein presumed to be the polymerase (L) (12).Inaddition,measlesvirus-infected cells
contain a21,000-daltonvirus-specific nonstructural protein, NS (12),which may beidenticalto theCprotein expressed fromthe P genefromanalternative reading frame (4).
Measles virus can cause acute as well as chronic infec-tions. Variousreportshavesuggestedthatchronic infections by measles virus may be caused by aberrant expression of some of these viral gene products (7, 14, 25). There is, however, little directevidencetosubstantiate these
hypoth-eses. A major obstacle in
studying
the different manifesta-tions of measles virusinfection isthe poorunderstanding of thefunctioning of itsgenes andgeneproducts. Experimen-tation is further complicated by thecomplexity
of the interaction among thedifferent viral proteinsaswellashost factors.Wewishtoapproach these problemsby studyingthe gene
products expressedfrom clonedgenes ofmeasles virus.To
facilitate these studies,a cDNAlibrarywas constructed by using a vector system which allows the expression of full-lengthcDNAclones ineucaryotic cells (20).
Africangreen monkey kidney (CV-1) cells were infected with Edmonston strain measles virus at a multiplicity of infection of 0.05. When 100% of the cells displayed
cytopathic effect,
total cellularRNA wasextractedfromthecultures by the addition of a solution containing 4 M
guanidine thiocyanate (Fluka),50 mMsodiumacetate, 1 mM
EDTA, 0.25% sodium sarcosyl sulfate, and 0.1 M
,B-mercaptoethanol (6). RNA waspurifiedby centrifugationat
30,000 rpm for 19 h at 20°C in a Beckman SW41 rotor
through a CsCl cushion with a specific gravity of 1.72,
dissolved in 300 ,ul ofTSE (10 mM Tris hydrochloride [pH
7.0],
0.2% sodium dodecyl sulfate, 1 mM EDTA), andprecipitated at-70°Cwith 3 volumesofabsolute ethanol in the presence of 0.3 M sodium acetate. Poly(A)+ RNA was
enriched by one passage through an oligo(dT)-cellulose column (1), precipitated with 3 volumes ofethanol in the
*Corresponding author.
presenceof0.3 Msodiumacetate,and storedprecipitatedat
-70'C until used. About 8% ofthe total RNA was
recov-ered.
For cDNA synthesis, 10 ,ug of poly(A)+ RNA from measlesvirus-infected cellswasheatedat65°C for3min and
annealed with 4 ,ug of oligo(dT)-tailed vector primer
pre-paredfrompcDV1plasmidaspreviously described(19,20).
Reversetranscriptionwascarriedout at42°C for30min in 30 ,ul of reaction mixturecontaining50 mMTrishydrochloride (pH 8.3), 8 mMMgCl2, 50 mM KCl,2 mM dithiothreitol, 2 mMeach dATP,dGTP, dCTP, and TTP, 100 U ofreverse
transcriptase from avian myeloblastosis virus (Bethesda
Research Laboratories), and 20 Uof RNasinRNase
inhibi-tor (Promega Biotech). The addition ofoligo(dC) residues [oligo(dC) tails]tothe 3' endof the cDNAwascarriedout at
37°C for5minbypublished proceduresin areaction volume of 30 ,ul (19). After
digestion
with enzyme(HindIII),
thecDNA vector molecules were
cyclized by ligation
with an oligo(dG)-tailedlinkerfragmentderived from the PL1 plas-mid, which contains the simian virus 40 (SV40) promoterregion (20).The RNAmoietyin the recombinant constructs was removed by RNase H and replaced with DNA by a DNA polymerase I reaction as previously described (20).
TheresultingcDNAlibrarywasstored inaliquotsat
-20°C.
Asmallportion of thecDNAlibrary (ca.
5%)
wasused totransform Escherichia coli HB101(17) and screenedforthe presence ofmeasles virus-specific sequences by hybridiza-tion (13). 32P-labeled
gel-purified
DNA fragments derived from cDNA clones representing portions of the measles virusNand Pgenes (kind gifts from S. Rozenblatt [3, 10,22]) were used as probes under hybridization conditions previously described (27). Three clones containing
N-specific
sequences andsix clones withP-specific sequences were identified. Together, these clones represented 12% of thetotalcDNA clonesscreened,indicatingthat themeasles virus-specific RNA represented a sizeable portion of theintracellular poly(A)+ RNA in the measles virus-infected cells.
Restriction enzyme analysis showed that all three
N-specificclones and one of theP-specificclonesmight contain full-lengthcDNAinserts. Digestion ofoneoftheN-specific
clones (pcD-N7) with
BamHI,
which released the cDNA insert from the vectorDNA,producedalongfragmentwithelectrophoretic mobility corresponding to ca. 1.8 kilobases
343
on November 10, 2019 by guest
http://jvi.asm.org/
ABCDE FG
2.27- 1.97--1.8--rn.
1
.35-1
.08--
0.87-
0.60-HI J KL
_ .
6--l.95
M N OP Q
- 0.88
0.74
0
u
-1
measles-cDNA
N probe
LXc"a
aXVI1
X .. if ...1.-measles^-cDNA
Eo 0-C
pcD-P8 poly(A)
P probe
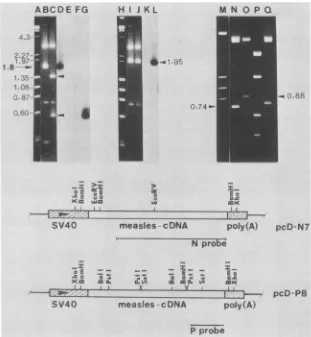
FIG. 1. Structuresof thefull-lengthcDNA clonesrepresentingthe Nand thePgenesof measles virus.pcD-N7andpcD-P8 DNAs (0.5 p.g) digested with restriction enzyme BamHI (lanes B and C, respectively) or XhoI (lanes I and J, respectively) were separated by
electrophoresisin 0.7%agarosegel and transferredontonitrocellulose filterpaper(23). Lanes:DandE,Southernblotofthematerials inlanes B andC,hybridizedwithanN-specificDNAprobe;Fand G, thesameblothybridizedwithaP-specificDNAprobe;Kand L,Southernblot
of theDNA in lanes Iand J,hybridizedwithaP-specificDNAprobe;N, pcD-N7 DNAdigested with EcoRV;0,P,andQ, products ofpcD-P8
DNAdigestedwithBalI,PstI,andSstI,respectively; A, H, and M,DNAsize markers.Aschematic representation ofthe pcD-N7and pcD-P8 clonesisdepicted, showing the locationsof theSV40promoterandpolyadenylation signals (hatched boxes) and theextentof thegel-purified DNAprobesused. Directionoftranscriptionisindicated byarrows.Otherarrows areexplainedinthetext.
(kb) (Fig. 1, lane B). Hybridization withanN-specific DNA probe confirmed that the 1.8-kb fragments in this clone contained N-related sequences (Fig. 1, lane D). Further restriction enzyme analysis (not shown) suggested that the 1.8-kbfragment spanned from the 3' terminus to aBamHI siteclosetothe5' terminusof theNgene(withrespect tothe [+]-strand mRNA [Fig. 1]).
With theP-specific clone(pcD-P8), similar digestion with BamHI produced a 1.4-kb fragment and a0.5-kb fragment
(Fig. 1, lane C, arrows). Only the 0.5-kb fragmenthybridized withtheP-specific DNA probe (Fig. 1, lane G). These results suggested the presence ofan internal BamHI site in theP
gene,andthe1.4-kbBamHIfragment inpcD-P8 represented sequences in the 5' region ofthe P gene which were not
recognized by the P-specific probe representing the 3'
se-quences (Fig. 1).
This interpretation proved tobecorrect. Digestion of the
cDNAcloneswith XhoIreleasedthecomplete inserts (1.95 kb) from both pcD-N7 and pcD-P8 DNAs(Fig. 1, lanesI and
J). The 1.95-kb fragment released from pcD-P8 contained P-related sequences which hybridized to the P-specific probe (Fig. 1, lane L).
The apparent sizes ofthe inserts in pcD-N7 and pcD-P8
correspondedwell with the apparentsizes of the correspond-ing intracellular RNAasdeterminedby gelelectrophoresis.
Totestif thesecDNA clonescontained thecomplete coding sequences of the N and Pgenes, we analyzed them further by restriction enzyme mapping. Digestion of pcD-N7 with
EcoRVproduced a0.74-kbfragment (Fig. 1, lane N),
signi-fying that thetwoEcoRVsitesinthe Ngene werepresentin
thisclone. Previous studies showed that the firstEcoRVsite waslocated in the 5'-untranslatedregion and the secondata
position 715 nucleotides from the AUG start site in the
coding region (21). These results indicated that pcD-N7 contained thecomplete codingsequences of the Ngene.
Similaranalysis showed that pcD-P8 containedtwoBalI
sites 0.88 kb apart (Fig. 1, lane 0). Since the 5'-proximal BalI siteis81 nucleotidesfromthestartofthecoding region (4), clone pcD-P8 contained at least most ofthe protein-codingsequences of the P gene up tothe 5'-proximalBalI
site. Based onthesedata and datanotshown,the organiza-tion of pcD-N7 and pcD-P8 is depicted in Fig. 1. The
restriction enzyme maps ofthese N-and P-specific cDNA
clones corresponded well with those of the previously
de-scribedcDNA clones ofthe measles virus (4, 21).
The pcD expression vector contains the SV40 early
pro-I_
Eo
Coix
±tl(A
poly(A)
°E E
SV40
O E =
Kco 0m
SV40
pcD-N7
on November 10, 2019 by guest
http://jvi.asm.org/
[image:2.612.165.476.64.401.2]345
1 3
.X Z ffi
6
aC
A BC D E F G
2.7 Kb- -I
1.85Kb-A
* 0 0
A B
C
D
E
F
G
-a28S
--
18S-1_ _
_
[image:3.612.128.463.70.288.2]B
FIG. 2. Expression ofN-andP-specificRNAin COS cellstransfectedwithcloned measlesgenes.COScellsweretransfected with 30,ug
of pcD-N7orpcD-P8 withoutcarrierDNAasdescribedin thetext.At 1day (31h)or2days (56 h)posttransfection,totalcellularRNAwas
extractedwith 4 Mguanidine thiocyanate and purified by centrifugation throughaCsClcushion(6). PurifiedRNA(20 ,ug)waselectrophoresed
in1%agarosegels in thepresenceof methyl mercurichydroxide (2). The resolvedRNAwastransferredontonitrocellulosefilterpaper(24)
andhybridized withanN-specific DNA probe (panelA)or aP-specificDNAprobe (panel B) under conditions previously described (27).As
control, total cellularRNAwaspreparedfrom COS cells infected with measles virusatatime when 10to 30% of the cultures showed cytopathic effect. Lanes:AandB, mock-transfectedCOScellsat1 and 2days,respectively;C andD,cellstransfected withpcD-N7at1 and 2 days, respectively; E and F, cells transfected withpcD-P8 at 1 and 2 days, respectively; G, measles virus (MV)-infected COS cells. Arrowheadsareexplained in thetext.
moterjuxtaposed with the inserted cDNA (Fig. 1). Splicing and polyadenylation signals are provided in the 5' and 3'
sequences flanking the cDNA insert. To testthe biological activity of the full-length cDNA constructions, pcD-N7 and pcD-P8wereintroduced intoSV40-transformed CV-1 (COS) cells (9) by DNA-mediated transfection.
Briefly, 1.8 x 106 COS cells were seeded onto 60-mm culture dishes and transfected with 10 or 30 ,ug of cloned
pcD-N7 and pcD-P8 DNA withorwithout carrier DNA(30 ,ug total), using the calcium phosphate coprecipitation method(11)asmodifiedbyWigleretal.(26) and Wongetal. (27). At 7 h posttransfection, cultures were shocked with growth medium containing 10% dimethyl sulfoxide for 15
min, replenished with fresh growth medium containing 10% fetalcalfserum,andmaintainedat37°C. Total cellular RNA was extracted at 1 day and 2 days posttransfection and analyzed by electrophoresis in denaturingagarosegels.
Cultureswhich received the N- and theP-specific cDNA clones expressed high levels of N- and P-specific RNA, respectively (Fig. 2A, lanes C and D; B, lanes E and F). In cellstransfected with clonepcD-P8, amajor RNAspecies of
ca. 2.5 kb was detected by hybridization with aP-specific DNAprobe butnotwithanN-specific probe (Fig. 2B, lanes E andF). This RNAspecieswasroughly 0.8 kb longer than theP-specific RNA in measles virus-infected COS cells (Fig. 2B, lane G). The longer length of the RNA in the transfected cells corresponded tothe additional 5' and 3' sequences in the cDNA construction, assuming that RNA transcription
wasinitiatedattheSV40promoterandterminated nearthe
SV40polyadenylation signal.
Asimilarpatternof RNAexpressionwasobserved in cells
transfected with pcD-N7 (Fig. 2A). The major species of N-specific RNA was 2.7 kblong (Fig. 2A, lanes C and D), alsoabout0.8 kblonger than the 1.85-kb N-specific RNA in
measles virus-infected cells (Fig. 2A, lane G, arrow). The size of the major species of N-specific RNA in the transfected cells was also consistent with the assumption
that it was derived from utilizing the SV40 promoter and
polyadenylation signals. Additional species of RNA both
longer and shorter than themajor specieswerepresentin all transformants. Since each transformed culturewasderived
from an independent event of transfection, it seemed
un-likely that the consistent appearance of discrete species of N- and P-specific RNA in the transformants was due to
randomaggregationordegradation. More likely, premature
termination and readthrough transcription might have pro-duced the aberrant RNAspecies. In this connection, previ-ous workers using this vector system have reported the presence ofseveral RNA species derived from differential utilization ofthetwosplicing signalsintheupstream vector sequences (20). We have not pursued these possibilities further.Inany case,theamountsofN- andP-specificRNA
in the transfected cells could beas highorhigher than that presentin thesamecellsacutely infected with measles virus. Totestif theobservedintracellularmeasles-specific RNA wastranslated intomeasles virus proteins, parallel cultures transfected with pcD-N7 and pcD-P8 were labeled with [35S]methionine for 23 h starting from 8 h or 1 day posttransfection.Labeledproteinswereimmunoprecipitated withantiseraagainst theNorPprotein and analyzed by gel
electrophoresis.
A single measles virus-specific protein ofca. 60,000 Mr
was detectedon day 2 in cultures transfected with pcD-N7
(Fig. 3, lane D, asterisk). Thisprotein comigrated with the N protein in measles virus-infected COS cells (Fig. 3, lane G). Moreover, a protein withidenticalelectrophoretic mobility
was immunoprecipitated from pcD-N7-transfected cultures
by monoclonal antiserum against the measles virus N protein '2.5Kb
-1.75Kb
VOL.57, 1986
on November 10, 2019 by guest
http://jvi.asm.org/
.Y z
0
3
a.
E
G H
200-
97.4-1K:
68-
-42-*
Co
.9 I
-0
a.o D
>
E a.
2
J K
LM NO P
,s4 *1w_ p
25.7
--P
18.4 -3
14.3 -
3
FIG. 3. Expression of N- and P-specific proteins in COS cells transfected with cloned measles genes. Parallel cultures of COS cells transfected with pcD-N7 and pcD-P8 as described in the legend to Fig. 2 were labeled with 50 ,uCi of[35S]methioninefor23 h starting at 8 h(lanes C, E, L, and N) or 33 h (lanes D, F, M, and0)posttransfection, such that the time of harvest corresponded to the time points used inthe RNA experiment shown in Fig. 2. Labeled proteins were immunoprecipitated and analyzed by electrophoresis in 6 to 18% gradient sodium dodecyl sulfate-polyacrylamide gels (15, 16). As a control, measles virus-infected COS cells were similarly labeled when 30% of the cellsdisplayed cytopathiceffect. Lanes: A andI,'4C-labeledprotein molecular weight markers (Bethesda Research Laboratories); B, J, and K,mock-transfected COS cells; C, D, E, and F, pcD-N7-transfected cells; L, M, N, and0,pcD-P8-transfected cells; G, H, and P, measles
virus-infectedCOS cells. Antisera used were against the total viral proteins (lanes C, D, G, J, L, and M) or were monoclonal antisera against the Nprotein (lanes B, E, F, and H) or against the P protein (lanes K, N,0,and P) (5,18). Due to the difference in avidity of the antisera, exposure timefortheproteins immunoprecipitated by the monoclonal antisera was longer. Asterisks are explained in the text.
(Fig. 3, lane F). This 60,000-Mr protein was notpresent in
themock-transfected cultures (Fig. 3, lanesB and J) nor in
the cultures transfected with the P-specific cDNA clone
(Fig.
3,lanes Land M).Similaranalysis of the cultures transfected with pcD-P8
revealed a measles virus-specific protein of ca. 70,000 to
75,000Mr
(Fig.
3, lane M).This proteinwas recognized by monoclonal antiserum against thePprotein (Fig. 3, lane0)
and
comigrated
with thePprotein in measles virus-infected COS cells(Fig.
3, laneP).These results provided direct evidence forthe biological
activity
of the full-length cDNA clones and confirmed theidentities ofthe clonedNand Psequences.
Previous studiesshowed that under certainconditions,the
measles virus P protein could exist as two electrophoreti-cally distinct species: P(75,000Mr) and P2 (70,000 Mr) (12).
Weobservedthatcloned pcD-P8DNAcouldoccasionallybe
expressed into two forms ofP-related proteinsin
transfec-tion experiments. COS cells infected with measles virus
producedtwospecies ofP-relatedproteins,PandP2,which
were immunoprecipitated by anti-measles virus serum as well asbymonoclonal antiserum againstthe Pprotein(Fig. 4, lanes A and F, respectively). In the same experiment,
COS cells transfected with pcD-P8 likewise produced two
P-related proteins which comigrated withP and P2. These proteinswere also recognized by the samepolyclonal and monoclonal antisera (Fig. 4, lanes B and G, respectively,
asterisks)and were notpresent inmock-transfectedcultures
(Fig. 4,lanesD,E, I,andJ).Theseresultsshowedthat the cloned P gene could be expressed into eitherone or both
speciesofP-relatedproteins (PandP2)intheabsence ofany other measles virusgene function. In contrast, the N-specific clonepcD-N7neverproducedmorethan asingle speciesof
N-specific proteinintransfectionexperiments.
Tryptic peptide analysis showed that P and P2 shared extensive homology (12). However, no precursor-product relationship was demonstrable, nor was significant differ-ence in theirextent ofphosphorylation detected (12). This
situation is reminiscent of that of the C and C' proteins of
Sendaivirus, which arethoughttobegenerated by alterna-tive translationinitiation from different AUG start sites on thesame readingframe ofa single speciesof mRNA (8).
Recently,athirdproteinof21,000daltons(protein C)was
found to be encoded by the measles virus P gene in an
alternativereading frame(4). Protein Cwas notrecognized bythe antisera used in this study. However, invitro
trans-I
.zn-f.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:4.612.163.484.74.383.2]MV
P8
mock
A
s
C
D
E
gm
IO
-200
gigs97
H
a..so-is
-~..I
~-
9.
w,tom
C*
-68
-42
-25.7
M\/
P8
mock
F
G
H
I J
u
-D - Z F s.*
2
_ I
I
I
i
FIG. 4. Expression oftwospeciesofP-relatedproteins by pcD-P8inCOS cells. COS cellsweretransfected with 10,ugofpcD-P8in the
presenceof 20 ,ugofsalmon spermDNAas acarrier. Transfected cellswere labeled with 50 ,uCiof[35S]methioninefor 24 h starting at
26 h (lanesB, D, G, and I)or 51 h(lanes C, E,H, andJ)posttransfection. Labeledproteins wereimmunoprecipitated andanalyzedby
electrophoresisin 10% sodiumdodecyl sulfate-polyacrylamide gels.Lanes:Aand F,measles virus(MV)-infectedCOScells; B,C, G,and H,COS cells transfected withpcD-P8; D,E,I,andJ, COS cells transfectedwith30 jig ofsalmonspermDNA. Lanes AthroughErepresent proteins immunoprecipitatedwithpolyclonalantiserumagainstmeasles virusproteins.LanesFthroughJrepresentproteins immunoprecipit-ated withmonoclonal antiserumagainstthePprotein.Molecularweightmarkers(x103)areshownonthe sideof eachpanel. Asterisksare
explained in thetext.
lationexperiments indicated thatprotein Cwasalsoencoded
by pcD-P8(unpublisheddata).
Theexperiments described here directly verified the iden-titiesof the N and P genesmapped nearthe 3' terminus of the measles virus genome. Aspointed outpreviously (20), the vector system used in this study is highly efficient in
obtaining full-length cDNAclones. We have sinceanalyzed over70additionalP-specific cDNA clonesfromthislibrary. Atotal of70% of the analyzed clones contained full-length cDNA inserts(unpublished data). Thus, this cDNA library couldprovide valuable information about theexact5' struc-tures of the measles virus mRNAs. Furthermore, statisti-cally this cDNA library likely contains additional functional viralgenesusefulforunderstanding the othergenefunctions
ofmeaslesvirus.Experiments toidentify these clonesarein progress.
WeespeciallythankCarol Miller,University of Southern Califor-nia School of Medicine, for providing the measles virus, her expertise,andherlaboratoryfacilities for theinitiationof thisstudy. We thankS.Rozenblattforprovidingthemeasles cDNAprobesand E.NorrbyandW.Bohn forprovidingthe monoclonal antisera. We also thank Tulan Do, Greg Wipf, and Kent Beech for technical assistance.
This work was supported by grant MV-238fromthe American Cancer Society, aBiomedical ResearchSupportgrant, andby the
Graduate School Research FundfromtheUniversityofWashington School of Medicine.
LITERATURE CITED
1. Aviv,H.,andP. Leder. 1972. Purification ofbiologicallyactive globin messenger RNA by chromatography on
oligo-thymidylicacid-cellulose. Proc. Natl. Acad. Sci. USA 65:1408-1412.
2. Bailey, J. M., and N. Davidson. 1976. Methylmercury as a
reversible denaturing agent for agarose gel electrophoresis.
Anal. Biochem. 70:75-88.
3. Bellini,W.J.,G.Englund,C. D.Richardson, and S. Rozenblatt. 1984. Positive identification of a measles virus cDNA clone
encodingaregionof thephosphoprotein. J. Virol.50:939-942. 4. Bellini, W. J., G. Englund, S. Rozenblatt, H. Arnheiter, and
C. D. Richardson. 1985. Measles virus P gene codes for two
proteins.J. Virol. 53:908-919.
5. Bohn, W., G.Rutter, and K. Mannweiler. 1982. Production of monoclonal antibodies tomeasles virus proteins by immuniza-tion of mice with heated anddetergent-treated antigens. Virol-ogy116:368-371.
6. Chirgwin, J.M., A. E. Przybyla, R.J. MacDonald, and W.J. Rutter. 1979. Isolation ofbiologically active ribonucleic acid from sources enriched in ribonuclease. Biochemistry 18:
5294-5299.
7. Choppin, P. W. 1981. Measles virus andchronic neurological diseases. Anal. Neurol. 9:17-20.
-200
-97.4
-68
-42
-
25.7
VOL.57,1986
on November 10, 2019 by guest
http://jvi.asm.org/
[image:5.612.148.451.76.378.2]8. Giorgi, C., B. M. Blumberg, and D. Kolakofsky. 1983. Sendai virus contains overlapping genes expressed from a single
mRNA.Cell35:829-836.
9. Gluzman,Y. 1981. SV40-transformed simian cells supportthe replication of earlySV40mutants.Cell23:175-182.
10. Gorecki, M., and S. Rozenblatt. 1980. Cloning ofDNA comple-mentary to the measles virus mRNA encoding nucleocapsid protein.Proc. Natl. Acad. Sci. USA77:3686-3690.
11. Graham, F. L., and A.vander Eb. 1973. Anewtechnique for
theassay of infectivity of humanadenovirus 5 DNA. Virology 52:456457.
12. Graves, M. C.1981.Measlesviruspolypeptides in infected cells studied by immune precipitationandone-dimensional peptide mapping. J.Virol. 38:224-230.
13. Grunstein, M., and D. Hogness. 1975. Colony hybridization: a
method fortheisolation of clonedDNAsthat containaspecific
gene.Proc. Natl. Acad.Sci. USA72:3961-3965.
14. Haase, A. T., D. Gantz, B. Eble, D. Walker, L. Stowring, P. Ventura, H. Blum,S.Wietgrefe, M. Zupancic, W. Tourtellotte, C. J. Gibbs, Jr., E. Norrby, and S. Rozenblatt. 1985. Natural historyof restricted synthesisandexpressionofmeaslesvirus
genesin subacutesclerosingpanencephalitis.Proc.Natl.Acad.
Sci.USA 82:3020-3024.
15. Laemmli, U. K. 1970. Cleavage ofstructural proteins duringthe assembly of the head ofbacteriophage T4. Nature (London) 227:680-685.
16. Laskey, R. A., and A. F. Mills.1975.Quantitative film detection of[3H]and["4C]inpolyacrylamide gels by fluorography.Eur.J. Biochem. 56:335-341.
17. Maniatis, T., E. F. Fritsch, and J. Sambrook. 1982. Molecular cloning,alaboratorymanual. ColdSpringHarborLaboratory,
ColdSpring Harbor,N.Y.
18. Norrby, E., C. Orvell, B. Vandvik, and J. D. Cherry. 1981. Antibodies against measles virus polypeptides indifferent dis-easeconditions. Infect. Immun. 34:718-724.
19. Okayama, H., and P. Berg. 1982. High-efficiency cloning of full-length cDNAs. Mol.Cell. Biol. 2:161-170.
20. Okayama, H., andP.Berg. 1983. AcDNAcloning vector that permits expression of cDNA inserts in mammalian cells. Mol. Cell. Biol. 3:280-289.
21. Rozenblatt, S., 0. Eizenberg, R. Ben-Lavy, V. Lavie, and W. Bellini. 1985. Sequence homology within the morbilliviruses. J. Virol. 53:684-690.
22. Rozenblatt, S.,C.Gesang,V.Lavie,andF. S. Neumann.1982. Cloning and characterization of DNA complementary to the measles virusmRNAencoding hemagglutinin and matrix pro-tein. J.Virol. 42:790-797.
23. Southern, E. M. 1975. Detection ofspecific sequences among DNAfragments separated by gel electrophoresis. J. Mol. Biol. 98:503-517.
24. Thomas,P.S. 1980.Hybridizationof denatured RNA and small DNAfragments transferredtonitrocellulose. Proc. Natl. Acad. Sci. USA 77:5201-5205.
25. Wechsler, S. L.,and H. C.Meissner. 1982. Measlesand SSPE viruses: similarities and differences. Prog. Med. Virol. 28:65-95.
26. Wigler,M., A.Pellicer,S.Silverstein,R.Axel,G.Urlaub, andL. Chasin.1979. DNA mediated transfer of the adenine phospho-ribosyltransferase locus into mammalian cells. Proc. Natl. Acad. Sci. USA 76:1373-1376.
27. Wong, T. C.,R. S. Goodenow, B. T. Sher, and N. Davidson. 1985. Thepromoterof the long terminal repeat of feline leuke-miavirus is effective forexpression ofa mouseH-2
histocom-patibilitygene inmouseand humancells. Gene 34:27-38.