0022-538X/84/060895-09$02.00/0
Copyright© 1984, American Societyfor Microbiology
Production of a
Monospecific Antiserum Against the Early Region
lA
Proteins
of
Adenovirus
12
and
Adenovirus 5 by an Adenovirus 12
Early Region
1A-1-Galactosidase Fusion Protein Antigen Expressed
in
Bacteria
MILLER
0.
SCOTT,'
DAVID KIMELMAN,2 DAVIDNORRIS,1t
ANDROBERT
P.RICCIARDI'*
The Wistar Institute
of
Anatomy andBiology, Philadelphia,
Pennsylvania 19104,1 andDepartment
of BiologicalChemistry,
Harvard
MedicalSchool, Boston,
Massachusetts 02115Received 13December1983/Accepted5March 1984
Antisera
wereprepared against the
amino acid
sequencesencoded within
theN-terminal half
of
theadenovirus
12(Adl2) early
region
1A(E1A)
gene. This wasaccomplished by
construction of a plasmid vectorwhich encoded theN-terminal
131amino acids of Adl2
ElAjoined in
frame
tothe
coding
sequenceof
,3-galactosidase. After induced synthesis
inEscherichia coli,
the Adl2ElA-p-galactosidase
fusionprotein (12-1A-FP)
wasextracted with
ureaand
used toraise antibodies
inrabbits.
The 12-1A-FPantiseraimmunoprecipitated
majorphosphoproteins
of 39,000 and 37,000 apparent molecularweights from
Adl2-transformed and infected cells. The
12-1A-FP antisera alsoimmunoprecipitated
ElAphosphoproteins from
Ad5-transformed and infected cells. Immunospecificity of the 12-1A-FP antisera
wasdemonstrated by the
ability of 12-1A-FP antigen
toblock immunoprecipitation of
ElAproteins. Furthermore,
ElAproteins
immunoprecipitated from in vivo-labeled
cellscomigrated with
thosetranslated in vitro
by
RNA thathad
been
hybridization selected
toElA
DNA.Characterization of
proteins encoded by regulatory
genesis critical for
acomprehensive understanding
of geneexpres-sion. The
early region 1A (E1A)
geneof adenovirus
plays
adynamic role during both cellular infection and
transforma-tion.
Analysis of several
ElA mutantsof adenovirus
5(Ad5)
has
revealed
thatthere
is atranscriptional dependence
of
early
viral genes upon aproduct of
theElA
gene(4, 24).
From
the ElA
geneof
AdS,
twomajor
RNAsof
12Sand 13S
are
transcribed, which
sharethe
same 5' and3' termini
butdiffer by
the amountof
intervening
sequenceremoved
by
splicing (5,
10,25,
41). The12S
and 13S mRNAsencode
acidic
proteins of
242and 288
amino
acids,
respectively
(32).
The
amino acid
sequencesof both ElA
proteins
areidentical
except
for
aninternal stretch
of 46 amino acids which is
unique
to thelarger ElA
protein (32). However, only
theproduct
of the
larger acidic
protein
is
required
for
modula-tion of
early
genetranscription
(28, 34).
Not
all
mutations in the
ElA geneof Ad5 exhibit
thesamepleiotropic effect. For example,
thedefects contained
in theframeshift
mutantsH5hrl and H5in500 and
the nonsense mutanthr440 each preventfull-length synthesis of
thelarger
ElA acidic protein (9, 34, 40). However,
theH5hrl
mutantdoes
notsynthesize
anE1B,
E2, E3,
or E4transcript
(4), whereasthe hr440
mutantexpressesE1B
and E4transcripts
(40),
and theH5in500
mutantproduces
wild-type
levelsof
E1B
andE3
transcripts
butlow
levelsof
E2 and E4transcripts (9). Taken altogether, these findings indicate
that theprotein product of the larger ElA transcript, in fact, does
modulate
expression of
otherearly genes, but that,depend-ing
upon thelocation
and natureof
themutation
in theElA
DNA,
manifestation of
thiseffect
will vary. These findingsraise
theintriguing possibility
thattheElA protein
productcontains
separate functional domains.*Correspondingauthor.
tPresent address: Department of Microbiology and Molecular
Genetics, Harvard Medical School, Boston,MA 02115.
How the ElA
protein activates expression of early
viral genesduring infection is unresolved. Feldman
etal.
(12)
propose
that the
ElA geneproduct indirectly
mediatesadenovirus transcription by interacting
with host cell
compo-nentsrather than
by
direct
recognition
of regulatory control
regions within the DNA. This they
arguesince
thepseudora-bies virus immediate
early
geneproduct
is aheterologous
activator of adenovirus
genetranscription in
theabsenceof
ElA
function.
It
is likewise
unclear whether the ElAprotein contains
transforming functions independent of modulation of
tran-scription during infection.
Inrodent
cells,
integration of both
ElA and
E1B
genes(0
to 11mapunits
[mu])
is
necessaryfor
complete transformation (18, 20, 44), but direct linkage
ofElA and
E1B genesis
notnecessary(43).
An argumentfor
an
independent transforming function of ElA
canbe made
from the analysis of
theElA
hr mutants hr440 and H5in500,which
are capable of transcribingE1B
yet are incapableof
transformation
(9, 40).In
general, antisera from animals bearing
adenovirus-induced
tumorsdo
notstrongly reactwith ElA
proteins, nor cansuch antisera be
used toidentify
themdefinitively.
These
limitations,
inaddition
to the lowconcentrations
ofElA
proteins produced in both transformed
and infectedcells,
havehampered
attemptstowards
thecharacterization
and
isolation of ElA proteins.
However,antisera
specific toAdS ElA proteins have recently been
produced by usingsynthetic peptides.
These antisera have been used to identify andlocalizeAdS ElA
proteins within cells by immunofluo-rescentstaining
of both the nuclear matrix and cytoplasmicregions
(13, 49).We report here the production
of
thefirst monospecific
antiserum
directed against theElA
proteins ofAdl2. Adl2
is
highly
oncogenic compared withAdS
(14, 15). In fact, thetumorigenic
potential ofAdl2
in rats has been recentlyattributed
to the ability of the larger of the twoAdl2 ElA
products
to suppress expression of class I majorhistocom-895
on November 10, 2019 by guest
http://jvi.asm.org/
patibility antigens
(6, 39). Ad5 ElA geneproducts
do notaffect
the level of class I antigensin Ad5-transformed
rat cells (6, 39). It isof
furtherinterest
that theElA
genesof
Adl2
are able to transcomplementAd5
duringinfection
(47).Similar
to the ElA geneof Ad5,
twomajor forms of
mRNAoriginate from region ElA of Adl2. The
Adl2 ElA mRNAsdiffer
internally from
one anotherby
utilizing
oneof
twodonor
splice sites but
share the same acceptorsplice site.
The 3' ends of each
Adl2 ElA
transcript
areidentical,
whereas the 5' ends are heterogeneous
since
two or moreinitiation
sites areutilized
(37,38).Prediction from
the DNA sequenceindicates that these
twoAdl2
ElA mRNAsencode
proteins
of 266 and 235amino acids
(33, 42).We
describe
the ElAproducts synthesized
inAdl2-transformed cells with
anantiserum directed
against
anE1A-,B-galactosidase
fusion
protein
antigen synthesized
inEsche-richia
coli.
We furtherdemonstrate
theability of this Adl2
ElA
monospecific antiserum
toimmunoprecipitate ElA
proteins from both
AdS-infected andtransformed cells
anddiscuss
theusefulness of
thiscross-reactive antiserum.
MATERIALS ANDMETHODS
Cells, viruses, and infection. The
Adl2-transformed
ham-stercell line
HA12/7
waspreviously described
(11,50).
TheAdl2-transformed BALB/c
mousecell line
M012/F10
wasobtained
from J.Williams, Carnegie
MellonUniversity,
Pittsburgh,
Pa., and
wasmaintained in minimal essential
medium
plus 5% fetal calf
serum. TheAd5-transformed
human cell
line
293 wasalso
previously
described (19). The
Ad5 virus
wasplaqued
on HeLacells and used
toinfect
HeLa
monolayer cells
at amultiplicity
of infection of
50in
the
absence
of drugs.
Growth and induction ofAdl2
ElA-Il-galactosidase
fusionprotein antigen
inE.coli.E.coli
LacIQ
W3110containing
theplasmid
pAd441, which encodes
an Adl2ElA-p-galacto-sidase
fusion
protein (12-1A-FP)
asdescribed
below,
was grown as anovernight culture in Casamino Acids
(Difco
Laboratories, Detroit,
Mich.)
medium
containing (per liter):
5
gof
Casamino
Acids,
5.8 gof
NaHPO4,
3.0 gof
KH2PO4,
0.5
gof
NaCl,
and
1.0 gof
NH4Cl.
Themixture
wasbrought
to
pH
7.2 to7.6
with
NaOH,
autoclaved,
and
supplemented
with
1ml
of sterile
0.2%
vitamin
B1,
25 mlof 20%
glucose,
1 mlof
1 MMgSO4, and 50
mgof
ampicillin.
Asample
of the
overnight
culture
(5
ml)
wasadded
to 500 ml of CasaminoAcids medium
ina2-liter
Erlenmeyer
flask
and growntoanA600
of 0.3
at37°C
on ashaker.
Theculture
wasinduced
tosynthesize
thefusion
protein by
theaddition of
119 mgof
solid
isopropyl-3-D-thiogalactopyranoside
(IPTG)
andcon-tinued
rapid shaking for
anadditional
45to 60min.Isolation of fusion
protein antigen
extractfromE. coli. Theinduced
E.coli isolates
werepelleted
by
centrifugation
at8,000
rpmfor 20
min at4°C.
Thepellet
was washed inphosphate-buffered
saline(PBS)
containing
25%
glucose
and wasrecentrifuged.
Thefusionprotein
wasextracted in 10 M urea asdescribed
by
Bikeletal.(7).
The 10 M urea extractwas
dialyzed
against
PBStwice
at roomtemperature(45
min eachtime) followed
by
exhaustivedialysis
at4°C.
Preparation
of Adl2 ElA antisera. New Zealand whiterabbits
were firstinjected
intradermally
with 300 ,ugof
the12-1A-FP
antigen
extractincomplete
Freundadjuvant
and thenboostedat2- to4-weekintervals
with 300 ,ugof
the 12-1A-FP inincomplete
Freundadjuvant.
Rabbitswere earbled 7 to 10days
afterinjection.
Sera were adsorbed with adialyzed
10M urea extractfrom
untransformedE.coli
cells(5
to 10mg ofprotein
extractper ml ofserum)
at0°C
for 1to2
days,
and theimmunecomplex
waspelleted
at40,000
rpmin a
Ty65
rotorfor
45 min at4°C.
The supernatant wasreadsorbed with
asonicated
extract derived from eitheruninfected
HeLa cells ornon-adenovirus-transformed ham-stercells (108 cells in 1 ml of PBSper 5 ml ofserum),
which wasincubated
at0°C
for
1to2days
and
repelleted
at40,000
rpm
for
45minat4°C.
This final supernatant,referred
to as the 12-1A-FPantiserum,
wasusedforimmunoprecipitation.
Labeling
of cells andpreparation
of cell extracts. Cellmonolayers
werelabeled
with[35S]methionine by
incubation
in
methionine-free medium for 1 h followedby
the additionof fresh methionine-free
mediumcontaining
50,uCi
of[35S]methionine (Amersham Corp., Arlington Heights,
Ill.)
per ml
and
continued incubation for
1to4 h.Cells
werealso
labeled with
carrier-free
32p;
(NewEngland
NuclearCorp.,
Boston,
Mass.) first
by
incubation in
phosphate-free
medium
for
1h and
thenby
theaddition of
freshphosphate-free
medium
containing
125,uCi of
32p,
permlfor
1 to4 h. The labeled cells wererinsed
inPBS,
removed with a rubberpoliceman,
pelleted,
and washed once in PBS. In all subse-quent steps,the buffers
and extract weremaintained
at4°C.
The washed
cell
pellet
wassuspended
in 3volumes
of 1mMMgCI2-1
mMKCl-10
mM Tris(pH 8.1), containing
100kallikren
IUof
aprotinin
per mland
1 mMphenylmethylsul-fonyl fluoride,
andkept
onice for
5 minfollowed by
rapid
freezing
in
adry ice-methanol bath (B. Atkinson, C. S.
Ernst,
B. E. D.Ghrist,
M.Herlyn,
D.Herly,
A. Ross, M.Blaszczyk,
Z.Steplewski,
and H.Koprowski, Cancer
Res.,in
press).
Thecell
suspension
wasthawed andcentrifuged
at40,000
rpmfor
45min
at4°C
in
aTy65
rotor.The supernatant(fraction I)
wasremoved
to a newEppendorf
tube.
Thepellet
waslysed
in
RIPAbuffer
(50
mMTris,
pH 7.2;
150 mMNaCl;
0.1%
[wt/vol]
sodium
dodecyl
sulfate
[SDS];
0.1%
[wt/vol]
sodium
deoxycholate
[NaDOC];
0.1%
[vol/vol]
Tri-tonX-100;
100kallikren
IUof
aprotinin
perml;
1 mMphenylmethylsulfonyl fluoride)
at aratio of
3 x107
cells
per mlof
RIPAbuffer.
Thelysed pellet
wasvortexed, kept
onice for 20 min,
and thenbriefly
sonicated and
centrifuged
at40,000
rpmin
aTy65
rotor at4°C
for
45min.This
superna-tant(fraction
II)
wasplaced
into
a newEppendorf tube,
andthe
pellet
wasdiscarded.
By this
procedure,
fraction
Irepresented
the
cell
cytosol,
and
fraction
IIrepresented
the
cell
nuclei, membranes,
and
organelles.
Fractions
Iand
II were eachprecleared by
theaddition of
150RI
of
a10%
suspension
of
Staphylococcus
aureus(Staph A,
BethesdaResearch
Labs, Bethesda,
Md.)
onice for
30 to 60 minfollowed
by centrifugation
inamicrofuge
at4°C
for
15min.The numbers
of trichloroacetic
acid-precipitable
counts weredetermined for both
precleared fractions. Fractions
I and IIwere either useddirectly
orstored
at-80°C.
Immunoprecipitation.
Immunoprecipitation
of[35S]methi-onine-labeled
proteins
wasperformed
with
30 x106
tri-chloroacetic
acid-precipitable
countsof
precleared lysate
fromfractionsI and II. For
32P-labeled proteins,
5 x106
to10 x
106
trichloroaceticacid-precipitable
countsof
lysate
from
eachfraction
were used.Separation
of
thelysate
into thesetwofractions
significantly
reduced the levelof
nonspe-cific
proteins
thatwereimmunoprecipitated.
The
precleared lysates
wereincubated
with 12-1A-FP antiseraat afinaldilutionof
1:20 onicefor 2 to24h.Fifty
microliters of
Staph
Awasaddedtotheimmunoprecipitation
reaction and
incubated
for 15to30minonice. TheStaph
Aantibody complex
waspelleted
for 1 min at room tempera-ture in anEppendorf
microfuge.
The supernatant wasdis-carded,
and thepellet
waswashedand vortexedfour
times in RIPA buffer(described above).
The SDSsample
buffer(0.065
MTris-hydrochloride [pH
6.8],
2%SDS,
10%
on November 10, 2019 by guest
http://jvi.asm.org/
ol, 5% 2-,-mercaptoethanol,
0.001%bromophenol blue [26]) was added to theStaph
Aantibody complex pellet, vortexedintermittently for
30min, andboiled for
5 min. TheStaph
A was pelleted at room temperature for 2 to 3min,
and thelabeled proteins in
the supernatant were size fractioned in15% SDS-polyacrylamide
gels (acrylamide-to-bis ratio, 30:0.8) asdescribed by
Laemmli (26) and visualized by fluorography (8). In some instances,protein
A (a20%
suspension [wt/vol] in PBS [Pharmacia
FineChemicals,
Inc.,
Piscataway, N.J.])
was usedin
placeof Staph
A. The procedure for usingprotein
Awasidentical
tothatfor
using
Staph
Aexcept that 100pAl
wasadded to theimmunoprecipi-tation reaction mixture
which
was then rotated at 4°Covernight.
Hybridizationselection and cell-free translation. RNA was phenol extracted
from transformed cell lines.
ForAd12-transformed
cells, vanadyl
ribonucleoside
complexes were usedin
thephenol extraction
(3). DNA restrictionfragments
generated from
plasmids
wereisolated by
the methodof
Vogelstein
andGillespie
(45).Hybridization
selection of
mRNAto
specific
DNArestriction fragments
wasperformed
as
described by Ricciardi
etal.
(35).Cell-free translation of
hybridization-selected
mRNAs wasin
therabbit
reticulocyte
system
(31).
Plasmids and DNA.
Plasmid
pLG400 (21) contains the LacI
and
LacZfusion
geneof
E.coli and
isreferred
tobelowas LacZ.Plasmid
pAd418, which contains
full-length
Adl2 ElA cDNAjoined
to the TAC promoter, wasprepared by
D.Kimelman,
L. A.Lucher, K. H.Brackmann,
M.Green,
and M.Ptashne(unpublished
data). Theplasmid
pLA-1contains
0 to 9.4 mu
of
Ad5 DNA and was obtained from F. Tamanoi, ColdSpring
HarborLaboratories,
Cold SpringHarbor,
N.Y.Plasmid
pSVG-12contained
theEcoRI
frag-ment
C
of Adl2 (0
to 16.5mu)
and wasderived
by inserting
this
Adl2
fragment within
the BamHI(linkered)
to theEcoRI site of
pSV-gpt
(29).RESULTS
Construction of the
Adl2
ElA-0-galactosidase
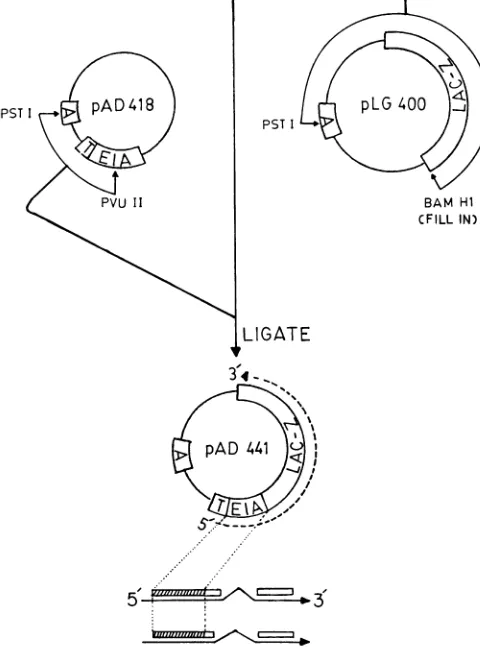
plasmid vector.Theplasmid pAd441 contains
sequences correspond-ing to theamino-terminal
halfof
theAdl2 ElAcoding region
which are
joined
5'andin-frame
tosequenceswhich
encodeP-galactosidase (Fig.
1). The
132amino
acids encoded within
theElA DNA
of
pAd441
aresharedcompletely by
the two viralElA proteins of
Adl2, which contain
235and
266amino
acids
(33,42) and
which are translatedfrom
overlapping
mRNAs
(37, 38)
(Fig. 1).
Preceding
the Adl2E1A-,B-galactosidase
fusedcoding
regions is
a 250-basepair (bp)
insert
(designated
Tin
Fig. 1) which contains the Lac
ribosomal
binding site
as well as theTAC
promoter. The TAC promoter isahybrid of
the Lac and TRP promoters and is 10times
morepowerful
than the Lacpromoter alone (2a). The pAd441 vector also contains the ampicillin resistance geneof
pBR322(designated
Ain Fig. 1).The
construction
of pAd441 was accomplished byisola-tion of
thePstI-PvuII
fragment
from pAd418 (Kimelman etal., unpublished
data), which contains the TAC promoter sequencejoined
tofull-lengthAdl2 ElA
cDNA (Fig. 1). TheElA
cDNAofpAd418 extends 4 bp 5' of the initiator AUG codon ofElA
and thePvuII
site is 400 bp downstream from the AUG. Toform
pAd441, we essentially joined thePstI-PvuII fragment
of pAd418 to thePstI-BamHI
fragment of pLG400 (21) which contained the LacZ gene encoding,B-galactosidase (Fig.
1). To achieve ligation of these two DNAfragments,
thePstI-BamHI
fragment ofpLG400 was derivedby
restriction ofpLG400 withBamHI
followed by a reactionwith the Klenow fragment of DNA polymerase to fill in protruding ends (46) and, finally, restriction by PstI. The pAd441 vector was
transfected into
a LacZ- strain of E. coli, and the isolated ampicillin-resistantcolonies,
which turn red on MacConkey-Lactose plates, indicated positiveexpression of ,B-galactosidase
(27).Expression of 12-lA-FP from E. coli. E. coli cells which
contained the
plasmid pAd441
wereinduced
to synthesize 12-1A-FPby the addition of IPTG to the growing culture. To assayfor induced synthesis,
wefractionated
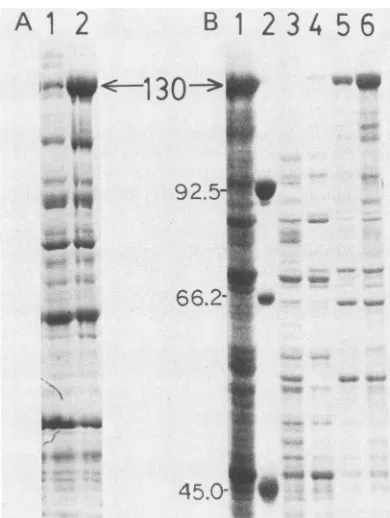
total E. coli protein in SDS-polyacrylamide gels (26), and the proteins were visualized by Coomassie blue stain. A protein of 130,000 molecularweight
(130K) was the only novelprotein synthesized at a high level in induced E. coli (Fig. 2, lane 2) compared with uninduced E. coli (lane 1). This 130K protein was the size predicted from the DNA sequences of Adl2 ElA and3-galactosidase contained within
pAd441. The N-terminal sequenceof
the 12-1A-FPantigen, purified from
apolyacrylamide gel,
wasanalyzed by automated Edman
degradation (23)
andindicated
that theinduced protein
PST I
5-FIG. 1. Construction ofthe Adl2 ElA-,B-galactosidase
expres-sion vector plasmid pAd441. The plasmid pAd441 encodes 132
amino acids correspondingto theamino-terminal end of theAdl2 ElAcodingregion.Thesesequencesarejoinedin-frametothegene
ofE.coli which encodesP-galactosidase(LacZ).The location of the
coding sequences ofAdl2 ElA inpAd441 with respect to
corre-spondingviral mRNAsis shownin the lowermost partof thefigure.
The pAd441vectoralsocontainsanampicillinresistance gene(A)
and a 250-bp sequence (T) containing the TAC promoter and a ribosomal binding site. Construction ofpAd441 was achievedby joiningthe smallerPstI-PvuIIfragmentofpAd418(Kimelmanetal.,
unpublished data) to the larger PstI-BamHI fragment ofpLG400 (21). Detailsof thisconstructionarediscussedin thetext.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:3.612.314.554.293.618.2]A 12
B
1 2
34
56
w<---130---9._
66.2-O.:W~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
45.0-FIG. 2. Fractionated E. coli proteins. (A)E. coli proteins
frac-tionated inapolyacrylamide gel showinginduction of 12-1A-FP. E.
coli cells containing the pAd44l expression vector (Fig. 1) were
induced to synthesize 12-1A-FP by the addition of IPTG as
de-scribed in the text. After induction,the E. colicells were pelleted
andlysedinSDS-Laemmlisamplebuffer(26),fractionated ina7.5%
SDS-polyacrylamide gel, and stained with Coomassie blue dye.
Lane1,noIPTGadded;lane2,IPTG added. Thearrowpointstothe
130K fusion protein. (B)E. coli proteins fractionated in an
SDS-polyacrylamide gel after induction of 12-1A-FP and extraction by
urea. E. coli cells containing the pAd44l vector (Fig. 1) were
inducedtosynthesize12-1A-FP.The E. ccli cellswerepelleted,and
theproteinswereextractedbyusingvariousureaconcentrationsas
described by Bikel et al. (7) and in the text. The urea-extracted
proteins were fractionated in a7.5% SDS-polyacrylamide gel (26)
and stained with Coomassie blue. Fractionation ofproteins from
induced E. ccli cellscontaining pAd44l wasasfollows: lane1,not
extractedbyurea;lane4,extracted in 2 Murea;lane5,extracted in
5M urea;and lane 6, extracted in 10 Murea. Lane3, 10 M
urea-extracted proteinsfrom induced E. ccli cells which donotcontain
pAd441. Lane 2, marker proteins as follows: phosphorylase B
(92.5K);bovineserum albumin(66.2K);andovalbumin(45K). The
position of the 130K fusion protein 12-1A-FP is indicated by the
arrow.
initiated with the same six N-terminal amino acids as viral Adl2 ElA (data not shown). The induced 130K fusion
protein represented about25% of the total protein of theE.
ccli isolates.
Extraction of thebacterial fusionproteinandproductionof
antisera. Extraction of the 12-1A-FP was accomplished by utilizing ureaasdescribed by Bikelet al. (7). As seen from
the Coomassie blue-stained SDS-polyacrylamide gel (Fig.
2B), extraction of the 12-1A-FP in 10 M urea(lane 6) was morethan twice as efficient as extraction in 5 Murea (lane 5), and extraction in 2M urea(lane 4) was hardlyeffective. The 130K proteincouldnot beextractedby 10Mureafrom
E. ccli isolates which were induced but didnot contain the
pAd44l expression vector(Fig. 2B,lane 3). Itisnoteworthy
that the 12-1A-FP failed to be extracted by nonionic deter-gents since >95% of the 130K protein remained in the cell pellet (datanot shown). Extraction by 10 M urea (Fig. 2B,
lane 6) resulted in a ca. 25% purification of the 12-1A-FP compared
with
total unextracted protein from induced E. coli samples(Fig. 2B,lane 1).Furthermore,
removal ofurea by dialysisin PBSreinstated
3-galactosidase
activity
of the 12-1A-FPcomparable with that
of
commercial
,B-galacto-sidase asmeasured by the
O-nitrophenyl-,-D-galactopyrano-sideassay(27;data not
shown).
Asmuch
as17mgof
12-1A-FP could be extracted from 1 liter of induced E.
coli cells.
The10M urea
extract
containing
the12-1A-FP
antigen
was dialyzed in PBS andintroduced into rabbits from which
antisera wereobtained.These
antisera,
referredto asthe 12-1A-FP antisera, were used toimmunoprecipitate
proteins
from
adenovirus-transformed
and-infected cells.
Bacterial fusion
protein antisera
immunoprecipitated
two majorElA proteins
fromAdl2-transformed
cells.The
12-1A-FP antisera were used to
identify
ElAproteins which
are synthesized in theAd12-transformed hamster cell
HA12/7,
which has been
reported
toexpress the
Adl2
genome extensively (1). The12-1A-FP
antisera
immunoprecipitated
two
major
proteins
withapparent
molecular
weights
of
39K and 37K from[35S]methionine-labeled HA12/7 cells
(Fig.
3A,lane1). Aminor
protein
of 36K was alsoimmunoprecipi-tated. In all
experiments with HA12/7 cells,
as well as otherAdl2-transformed hamster
and mouse cells,only
the 39K and37Kproteinswereconsistently
detectedin
both
thecell
cytosol fraction (fraction I) and the cellfraction
containing
nuclei,
membranes,
andorganelles (fraction
II). To prove thatthese proteins
wereimmunospecific precipitation
prod-ucts ofthe
12-1A-FP antisera,
weadded
12-1A-FP
antigen
with the
antisera
to thelabeled
celllysates.
Immunoprecipi-tation
of the 39K,37K,
and 36Kproteins
wasinhibited
asthe amount of12-1A-FP antigen
in thereaction increased
(Fig.
3A,lanes 2to4).
Indeed,
the profile oflabeled proteins
from
a
completely inhibited
immunoprecipitation reaction
(Fig.
3A, lane 4) was
identical
to one in whichonly
preimmune
serum
wasadded
to the[35S]methionine-labeled
cell
lysate
(Fig. 3A, lane 5). The series of
high-molecular-weight
bands (Fig.3A, lane1),which
werealso inhibited by
theaddition of
12-1A-FP
antigen (lanes
3 and 4),likely
representstightly
associated
nonspecific cellular proteins
that arecoprecipitat-ed with the 39K and 37K ElA
proteins.
We
further
established
thatfor
eachprotein
immunopreci-pitated from
Adl2-transformed cells by the 12-1A-FP
anti-sera, there was a
corresponding translation
product
synthe-sized
in vitro by RNAthat had
been
hybridization
selected
to Adl2 ElA DNA. When total
cytoplasmic
RNAfrom
either
theAd12-transformed
hamstercell
HA12/7
or mouse cellM012/F10
wasfirst
hybridization
selected to DNAwhich
contained sequences exclusive
toAd12
ElA(pAd441
DNA;
Fig. 1) andwas
then translated in acell-free system,
three proteins
of 39K, 37K, and36K
weredetected
(Fig.
3B,
lanes
8and 1,respectively).
Inaddition,
aminor ElA
protein
of 22K was also detected as an in vitro translation
product.
For
comparison, mRNA
fromHA12/7 cells
washybridiza-tion
selected
to aPvuII fragment of
Adl2 DNA(1,906
to3,623 bp)
which
exclusively contained
E1B sequences. Pro-teinstranslated
fromhybridization-selected
E1B mRNAs in thecell-free
system wereof
58K and 28K(see
onlonger
exposure)
and of19K
and 17K apparent molecularweight
(Fig. 3B, lane 7).
Both
Adl2 ElA
and E1Bproteins
weretranslated
frommRNA
that hadbeen
hybridization
selected to theentire
transforming region
(EcoRI-C;
0to16.5mu)
of
Adl2
(Fig. 3B, lane 6).Most
importantly,
the Ad2ElA
proteins
produced
invivocomigrated
in the sameSDS-polyacrylamide
gel
with thecorresponding
ElA
proteinssynthesized
in vitro. This wason November 10, 2019 by guest
http://jvi.asm.org/
[image:4.612.83.278.73.332.2]5
B
1
2 3
4 5
6
7
8
W. w- I
I
~~~EU
W-58
3
9U
--4-
o,39
37"wl
_-
"*
*a-37
386'
36
%.,...,..
19
17
FIG. 3. Fluorograph of [35S]methionine-labeled proteins from Adl2-transformedcells. (A) Proteinsimmunoprecipitated by 12-1A-FPantisera in the presence of the 12-1A-FP-antigen.The HA12/7-transformed cells were labeled with 50
1±Ci
of[35S]methionine
per ml for 2 h, and proteins of the cell lysate (fraction I, cytosol) wereimmunoprecipitated with12-1A-FP antiseraasdescribed in thetext.
Where indicated, increasing amounts of theextractcontainingthe 12-1A-FP antigen (Fig. 2B, lane 6)wereincubated for10min with the 12-1A-FP antisera before the immunoprecipitation reaction. Labeled proteins were separated in 15% SDS-polyacrylamide gels (26). Immunoprecipitationof
[35S]methionine-labeled
proteinsfrom HA12/7 cells with12-1A-FPantiserawas asfollows: lane1, without 12-1A-FP antigen; lane 2, with 2 ,ug of 12-1A-FPantigen;lane3,with5 ,ug of12-1A-FP antigens;lane4, with10
pLg
of 12-1A-FPantigen. Lane 5, immunoprecipitation of[35S]methionine-labeled
proteins with preimmune sera and no 12-1A-FP antigen. The apparent molecularweights of the proteinsareshownattheleft. (B)Compari-son ofElA proteins translated in vitro by hybridization-selected RNAwithproteins labeled in vivo andimmunoprecipitated bythe 12-1A-FPantiserum. Extracted RNA fromAdl2-transformed cells was hybridization selected to the specific restriction fragments containing Adl2 DNA (35) and in vitro translated in the rabbit reticulolysate cell-free system (31). Labeledproteinswereseparated in a15% SDS-polyacrylamide gel (26). The Adl2-transformed cells were labeled with 50 ,Ciof
[35S]methionine
per mlfor 1 to4h, and invivo-labeled proteins of the cell lysate(fractionII)were immuno-precipitated with the 12-1A-FP antiseraas described in the text.[35S]methionine-labeled
proteins translated in vitro by RNA from HA12/7 cells were hybridization selected to the following: Adl2 EcoRI-CDNA(0to 16.5 mu;ElA + E1B)(lane 6); anAdl2 PvuIl fragment (1,906 to 3,623 bp;E1B)(lane 7);pAd441 DNA (404 to 911 bp; E1A) (lane 8); and RNA from M012/F1O cells hybridization selectedtopAd441DNA(E1A)(lane 1). Proteins translated in the cell-free system with RNA from Ad5-infected HeLa cells isolated during late infection serve as molecular weight markers (lane 5).Lane 4, cell-free reaction with no added mRNA. [35S]methionine proteins frominvivo-labeledM012/F1Ocells were
immunoprecipi-tatedby12-1A-FPantisera (lane 2) andpreimmune sera (lane 3). The apparent molecular weights are shown at the left and right. Lane 1
wasoverexposed.
true
for
severalAdl2-transformed
hamster and mouse cell lines. For example,analysis of
theM012/F1O
cells shows that the 39K and 37KElA
proteins translated in vitro fromhybridization-selected
RNA (Fig. 3B, lane 1) comigratedwith proteins of thesame size that had been labeled in
vivo
andimmunoprecipitated by the 12-1A-FP antisera (lane 2); theminor 22K ElA protein detectedas atranslation product was not observed byimmunoprecipitation. It is also impor-tantthat the Adl2 ElAproteins from the M012/F1O-trans-formed mouse cells (Fig. 3B, lanes 1 and 2) are the same apparentsize asthose from the HA12/7-transformed hamster cells(lane 8).
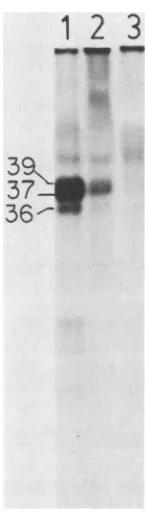
Adl2 ElAproteins were phosphorylated in vivo. The Adl2 ElA proteins which had been immunoprecipitated from [35S]methionine-labeled, transformed cells were alsoshown tobephosphorylated in vivo. Phosphoproteins of 39K, 37K, and 36K wereimmunoprecipitated from 32P-labeled HA12/7-transformed cells by the 12-1A-FP antisera (Fig. 4, lane 1).
No
phosphoproteins
were immunoprecipitated by preim-mune sera (Fig. 4, and lane 3). A partial inhibition of theimmunoprecipitation
reaction by coaddition of 12-1A-FP antigen extract with 12-1A-FP antisera to the labeled celllysate helped
todramatize that the 37K immunoprecipitation product is the predominant Adl2 ElA phosphoprotein inHA12/7-transformed
cells (Fig. 4, lane 2). Completeinhibi-tion of the
immunoprecipitation reaction (Fig. 4,
lane2)
was achievedby using
a greater amount of 12-1A-FP antigen (data not shown). In similar experiments (datanot shown), the Adl2 ElAproteins
from infected cells werealso shown to bephosphorylated.12-lA-FP antisera
immunoprecipitated
ElAproteins
from Ad5-transformed and infected cells.The
ElAcoding region
1
2 3
39<
37
36-FIG. 4. Autoradiograph of
32P-labeled
proteins from Adl2-trans-formed cellsimmunoprecipitated by the 12-1A-FP antisera.HA12/7-transformed cells were labeled with
32p
(125±Ci/ml
for 4h),andthecell extract was fractionated as described in the text. Fraction I
lysate(cytosol) was used here. Labeledproteinswereseparatedin a
15% SDS-polyacrylamide gel (26) and visualized on Kodak XAR
film.Immunoprecipitationof
32P-labeled
proteinswasbythe follow-ing: the 12-1A-FP antiserum (lane 1); the 12-1A-FP antiserum incubatedin thepresence of 10 ,ug ofthe12-1A-FP antigenextractasdescribedin thelegendtoFig.3A(lane 2); preimmunesera(lane 3).
Theapparent molecularweightsof theimmunoprecipitated proteins
areindicatedtotheleft oflane 1.
Al
2 3 4
367-
_36
.*on November 10, 2019 by guest
http://jvi.asm.org/
[image:5.612.58.295.74.319.2] [image:5.612.397.471.371.627.2]of
Adl2
contained in the vector pAd441 (Fig. 1) shares limited homologywiththecorresponding N-terminus coding region of Ad5 ElA. A direct comparison ofthe aminoacid sequencesofAdl2 ElA(33, 42)andAd5ElA (32)indicates that thehighest homologyis between amino acid residues 40 through 80 of both AdS and Adl2 ElA in pAd441. On the basis of this homology, the12-lA-FP
antiseraweretestedfor their ability to immunoprecipitate Ad5 ElA proteins. Ly-sates were prepared from 293 cells that had been labeled with[35S]methionine.
The 293 cells are human embryonic kidney cells that were transformed by AdS DNA (19) and expressAd5
ElA andE1Bgeneproducts (2, 34).Reaction of the 12-1A-FP antisera with either cell fraction I lysate (cytosol) or fraction II lysate (nuclei, membranes, and organelles) resulted in the immunoprecipitation of two groups ofproteins ranging from 38 to 36K and 35 to 33K (Fig.5A,lanes 2 and 4, respectively). Preimmune sera failed to immunoprecipitate these Ad5 proteins (Fig. 5A, lanes 3 and 5). For comparison, the corresponding ElA proteins translated in vitro by RNA from 293 cells that had been hybridization selected to a restriction fragment which con-tainedAd5 EIADNA are shownin Fig.5A, lane 1,and are indicated by both sets of arrows. In a previous study (34), the ElA proteins translated in vitro from hybridization-selected RNAs were reported as 51K and 48K. In this experiment, the apparent molecularweights of the two Ad5 ElA proteins are dramatically smalleras aresult of altering the ratio of bis toacrylamide.From the DNA sequence, the Ad5 ElAgene is known to encode only two majorElAproteins (32). Furthermore, by hybridization selection and cell-free translation, each ElA mRNA from 293 cells appears to encode a single protein as shown previously (34)and above (Fig.5A,lane 1). Thus,the several
[35S]methionine
products synthesized in vivo (Fig. 5A, lanes 2 and 4) could represent a population ofvarious modified forms ofElA
proteins in 293 cells. Indeed, lysates prepared from 293 cells that had been labeled with32P and reacted with the 12-1A-FPantisera appeared to immunopre-cipitate only two Ad5 ElA proteins (Fig. 5B, lane 4; upper and lower arrows). These ElAphosphoproteinswere detect-ed in both fractions I andII of the cell lysate (Fig.5B, lanes 1 and 4, respectively) and failed to be immunoprecipitated in the presence of 12-1A-FP antigen (Fig. 5B, lanes 2and 5) or when preimmune serum was substituted for the 12-1A-FP antiserum (Fig. 5B, lanes 3and 6).These results demonstrated that antibodies directed against theAdl2 ElAproteins are capable ofstrongly cross-reacting with Ad5 ElA proteins. Offurther interest, ElA proteins from Ad2-transformed hamster cells were also immunoprecipitated by the 12-1A-FP antisera (data not shown).
In addition, the12-1A-FPantisera wereused to immuno-precipitate the ElAproteins fromAd5-infected cells. In this experiment, HeLa cells were infected at a multiplicity of infection of50in theabsence ofdrugs andwerelabeled at 8 h postinfection with [35S]methionine or
32p,.
When the [35S]methionine-labeled proteins from the cytosol fraction (fractionI) were incubatedwith12-1A-FP antisera, proteins of 38K and 36K were immunoprecipitated (Fig. 6, lane 4; upper and lower arrows,respectively).Theseinvivo-labeled proteins comigrated with thetwoknown AdS ElA proteins translatedinvitro byhybridization-selected RNA (34; Fig. 6, lanes 5 and 9). Furthermore, the [35S]methionine-labeled proteins werenotimmunoprecipitated from uninfected cells (lane 1), by preimmune antisera(lane 2), or inthe presence of the12-1A-FPantigen (lane3). Moreover, 32P-labeled ElAA
A123l
1
2 3
4
5
_
{36-38
{33-35
B
1
2
3456
_"
WV,
3
FIG. 5. Proteins from theAdS-transformed293cells immunopre-cipitated with the 12-1A-FP antiserum. (A) Fluorograph of [35S]methionine-labeled proteins. 293 cells were labeled with
[35S]methionine
from which the fractionated cell lysates were im-munoprecipitated with the 12-1A-FP antisera. Labeled proteins werefractionated in a 15%SDS-polyacrylamide gel (26).Immuno-precipitation of
[35S]methionine-labeled
proteins from fraction Ilysate (cytosol) is shown as follows: lane 2, by 12-1A-FPantisera; lane 3, by preimmune sera. Immunoprecipitation of
[35S]methion-ine-labeled proteins from fraction II lysate (membrane, nuclei, and organelles) is shown as follows: lane 4, by12-1A-FPantisera;lane5, bypreimmune sera. Lane 1, labeled proteins translated fromRNA
isolated from 293 cells that had been hybridization selected to a
fragment ofAdSDNA(O to 9.4 mu;ElAand E1B) and translated in a cell-free system as previously described (35). The arrows to the left of lane 1 identify the positions of the larger (upper arrows)and smaller (lower arrows) ElA proteins translated in vitro, and the apparent molecular weights of immunoprecipitated proteins are
shown at the right. (B)Autoradiograph of
32P-labeled
proteins.The293 cells were labeled with
32p,
and thecell lysate wasfractionedasdescribed in the text. The labeled proteins were separated ina15% SDS-polyacrylamide gel and visualized on Kodak XAR film.
32p_
labeled proteins from fraction I lysate(cytosol) were immunopreci-pitated by the following: the 12-1A-FP antisera (lane 1);the 12-1A-FP antisera incubated in the presence of 10 ,ug ofthe 12-1A-FP antigen extract as described in the legend to Fig. 3A (lane 2); preimmune sera (lane 3). Lanes 4, 5, and 6 are the sameaslanes1, 2, and 3,respectively, except that fraction II lysate (membrane, nuclei and organelles) was used in place of fraction I. Twice as many trichloroacetic acid-precipitable counts were used in lanes 1 to 3compared with lanes 4 to 6. The positions of theimmunoprecipitated proteins are indicated by the upper and lower arrows.
proteins in both fraction I (cytosol) and fraction II
(nuclei,
organelles, and
membranes)
lysates
ofAdS-infected
cells (Fig. 6, lanes 6 and 8,respectively)
comigrated
with[35S]methionine-labeled
ElAproteins
(Fig.
6,
lanes 4 and9).
These AdS ElA
phosphoproteins
were notimmunoprecipi-tated from uninfected cells
(Fig. 6,
lane7).
These results againdramatize thehighly
specific
and strongcross-reactive nature of the 12-1A-FP antisera toimmunoprecipitate
ElA proteins ofAdS.
DISCUSSION
TheElA geneof adenovirus
performs
acrucialfunction
in both infected and transformed cells.During
infection of human cells, a product of the ElA gene modulatesexpres--,,. W1---w
]P.>400
a
)0.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:6.612.324.558.78.287.2]12
3 4 5 6 7 8 9
;3
~~~~4<
FIG. 6. Fluorograph of ElA proteins from AdS-infected cells labeled with
[35S]methionine
or32Pand immunoprecipitatedbythe12-1A-FPantisera.Ad5ElA and E1B proteins translatedin vitroby
hybridization-selected RNAswereusedtocompareand confirmthe
identity of the immunoprecipitated products. Labeled proteins were
separated ina 15% SDS-polyacrylamide gel (26). Extracted RNA
from AdS-transformed293 cellswashybridization selected (35)toa 0to9.4mufragment(ElAand E1B)ofAdS DNA,derivedfromthe
pLA1 plasmid, andin vitro translated in the rabbit reticulocyte cell-free system (31). HeLa cells were infected at a multiplicity of
infection of 50 in the absence of drugs andwerein vivolabeledat8h postinfection with [35S]methionine or32Piasdescribed in the text.
The cellextractswerefractionated intoacytosol fraction(fractionI)
lysateandanuclei, membrane, and organelle (fractionII)lysate,as
described in the text, and immunoprecipitated with the 12-1A-FP
antisera.Immunoprecipitated[35S]methionine-labeledproteinswere
fromthecytosolfraction of the following: uninfected HeLa cells by the 12-1A-FPantisera (lane 1); infected HeLa cells by preimmune
sera(lane 2);infectedHeLa cellsby12-1A-FP antisera incubatedin thepresenceof10
,ug
of12-1A-FP antigenextractasdescribedinthelegend to Fig. 3 (lane 3); infected HeLa cells by the 12-1A-FP antisera (lane 4). Immunoprecipitated 32P-labeled proteins were
from the following:the cytosol fractionof infectedHeLacells bythe 12-1A-FPantisera (lane 6); the cytosol fractionof uninfected HeLa
cellsby the 12-1A-FPantisera (lane7); thenuclei, membranes,and
organelle fraction of infected HeLa cells (lane 8). [35S]methionine
proteins translated in vitro by RNA from 293 cells hybridization selectedtoa0to9.4murestriction fragmentof AdS DNA (lanes5
and 9). The upper and lower arrows point to the ElA proteins
synthesized in vitro (lanes5 and 9) as previously determined by
hybridization selection (35).
sion of other early viral genes(4, 24, 30). This product isthe larger of the two acidic proteins encoded by Ad5 ElA (28, 34). Events leading to complete cellular transformation of
rodent cellsrequire both adenovirusElAandE1Bgenes(18,
22, 44). Characterization and comparison of the proteins of
Ad5
and Adl2areimportant sincethese divergentstrainsareable to transcomplement one another during infection (47)
and since the ElA gene product of Adl2, but not AdS,
appearstosuppressclassImajor histocompatibility antigens
in
ratsto atleast
paitially
accountfor the
highly
tumorigenic
potential of
Adl2 and the
weakly tumorigenic potential
ofAd5
(6, 39).
In
this
study,
wereport theproduction
of the firstAdl2
ElA
monospecific antiserum. This antiserum
(12-1A-FP
antiserum)
is
specifically
directed
against
the N-terminal half
of the ElA
proteins
of
Adl2.Moreover,
the12-1A-FP
antiserum
strongly
cross-reactswith AdS ElA
proteins.
Forboth
transformed and infected cells of Adl2
andAd5,
respectively,
we report thatthe
12-1A-FP antiserum
isim-munospecific for ElA
proteins
by
(i)
the
ability
of12-1A-FP
antigen
toblock
immunoprecipitation
ofElA
proteins
and(ii) the
comigration
of
immunoprecipitated
ElA
proteins
with
ElA proteins translatedin
vitrofrom
hybridization-selected
mRNAs. Furthermore,the
12-1A-FPantiserum
immunoprecipitated
phosphorylated ElAproteins
from both
transformed and infected cells of
Adl2
andAdS,respective-ly.
The 12-1A-FP antiserum appeared to react stronglywith
ElA
proteins,
presumablysince it is
polyclonal
for
multiple
determinants within the N-terminus.
In
the Adl2-transformed hamster
cell line
HA12/7,
the12-1A-FP
antiserum
specifically immunoprecipitated
twomajor
ElA
proteins of
39K and 37K,respectively.
Inaddition,
aminor ElA
proteinof
36K was observed.Since
two openreading frames
arepredicted from
the DNA sequence ofAdl2
ElA (33, 42), it was reasonable to postulate that theminor
ElAprotein
(36K)could be
generated
from
aproteo-lytic cleavage of
one orboth major
ElA proteins (39K and37K). However,
mRNAfrom HA12/7 cells that
hadbeen
hybridization selected by
Adl2
ElA DNAand translated in acell-free
systemalso synthesized two major proteins andoneminor protein
which comigrated with eachof
therespective
immunopreciptated
ElAproteins. Thus, there is either
a susceptible proteolytic cleavagesite in
one or both of the majorAdl2
ElA proteins which is recognized in vivo and in vitro, or, less likely, use of a different combination ofreading frames or an internal start site accounts for the appearanceof
thethird
andunpredicted
ElAprotein
of
Adl2.Tryptic
and N-terminal amino acid sequence analysis ofthese
pro-teins
willbe
necessary to determine theorigin of
the 36K protein.It
is important
to notethatthe apparent molecularweights
of
theAdl2
ElA proteins reported here exceed thecoding
capacity of the
Adl2
ElA transcripts (37). This was not surprising since the highly acidic ElA proteins of AdS are known to migrate aberrantly inSDS-polyacrylamide
gels(34, 50).
Infact,
we haveobserved
thatlowering
thebis/
acrylamide ratio causes the Adl2 ElA proteins to migrate as proteins which are almost double in molecular weight from their predicted sizes of 29K and 26K, respectively (33, 42).
According to the DNA sequence (33, 42), the two major
Adl2
ElA proteins are 266 and 235 amino acidresidues
in length. The12-1A-FP
antiserum recognizes the N-terminal 131 amino acids of eachAdl2ElA
protein since this stretch is encoded by the bacterial expression vectorpAd441,
used in the production of antigen. The12-1A-FP
antiserum
isalso capable of immunoprecipitating AdS ElA proteins probably by virtue of amino sequence acidhomology
toAdl2
ElA, which almost exclusively resides betweenresidues
40 and 80.The 12-1A-FP antiserum immunoprecipitated two ElA proteins from
Ad5-infected
cells, when labeled with either[35S]methionine
or32p.
In contrast, the 293 cells, which constitutively synthesize ElA mRNAs (2),appeared
to contain several forms of each ElA protein when the cells were labeled with[35S]methionine
butfewer forms when
on November 10, 2019 by guest
http://jvi.asm.org/
[image:7.612.98.261.78.340.2]theywerelabeled with
32p.
As observed hereand alluded
toelsewhere (16 and 36), differences in the number
of
ElA proteins seen inAd5-infected
and transformed cellscould
reflect a distinction between their respective
pool sizes of
phosphorylatedandunphosphorylatedformsaswellas
their
cellular
half-lives.
Rowe etal.
(36)
recently reported four
major and four minor forms of ElAproteins from
AdS-infected cells in two-dimensionalpolyacrylamide
gels after
immunoprecipitation by an antipeptide serumdirected
against the
carboxyl terminus. Several
cautions about
at-tempting to interpret the
actual number
of cellular ElA
proteinsby
immunoprecipitation with
monospecific
antisera
are suggested by the resultsof
theseexperiments.
Factors such as the time point ofinfection,
the typeof infected
or transformed cell, andthe
typeof gel matrix used in the
analysis (e.g., the bis/acrylamide ratio in
SDS-polyacryl-amide gels) may influence the
number of detectable ElA
proteins which are incompletely modified or partially de-graded.The potential uses of the 12-1A-FP monospecific
antisera
are (i) toimmunoaffinity purify cellular Adl2 and Ad5 ElA proteins, (ii) to discern whether the N termini of Adl2 andAd5
ElAproteins perform functionswhich areindependent
of the carboxylterminus, (iii)to
determine
specific
biochem-ical featuresof theElA proteins (e.g.,bindingto DNA),and
(iv) to determine whether proteins immunoprecipitatedby
tumor-bearing antisera areElA proteins (1, 17, 37, 48).ACKNOWLEDGMENTS
We thank Jim Williams from Carnegie Mellon University, Pitts-burgh, Pa., for making available the Adl2-transformed mouse cell line M012/F10; W. Doerfler from the University of Cologne, Germany, for the Adl2-transformed hamster cell line HA12/7; Barbara Ghrist and Agata Giallongo of The Wistar Institute for advice on immunoprecipitation schemes; and Angela Varrichio (The Wistar Institute) for help with the N-terminal sequence analysis.
This work was supported by Public Health Service grant CA 29797 from the National Cancer Institute and the Ruth Estrin Goldberg Memorial for Cancer Research to R.P.R.
LITERATURE CITED
1. Achten, S., and W. Doerfler. 1982. Virus-specific proteins of
adenovirus type 12-transformed and tumour cells as detected by immunoprecipitation. J. Gen. Virol. 59:357-366.
2. Aiello, L., R. Guilfoyle, K. Huebner, and R. Weinmann. 1979.
Adenovirus 5 DNA sequences present and RNA sequences transcribed in transformed human embryo kidney cells
(HEK-AdS or 293). Virology 94:460-469.
2a.Amann, E., J. Brosius, and M. Ptashne. 1983.Vectorsbearing a hybrid trp-lac promoter useful for regulated expression of
cloned genes in Escherichia coli. Gene 25:167-178.
3. Berger, S. L., and C. S. Birkenmeier. 1979. Inhibition of
intractable nucleases with ribonucleoside-vanadyl complexes: isolation of messenger ribonucleic acid from resting
lympho-cytes. Biochemistry 18:5143-5149.
4. Berk, A. J., F. Lee, T. Harrison, J. Williams, and P. A. Sharp. 1979. Pre-early adenovirus 5 gene productregulatessynthesis of early viral messenger RNAs. Cell 17:935-944.
5. Berk, A. J., and P.A. Sharp. 1978. Structure of the Ad2 early mRNAs. Cell 14:695-711.
6. Bernards, R., P.I. Schrier, A. Houweling, J. L. Bos, A.J. van der Eb, M.
ZiJlstra,
and C. J. M.Melief. 1983.Tumorigenicity of cells transformedbyadenovirus type 12 byevasionofT-cellimmunity. Nature (London) 305:776-779.
7. Bikel, I., T. M. Roberts, M. T. Blandon, R. Green, E. Amann, and D. M. Livingston. 1983. Purification ofbiologicallyactive Simian virus 40 small tumor antigen. Proc. Natl. Acad. Sci.
U.S.A. 80:906-910.
8. Bonner, W. M., and R. A. Laskey. 1974. A film detection
method for tritium-labeled proteins and nucleic acids in poly-acrylamide gels. Eur.J. Biochem.46:83-88.
9. Carlock, L. R., and N. C. Jones.1981.Transformation-defective
mutant of adenovirus type 5 containing a single altered ElA mRNAspecies. J.Virol. 40:657-664.
10. Chow, L. T., T. R. Broker, and J. B. Lewis. 1979. Complex splicing patterns ofRNAsfrom the early regions of Ad2.J.Mol. Biol. 134:265-303.
11. Esche, H., and B.Siegmann. 1982. Expression of early viral gene products in adenovirus type12-infected and transformed cells. J. Gen. Virol. 60:99-113.
12. Feldman, L. T., M. J. Imperiale, and J. R. Nevins. 1982. Activation of early adenovirustranscription by the herpesvirus immediate early gene: evidence for a common cellular control factor. Proc. Natl. Acad. Sci. U.S.A. 79:4952-4956.
13. Feldman, L. T., and J. R. Nevins. 1983. Localization of the adenovirusElA protein, a positive-acting transcriptional factor in infected cells. Mol. Cell. Biol. 3:829-838.
14. Freeman, A. E., P. H. Black, E. A. Vanderpool, P. H. Henry, J. B. Austin, and R. J. Huebner. 1967. Transformation of pri-mary ratembryo cells byadenovirus type 2. Proc. Natl. Acad. Sci. U.S.A. 58:1205-1212.
15. Freeman, A. E., P. H. Black, R. Wolford, and R. J. Huebner.
1967. Adenovirus type 12-ratembryo transformation system. J. Virol. 1:362-367.
16. Gaynor, R. B., A. Tsukamoto, C. Montell, and A. J. Berk. 1982. Enhanced expression ofadenovirus transforming proteins. J. Virol. 44:276-285.
17. Gilead, Z., Y.-H. Jeng, W. M. S. Wold, K. Sugawara, H.M. Rho, M. L.Harter, and M.Green. 1976.Immunological identifi-cation of two adenovirus 2 induced early proteins possibly involved in celltransformation. Nature (London) 264:263-266. 18. Graham, F. L., P. J. Abrahams, C. Mulder, H. L. Heijneker, S. 0. Warnaar, F. A. J. DeVries, W. Fiers, and A. J. van der
Eb. 1974. Studies on invitrotransformation by DNA and DNA fragments of human adenoviruses and simian virus 40. Cold Spring Harbor Symp. Quant. Biol. 39:637-650.
19. Graham, F. L., J. Smiley, W. C. Russell, and R. Nairn. 1977. Characterization of a human cell linetransformed by DNA from human adenovirus type 5. J. Gen. Virol. 36:59-72.
20. Green, M., W. S. M. Wold, and W. Buttner. 1981. Integration andtranscription of group C human adenovirus sequences in the DNA of five lines oftransformed rat cells. J. Mol. Biol.
151:337-366.
21. Guarente, L., G. Lauer, T. M. Roberts, and M. Ptashne. 1980. Improved methods for maximizing expressionofacloned gene: abacterium that synthesizes rabbit ,-globin. Cell 20:543-553.
22. Houweling,A., P. J. vandenElsen, and A. J. van derEb. 1980. Partialtransformation of primary rat cellsby the leftmost4.5% fragment ofAdenovirus 5 DNA. Virology 105:537-550. 23. Hunkapillar, M. W., and L. E. Hood. 1978. Direct
microse-quence analysis ofpolypeptides using an improved sequenator, anonprotein carrier (polybrene), and high pressureliquid chro-matography. Biochemistry 17:2124-2133.
24. Jones, N., and T. Shenk. 1979. Anadenovirus type5early gene function regulates expression of other early viral genes. Proc.
Natl. Acad. Sci. U.S.A. 76:3665-3669.
25. Kitchingman, G. R., and H. Westphal. 1980. The structure of
adenovirus 2 early nuclear and cytoplasmicRNAs.J.Mol. Biol. 137:23-48.
26. Laemmli, U. K. 1970. Cleavage of the structuralproteins during the assembly of the head ofbacteriophage T4. Nature(London) 227:680-685.
27. Miller, J. H. 1972. Experiments in molecular genetics. Cold SpringHarbor Laboratory, Cold Spring Harbor, N.Y.
28. Montell, C., E. F. Fisher, M. H. Caruthers, and A.J. Berk.
1982.Resolving the functions of overlapping viral genes bysite
specific mutagenesis at a mRNA splice site. Nature (London)
295:380-384.
29. Mulligan, R. C., and P. Berg. 1981. Selection for animal cells that express the Escherichia coli gene coding for
xanthine-guanine phosphoribosyltransferase. Proc. Natl. Acad. Sci. U.S.A. 78:2072-2076.
on November 10, 2019 by guest
http://jvi.asm.org/
30. Nevins, J. R. 1981. Mechanism of activation of early viral transcription by the adenovirus ElA gene product.Cell 26:213-220.
31. Pelham, H. R. B., and R. J.Jackson. 1976. An efficient mRNA-dependent translation system from reticulocyte lysates. Eur. J. Biochem. 67:247-256.
32. Perricaudet, M., G. Akusjarvi, A.Virtanen,and U. Pettersson. 1979. Structure of two spliced mRNAs from the transforming region of human subgroup C adenoviruses. Nature (London) 281:694-696.
33. Perricaudet, M., J. M.le Moullec, and P. Tiollais. 1980. Struc-ture of two adenovirus type 12 transforming polypeptides and their evolutionary implications. Nature (London) 288:174-176.
34. Ricciardi, R. P., R. L. Jones, C. L. Cepko, P.A.Sharp, and
B. E. Roberts. 1981. Expression of early adenovirus genes requires aviral encoded acidic polypeptide. Proc. Natl. Acad. Sci. U.S.A. 78:6121-6125.
35. Ricciardi, R. P., J. S.Miller, and B. E. Roberts. 1979. Purifica-tion and mapping of specific mRNAs by hybridizaPurifica-tion-selecPurifica-tion and cell-free translation. Proc. Natl. Acad. U.S.A. 76:4927-4931.
36. Rowe, D.T., S. Yee, J. Otis, F. L. Graham, andP. E.Branton. 1983.Characterizationof human adenovirus type5early region 1Apolypeptides using antitumor sera and an antiserumspecific for the carboxy terminus. Virology 127:253-271.
37. Saito, I.,K. Shiroki, andH.Shimojo. 1983.mRNAspecies and proteins of adenovirus type12transforming regions: identifica-tion ofproteins translated from multiple coding stretches in2.2
kb region E1BmRNA invitro andin vivo. Virology
127:272-289.
38. Sawada, Y., and K.Fujinaga. 1980. Mapping of adenovirus 12
mRNA's transcribed from the transforming region. J. Virol. 36:639-651.
39. Schrier,P.I.,R.Bernards,R. T.M. J. Vaessen,A.Houweling,
and A.J. van der Eb. 1983. Expression of class I major histocompatibility antigens switched off by highly oncogenic adenovirus 12 in transformed rat cells. Nature (London) 305:771-775.
40. Solnick, D., and M. A. Anderson. 1982. Transformation-defi-cient adenovirus mutant defective in expression of region 1A
but not region 1B.J. Virol. 42:106-113.
41. Spector, D. J., M.McGrogan, and H. J. Raskas. 1978. Regula-tion of the appearance of cytoplasmicRNAs from region1of the adenovirus-2 genome. J. Mol. Biol. 126:395-414.
42. Sugisaki, H., K.Sugimoto, M. Takanami, K. Shiroki, I. Saito,H.
Shimojo, Y. Sawada, Y. Uemizu, S. Uesugi, and K. Fujinaga. 1980. Structure and gene organization in the transforming HindIII-G fragment of Adl2. Cell 20:777-786.
43. van denElsen, P.,S. de Pater, A. Houweling, J. van der Veer,
and A. van der Eb.1982. Therelationship between region ElA andE1B of human adenoviruses in cell transformation. Gene
18:175-185.
44. van der Eb, A. J., C. Mulder, F. L.Graham, andA.Houweling. 1977. Transformation with specific fragments of adenovirus
DNAs. I. Isolation of specific fragments with transforming activity of adenovirus 2 and5DNA. Gene2:115-132.
45. Vogelstein, B., and D. Gillespie. 1979.Preparative andanalytical purification of DNA from agarose. Proc. Natl. Acad. Sci. U.S.A. 76:615-619.
46. Wartell, R. M., and W. S. Reznikoff. 1980. Cloning DNA restriction endonuclease fragments with protruding single-stranded ends.Gene 9:307-319.
47. Williams, J., Y. Ho,and R. Galos. 1981. Evidencefor functional relatednessofproductsencodedbythetransformingsequences of human adenovirus types 5 and 12. Virology 110:208-212. 48. Wold, W. S. M., G. Chinnadurai,M.Green, and S. Mak. 1979.
Identification of adenovirus type 12 candidate transformation proteins by radioimmunoprecipitation with antisera to EcoRI-C-fragment transformed cells. Virology 94:208-213.
49. Yee, S.-P., D. T. Rowe, M. L. Tremblay, M. McDermott,and P.E. Branton. 1983. Identification of human adenovirus early region 1 products by using antiseraagainst synthetic peptides corresponding to the predicted carboxy termini. J. Virol. 46:1003-1013.
50. Zur Hausen, H. 1973. Interaction of adenovirus type 12 with hostcell chromosomes. Prog. Exp. Tumor Res. 18:240-259.
![FIG.5.cipitatedlaneine-labeledbylysateorganelles)precipitation[35S]methionine-labeled[35S]methioninemunoprecipitatedwereaisolatedfragmentleftdescribed293apparentsmallerlabeledandantigenshownSDS-polyacrylamideandtrichloroaceticpitatedcomparedpreimmuneFPprot](https://thumb-us.123doks.com/thumbv2/123dok_us/1428256.95450/6.612.324.558.78.287/cipitatedlaneine-labeledbylysateorganelles-precipitation-methionine-labeled-methioninemunoprecipitatedwereaisolatedfragmentleftdescribed-apparentsmallerlabeledandantigenshownsds-polyacrylamideandtrichloroaceticpitatedcomparedpreimmunefpprot.webp)
![FIG. 6.labeled Fluorograph of ElA proteins from AdS-infected cells with [35S]methionine or 32P and immunoprecipitated by the](https://thumb-us.123doks.com/thumbv2/123dok_us/1428256.95450/7.612.98.261.78.340/fig-labeled-fluorograph-proteins-infected-cells-methionine-immunoprecipitated.webp)