JOURNALOFVIROLOGY, June 1973,p.823-831
Copyright6 1973 AmericanSocietyforMicrobiology
Vol.11, No.6
Printed inU.SA.
Formation of Influenza Virus Proteins
HANS-DIETER KLENK AND RUDOLF ROTI
Institut furVirologie, JustusLiebig-Universitdt, 6300 Giessen, Germany Received forpublication 1 February 1973
Eight virus-specific proteins have been found in chicken embryo fibroblasts
infected with fowlplague virus. Among themaretwoglycoproteins whicharethe
constituentsofthe hemagglutininonthe virus particle.Theyarederived froma
largeprecursor glycoprotein by cleavage ofa covalentlinkage. The reaction can
be blocked by theprotease inhibitordiisopropylfluorophosphate and the amino
acid analoguefluorophenylalanine. This indicates thatapeptide bond is cleaved.
If infected cells are kept at 25 C, a temperature at which virus maturation is
inhibited, theprecursorglycoprotein is cleavedatasignificantlyslower ratethan
at37 C. It appears, however, that areducedsynthesis of the carbohydrate-free
envelope protein is responsible for the block of virus maturation at25 C rather
thanthelower cleavage rate of the precursor.
The protein constituents of influenza virus consist ofcarbohydrate-free polypeptidesand of
glycoproteins. One of the carbohydrate-free polypeptides forms the protein subunit of the nucleocapsid (10, 16), anotherone isa
constit-uent of the viral envelope where it coats the inner side of the lipid layer (2, 20, 21). The glycoproteins are located on the outside of the lipid layerasconstituentsofthe surface projec-tionsorspikes (2, 12, 20).
Inaddition tothe constituent proteins of the virion, nonstructural polypeptides have been described in influenza virus-infected cells (4, 13,
15, 22). Some ofthese nonstructural proteins
are precursors of the viral hemagglutinin, and
theiranalysis throwssomelightonthesequence
ofeventswhichareinvolvedinthebiosynthesis
ofthisenvelopeconstituent. The first identifia-blestepistheglycosylationof apolypeptideto a
largeglycoprotein (13). Itwassuggestedfirstby Lazarowitz et al. (15) that this glycoprotein is cleaved into two smaller glycoproteins. This
concept has been confirmed in the present
communication by fowl plague virus (FPV)
grown in chicken embryo fibroblasts.
Experi-ments will be described which give further
information about the appearance of virus-specific material ininfectedcells, and
particu-larly aboutthenatureof the cleavage reaction.
MATERIALS AND METHODS
Virus and cells. The Rostock strain of FPV was used in all experiments. It was grown in primary cultures of chicken embryo fibroblasts (23). Virus stocksweregrown inembryonated eggs.
Virus assay. Virusyields were assayed by hemag-glutination titrations (3) and plaque assays (23).
Chemicals andisotopes. Reagents for polyacryla-mide gels and diisopropylfluorophosphate were ob-tained fromServa,Heidelberg, Germany; DL-p-fluoro-phenylalanine from Calbiochem, LosAngeles, Calif. Protein [U-_4C]hydrolysate, L-[U-14C]valine (300 mCi/mmol),L-[4, 5-3H]leucine (1.0Ci/mmol),L-
[2,3-3H]valine (1,5Ci/mmol), L-[3,5-3H]tyrosine (1,0Ci/ mmol), and D-[1_-3Hjglucosamine hydrochloride (1,1 Ci/mmol) werepurchased fromAmersham, Bucking-hamshire, England.
Labeling of infected cells. Confluent monolayers
ofinfected cellswereinoculatedat amultiplicityof10 to 50PFU/cell.Aftera30-minadsorption period,the inoculum was removed and replaced by minimal medium (6). When experimentswerecarriedoutat 37 C, the mediumwasremoved andreplaced bymedium containing radioactive isotopes 4 h after infection. When experiments were carried out at 25 C, the medium was replaced 24 h after infection.
Radioac-tive isotopes were used at the following
concentra-tions: [14C
]protein
hydrolysate,5 sCi/ml; [14C]valine,5 MCi/ml; ['H]amino acids, 50MCi/ml;
[3H]glucosa-minehydrochloride,50,Ci/ml.The medium
contain-ingtheisotopeswasleftonthe cells for10or60min.
When theexperimentwas stopped atthis point, the
monolayer waswashed three times with cold
phos-phate-buffered saline (PBS) (5), scraped offwith a
rubber policeman in 1 ml ofPBS, and storedat -20 C.When thepulsewasfollowedbyachaseperiod,the radioactive medium was removed and the cellswere
washed three times with mediumcontaininganexcess
of thecoldanalogueoftheradioactive label (10 gmol of aminoacid/ml) andincubated inthismedium. At the endofthechaseperiod the cellswerewashedand scrapedoff asdescribed above.
Polyacrylamide gel electrophoresis. The cells werehomogenized by sonic treatment. Samples of 100
juliterswereused for gelelectrophoresis. Proteins were
dissociatedwithsodiumdodecyl sulfate and mercap-toethanol and separated by electrophoresis on 10% 823
on November 10, 2019 by guest
http://jvi.asm.org/
acrylamide gels as described (1). The slicing and processing of gels for the determination of radioactiv-ity by liquidscintillation has been reported previously (11).
RESULTS
Viral proteinsynthesis in infected cells at 37 C. We reported previously that
polyacryla-mide gel electrophoresis reveals eight fowl plague virus-specific polypeptides in chicken embryofibroblasts (13). Sixofthese are
struc-tural polypeptidesofthe virion.They comprise
two hemagglutinin glycoproteins,
HA,
(mol wt49,000) and HA2 (mol wt 32,000), glycoprotein
NA (molwt 45,000) which has been attributed
totheneuraminidase,acarbohydrate-free
enve-lope polypeptide M (mol wt 26,500), the
nu-cleocapsid protein NP (mol wt 60,000), and another internal protein P (mol wt 83,500). In
addition to these structural polypeptides,
in-fected cells contain two virus-specific proteins whichare notpresent inthematurefowlplague
virion: protein NS (molwt 25,000) is found in
largeamounts in thenucleus, proteinHA (mol wt76,000)hasbeensuggestedtobeaprecursor of the hemagglutinin glycoproteins
HA,
and HA2 (15). This pattern obtained 4 h after infection in a 1-hpulse with radioactive aminoacids comprises all fowl plague virus-specific
proteins which we were able to detect in
in-fected chickenembryofibroblasts.
The pattern is different ifthe cells obtain a
pulse ofonly 10 min in length (Fig. 1). Ifthe
pulse is not followed by a chase period, the
carbohydrate-free polypeptides P, NP, M, and NSare clearlyrecognizable. Theprotein show-ing the highest incorporation of radioactivity
under these conditions is proteinHA. Proteins
HA1and HA2 cannotbedetected.
There is a shift in the virus-specific protein
pattern ifthe 10-min pulseisfollowed
by
chaseperiodsofincreasing length(Fig. 1and2).The radioactivity in proteins P, NP, and M remain
constant and in the same relative proportions
independently ofthelengthofchase. However,
afterachaseof5min,peakHAisdecreased and
peaks HA1 andHA2aredetectable. If the chase period is extended to 60 min
peak
HA hasalmost disappeared, whereas the label in
peak
HA1
and that inHA2have increasedconcomi-tantly.Theseresults indicatethat
protein
HA is lessstablethantheothervirus-specificproteinsand thatproteins HA1andHA2aretheresult of
the breakdownofprotein HA.
Viral protein synthesis at 25 C. In FPV-infected cells which are kept at 25
C,
virus-specific components such as RNA
polymerase,
ribonucleoprotein antigen, hemagglutinin, and
neuraminidase are synthesized, even though at
aslowerrate than at 37 C. However, infectious
virions are not released. These data suggested
that, at 25 C,virus maturation is inhibited at a
late stage(18).
It was ofinterest which ofthe virus-specific
proteins wereformedat 25C.Cells were kept at
this temperature for 24 h after infection and
then labeled in a 1-h pulse with radioactive amino acids. At the end of the experiment the
cells contained a considerable amount of
he-magglutinin and they showed hemadsorption
and agglutinability by concanavalin A (17).
However, neither hemagglutinin nor infectious
particleswerereleasedintothemedium.Figure 3shows a comparison ofFPV proteins
synthe-sized at 25 C 24 h after infection with those
synthesized at 37 C 4h after infection. In both
cases the proteins have been labeled in a 1-h
pulse with radioactive amino acids. Several
striking differences in the polypeptide profiles
at thesetwotemperatures can beseen. At 25C
protein HA shows the highest incorporation of
radioactivity. Ascan beseen fromdouble-label experiments with radioactive amino acids and
glucosamine,proteinHAisglycosylatedat 25 C
(Fig. 4). The other FPV-specific glycoproteins,
HA1
andHA2, are missing at 25C.However,ifthepulseat 25C isfollowed by a
chase periodatthesametemperature, it canbe
seen that proteins
HA1
and HA2 are being formed at the expense of protein HA (Fig. 5).Similar results were obtained ifthe chase was
carriedout at 37 C (Fig. 6). These experiments indicate that HAisconvertedinto
HA1
andHA2 at 25 C, but at a slower rate than at 37 C.Therefore, the formation of the hemagglutinin glycoproteins does not seem to be basically altered at 25 C.
A striking difference, however, between the
polypeptide profilesat 25 C andat37Cwasthe
apparent absence of protein M at the lower
temperature. Whereas, at 37C, proteins M and
NS can be identified as a double peak or as a
peak and a shoulder, at 25 C protein M could
notbediscriminatedinthepolypeptide profile.
Thefastest migratingpeakclearly corresponds
to proteins NS (Fig. 4). This was a constant findingin many experiments.
However, it is not clear from the data
pre-sented here whether protein M is completely
missingorwhethersmallamounts arehiddenin a relatively large NS peak. Future studies
employingother means ofdistinguishing these polypeptides, such as peptide mapping or im-munological assays, shouldthrowmorelighton
thisproblem.
on November 10, 2019 by guest
http://jvi.asm.org/
INFLUENZA VIRUS PROTEINS
3H
dm
NP
I
HA1
NS
'I
HA2
I
V0
20
301.0
50W6
70
80
90100
Fraction
number
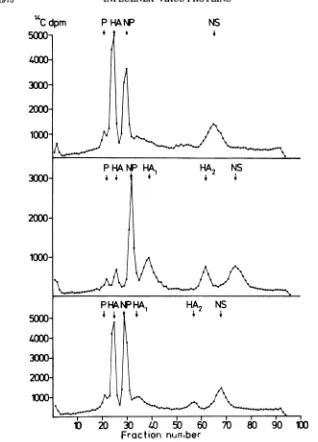
FIG. 1. Polyacrylamide gel electrophoresis ofFPVproteinssynthesized in chicken embryo fibroblasts. Cells
werelabeled4hpostinfectionbya10-minpulsewith [3H]leucine. Top, Cells were scraped off immediately after thepulse;bottom, the pulse was followed by a chase period with cold leucine for 5 min. Migration in this and all subsequent gels is from the left to the right. Arrows indicate the position of polypeptides. The neuraminidase glycoprotein has not been marked on this and most of the other figures in this paper. There is evidence that it migrates in front of theHA1peak (13).
Inhibition of the cleavage of glycoprotein from glycoprotein HA. Since the sum of the HA. The data described above conform with molecularweightsofHA1 and HA2issimilarto those of Lazarowitz et al. (15). They suggest that of HA, a cleavage reaction might be
that glycoproteins
HA,
and HA2 are derived involved.Thelinkage which is affectedby this825
VOL.11,1973
on November 10, 2019 by guest
http://jvi.asm.org/
[image:3.493.84.416.58.556.2]PHA NPH1
2M
NSf
30 40 50 6070
Fraction number
FIG. 2. Polyacrylamide gel electrophoresisofFPVproteins. Cellswereexposed toapulse with [3H]leucineas describedinFig.1.Top,Thepulsewasfollowed bya15-minchaseperiod; bottom,thepulsewasfollowed bya 60-minchaseperiod.
cleavagemustbeof covalentnatureandcannot
be a disulfide bond, because protein HA is
stable underthe conditions of the
electrophore-sis employedinthis study, i.e., inthepresence
ofSDS andareducingagent.Thesefindingsare
compatible with the hypothesis that the
cova-lentlinkage which iscleaved is apeptide bond
and that proteolytic enzymes are involved in
this reaction.
To testthis hypothesis, proteaseshave been
blockedby the useof theinhibitor
diisopropyl-fluorophosphate (DFP). This substance had
beenemployed successfully in the elucidation of
large polypeptides as precursors of poliovirus
proteins (9). The effect ofDFPonthesynthesis
of influenza virus proteins is shown in Fig. 6.
Infectedcellswerekeptat25C for 24 h and then
labeled for 1 h with radioactive amino acids.
Theonlyglycoprotein labeled under these
con-ditions is protein HA. If the pulse at 25 C is
followed by a chase period at 37 C, HA
de-creaseswithaconcomitantincrease of
HA,
andHA2. However, if DFP is addedtothe medium
during the chaseperiod, the shift of the
radioac-tivity from HA to
HA,
and HA2 is clearlyinhibited. Thisprovesthat HA isconverted into
HA,
and HA2by cleavage ofapeptide bond.Further evidence for suchaprecursor-product
relationship stems from experiments with an
amino acid analogue. It has been known for
some time that, under appropriate conditions,
L-fluorophenylalanine inhibits the formation of 3H dpm
on November 10, 2019 by guest
http://jvi.asm.org/
[image:4.493.92.407.50.455.2]INFLUENZA VIRUS PROTEINS
PIHA
4H
I
NPHAl
I
3H
dpm
4
4
30
40
50
HA2
M
NS
14C
dpm
[image:5.493.52.449.49.332.2]Fraction
number
FIG. 3. Polyacrylamide gel electrophoresis ofFPVproteins synthesized at 25 C (@-*). The cells were
labeled for 1 h with [3H]leucine, ['H]tyrosine, and [3H]valine. For comparison, virus-specific proteins synthesized at 37Candlabeledfor1h with ['4Clproteinhydrolysateweresubjectedtocoelectrophoresisonthe samegel(0--0).
14C
dPm
P
HA
NP
NS
U0
20 30
40Q50
60 70
8090
Fraction number
FIG. 4. Polyacrylamide gel electrophoresis ofFPVproteins synthesizedat25 C. Cellswereexposed for1hto
apulsewith ['4CJaminoacids(@-@)and [3HJglucosamine hydrochloride(0--O).
fowl plague virions and biologically active he-magglutinin although nucleocapsid protein is
synthesized as detected by immunological methods (19, 23). Polyacrylamide gel electro-phoresis reveals that, in the presence of
L-fluorophenylalanine (FPA) only proteins P, HA,
and NPare labeled (Fig. 7).Proteins HA1 and
HA, cannot be detected. On the gel shown in
Fig. 6, proteinsM and NS cannot be detected either. The absence of these latter proteins, however, was not a constant finding. This
experiment indicates that, in the presence of 827
VOL.11,1973
- -r-
on November 10, 2019 by guest
http://jvi.asm.org/
[image:5.493.60.447.381.572.2]P
HA NP
NSI
i
I$I
10
20
30
40
50
60
70
80
90
100
Fraction
number
FIG. 5. Polyacrylamide gel electrophoresis ofFPVproteins synthesized at 25 C. Cells were labeled 24 h postinfection bya1-hpulsewith [3H]leucine. Top,Cellswerescraped offimmediately afterthepulse;bottom, thepulsewasfollowed bya3-hchase at 25 C.
FPA, cleavage of protein HA is inhibited. The
cleavage reaction appears to be completely
blocked, not simply retarded, because even
after chase periods of several hours
HA,
andHA2cannot be detected. DISCUSSION
Theprofiles of fowl plague virus-specific
pro-teinsreported in this and inaprevious
commu-nication (13) agree well with the findings of
othergroups. Lazarowitzetal. (15)found eight
virus-specific polypeptides in cells infected with
the WSN line of influenza A virus, each of
which can be coordinated to a corresponding
FPV protein. The polypeptide pattern ofFPV
reported bySkehel (22) is, in general, similarto
ourresults, although thereare some
discrepan-cies inthe number aswell asintheproportions
of the various proteins. Skehel described two
additionalpolypeptides (P1 and 9 accordingto
hisnomenclature) whichwehavenotbeen able
to detect. It remains to be seen whether this
discrepancy is duetodifferences inthelabeling
procedures or in the electrophoresis systems.
There are striking differences in the relative
amounts ofradioactivity incorporated into the
various polypeptides between our data and
Skehel's.Theycanbeexplainedbythedifferent
labeling techniques. Whereas Skehel used
35S-methionine as protein label in most of his
experiments, we employed 3H- or "4C-amino
acids. Since methionine isincorporated
prefer-entially into the carbohydrate-free polypeptides
(15), this methodappearstobe lesssuitable for
theanalysis ofthe influenza virusglycoproteins.
Three of the four influenza virus-specific
glycoproteins (HA, HA1, HA2) werefound to be
associatedwith the viralhemagglutinin (8, 14).
There is evidence for the presence of another
glycoprotein which migrates as a shoulder on
3H
dpnm
on November 10, 2019 by guest
http://jvi.asm.org/
[image:6.493.92.442.56.419.2]INFLUENZA VIRUS PROTEINS
14C
dpm
PHANP
NS
mrn-
I
+ 4PHIA
4'4
HA2
4NS
HA2
NS
4 4
20
30
4050
e [image:7.493.77.395.55.497.2]Fraction number
FIG. 6. Polyacrylamide gel electrophoresis ofFPVproteins synthesizedat 25 C. Cells were labeled 24 h
postinfectionbya1-hpulsewith [14C]valine. Top,Cellswerescraped off immediatelyafterthepulse; middle,
thepulsewasfollowed bya1-h chaseat37C; bottom,thepulsewasfollowed bya1-h chase at 37 C, and the
mediaaddedtothecellsduringthe chaseperiodcontaineddiisopropylfluorophosphate (20 Mmol/ml).
the frontslopeofthebroad HA1 peak and which tentatively has been assignedtoasubunit of the viral neuraminidase (13). This assignation
seems to be justified by comparing the gel
patterns of fowl plague virus and other
influ-enzastrains (7). However, because of the poor
resolution and possibly the small amount of
protein NA, little information on the viral neuraminidase can be obtained from the data presented inthis publication.
The observations of Lazarowitz et al. (15)
suggested that glycoproteins HA1 and HA2 are
formed by cleavage of the large glycoprotein HA.Theirfindingsareconfirmedby the results
ofourpulse-chase experimentsat25 C as well as at37 C which also substantiate that a precur-sor-product relationship exists between HA and
HA,
and HA2. The block ofthisreaction by aprotease inhibitor proves that it is the cleavage
ofapeptide bond by which HA is converted into
HA1 andHA2.
Further evidence for proteolytic cleavage of
829
VOL.11,1973
on November 10, 2019 by guest
http://jvi.asm.org/
PFHA
NtPLNA
I,
II
I II I
I
HA2
M NSlMNS
14C
dpm
10
30
40
50
60
70
80
90
[image:8.493.57.449.61.302.2]Fraction number
FIG. 7. Polyacrylamide gelelectrophoresisofFPVproteins synthesizedin thepresenceof p-fluorophenylala-nine. Two hourspostinfection theamino acid analogue (300 ug/ml) was added tothe medium. Four hours
postinfection the cells were exposed to a pulse with [3H]valine for 1 h (a-_). During the pulse
p-fluorophenylaianine waspresent in the medium. Inacontrolexperiment, infected cells weregrown in the
absence ofp-fluorophenylalanine, labeled4hpostinfection for1hwith [14C]valine,andsubjectedto
coelectro-phoresison thesamegel(0- -0).
HAstemsfrom theexperimentswith FPA. Itis
known that this amino acid analogue is incor-porated into
poliovirus-specific
precursorpro-teins, thereby
blocking
theircleavage
(9).Ac-cordingly, HA is still
being
formed in the presence ofFPA, but cleavage asindicated bythe appearanceof
HA,
andHA2isinhibited.Itappearsthat, bytheincorporationofthe amino acid analogue, the configuration of the HA
polypeptide chain is altered insuchawaythat
it can no longer be properly cleaved. This
concept is also supported
by
thefinding
that HA synthesized in thepresence of FPAcan no longer exerthemagglutination
(20). Ourfind-ings agree with those of Lazarowitz et al.
(Virology, in press), who demonstrated the proteolytic nature ofthe cleavage with trypsin invitro. These authors also havegoodevidence that the enzymes involved are host cell
speci-fied.
At 25 C cleavage ofglycoprotein HA takes
placeat asignificantlyslowerratethanat37C.
This, however, does not
explain
the block of virusmaturation at25C, because thereisgoodevidence that thecleavagereaction isnot
essen-tial forvirusassembly.Thisissuggested
by
thefinding that large amountsof
glycoprotein
HAare present in mature virions of several
influ-enzastrains (2, 8, 12, 20). Furthermore,
experi-ments ofLazarowitz et al. (Virology, in press)
show that cleavage of glycoprotein HA is not required for the expression of the biological
properties ofthe virion, including hemaggluti-nation, butthat it appearstobe a nonessential
result of eventsoccurring in infected cellswhich
are undergoing cytopathic effects. At 25 C the
hemagglutinin glycoproteins are not only
prop-erly formed,buttheyarealsoincorporated into
the plasma membrane ofthe cell. This can be shown by a positive hemadsorption reaction (unpublished results) and byagglutinability of infected cells withconcanavalin A (17).
There-fore, the block of virus maturation at 25 C appears to be neither at the stage of the biosynthesisofthehemagglutinin glycoproteins
nor at the stageoftheirincorporation into the
cellsurface.
Our data suggest that at 25 C the
carbohy-drate-free envelope protein M is made insuch
small amounts that it cannot be detected on
polyacrylamide gels as a distinct peak in the polypeptide profile. Evenat 37Cthis proteinis
found in the infected cell in relatively small
amounts compared to those in thevirion, and
therefore it hasbeen suggested that the
synthe-sis ofproteinM mightbetightlycontrolled and
3H dpm
on November 10, 2019 by guest
http://jvi.asm.org/
INFLUENZA VIRUS PROTEINS
arate-limiting stepin virusreplication (13, 15). These findingsarecompatible with the
hypoth-esis that at 25 C the synthesis of protein M is reduced to a rate which no longer permits proper envelope assembly and, therefore, virus maturation.
ACKNOWLEDMENT
We thankElfriede Otto and Michaela Orlich for excellent technical assistence. This workwassupported by
Sonderfor-schungsbereich47(Virologie).
LITERATURECITED
1. Caliguiri, L. A., H.-D. Klenk, and P. W. Choppin. 1969. The proteinsof the parainfluenza virus SV5. I. Separa-tion of virionpolypeptides by polyacrylamide gel elec-trophoresis. Virology 39:460-466.
2. Compans, R. W., H.-D.Klenk, L. A.Caliguiri, and P. W. Choppin. 1970.Influenza virus proteins. I. Analysis of polypeptides of the virion and identification ofspike glycoproteins. Virology42:880-889.
3. Davenport, F. M., R. Rott, and W. Schafer. 1960. Physical and biological properties of influenza virus components obtained after ethertreatment. J. Exp. Med. 112:765.
4. Dimmock, N. J., and D. H. Watson. 1969. Proteins specified by influenza virus in infected cells: analysis by polyacrylamide gel electrophoresis of antigensnot presentin the virus particle. J. Gen. Virol.5:499-509. 5. Dulbecco, R., and M. Vogt. 1954.Plaqueformationand isolation of pure lines with poliomyelitis viruses. J. Exp. Med.99:167-182.
6. Eagle, H., and K. Habel. 1965.The nutritional require-mentsfor the propagation ofpoliomyelitis virus by the HeLa cell. J. Exp. Med.104:271.
7. Gregoriades, A. 1972. Isolation of neuraminidase from the WSN strain of influenzavirus.Virology49:333-336. 8. Haslam, E. A., A. W. Hampson, J. A. Egan, and D.0.
White. 1970. Thepolypeptides of influenza virus. II. Interpretation of polyacrylamide gel electrophoresis patterns.Virology42:555-565.
9. Jacobson, M. F., J. Asso, and D. Baltimore.1970.Further evidence on the formation of poliovirus proteins. J.
Mol. Biol. 49:657-699.
10. Joss, A., S. S. Gandhi, A. J. Hay, and D. C. Burke. 1969. Ribonucleic acid andprotein synthesis in chickembryo cells infected with fowl plague virus. J. Virol. 4:816-822.
11. Klenk, H.-D., L. A. Caliguiri,and P. W.Choppin.1970. The proteins of the parainfluenza virus SV5.II. The carbohydrate contentandglycoproteins of the virion. Virology 42:473-481.
12. Klenk, H.-D., R. Rott, and H. Becht. 1972. On the structure of the influenza virus envelope. Virology 47:579-597.
13. Klenk, H.-D., C. Scholtissek, andR. Rott. 1972. Inhibi-tion ofglycoprotein biosynthesisof influenza virusby D-glucosamine and 2-deoxy-D-glucose. Virology 49:723-734.
14. Laver, W. G. 1971. Separationof twopolypeptidechains from the hemagglutinin subunit of influenza virus. Virology 45:275-288.
15. Lazarowitz, S. G.,R. W.Compans,and P. W.Choppin. 1971. Influenza virus structural and nonstructural proteins in infected cells and theirplasmamembranes. Virology 46:830-843.
16. Pons, M. W., I. T. Schulze, and G. K. Hirst. 1969.
Isolation and characterizationof theribonucleoprotein ofinfluenza virus. Virology 39:250-259.
17. Rott, R., H. Becht, H.-D. Klenk, and C. Scholtissek. 1972. Interactions of concanavalin A with the
mem-brane of influenza virus infectedcells and with
enve-lopecomponentsof the virusparticle. Z. Naturforsch. 27:227-233.
18. Rott, R., and C. Scholtissek. 1968.Biochemical studies of influenza virus multiplication at reduced tempera-tures.J. Gen.Virol.3:239-245.
19. Scholtissek, C., and R. Rott. 1961. Zusammenhange zwischen derSynthesevonRibonucleinsaure and
Pro-tein bei der Vermehrung eines Virus der
Influenza-gruppe(Virus der klassischenGefluigelpest). Z. Natur-forsch. 16:663-673.
20. Schulze, I. T. 1970. Thestructureof influenza virus. I.
Thepolypeptides of the virion. Virology 42:890-904. 21. Schulze, I. T.1972.Thestructureofinfluenza virus. H. A
model based onthe morphology and composition of subviralparticles. Virology 47:181-196.
22. Skehel, J. J. 1972. Polypeptide synthesis in influenza virus-infected cells. Virology49:23-36.
23. Zimmermann, T., and W. Schifer. 1960. Effect of
p-fluorophenylalanine on fowl plague virus
multiplica-tion.Virology 11:676.
VOL. 11, 1973 831
on November 10, 2019 by guest
http://jvi.asm.org/

![FIG. 2.described60-min Polyacrylamide gel electrophoresis of FPV proteins. Cells were exposed to a pulse with [3H]leucine as in Fig](https://thumb-us.123doks.com/thumbv2/123dok_us/1594051.112165/4.493.92.407.50.455/described-polyacrylamide-electrophoresis-proteins-cells-exposed-pulse-leucine.webp)