0022-538X/87/051672-06$02.00/0
Copyright C) 1987, AmericanSocietyforMicrobiology
The
preSI
Protein of
Hepatitis B Virus Is Acylated at Its
Amino
Terminus with
Myristic
Acid
DAVID H.
PERSING,l
HAROLD E. VARMUS,"2 AND DONGANEM23*Departments
of
Microbiology
andImmunology,2 Biochemistry
andBiophysics,'
and Medicine,3 University of California MedicalCenter,
SanFrancisco,
California
94143Received 31 October1986/Accepted19January1987
The
preS/S
codingregion ofhepatitis
B virusencodestwopolypeptides
(preSl
andpreS2)
that arelarger
in size but less abundant than themajor
viral surfaceantigen (S) protein. Unlike the preS2 and S proteins, the preSl protein ispreferentially
localized oncirculating
virusparticles
but is not efficiently secreted from mammaliancels
in culture. To search for differences inprotein processing
thatmight
relate to these properties, we determined whether any of thehepatitis
B virus surfaceproteins
areacylated
withlong-chainfatty
acids. Transfected COS cells expressing all three proteins were incubated with 3H-palmitate or3H-myristate,
and the cell extractswereexaminedby
immunoprecipitation.
While none of theseproteins was labeled with3H-palmitate,
thepreSl protein
but not thepreS2 or S protein incorporated 3H-myristate via a hydroxylamine-resistant amide linkage.Comparison
oftheN-terminal amino acidsequences ofhepadnaviral
preSl proteins with those of known
myristylated proteins
suggests that this unusual modification may be a common feature of allhepadnaviral
preSl proteins.
Recent
studies of
hepatitis Bsurface
antigen (HBsAg)
purified from
the serumof infected
patients
orfrom
mam-malian
cellstransfected with viral DNA haveidentified
twoHBsAg-related species,
designated thepreSl
andpreS2
proteins,
inaddition
to the more abundantsurface antigen
(S)
protein
(12, 14, 23, 25). Thecoding region for
thepreS
and S
proteins comprises
asingle 1,182-base-pair
openread-ing frame with three translation initiation codons
thatdirects
the
production of
threeproteins with
a commoncarboxy
terminus: the
major viral S protein
(226amino acid
residues),
the
preS2
protein
(281amino acids),
and the380-amino-acid
preSl protein
thatis
the productof
theentire
openreading
frame (Fig.
1). Thedistribution of
thesepolypeptides
is notequivalent
among thecirculating forms of
HBsAg; thepreSlprotein is found in
higher abundancein viral particles
andfilaments
than inthe more numerous 22-nm subviralparti-cles, while
the converseis
true of the moreabundant preS2 andS
proteins
(12). These findings suggest that preSldeterminants
maybe important participants in virusassem-bly
orinfectivity
or both.In a
previous
report, we examined the expression of the S andpreSl proteins
by transfection of viralDNA
into cul-turedmammalian
cells and injection of synthetic mRNAsinto
Xenopus oocytes (25). The findings indicated that thepreSl
proteins, unlike their preS2 and S protein counter-parts, are not secreted into the culture medium despite the presenceof
the secretory information contained in the S-specific domains. Furthermore, when the S andpreSl
proteins
aresynthesized together, secretion of the S proteins is strongly andspecifically inhibited. This suggests that someelement
of thepreSl
proteins, whether alone or in a mixed aggregate of S and preS protomers, is inhibitory to thesecretion
of HBsAg polypeptides.In
seeking
differences in protein processing that mightexplain
the unexpected secretory properties of thepreSl
proteins,
we examined the preS and S proteins for the presence of long-chain fatty acids, a knownposttranslational modification of viral envelope glycoproteins. To obtain*
Corresponding
author.expression
of the S andpreS proteins
in mammaliancells,
we
constructed
plasmid pSV45H (25),
which contains thecontiguous
preS/S coding region
downstream of the simianvirus
40(SV40)
early
promoter(Fig. 1A).
Thisplasmid
was then used to transfectCOS7 cells. At 48 h followingtrans-fection, the medium
wasremoved from
the transfected cells andreplaced
with medium containing3H-palmitate,
3H-myristate,
or35S-methionine.
35S-methionine
labeling
wasperformed
inaccordance
withapreviously
described proce-dure(25).
For3H-myristate
and3H-palmitate labeling,
the cells werelabeled
with 1mCi of
theappropriate fatty
acid per mlfor
20h(2),
andcell extracts were then examinedby
immunoprecipitation with
antibody
to HBsAg (25). Asex-pected,
35S-methionine-labeled
cells transfected withplas-mid
pSV45H produced, in
addition
to the Sproteins of
24and
27kilodaltons
(kDa), both
sets ofpreS-encoded
polypeptides: the preSl proteins of
39 and 42kDa and the
preS2
proteins of
31, 33, and 36 kDa(Fig.
1B, lane M). (In each case, the mostrapidly migrating band corresponds
to theunglycosylated form,
and theslower speciescorresponds
tothe
glycosylated
form(s) of
theindicated protein.) Noneof
the S
protein species
incorporated 3H-palmitic acid; labeled
extracts
from
aduplicate
set of transfected cells (Fig. 1B, lanes 5 and 6) as well as cell extracts from untransfected cells(Fig.
1B, lane 4)yielded
no immunoprecipitable species.However,
when3H-myristate
wasused for labeling, a dou-bletmigrating
in the position of thepreSl
proteins was observed(Fig.
1B, lanes 2 and 3). Despite the contempora-neousproduction
of the preS2 and S proteins in these cells,only
thepreSl proteins became
labeled in the presence of3H-myristate.
To test whetherthe inability todetect a palmitate-labeled
species
was dueto inefficient labeling with this fatty acid, weexamined
the unprecipitated cell extracts from palmitate-andmyristate-labeled
cells. In the presence of3H-palmitate,
many labeled protein species wereobserved (Fig.
1C,
lanes 4 to6).
Bycontrast,3H-myristate
efficiently labeled only a fewcellularproteins (Fig. 1C, lanes 1 to 3), in keeping withobservations
in othersystems (15).1672
on November 10, 2019 by guest
http://jvi.asm.org/
A.
preSi preS
SV40 promoter _ =
P H Bst
B.
Myr
s
Bg
C.
Pal
Myr
Pal
I
I-Fr "I 1
X.-A4
68
preSlC1
preS2:
_0<-_-. 4
M 1 2 3 4 5 6
Immune
precipitates
45
26
1 2
3
4 5 6Total
protein
FIG. 1. (A) Expression of the HBV preS and S proteins from anSV40vector. The construction of plasmid pSV45H has been described elsewhere (25). Shown is the 342-base-pairPvuII(P)-HindIII (H) fragment ofSV40,containing theSV40replication origin and theSV40early promoterinserted upstream of the HBV preS/S coding region. The asterisks indicate the approximate transcription initiation sites of the HBV preS2/S promoter. The arrows indicate the probable sites of translation initiation for the preSl, preS2, and S gene products. (B) Immunoprecipitation of35S-methionine-, 3H-myristate-,and3H-palmitate-labeled cell extracts from HBsAg-producing COS cells. Plates (60 mm)of COS7 cells were transfected with pSV45H in the presence of DEAE dextran byaprocedure previously described (8) and labeled with 35-methionine,
3H-myristate,
or 3H-palmitate. Cell lysates were prepared and immunoprecipitated as described elsewhere (25). Im-munoprecipitated samples were subjected to sodium dodecylsulfate-polyacrylamide gel electrophoresis (SDS-PAGE) on a 12% polyacryl-amidegel. The gel was then fixed, fluorographed, dried, and exposed to film for 6 days. Shown are immunoprecipitations of 35S-methionine-labeled transfected cells (lane M), transfected (lanes 2 and 3) and untransfected (lane 1) cells 35S-methionine-labeled with3H-myristate,andtransfected (lanes 5 and6) and untransfected (lane 4) cells labeled with 3H-palmitate. (C) SDS-PAGE ofHBsAg-producing COS cell extracts labeled with3H-myristate
or3H-palmitate.A4-pl
quantity of each of the3H-labeledcell extracts used in the above-describedimmunoprecipitations(0.8% of the immunoprecipitated volume) was subjected to SDS-PAGE on a 12% polyacrylamide gel. The gel was then fixed, fluorographed, dried, andexposedtofilmfor3days.The sources of the unprecipitated cell extracts shown in lanes 1 to 6 are as in panel B. The numbers between panels B and C represent kilodaltons.The
inhibitory
propertyof
preSlprotein
expression
on Sprotein secretion has recently
beendocumented
in a varietyof
experimental
systems(4,
5, 25, 32).
Since we andothers
have observed
theinhibitory effects of preSl protein
expres-sion
onS
protein secretion in
Xenopus oocytes, we wereinterested in whether myristylation of
thepreSl protein
occurs
in this
system(25, 32). Plasmids containing
the S orpreS/S
coding region downstream from
theSP6
promoter wereconstructed, and synthetic
mRNAs weresynthesized
from
theappropriate template by
invitro
transcription with
SP6
polymerase. Injection of
Xenopus oocytes was carried out aspreviously described (24, 25;
K.Simon,
E. Perrara,and
V.Lingappa, submitted for
publication). Briefly,
a40-nlvolume containing
60 to 120 ngof
transcript
intranscription
buffer
wasinjected into each of
15freshly dissected
oocytes,after which the
transcript
wasallowed
toequilibrate
for 12 h.The medium
wasthenremoved and
replaced
withmedium
containing
5mCiof
35S-methionine
or2mCi of3H-myristate
per
ml, followed by incubation
at18°C
for 24 h. Cellhomogenates
wereprepared
asdescribed
previously (25)
and spunin
anEppendorf centrifuge
for
10min,
and theresulting
supernatants were removed for
immunoprecipitation
withantibody
toHBsAg
(Calbiochem-Behring).
As inmammaliancells,
thepreSl
proteins,
butnottheSproteins,
werelabeledwith
3H-myristic
acid,
whether or not S mRNA wascoinjected (Fig.
2,
lanes 1 and3,
and datanotshown).
The
failure of
S andpreS2 proteins
to belabeled with
3H-myristate
suggested
that thepreSl domain itself
wasthesite
of
fatty acid addition.
Tofurther
examine
thispossibil-ity,
weconstructed
afusion
geneencoding
ahybrid
protein
consisting of
thefirst
110amino acids of the
preSl
protein
fused
to the Nterminus
of
chimpanzee a-globin,
anonmyristylated
cytoplasmic protein (22).
Invitro
tran-scripts encoding
thisprotein
wereinjected
into Xenopus oocytes in the presence of3H-myristate,
and thelabeled
translation
products
wereimmunoprecipitated
withanti-globin
antiserum(Calbiochem).
Aspredicted,
thehybrid
protein
waslabeled
efficiently
with3H-myristate
(Fig.
2,
lane 4). Thisresult
indicates that theC-terminal
278-amino-acid
residues
of
thepreSl
protein, including
S andnearly
allof
the
preS2 domains,
aredispensable
for
acylation.
It
is
interesting
to notethat
themyristate-labeled
hybrid
protein
appeared
asadoublet
of
22and
26 kDa. Thereason forthepresence
of themultiple
species
ofthefusionprotein
is unclear. The
preSl
region
of
the aywhepatitis
B virus(HBV)
subtype
used in theseexperiments
contains
twoin-phase
ATGcodons
separated by
10codons;
independent
initiation
ateach ATG is thusunlikely
to account for theobserved 3-kDa
difference between the twospecies.
Alter-natively,
thebandscould
resultfrom
posttranslational
proc-essing
events(e.g.,
proteolysis,
glycosylation,
orothermod-ifications).
Theshift in molecular
weight
isprobably
notdueENE
mmmmmmm.-r32
on November 10, 2019 by guest
http://jvi.asm.org/
[image:2.612.126.503.72.308.2]tothe presence of the high-mannose oligosaccharide moiety; under conditions ofendoglycosidase H digestion, in which theglycosylated forms of the S andpreS proteinsare fully sensitive, the slower-migrating form of the hybrid
protein
remains resistant (D. H. Persing, unpublishedobservation). Further studies are in progress to better characterize the modifications that may account for the apparent shift in mobility.
Tobe certain that the labeled moietyon the
preSl
protein was not derived from the conversion ofmyristateto palmi-tate or another novel metabolite, we performed acid hy-drolysis on preparations of gel-purified proteins. Poly-acrylamide gel slices containing 3H-labeled proteins were obtained fromimmunoprecipitation
of3H-myristate-labeled COS cells transfected withpSP45H (recovered from atotal of fourimmunoprecipitations), 3H-myristate-labeled
oocytesinjected
withsynthetic
preSl
mRNA(two immunoprecipita-tions), and3H-myristate-labeled
oocytes injected withsyn-thetic
mRNAencoding
thepreSl-globin hybrid protein
(oneimmunoprecipitation).
The slices wererehydrated, placed
in 50 ml of 100% dimethyl sulfoxide for 2 h to remove thediphenoxazole fluor,
and then incubated in 50mlofdistilled waterfor 1 h to remove thedimethyl sulfoxide.
The slicesA.
Direction of Migration - o
preS1- 2 L3
Std.
B.
0
C:
0
C)
C3)
c3
C-C)
9
KMyristate
Palnmitate
100 1
75 I -i
50 -!
25 l
m 4 HAcleavage LZ - HAcleavage
1 2 3 4
preSil
-68
-45
-26
FIG. 2. Production of3H-myristate-labeled preSl proteins and preSl-globinfusion proteins by injection of syntheticmRNAsinto Xenopus oocytes. The construction of plasmids pSP24H and pSP45H (not shown) has been described elsewhere (25). To
con-structplasmid pSP45GLO (also not shown), wefirst inserted the
2.8-kilobaseBglII fragment ofHBVintothe BamHIsite of pSP125E (a plasmid containing the chimpanzee a-globincodingregion
down-streamof theSalmonella phageSP6 promoter)and then screened resultant clones for orientation by restriction analysis. A clone bearing thepreSl region in the correct transcriptional orientation
wasdigested withNcoI and treated briefly with Bal3l to provide bluntends in allthreepossiblereading framesnearthe 5'end of the
at-globin coding region. Next, the exonuclease-treated clone was
digested with EcoRI to remove most of the HBV sequences
up-stream of the a-globin coding region, with the exception of the coding regions for preSl and the first few nucleotidesof preS2. The preSl regionwasthen fusedtoglobinsequencesby theaddition of
T4DNAligase. preS and S proteinswereproduced by injection of
synthetic mRNAs ofeach polypeptide intoXenopus oocytes (25;
Simon et al., submitted) in the presence of 35S-methionine or
3H-myristate and detected by immunoprecipitation. Shown are
immunoprecipitates of 35S-methionine-labeled oocytes coinjected
with equal amounts of S- and preSl-specific mRNAs (lane 1),
uninjectedoocyteslabeled with3H-myristate (lane2),
3H-myristate-labeled oocytes coinjected with equal amounts ofS- and preSl-specificmRNAs(lane 3), and ananti-globin immunoprecipitate of
3H-myristate-labeledoocytesinjectedwith RNA encodinga
preSl-globin hybrid protein (lane4). Samples were subjectedto sodium
dodecyl sulfate-polyacrylamide gel electrophoresis on a12%
poly-acrylamide gel, which was then fixed, fluorographed, dried, and
exposed to film for 4 days. The numbers at the right represent
kilodaltons.
HBV preSl VSV G
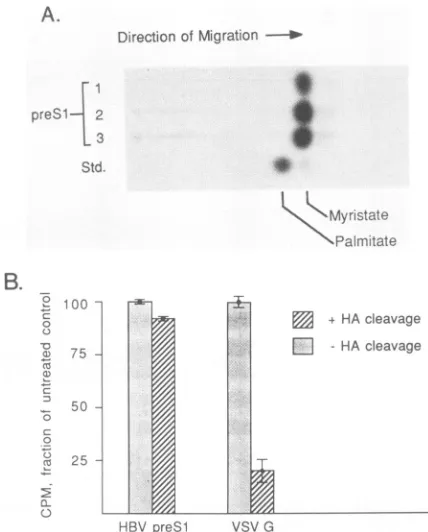
FIG. 3. Confirmation of the labeled moietyas an amide-linked myristic acid residue. (A) Thin-layerchromatographic analysis of acid hydrolysates of preSl proteins. Gel slices containing the electrophoretically resolved, 3H-myristate-labeled preSl proteins produced in COS cells and Xenopus oocytes wererehydrated and suspended in 6 N HCl. After incubation in 110°C for 24 h, the liberated products wereextracted intononpolarsolvents and ana-lyzed byreversed-phasethin-layer chromatography. Shownarethe chromatographed acid hydrolysis productsof3H-labeledgelslices from oocytesinjected withsynthetic preSlmRNA(lane 1), oocytes injected withRNAencodingthepreSl-globinfusionprotein(lane 2), andpreSl proteins precipitatedfromHBsAg-producing COS cells (lane 3). Lane std. contains the authenticpalmitate and myristate standards. (B) Hydroxylamine (HA) sensitivity of the preSl-myristate linkage. Two identical polyacrylamide gels containing 3H-myristate-labeledHBVpreSl proteinsaswellasVSV Gprotein labeled with 3H-palmitate were run. One gel was fixed and fluorographedimmediately after electrophoresis, and the otherwas soaked for 1 h in 1 M hydroxylamine (pH 8) fixed, and fluoro-graphed. The gelswere exposedtopreflashed film for4days, and the3H-labeledprotein bandswereexcisedfrom thegel and counted in a liquid scintillation counter(36). The results are shownin the histogram as a percentage ofresults for the untreated control; 2 standard deviations are indicated at the top of each bar. Actual counts per minute (+2 standard deviations) obtained from the sampleswere asfollows:7,189±47for untreatedpreSl; 6,988±44 forhydroxylamine-treated preSl; 1,102± 33for untreatedVSV G protein; and222 ± 37for hydroxylamine-treated VSV G protein.
werethen
incubated
in 0.3 ml of 6 NHCl at 110°C for 24 h. Both theacid hydrolysate
and the polyacrylamide pellet were extracted three times with 0.5 ml of toluene, and reversed-phasethin-layer chromatography was performed as describedby
Buss and Sefton (2).Figure
3 shows the results of this analysis. Authenticmyristic
acid runs ahead of palmitic acid on thin-layerchromatograms
because of its shorter carbon-chain length. Theacid
hydrolysis products from purifiedpreSl
proteinsproduced
in COS cells or oocytes are shown in Fig. 3A, lanes 1 to 3. All of the acid cleavage productsmigrated with themobility
expectedfor
myristic acid. Small amounts of aon November 10, 2019 by guest
http://jvi.asm.org/
[image:3.612.325.539.66.332.2] [image:3.612.128.227.314.429.2]faster-migrating
species may have been an artifact ofhydro-lysis
or a 12-carbon metabolite of the labeled myristic acid.To
assess the nature of the covalent bond between thepreSl
protein and3H-myristic
acid, we examined thehydroxylamine
sensitivity of the linkage. Whereas3H-palmitate
andother
acyl groups are typically bound topolypeptide
backbones via an ester linkage,3H-myristate
isusually
bound via anamide
bond. Because hydroxylamineefficiently
cleaves ester and thioester linkages but leavesamide bonds intact,
treatment with this agent distinguishesbetween the
two chemically distinct types of linkages.3H-myristate-labeled preSl
proteins were immunoprecipitatedfrom HBsAg-producing
COS cells as described above. For use as a positive control,3H-palmitate-labeled
vesicularstomatitis virus (VSV)
G protein was produced by infectingCOS cells with
20 PFUof VSV
Indiana (a generous gift fromJudy White)
per cell. The3H-palmitate-labeled
VSV Gprotein and
the3H-myristate-labeled
HBVpreSl
proteins were run in adjacent wells of identical 12% polyacrylamidegels. After electrophoresis,
onegel
was treatedfor
1 h at20°C with
1 M hydroxylamine (pH 6.8), fixed, andfluorographed
(15);
the other
wasfixed and fluorographed
immediately for
use as a control(Fig.
3B). Whereas there was nosignificant
decreasein
the level of label associatedwith
thepreSl proteins following hydroxylamine
treatment,the label bound
tothe VSV G
protein
decreasedfourfold
following
such
treatment. Therelative resistance of the
preSl protein
tohydroxylamine
treatment suggests that3H-myristic
acid is bound
tothis
protein
via
ahydroxyl-amine-resistant amide
link,
asfound
for other
myristylated
proteins
(15).
A
small
number of cellular and viral
proteins
areknown
tobe
modified by
myristate
addition;
modification
usually
occursat
the
Nterminus of
theprotein (1, 2, 3, 13, 17, 27).
Comparison
of
theN-terminal
amino
acid
sequencesof these
proteins
reveals that the N-terminal
methionine
residue is
invariably
followed
by
aglycine (Fig. 4A).
Following
cleav-age
of the
methionine,
anamide
linkage
is
formed between
the
COOH
groupof
myristic
acid and the
exposed
amino
group
of the
glycine.
Examination of
theN-terminal residues
predicted by
the mammalian and avian
preS
openreading
frames reveals
asimilar
sequencemotif
(Fig.
4B).
The
conservation
of this
structure acrossconsiderable
evolution-ary
distances
suggests thatpreSl
myristylation
is
likely
tobe a commonfeature
of
thehepadnaviruses.
The
preSl
proteins
differ
from
previously
described
myristylated
proteins
in
oneimportant
respect.Other
myristylated
proteins
arethought
tobe
synthesized
onfreepolyribosomes
in
thecytoplasm;
in
onewell-studied
case,myristate
addition
tothe
Nterminus of
theviral
transform-ing
protein
pp60Osc
has been shown
tobe
important
in thesubsequent
affiliation
of the
protein
with the inner(cytoplas-mic)
face
of the
plasma
membrane
(6,
7). By
contrast,
thepreSl
proteins,
like their S
protein
counterparts,
areinitially
synthesized
astransmembrane
glycoproteins
spanning
themembrane
of the
endoplasmic
reticulum
(B.
Eble,
V.Lingappa,
and
D.Ganem,
unpublished data).
We presumethat,
as in other cases,myristylation
occurswhile
thegrowing
polypeptide
is
in thecytoplasm.
The
presenceof
N-terminal
myristate
does
not appear toprevent
themem-brane
translocation of
downstreamprotein domains,
sinceglycosylation
of
the Sdomain of
thepreSl
protein
(12;
K.Simon,
V.Lingappa,
and D.Ganem,
manuscript
in
prepa-ration)
proceeds
withnormal
efficiency.
What
arethe
biological
roles of
thepreSl
proteins,
and
how
might
myristylation
influence
them? Recentfindings
from this
(23, 24;Simon et al., in preparation) and other (12, 19, 20)laboratories
have provided new insights into the structure of thepreSl
proteins and their disposition in HBsAg particles. Studies by Heerman et al. (12) indicatethat
preSl polypeptides are preferentially localized on virions. At least somepreSl-specific
epitopes areimmuno-genic
(19, 24) and are exposed on the virion surface (12). Inaddition,
a rolefor
someof
these determinants in host cellbinding
hasrecently been suggested (20). Interestingly,
however, the
extreme Nterminus
of the protein is notreactive with specific antipeptide
antisera (20; D. Persing,unpublished data),
afinding
that could result either frominterference by the acyl
groupitself
orfrom
aresulting
insertion of
the Nterminus into
the lipid of the envelope.Such
insertion could be important for the
correctdisposition
of
theexposed
regions of
themolecule.
Like the surface glycoproteins of
mostenveloped viruses
but
unlike
the Sproteins,
thepreSl proteins
are notsecreted
from the cell
(4, 5, 25, 32).Like other envelope proteins,
they
areonly exported from cells
as partof
avirion
(al-though, unlike
theseproteins, their intracellular
locale isprimarily
theendoplasmic reticulum
ratherthan
theplasma
membrane).
Theseobservations
areconsistent
with arole
for
thepreSl
proteins
in
virion
assembly.
For mostbudding
viruses, interactions between
thecytoplasmic domains of
envelope
proteins and
theirnucleocapsid
ormatrixproteins
are
believed
toplay
arole in the
targeting of virion
compo-nents tothe
appropriate membrane for
morphogenesis
and
perhaps in
thetriggering of the budding
eventitself
(26, 31,
37). Membrane
interactions mediated
by
N-terminal
myristic
acid could be
important
in
anchoring
the Nterminus
of
thepreSl
protein in
themembrane of the
endoplasmic
reticulum
A.
Known
myristylated
proteins
MLV
pl5g9
pp60v-src
PK-A
pp56lck
CALCINEURIN B
iaQTVTTPLSLTLGHW
IESSKSKPKDPSQRRR
| NAAAKKGSEQSVK
NCVCSSNPEDDWME
iNEASYPLE
B.
Hepadnavirus preSl
proteins
GSHV WHV DHBV HBV
(ayw)
HBV(adw2)
MNNIKVTFDPNK
FINNIKVTFNPDK
EMHPALSMDVR
EQNLSTSNPLG
FMGGTSSLPALG
3FIG. 4. N-terminal amino acidsequence
comparisons
of knownmyristylated proteins
(A)
andhepadnavirus
preSl proteins
(B).The N-terminalmethionine-glycine
motif,
thought
tobeamyristylation
targetsequence, is boxed. Abbreviationsin
panel
A: MLVp159'9,
the15-kDagagprotein
of murine leukemiavirus(13, 29);
pp60v-src,
the
transforming
protein
ofRSV(Schmidt-Ruppin
A)
(2,
22); PK-A, thecatalytic
subunit ofcyclic
AMPprotein
kinase(3, 30);
pp56lck,
the56-kDatyrosine
kinase foundinLSTRA T-celllymphoma
cells(17, 18);
and calcineurin B, thecalcium-binding
B subunit of calcineurin(1).
Abbreviations inpanel
B: GSHV,ground squirrel
hepatitis
virus(28);
WHV,
woodchuckhepatitis
virus(10);DHBV, duckHBV(16);
HBV(ayw),
theaywsubtype
of human HBV(11); andHBV(adw2),
theadw2subtype
ofhuman HBV(35).
on November 10, 2019 by guest
http://jvi.asm.org/
[image:4.612.356.528.412.603.2](thereby preventing
its spontaneoussecretion)
orinorient-ing
thepreSl domain
to allowinteractions
with other viralcomponents.
Furtherstudies
will benecessarytoexplore
the roleof
thismodification
in the structure,assembly, and
infectivity
of
hepadnaviruses.
We thank K. Simon, V. Lingappa, and J. Buss for
helpful
discussions and J.Marinos for excellent manuscript preparation.This work was supported by Public Health Service grant Al 18782 from the NationalInstitutes of Health, by Medical ScientistTraining Program award GM07618 (to D.H.P.), by an American Cancer Societyprofessorship(to H.E.V.), andbyaJohnL.andGeorgeH. HartfordFoundation fellowship (toD.G.).
LITERATURECITED
1. Aitken, A., P. Cohen, S. Santikarn, D. H. Williams, A. G. Calder, A. Smith, and C. B. Klee. 1982. Identification ofthe NH2-terminalblocking group of calcineurin B as myristic acid. FEBS Lett. 150:314-318.
2. Buss, J. E., and B. M. Sefton. 1985. Myristic acid, a rarefatty acid, isthelipid attached to the transforming protein ofRous sarcomavirus and its cellular homolog. J. Virol. 53:7-12. 3. Carr, S. A., K. Biemann, S. Shoji, D. C. Parmelee, and K.
Titani. 1982. n-Tetradecanoyl is the NH2-terminal blocking group of the catalyticsubunit ofcyclic AMP-dependentprotein kinase frombovine cardiac muscle. Proc. Natl. Acad. Sci. USA 79:6128-6131.
4. Cheng,K.C., G.L.Smith,and B. Moss. 1986. Hepatitis B large surfaceprotein is not secreted but is immunogenic when selec-tively expressed by recombinant vaccinia virus. J. Virol. 60:337-344.
5. Chisari,F. V., P. Filipi, A.MacLachlan, D. Milich, M. Riggs, S. Lee, R. D. Palmiter, C. A. Pinkert, and R. L. Brinster. 1986. Expression of hepatitis B virus large envelope polypeptide inhibits hepatitis B surface antigen secretion in transgenic mice. J. Virol.60:880-887.
6. Courtneidge, S. A., A. D. Levinson, and J. M. Bishop. 1980. The protein encoded by the transforming gene of avian sarcoma virus (pp60src) and a homologous protein in normal cells
(pp60Pr,-osrc)
are associated with the plasma membrane. Proc. Natl. Acad. Sci. USA77:3783-3787.7. Cross,F. R., E. A.Garber, D. Pellman, and H. Hanafusa. 1984. A short sequence in the pp6Osrc N terminus is required for
pp6Osrc
myristylation and membrane association and for cell transformation. Mol. Cell. Biol. 4:1834-1842.8. Crowley, C. W.,C.-C. Liu, and A. D. Levinson. 1983. Plasmid-directed synthesis of hepatitis B surface antigen in monkey cells. Mol. Cell. Biol. 3:44-55.
9. Eble, B. E., V. R.Lingappa, and D. Ganem. 1986. Hepatitis B surface antigen: an unusual secreted protein initially synthe-sized as a transmembrane polypeptide. Mol. Cell. Biol. 6:1454-1463.
10. Galibert, F., T. N. Chen, and E. Mandart. 1982. Nucleotide sequence of a clonedwoodchuck hepatitis virus genome: com-parisonwith thehepatitis Bvirus sequence. J. Virol. 41:51-65. 11. Galibert, F., E. Mandart, F. Fitoussi, P. Tiollais, and P. Charnay. 1979. Nucleotide sequence of hepatitis B virus genome (subtype ayw) cloned in E. coli. Nature (London) 281:646-650.
12. Heermann,K. H., U.Goldmann, W. Schwartz, T. Seyffarth, H. Baumgarten, and W. H. Gerlich. 1984.Large surface proteins of hepatitis B virus containing the pre-S sequence. J. Virol. 52:396402.
13. Henderson, L. E., H. C. Krutzsch, and S. Oroszlan. 1983. Myristylaminoterminal
acylation
ofmurine retroviral proteins:anunusual post-translational proteinmodification. Proc. Natl. Acad. Sci. USA80:339-343.
14. Machida, A., S. Kishimoto, H. Ohumura, H. Mliyamoto, K. Baba, K. Oda, T. Nakamura, and Y. Miyakawa. 1983. A hepatitisB surface antigenpolypeptide (P31) with the receptor forpolymerized human as well aschimpanzee albumins.
Gas-troenterology85:268-274.
15. Magee,A.I.,and S. A.Courtneidge. 1985. Twoclassesof
fatty
acylated proteins exist in
eukaryotic
cells. EMBO J. 4:1137-1144.16. Mandart, E., A. Kay, and F. Galibert. 1984. Nucleotide
se-quenceofacloned duck
hepatitis
Bvirus genome:comparison
with woodchuck and human hepatitis B virus sequences. J. Virol. 49:782-792.
17. Marchidon, G. A., J. E.
Casnellie,
K. A. Walsh, and E. G. Krebs. 1984.Covalently
boundmyristic
acid in alymphoma
tyrosine protein kinase. Proc.
Natl.
Acad. Sci. USA 81:7679-7682.18. Marth, J. D., R.Peet,E.G.Krebs,and R. M.
Pearimutter.
1985. A lymphocyte-specific tyrosine kinase gene isrearranged
and overexpressed in the murine T celllymphoma
LSTRA. Cell 43:393-404.19.
Milich,
D. R., G. B. Thornton, A. R. Neurath, S. B. Kent, P.Tiollais, and F. V. Chisari. 1985. Enhanced
immunogenicity
of the pre-S region of hepatitis B surfaceantigen.
Science 228:1195-1198.20. Neurath, A. R.,S. B. H. Kent, N. Strick, and K. Parker. 1986. Identification and chemical synthesis of a host cell receptor binding siteon hepatitis B virus. Cell 46:429-436.
21. Patzer, E. J., G.
R.
Nakamura, C. C.Simonsen,
A. D.Levinson, and R. Brands. 1986. Intracellular assembly and packaging of hepatitis B surface antigen particles occur in theendoplasmic
reticulum. J. Virol. 58:884-892.22.
Peliman,
C., E. A. Garber,F. R. Cross, andH.Hanafusa.1985. AnN-terminalpeptide frompp6Osrc
candirectmyristylation
and plasma membrane localization when fused to heterologous proteins. Nature (London) 314:374-376.23. Persing,D.H.,H.E. Varmus,and D.Ganem. 1985.Aframeshift mutation in the pre-S region of the human hepatitis B virus genome allows production of surface antigen particles but eliminates binding to polymerized albumin. Proc.
Natl.
Acad. Sci. USA 82:3440-3444.24. Persing,D.H.,H.E. Varmus, and D. Ganem. 1986. Antibodies to pre-S and X determinants arise during natural infectionwith ground squirrel hepatitis virus. J. Virol. 60:177-184.
25. Persing, D. H., H. E. Varmus, and D. Ganem. 1986. Inhibitionof secretionof hepatitis B surface antigen by a relatedpresurface polypeptide. Science 234:1388-1392.
26. Rodriguez Boulan, E., and D. D. Sabatini. 1978. Asymmetric budding of viruses into epithelial monolayers: a model system for study of epithelial polarity. Proc.
Natl.
Acad. Sci. USA 75:5071-5075.27. Schulz, A. M., L. E. Henderson, S. Oroszlan, E. A. Garber, and H. Hanafusa. 1985. Amino terminal myristylation of theprotein kinase
pp6osrc,
a retroviral transforming protein. Science 227:427-429.28.
Seeger,
C., D. Ganem, and H. E. Varmus. 1984. Nucleotide sequence of an infectious molecularly cloned genome of ground squirrel hepatitis virus. J. Virol. 51:367-375.29. Shinnick, T. M., R. A. Lerner, and J. G. Sutcliffe. 1981. Nucleotide sequence of Moloney murine leukemia virus. Nature (London) 293:543-548.
30. Shoji, S., D. C. Parmelee, R. D. Wade, S. Kumar, L. H. Ericsson, K. A. Walsh, H. Neurath, G. L. Long, J. G. DeMaille, E. H. Fischer, and K. Titani. 1981. Complete amino acid se-quence of the catalytic subunit of bovine cardiac muscle cyclic AMP-dependent protein kinase. Proc. Natl. Acad. Sci. USA 78:848-851.
31. Simons, K., H. Garoff, and A. Helenius. 1982. How an animal virus gets into and out of its host cell. Sci. Am. 246:50-66. 32. Standring, D. N., J. S. Ou, and W. J. Rutter. 1986. Assembly of
viral particles in Xenopus oocytes: pre-surface-antigens regulate secretion of the hepatitis B viral surface envelope particle. Proc. Natl. Acad. Sci. USA 83:9338-9342.
33. Stibbe, W., and W. Gerlich. .1983. Structural
relationsbips
between minor and major proteins of hepatitis Bsurfacei
anti-gen. J. Virol. 46:626-628.34. Tiollais, P., A. Dejean, C. Brechot, M. Michel, P.
Sonigo,iand
S. Wain-Hobson. 1984. Structure of hepatitis B virus,-DNA,
p.on November 10, 2019 by guest
http://jvi.asm.org/
49-65.InG.N.Vyas, J. L.Dienstag, andJ. H.Hoofnagle(ed.), Viral hepatitis and liver disease. Grune & Stratton, Inc.,
Orlando, Fla.
35. Valenzuela, P., P. Gray, M. Quiroga, J. Zakdivar, H. M. Goodman,and W.J. Rutter. 1979. Nucleotidesequenceof the
genecoding for the major protein of hepatitis B virus surface
antigen. Nature (London) 28Q:815-819.
36. Walter,P.W.,R.C.Jackson, M. M. Marcus, V. R. Lingappa, and G. Blobel, 1979. Tryptic dissection and reconstitution of translocation activity fornascentpresecretory proteinsacross
microsomal membranes. Proc. Natl. Acad. Sci. USA 79:1795-1799.
37. Wiley, D. 1985. Viral membranes, p. 45-68. In B. N. Fields (ed.),Virology. Raven Press, Publishers, New York.