JOURNAL OF VIROLOGY, Sept.1979,p.718-732 Vol.31, No. 3 0022-538X/79/09-0718/15$02.00/0
Structural
Changes
in
Simian Virus
40
Chromatin
as
Probed
by Restriction Endonucleases
GEORGE L. LIGGINS,t MICHELE ENGLISH,4 ANDDAVTD A. GOLDSTEIN§*
Receivedforpublication13June1979
The structure of simian virus 40 (SV40) chromatin wasprobed by treatment withsingle-andmultiple-sitebacterial restriction endonucleases. Approximately the same fraction of the chromatin DNAwascleavedbyeach of threedifferent single-site endonucleases, indicating that the nucleosomes do not have unique positions with regard tospecific nucleotide sequences within the population of chromatin molecules. However, theextentofdigestionwasfoundtobestrongly influenced by saltconcentration. At 100 mMNaCl-5mM
MgCl2,
only
about 20% of the simian virus 40 (SV40) DNA I in chromatinwasconvertedtolinearSV40 DNA III. In contrast, at lower concentrations of NaCl (0.05 or 0.01M),
an additional20 to30% of the DNA wascleaved.Theseresults suggest thatat100 mMNaClonly theDNA betweennucleosomeswasaccessibletotherestriction enzymes,whereas atthelower saltconcentrations, DNA within thenucleosome regions becameavailable for cleavage. Surprisingly, whenSV40 chromatin was digested with multiple-site restriction enzymes, less than 2% of the DNA wasdigested to limit digest fragments, whereas only a small fraction (9 to 15%) receivedtwo or more cuts. Instead, theprincipaldigestfragmentwasfull-length linear SV40DNA III. The failureto generatelimit digestfragmentswas not a consequence ofreduced enzymeactivity in thereaction mixtures orofhistone exchange. When the position of the principal cleavage site was mapped after HpaIdigestion,it wasfound that this sitewas notunique.
Nevertheless,
allsites were not cleaved with equal probability. An additional finding was that SV40 chromatin containing nicked-circular DNA II produced by random nicking of DNA I was also resistant to digestion by restriction enzymes. These results suggest thatthe initial cut which causes relaxation of topological constraint in SV40 chromatin DNA imparts resistance to further digestion by restriction enzymes. We proposethatthismaybeaccomplished byeither"winding" ofthe internucleosomal DNA into the body of the nucleosome, or as suggested by others,bysuccessiveright-hand rotation of nucleosomes.Polyomaandsimian virus 40(SV40) DNA can beisolatedfrominfected cellsandpurified virus in association withcellularhistones (13, 16, 17, 19, 26, 30, 32-34). In the electron microscope, these nucleoprotein complexes are seen as cir-cularmolecules havingnucleosomes character-istic of cellular chromatin (11, 21, 58). These molecules have been designated as SV40 mini-chromosomes or, morerecently, SV40 chroma-tin.
To understand the arrangement of nucleo-somes in relation to the DNA base sequences, SV40chromatin has been used as a substrate for
tPresent address:Hyland Division, Travenol Laboratories, Inc.,Round Lake, IL 60073.
tPresent address: Department of Biology, University of California,SanDiego,CA 92093.
§Presentaddress:DepartmentofMicrobiology, The Med-icalCollegeofPennsylvania,Philadelphia, PA 19129.
bacterial restriction endonucleases. The as-sumption inherent in this approach has been that DNA cleavage sites which are covered or are strongly associated with histones will be resistant to the endonucleases. Several groups have studied SV40 chromatin, but the conclu-sionsarrived atby the various workers have not always been in agreement. Polisky and Mc-Carthy(42) foundthatonly20% of the DNA in SV40 chromatin was protected from nuclease digestion and that the protected regions were randomly distributed. However, the SV40 nu-cleoproteincomplexwhichtheystudied was pre-pared by alkaline degradation of virions, and thistreatment hasbeen shown to cause partial removal ofnucleosomes anddegradationof his-tones (3, 7, 34, 56). In contrast, Cremisi et al. (11),who studiedthe more"native"intracellular
718
on November 10, 2019 by guest
http://jvi.asm.org/
form of SV40chromatin, found that 73 to 85% of the DNAwas resistant to digestion by EcoRI. Theysuggested that the cleavage was occurring only in the internucleosomal DNA regions. Nevertheless, Persico-Dilauro et al. (41) found that 50% of DNA within intracellular SV40 chro-matin was cleaved by both staphylococcal nu-cleaseand EcoRI nuclease. From their analysis ofthe HindlIl digest patterns, they proposed that thenucleosomes are distributed randomly in the population of chromatin, but are not randomly arranged within the SV40chromatin molecule. Ponder and Crawford (44) on the other hand, proposed that the arrangement of nucleosomes is not completely random within the wholepopulation,but that eachnucleosome canoccupy oneofalimited numberofpositions. In this paper, we describe the results of our experiments using intracellular SV40chromatin purified by hydroxyapatite chromatography(34) as asubstrate for bacterial restrictionenzymes. TheSV40chromatin purified by this procedure hasa ratio ofproteinto DNA greaterthan one (22,34) and hasaboutthesamecomplementof histones asis foundinwholevirionsand in
Hi-depleted cellular chromatin. In our study, we found that thesusceptibility of SV40 chromatin DNA torestriction endonucleaseswasstrongly affected by the exact experimental
conditions,
particularly
thesaltconcentration, andthisfind-ing may provide a partial explanation for the discrepancies in some of the previous studies. Anadditional
significant
findingwasthat relax-ation of thetopological
constraintsinSV40 chro-matinDNA Ialteredthat structureof theSV40 chromatin such that the chromatin DNA be-camehighly resistanttothe action ofthe restric-tionenzymes.MATERIALS AND METHODS
Cells and viruses. Plaque-purified SV40 virus
(strainRH911)waspropagatedinthe TC7subline of
CV-1aspreviouslydescribed(22, 34).
PurificationofSV40 chromatin. Theprocedure
usedtopurifySV40 chromatin from Tritonlysatesof
infected cellswasessentially thatdescribed in detail
inapreviouspublication(34), with thefollowing
mod-ifications.SV40 chromatinwaseluted from
hydroxy-apatite columns with 0.28 M phosphate buffer, pH
6.8-0.1% Triton X-100afterwashingextensivelywith
0.22 Mphosphate buffer, pH 6.8-0.1%Triton X-100.
Columns and buffers weremaintained at 8 to 100C.
The SV40 chromatinwhich elutedfrom thecolumns
wasconcentrated andthensedimentedat36,000 rpm
in a5 to20%(wt/wt) sucrosegradientcontaining0.1
MNaCl-0.025%Triton X-100-0.001 MEDTA-0.01M
Tris-hydrochloride(pH7.9) for2.5hat40C.Onlythe
leading two-thirdsportion oftheSV40chromatinpeak
was pooled. These fractions were concentrated by
vacuum dialysis and then dialyzed into the appropriate reaction buffer.
The average buoyant density in CsCl gradients of
theSV40chromatin purified by the above procedure
and fixed with glutaraldehyde was 1.44 g/cm3. The
histone complement of thisform of SV40 chromatin is
similar to that found in virions in that it contains
aboutequimolar amounts of H2A, H2B, H3, and H4
histones but is devoid ofHihistone (34). Nonhistone
proteinscould not be detected in the SV40 chromatin
evenafter the proteins were labeled in vitro with125I
to a high specific activity (G. L. Liggins and D. A.
Goldstein,unpublisheddata).
Radioactivelabelingof SV40 chromatin DNA.
Forthe routine preparation of SV40 chromatin DNA,
the DNAwaslabeled at low specific activity to assist
intheidentification and quantitation of the chromatin
duringpurification. ['4C]thymidine, (0.05 to 0.1 ,iCi/
ml; New England Nuclear Corp.) was added at 24 h
after infection and not removed until thecells were
lysed at72to90h afterinfection.
Forlabeling of SV40 chromatin DNA to high
spe-cificactivity with32P,[32P]orthophosphate(25-50,Cil
ml; New England Nuclear)wasadded in
phosphate-free growth medium (Eagle minimum essential
me-dium containing 2.5%dialyzed horseserum) at24h,
and theradioactivitywasremovedjust before Triton
lysisofcellsat 72hafter infection. Naked32P-labeled
SV40 DNA was prepared from purified SV40
chro-matin by removal ofprotein. For this purpose,32p
labeledSV40 chromatinwasincubatedat40°C in 0.1%
sodium dodecylsulfate (SDS) with50ygof pronase
perml for1h.Thereafter, theDNA wassedimented
for2hat45,000 rpm ina 5 to20%sucrosegradientat
roomtemperatureinanSW50.1 rotor(22).39P-labeled
SV40 DNApreparedin this way hadaspecificactivity
of 1x 105to2x 105cpm/jLgof DNA.
Bacterial restriction endonucleases. Restriction
endonucleases R-EcoRI, R-HpaI, R.HpaII, R.
BamHI and R*HindIII were purchased from
Be-thesda Research Laboratories, Bethesda, Md. R.
HindII + III was obtained from Mary Gutai and
Michael Chen. The locations of thecleavagesites for
the variousenzymes usedinthisstudyare depicted
ontheSV40 DNA mapinFig.1.Enzymeswerestored
at-20°C.Unless otherwisespecifiedinthetextorin the legendsto the figures, SV40DNA or chromatin
wasincubated with the enzymesat37°Cinthe
"stan-dard" reactionmixturecontaining10mM
Tris-hydro-chloride (pH7.5), 0.025% TritonX-100,100mMNaCl,
5mMMgCl2and2mM
/3-mercaptoethanol.
All bufferswere autoclaved before use, and all reaction were
carriedoutinsterile vials and tubes. Reactionswere
terminatedbySDS(finalconcentration, 0.1%)-EDTA
(0.05M) and heatedat56°Cfor5min.
Polyacrylamide and agarose gel
electropho-resis.For theseparationofthe DNAfragments
gen-eratedbydigestionofSV40 chromatinwith restriction
enzymes,theproteinwasfirstremovedbyheatingthe
chromatinat56°Cfor10min in0.35%SDS,and the
DNAwaselectrophoresedonverticalagaroseor
poly-acrylamideslabgels.Sampleswereadjustedtoa
con-centrationof 5%glyceroland0.02%bromophenolblue
in avolume of50
1IL
andloadedontoslots of slabgels.on November 10, 2019 by guest
http://jvi.asm.org/
J. VIROL.
720 LIGGINS, ENGLISH, AND GOLDSTEIN
EcoRI I
Hl
LMC*iD E KF
Hind M BED C A B
Hin d +-A
HindM
HpaI
A D C E FA
A C B A
Hpao
BamHI -
--MapUnits 0 .2 .3 4 5 .6 .7 .8 .9 1.0
FIG. 1. Restriction enzymecleavage mapof SV40 DNA. The cleavage mappositions assigned to the various enzymes are shown relative to the single cleavage sitefor EcoRI. There are4 cleavage sites forHpaI (12, 50, 66), 6sitesfor HindIII (66), 13 sites
forHindII+III(12, 66), 1siteforHpaII (50, 66), and
1 sitefor BamHI (R. Roberts,personal communica-tion).
Theagarosegelswere 1.4%agarosein 40 mMTris, 5 mM sodiumacetate,and 1mMEDTA(pH 7.8). The
polyacrylamidegelelectrophoresisbufferwas40mM
Tris-20 mM sodium acetate-2 mM sodium EDTA (pH 7.8). Thedimensions ofthegels, the time of the runs, and other conditions varied for each enzyme
reaction. These detailsaregiven in thetextorinthe
legendstothefigures. In allexperiments, bufferswere
circulated between thereservoirs during theruns.
Forradioautographic analysis of 32P-labeled DNA
digest fragments, slab gels were removed from the
glass plates at theconclusion of the electrophoresis, thewetgelswerewrapped tightly in Saran Wrapto preventdehydration, and thegelswereplaced in
con-tactwithmedical X-ray film. Filmsweredeveloped8 hto 5daysafterexposure. Forquantitative analysis, theexposed filmswerescanned witharecording
den-sitometer. For analysis of DNA digests not labeled with 32p, gelswereplaced inan aqueous solutionof
ethidium bromide (2 ytg/ml) at 4°C for at least 4 h. Thewetgelswereilluminated with UV light,and the fluorescent DNA bands were photographed with a
Polaroidcamera.
Forremoval of DNA fragments fromagarose gels,
gel slicesweredissolvedin2volumes of5Msodium perchlorate at 65°C, and the dissolved agarose was
passed through small columns of hydroxyapatite at
65°C. Columnswerewashed with 10volumes of0.01 M sodium phosphate buffer (pH 6.9),and the DNA
waseluted with0.4 M sodiumphosphate buffer (pH
6.9). DNA wasdialyzed against 0.1 M NaCl-0.01 M Tris (pH 7.9)-0.001 M EDTA, and the DNA was
concentrated byethanol precipitation.
RESULTS
Theoretical considerations. The generally
accepted notion that histones do not bind to DNAwithbasesequencespecificityhasa
theo-retical basis in thefact that the histones as an
evolutionary class of proteins are very highly
conserved. Nonetheless, this does not rule out
thepossibilitythat in vivo there is
"phasing"
of nucleosomeswith respectto aspecific
DNA base sequence as, forexample,
the sequence at the origin ofreplication.
If nucleosomes doindeed associatewith the DNAat aregion
suchasthe origin qfreplication
(20, 49), eachsubsequent
nucleosomecouldbe "in
phase"
with thecorre-sponding nucleosome in all other SV40 chro-matinmolecules. The
assumptions
inthis argu-ment are that (i) nucleosomes do not movelaterally
along the DNA backbone in vivo and(ii)thefirstnucleosome addedtothe DNAafter initiation ofreplication establishes the natural spacing of nucleosomes inherent in chromatin. In a casewhere thenucleosomeswould be in phase, digestion by a restriction endonuclease withasingle cleavagesiteshould resultin either cleavage ofallthe molecules (e.g., site not pro-tected byanucleosome) or
complete
resistance (siteprotected
byanucleosome). If the nucleo-somesare notinphasewithinsuchapopulation
but randomly arranged with respecttoa given base sequence, thenan intermediate degree of digestion would be
predicted.
In this case, the resistant fraction would reflect the degree towhich the histones protect the DNA. Neverthe-less, partial digestion might result from other causessuchasreassortmentduring isolation or
digestionorboth,orduringother
perturbations
which would allow access tosites in some, but
not all molecules. Of course, the above predic-tionsare
justified provided
nounforeseen inter-actions take place between the endonuclease andthe histones during treatmentof the SV40 chromatin. However, if the same fraction of chromatin molecules was resistant to cleavage after digestion by other single-site restriction enzymes,thenastrongargumentcouldbe made foranonspecific arrangementof histones. Fur-thermore, chromatin with such a distribution of histonesshouldyield equimolaramountsof each limit digest DNA fragment after cleavage by multiple-site restriction enzymes. It should be pointed out that others (11, 41-44) have pre-sentedsimilararguments inprevious studies on thearrangement of histonesinSV40 andpoly-omachromatin.
Digestion of naked SV40 DNA and SV40 chromatin with single-site restriction en-donucleases.SV40 chromatin and naked SV40 DNAwereincubatedseparately with
EcoRI
and treated with SDS to stop the reaction and toremove the protein from the DNA, and the
reaction products wereanalyzed by electropho-resis on agarose slab gels (Fig. 2). Bands are observed at threepositions; the top, middle, and lower bandscorrespond to nicked-circular SV40 DNA (DNA II), linear DNA (DNA III), and
on November 10, 2019 by guest
http://jvi.asm.org/
[image:3.507.69.259.72.200.2]supercoiled DNA (DNA I), respectively. Treatmentof naked control supercoiledDNA
wascharacterized by the conversion of DNA I
tolinear DNA III molecules bywayofa
short-lived,nicked-circularDNAIIintermediate (Fig.
2a).The conversionsequencehasbeennotedby
others(37, 47). Theintermediate doesnotresult from endonuclease contamination, since
pro-longed incubation with EcoRI didnotresult in furtherdegradation of thelinearform and alkali
denaturation of the linear molecule failed to
reveal significant amounts of single strands
shorterthanunit-length SV40 DNA.
Incontrast tonakedSV40 DNAI,whichwas
entirelyconvertedtoDNA III, SV40 DNA I in chromatin was partially resistant to digestion
evenafterprolonged incubation with theenzyme
DNA 11
DNA III
DNA I
DNA 11
DNA III
DNA I
Controls1 3 6 9 12 15 30 45 60
MINUTES AT 37`C
FIG. 2. Kinetics of digestion ofSV40 DNA and SV40 chromatin by EcoRIat 100 mMNaCl. SV40 DNA (5.5pg) andSV40chromatin (8.3pg) were in-cubatedseparatelyinavolumeof120,ul ofthe "stan-dard" reaction buffer (see text) to which 2.5 Uof EcoRIwasadded. At the times indicated, 10,Il of reaction mixture wasremoved and diluted
immedi-atelyinto30,l ofasolutioncontaining10mM
Tris-hydrochloride (pH 7.5),50 mMEDTA,and 0.1% SDS
tostopthereaction,and thesampleswereanalyzed
byagarosegelelectrophoresis. Thedigestion
prod-ucts werelocatedby stainingthegelwith ethidium bromideatafinalconcentrationof2pg/ml. (a)SV40
DNA; (b) SV40 chromatin. The controls represent
portions ofSV40DNA andSV40chromatinremoved
fromthe reactionmixturebeforeadditionofthe en-zymeandincubatedat0°C (slot 1)orat37°C(slot 2)
for60min.
(Fig. 2b). A large fraction remained as DNA II andwas notconverted to DNA III. To demon-stratethat the partialdigestionpatternobserved with DNA from chromatin was not due to lim-itingenzyme orinteraction of the histones with enzyme, additional reactions were carried out in the presence of a large excess of EcoRI. Even underthesereaction conditions, only 23% of the DNA in chromatin was converted to DNA III (Table 1),whereas control naked SV40DNA I wasentirely converted to DNA III within 1min (data not shown). In addition, the amount of DNA IIinitiallypresent inSV40chromatin did notdecrease,but
actually
increased slightly (Ta-ble 1).In view of the finding that treatment with EcoRI resultedin only partial cleavage ofthe SV40 chromatin, the tentative conclusion would be thatnucleosomesare not inphase within the population. However,theexperimentsdescribed abovegive little informationas to the distribu-tion or coverage by nucleosomes within any givenSV40 chromatinmolecule, since as pointed out above, partial cleavage could be a conse-quenceofanumber of factors.
To further investigate this distribution, SV40 chromatin was digested with twoother restric-tionenzymes, BamHIandHpaII, each of which cleaves SV40chromatin at adifferentsingle site. These sitesarelocatedatfractional-lengthmap positions0.14(61) and0.735(50) for BamHI and HpaJJ,
respectively,
relativetothesingle EcoRI site, whichisarbitrarily assignedthe zero posi-tion(12).Separate kinetic experiments were done on thesamepreparation of
32P-labeled
SV40chro-TABLE 1. EcoRItreatmentofSV40 chromatin in
100mMNaCla
Distributionof DNA(%)b Tine(min)
DNA I DNA II DNA III
0 91 9 0
1 80 16 4
3 71 14 15
6 68 13 19
15 66 13 21
60 62 15 23
aFive units of theEcoRIwerereacted with 380ng
of3P-labeled SV40chromatin. The reactionwasina
volume of60
Id
ofthestandard buffer (seetext).Atthe timesindicated, 10
pl
wasremoved fromthere-action mixture, and the samples were analyzed by
agarose gel electrophoresis. Under these conditions,
naked SV40 DNA I usedas a control was
quantita-tively convertedtoSV40DNA III within 1 min.
b Valueswerecalculatedfromdensitometertracings
ofexposed X-rayfilmsplacedincontactwithagarose
slabgels.
w
on November 10, 2019 by guest
http://jvi.asm.org/
[image:4.507.264.454.466.556.2]722 LIGGINS, ENGLISH, AND GOLDSTEIN
matin by using each of the three enzymes.Before digestion, theSV40chromatinpreparation
con-tained90% DNA I and 10% DNA II. The results ofonesuchexperimentaresummarized in Table 2.Only 19 to 22% of the DNA inSV40chromatin was converted to DNA III by each of the
en-zymesunder the conditions of this reaction (100 mM NaCl, 5 mM MgCl2, 0.025% Triton X-100, 10mMTris, pH 7.5,at37°C).The salientfeature of thisexperiment is that all three enzymes gave approximately the same degree of double-stranded cleavage, even though the respective sensitive sites are located at considerable dis-tances apart on the genome. Therefore, these results indicate that there isnopreferential dis-tribution ofnucleosomes within any oneregion of the genome. Instead, it appears that the nu-cleosomes are arranged along the DNA inde-pendentof nucleotidesequence.
Thevalue of19to22%for thefraction suscep-tible to double-stranded cleavage agrees well with the findings of Cremisi et al. (11), who digested SV40 chromatin with EcoRI and ex-amined the distribution of linear and circular molecules in the electron microscope. These workers interpreted thecleavage as representing digestion in the naked DNA "bridges" or "spacers"extending between nucleosomes, since
asimilar sensitive region is found upon digestion ofcellular chromatin (1, 10, 39, 52, 55, 57).
Effect of salt concentration on endonu-clease cleavage of SV40 chromatin. In an
[image:5.507.263.453.202.463.2]electron microscopic study, Griffith (21) found that the SV40 chromatin went from a highly condensed circular structure with a contour length of about 210 nm to an extendedbeaded form with a contour length of about 514 nm when the salt concentration was lowered from
TABLE 2. Comparison of EcoRI, HpaII,and BamHI treatmentofSV40chromatinin100mM
NaCla
Restriction endo- Distribution of DNA(%) nuclease DNA I DNA II DNA III
EcoRI 65 16 19
HpaII 70 14 16
BamHI 66 12 22
a:32P-labeledSV40 chromatin was incubated in
sep-aratereactionmixtures with either 4 U of EcoRI, 2 U ofHpaII, or2U of BamHI in the standard reaction buffer. As acontrol,32P-labelednakedSV40 DNA was incubated at the same ratio of DNA to enzyme. Under these conditions, the control DNA was converted
quantitativelytoDNA IIIwithin 3 min.
bValueswerecalculated from densitometer tracings
of the 20-min reaction products, at which time the reactions were 90%complete.
J. VIROL.
0.15 to 0.015 M NaCl. On the basis of this
observation, it might be predicted that such a
configurational change would cause the SV40 chromatintobemoresusceptibleto endonucle-asecleavage.
Figure 3 shows the kinetics of cleavage of
SV40 chromatin upon digestion with EcoRI
(Fig. 3a) orBamHI (Fig. 3b) in the presence of 50mMNaCl-5 mM MgCl2. Naked SV40 DNA
I is
completely
converted to DNA III in 1 min(as indicated by the arrow in Fig. 3), whereas the reaction of enzyme with chromatin is not
100-
I0
'(a)
75-enX
w U
-J 50
-0
0
Oj
25-5z
0
a IOO-
(b)
0
7575
z w
25
-0
1 2 3 5 10 15 20
[image:5.507.59.250.487.560.2]MINUTES AT 37°C
FIG. 3. Kinetics of SV40 chromatin digestion by
EcoRI and BamHIat50mMNaCl.32P-labeledSV40
DNA and32P-labeled SV40 chromatin, each
contain-ing 55,000 cpm (1.52 x 105 cpm/lg of DNA), were
mixedseparately with either: (a)8UofEcoRIor(b)
4 Uof BamHI in reaction mixtures containing 50
mMNaCl,10mMTris-hydrochloride (pH 7.5),5mM
MgCl2, 0.025% TritonX-100, and2mM
/3-mercapto-ethanol. Reaction mixtures were incubated at 37°C and,atthetimesindicated, sampleswerewithdrawn and thereactionswereterminated.SV40 naked DNA I(arrow) wascompletelyconverted to DNA III within 1minunder these conditions.Sampleswereanalyzed
byelectrophoresisin 1.4% agarosegels, and the
diges-tionproductswere revealedbyexposure ofthe slab
gels toX-ray films. The percentages of the various
forms of SV40DNA were calculatedfromthe areas
under thepeaksobtainedfromdensitometric tracings
ofthefilms. Curvesaredrawn in thisfigureonly for
DNAfromSV40 chromatin. Symbols:U, DNA I; A, DNAII;0,DNAIII.
on November 10, 2019 by guest
http://jvi.asm.org/
complete untilsometime between5and 10min. It is clear thatthere is asignificant increase in the susceptibility of SV40chromatinupon low-ering the saltconcentration. Whereas 19 to 22% of themoleculeswereconvertedtoDNA III at 100 mM NaCl (Table 2), a decrease in concen-tration to 50 mM increasedyieldsof DNA III to 47 to 50% (Fig. 3). There was no significant increase in yields of DNA III when the NaCl concentration was lowered even further to 10 mM (datanotpresented).
Examination of the kinetic data in Fig. 3 re-veals that digestion is atwo-phase process. Dur-ing the firstminute, 25% of the chromatin DNA is digested, whereas the remaining 25% is cleaved during thesubsequent 10-min period.The first fraction,presumablycorresponding to the inter-nucleosomal fraction,appears tobe quite acces-sibletothe enzyme,sincecontrol nakedDNA is
also
completely digested
during the first minute.This isconsistent with the electronmicroscopic data ofGriffith (21),which showed naked inter-nucleosomal DNA regions exposedatthelower salt concentrations. The additional25% which is cleaveduponprolonged incubationsuggests that regions presumablynear, orperhaps within the nucleosome, are lessaccessible for cleavage by the enzyme. Importantly, the total susceptible fraction of50% corresponds wellwith the esti-mate of coverage as determined by digestion with nonrestricted endonucleases (8, 9, 32, 41). Resistance ofSV40 chromatin containing DNA II to cleavage by single-site endonu-cleases. In most ofthe experiments described above,uponpurification from cellextracts, chro-matin preparations contained about90% DNA I andabout 10% DNAII. Since the fraction of DNA IImoleculesremainedapproximately con-stant orincreased uponendonucleasetreatment (Fig. 2band 3;Tables1and2),it appearedthat the chromatincontainingDNA II wasresistant todouble-strandedcuts atall of thesalt concen-trationstested.
Todetermine whetherthisresistancewas per-haps a property of DNA II chromatin derived fromanin vivo mechanism (14)or,instead,was a general characteristic ofDNA II chromatin generated byrandomsingle-strandcutsofDNA Ichromatin, thefollowing experimentwasdone. SV40 chromatin labeled with 32Pwasstored for several weeks at
40C,
duringwhich time random single-strand nicks were introduced by 32P de-cay. This chromatin and nakedSV40DNA pre-pared from the chromatin were used as sub-strates in separate reactions with BamHI and EcoRI. Figure4showsresults of anexperiment inwhich about 25% of the chromatinemployedas asubstratefor theenzymeBamHIcontained
Naked DNA Minichromosomes
DNA E - _ me_ SW
DNA
MimpqnO
- _
amDNAI _i
_1U
_ _m0 1 3 5 15 0 1 3 5 15
MINUTES AT370C
FIG. 4. Resistance of SV40 chromatin DNA II to
BamHI.32P-labeled DNA and chromatinwerestored
at4°C for4weeks,during which time about25%of
theDNAwas found to be in the linearDNA III form,
presumably converted to this form by decay of the32p.
SV40 chromatin and DNAwerereactedwith BamHI
under conditions similar to those described in the
legendtoFig. 3, except that theNaCl concentration
was 10mM insteadof50mM.
DNA II. In contrast to naked DNA, in which DNA I and DNA II areboth rapidly converted toDNAIII, the fraction initiallypresent as DNA II remained essentially unchanged. This SV40 chromatin preparationrepresents onlyoneofa number of different SV40 chromatin prepara-tions inwhichthe amountofSV40 DNA IIdid notdecreasesignificantlyupon incubation with thesingle-site restriction enzymes.
Digestion of SV40 DNA and SV40 chro-matin with multiple-site restriction endo-nucleases. The apparent resistance of the nicked-circular form of SV40 chromatinto cleav-age suggested that, upon relaxation of the closed-circular DNA, a conformational change mayhaveoccurred,causing theDNA tobecome resistant to digestion by restriction endonucle-ases.Thus itwould followthat,
regardless
of the number ofpotentialrestriction enzymecleavage sites, theremaining sitesmight be rendered in-accessible afterreceiving thefirstnickorchop.
To testthis
hypothesis,
32P-labeled SV40 chro-matin and 32P-labeled naked SV40 DNA were reacted separately with themultiple-site
en-zymesHindll+IIIforvarious timeperiods,and the resultant
fragments
wereseparated
on 4% acrylamide slabgels.
The location ofcleavage
sites in SV40 DNA for
HindII
+ III and the otherenzymesused in thisstudy
areshown in Fig. 1. The digest patterns areshowninFig.
5.SV40 DNAyielded 11
fragments
within 15min after exposure to the enzymes. Not shown aretwovery smallfragments,30and20base
pairs,
respectively, which run off the
gels
undercon-ditionsrequiredforresolution of the other
frag-ments (66). The naked SV40 DNAused inthis
experiment
wasobtainedby
theremoval ofpro-tein from intracellular SV40
chromatin,
but iton November 10, 2019 by guest
http://jvi.asm.org/
[image:6.507.258.453.75.178.2]724 LIGGINS, ENGLISH, AND GOLDSTEIN
~*
--f,/ I :1.."
,
I-
tl.. .,4'...11 11-1-I
.z '.7!..
FIG. 5. HindII+IIIdigestion ofSV40 naked DNA andSV40chromatin.Naked32P-labeled SV40 DNA (1.3 jig)and32P-labeledSV40chromatin (2.5jug)were
incubatedseparatelyat37°C with HindII+III (5 U)
in a volumeof 250 ,ul ofabuffercontaining50mM
NaCl,6mMTris, 6 mMMgCl2, 0.6 mM
,8-mercapto-ethanol, 0.6%glycerol, and 0.01% TritonX-100 (pH 7.5). At the times indicated, 50-jilsamples were
re-movedfrom each tube, the reactionswereterminated,
and the samples were electrophoresed on 4% poly-acrylamide slab gels (40 by0.3cm)for21.5h at100 V,53mA.To locatethe DNA bands, gelswereplaced
incontactwith X-ray filmsfor 72 h.
gave the same digestion pattern as DNA from
purified virus.
In contrast tothe 11 fragments of the naked
SV40 DNA digest pattern, the majority ofthe
DNA in chromatin was resistant to multiple cleavagebyHindII+ III(Fig.5). Even4h after incubation,mostof theSV40DNA in chromatin
had received only one cut and migrated to a
position characteristic of linear full-lengthDNA.
Furthermore, asignificant fractionwasnot
sus-ceptible to cleavage and, instead, failed to
mi-gratein the
gel.
Only
about 10% of the molecules received more than one chop, as indicatedby
the mixture of incomplete digest
products,
whereaslessthan 1%correspondedtoanyof the 11 limit digest
products-in
this case,only
tofragment A.Ifallthecleavagesiteswereequally susceptible,onthe average, theSV40 chromatin molecules shouldhavereceivedsixtosevencuts
under these reaction conditions. The fact that most ofthe cleavage sites in SV40 chromatin were protected from
digestion
after the initial cut provided additional evidence that arear-rangement or a
configurational change
orboth tookplace suchthatpotentially susceptible
sites became coveredby
histones. To what extentthese
configurational changes
could be influ-enced by the reaction conditions was nextex-amined.
It is known that rearrangement andexchange of histonescan occuratcertainsalt anddivalent ionconcentrations (8). Thus,oneofthefactors studiedwastheeffect of
magnesium
concentra-tion onthe
susceptibility
ofSV40 chromatin toHindII + III. Inthe
experiment just
discussed (Fig. 5), the concentration ofmagnesium chlo-ridewas6mM.Shown inFig.6 arethedigestion products of SV40 chromatin DNA at 0, 1, 20, and100mMmagnesiumconcentrations. At very low concentrations ofmagnesium (less than 1mM), control naked SV40 DNA was only par-tially convertedtolimitdigest products,sothese reactions with SV40 DNA and chromatin are not shown (also, thereactions had to be incu-bated forlong periodsat37°C,conditions which could cause degradation and denaturation of chromatin).
Atall of themagnesiumconcentrationstested, SV40DNA inchromatinwaseither resistantor was converted mostly to linear DNA III, indi-cating that the majority of the molecules had
one cut or less. An interesting feature of this
experimentwasthatSV40 chromatinDNA was digestedto a greater extent at the lower mag-nesium concentration (1mM), whereas the op-positewas truefor the control nakedSV40 DNA (e.g.,Fig. 6, slots3and4 compared with slots 7 and 8). These reactionswerecarriedout at37°C for4.5 h, so it isdifficult to conclude that this effectwassolely the result ofa magnesium-de-pendent configurational change, since it may
have been caused by partial denaturation or
degradation ofproteins. However, it is the op-posite effect that would be predicted for ex-changeorremoval of histones, since these
phe-nomenaappeartoincreasewith increasing
con-centration of divalent cation (8). It should be noted thatat100mMmagnesium concentration, both theSV40DNAand chromatinwere resist-ant, probably due to the aggregation of DNA,
J. VIROL.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:7.507.70.244.74.428.2]1
l 2 3 4 5 6 .7 6 9
19
RI3...
_0
to
g4
1_
_I
_ _ _
C D
E
F
G-H
K-FIG. 6. Effect of magnesium concentration onthe
digestionof SV40 DNA and chromatin byHindII+
III. Naked 32P-labeled SV40 DNA or 32P-labeled SV40chromatinwereincubatedat37°C for4.5 hwith
HindII+ III(2 U) inavolumeof72,l ofa buffer
containing80 mMNaCI,8mMTris,
mM/3-mercap-toethanol, 0.02% TritonX-100(pH 7.5), andvarious
concentrationsofMgC42.Reactionswereterninated, and 50-pi samples were electrophoresed on a 4%
polyacrylamideslabgel for19.5hat120V, 48 mA.
Each slot represents a different reaction mixture. Slots1through6,SV40 DNA;slots7through9, SV40
chromatin; slot 10, SV40DNAplus EcoRIcontrol.
Slot 1, 0 MMgCl2, no enzymes; slot2, 0MMgC12, enzymes;slots 3 and7,1mMMgCl2, slots4and8, 20
mMMgCl2;slots 5and9, 100mMMgCI2;slot 10,5
mMMgC12.
chromatin,enzyme,or acombination of these.
Further attempts were made to detect the
removalorexchangeofhistones underthe
stan-dardenzymereaction conditions by sedimenta-tion analysis. 3H-labeled SV40 chromatin was mixed with32P-labeled SV40 naked DNA at a weight ratio of chromatin to DNA of 16 to 1, incubatedfor 1 h in 50 mMNaCl-6mM
MgCl2-1mM
,6-mercaptoethanol-0.01
MTris (pH7.5)-0.01% Triton X-100, andsedimentedin asucrose gradientcontaining the same buffer. Except for asmallshoulder on the leading edge of the DNA peak, therewasno evidence of a change inthe sedimentationpatternof theDNA or chromatin (Fig.7).Thus, the conclusion from the two afore-mentioned experiments is that protein exchange is not adeterminingfactor inthe acquisition of resistance of SV40 chromatin upon the initial cleavage of SV40chromatin.
Reactivity of restriction enzymes with SV40 chromatin. Another possibleexplanation for the inability to generate limit digest frag-ments by multiple-site restriction enzymes is thatthechromatin or some other component in the reaction mixturemay cause adrastic reduc-tion or change in the enzyme activity so that only sufficient activity remains to generate a
^-- 4
0
x3 "2
0.
=u
4^
0
3,X 2 0
a.
cm
I M
4^-..
0
3 X CL
0~
,.4
0 10 20 30 40
Fraction Number
FIG. 7. Sedimentation ofmixturesof SV40 chro-matin withSV40DNAathighratiosofchromatinto DNA.A2-pgamountofSV40 [3HJchromatin(20,000
cpm/pg)wasmixed with 0.125pgof32P-labeledSV40 DNA (122,000 cpm/pg) in a volume of50 ,ul and incubated as described in the text. Samples were
layered onto 5 to 20% sucrose gradients and sedi-mentedfor2 h at36,000rpm in a SpincoSW50.1 rotorat4°C. Radioactivity representstrichloroacetic acid-insolubleproducts. (a)DNAalone;(b)DNAplus chromatin.Sedimentationwasfrom righttoleft. A
on November 10, 2019 by guest
http://jvi.asm.org/
[image:8.507.54.247.79.472.2] [image:8.507.278.425.332.566.2]726 LIGGINS, ENGLISH, AND GOLDSTEIN
single cleavage inmost of the molecules. Con-sequently, SV40 chromatin and naked SV40 DNA were incubated togetherin various
com-binations withHindIII to determinetheextent
of enzyme activityduring the courseof the
re-action. Such an experiment is shown in Fig. 8 which shows the reaction of 3H-labeled SV40 chromatin and SV40 [32P]DNA with HindIII endonuclease. HindIII cleaves SV40 DNA into six fragments (12, 66). Incubation of HindIII with SV40 chromatin for 2 h (slot 2) did not
generatelimitdigestfragments.Instead,the
ma-jor fraction was cleaved only once toyield full-length linear DNA (position of RI marker), whereas the remainder of the DNA either did not migrate into the gel (this fraction was a
mixture of closed-circular and nicked-circular
DNA)or wasconverted intopartial digests (8%).
The addition of fresh enzyme after 2 h and incubation foran additional 2hdid not signifi-cantlychange thedigestpattern (slot3,Fig. 8). Todemonstratethat the enzymewasindeed in
excess and still active after 2 h of incubation,
[32P]DNAaddedtothe SV40 chromatin reaction mixture at this time and incubated for2hmore
wascompletelydigested(slots4and 4B). When SV40 DNA and chromatin were mixed before the additionof enzyme, thecompositeresultwas
the sameasthat obtained when DNA and chro-matin were incubated separately (slots 7 and 7B).These experiments show that the resistance of the SV40 chromatin to limit digestion by HindIIIdoesnotresult fromachangeor
reduc-tion in enzymeactivity.
Initial cleavage sites of HpaI in SV40 DNA chromatin. Since the majority of the SV40 chromatin molecules could only be cut
once by the multiple-site enzymes, it was of interest to ascertain whether or not the initial cleavageoccurred at asingle site or at preferred sites on the SV40 genome. The position of the initial cleavage site was determined by restric-tion enzyme mapping. To simplify the analysis, themultiple-site restrictionendonuclease HpaI
was used to generate linear DNA III, since it chops SV40 DNA at only four positions along the genome (12, 50, 66). Two of these sites are situated very close together, being only 0.004 map units apart (66), and for the purposes of mapping, theycanbetreated as a single site.
A time course ofdigestion of control naked SV40 DNA by HpaI is shown in Fig. 9. Limit digestion generated by cleavage at all sites, i.e.,
0.755(actually,0.753and0.757), 0.375, and 0.175
produces three major fragments: (A) 41% frag-ment; (B) 38%fragment; and (C) 20% fragment (thepercentages denote the fractional length of the DNA). Incomplete digestion caused by
Rl-
A-
B-
C-
[image:9.507.266.458.74.300.2]D--E
-.FIG. 8. HindIII digestion of mixtures of SV40
chromatin and nakedSV40 DNA. SV40 chromatin
and32P-labeled nakedSV40DNA wereeither incu-batedseparatelyormixedtogetherinapproximately
equalamounts(2 ,ug) anddigested with HindIII(2 U)
in 100jil ofabuffercontaining 85 mMNaCl,8.5mM
Tris, and 15 mM MgCl2 (pH 7.5). Reactions were
terminatedafter4h. Samples were electrophoresed
on a4%polyacrylamideslabgel (0.15by10cm)at60
Vand70mAfor9h. Slot1, chromatin,noenzyme;
slot 2, chromatin with HindIII; slot 3, chromatin,
HindIII (2 U), 2 h, additional HindIII (2 U) added
and incubated for 2 more h; slot 4, chromatin,
HindIII2h, DNA added and incubatedfor2
addi-tional h; slot 5, DNA, HindIII; slot 6, DNA, no
enzyme; slot 7, chromatin, DNA, no enzyme, 2 h,
HindIII added and incubated for2 additional h.
Slots 1 through 7 show ethidium bromide-stained
gels. Slots 4b through 7brepresentautoradiograms
of slots4 to 7.
cleavage at onlytwo sites yields the following "partial" digest fragments: an 80% fragment (0.175 and 0.375); a 62% fragment (0.755 and 0.375);and a58%fragment (0.755and0.175).In
Fig. 9, the 58% and 62%fragmentsare notclearly resolved.
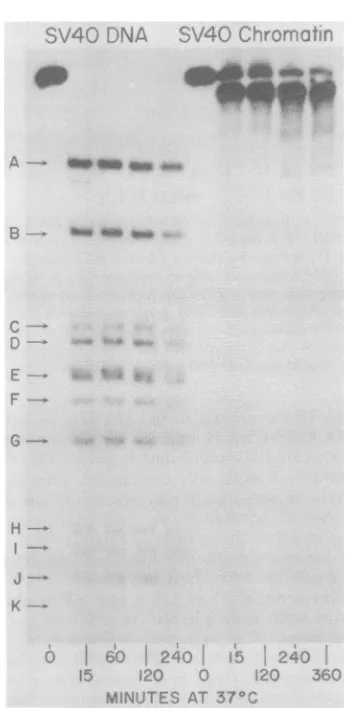
Figure 10 shows a time course of SV40 chro-matindigestionby HpaI. Unlike the SV40 DNA, which is cleaved to the limit digest fragments within1 h, digestion of SV40 chromatin results inless than 3% of any limit digestfragment, even after prolonged incubation in a vast excess of enzyme for6h.Instead, the major digest product
was SV40 DNA III. Thisis consistent with the previous data obtained after digestion with HindIII and HindIl + III.
J. VIROL.
.91LAINk
on November 10, 2019 by guest
http://jvi.asm.org/
STRUCTURAL CHANGES IN 727
SV40 DNA SV40 Chromatin
DNA 11 am
_M
-DNA III
80% fragment. DNA 58% fragment
A_
B-,.U_4_._ _
DNA 11
El_
DNA III
S*m_ *. ,,
.~~~ ~ ~ ~~~~z
0 p n
:..*.#.
S.
DNA I
A
B
W
C-s
0.25 3 5 15 20 30 60
Minutes
FIG. 9. Kinetics of SV40 DNA digestion by HpaI.
32P-labeled SV40 DNA (2 pg) was incubated with
HpaI(2U) inavolumeof 50 ,il ofabuffer containing
50mMNaCl, 6mMKCI, 15 mM Tris, 5 mMMgCl2,
1 mM,8-mercaptoethanol, and 1%glycerol (pH 7.5)
andincubatedat25°C. At the times indicated, 5,ul
was removed and the reactions were terminated.
Sampleswereelectrophoresedat100V,48 mA on a
1.4% agaroseslab gel (0.3 by30cm)at48 mA, 100V
for7h. DNAfragmentsweredetectedby
autoradiog-raphy. FragmentsA, B, and C represent the limit
digestfragments forHpaIdigestion ofSV40DNA,
whereas 80% and 58% representpartialdigest
frag-mentsproducedby cuttingthe genomeatonly2sites.
There is also a 62%fragmentwhich isnotresolved
fromthe58%fragmentonthisgel.
About 12% of the chromatin DNAappeared
as
partial
digestfragments-a
62%fragment
anda58%
fragment.
These partialdigest fragments
migrateveryclose-totheposition ofDNA Iand are notresolved in these gels. However,no sig-nificant amount ofDNA could be detected at the position of the 80%partial
digest fragment.
Since
cleavage
at 0.755 is required toproduce
the62and 58%
fragments,
itappeared that0.755 was moreprobable
as afirst(orsecond)
cleavage
site.
Todeterminethesite(s) ofthe firstcleavage, linear DNA III was eluted from the gelsafter digestionofSV40 chromatin by HpaI, andthe DNA IIIwasredigestedwithrestrictionenzymes whichcleaveSV40DNAat
only
onesite. Since thecleavage positionfor eachsingle-siteenzyme isknown,thepositionof the first sitecleavedby HpaI can be calculated from the size of theC
Control 1 5 15 30 120 360
Minutes at 370C
FIG. 10. HpaI digestion of SV40 chromatin.
32p-labeledSV40 chromatin
(3jg)
wasincubated at 37°CwithHpaI(1.5U) in 175IlIofabuffer containing75
mMNaCl,3mMKCI,0.7mM/3-mercaptoethanol,3
mMMgCl12, 13mM Tris(pH 7.5), 0.02% Triton X-100,
and 0.02mM EDTA. At various times between 1 and
120min, 25-,dsampleswereremoved andthe
reac-tionswereterminated.After120min, 1.0 Uof HpaI
enzymewasadded,andthe remainderofthe reaction
mixture wasincubatedfor anadditional4h. Asa
control,naked32P-labeled SV40DNAwasincubated
under similar reactionconditionsfor120min.
Sam-pleswereappliedto a/.4% agarose slabgel (22by0.3
cm)at45mA,100Vfor6h. DNAdigestion products
were detected byexposure ofthe slabgels toX-ray
films.
resulting DNA fragments. As a
control,
nakedSV40DNAwas
partially digested
withHpaI
tothestagewhere theDNAhad
only
asingle
chop. HpaI-derived DNA III from naked SV40 DNAandSV40 chromatinwere redigested with BamHI,EcoRI, andHpaII (Fig. 11). Inspection of therightpanelofFig.11shows thatthe initial cleavage of the SV40 chromatinby HpaI
wasnot at aunique site, forhad thisbeenthe case, redigestion with BamHI should have
yielded
onlytwodistinctfragments.
Instead,
redigestion
yieldedsixfragments,five shown in
Fig.
11(right panel,slotc)andonevery smallfragment
which can only be resolved on ahigher-percentage
u
0:00
on November 10, 2019 by guest
http://jvi.asm.org/
[image:10.507.54.245.78.315.2] [image:10.507.268.439.79.366.2]728 LIGGINS, ENGLISH, AND GOLDSTEIN acrylamide gel (notshown in Fig. 11). Indeed, comparison of the naked SV40 DNA III and SV40 chromatin DNA IIIdigestionpatterns
by
BamHI shows that thesepatterns arequite sim-ilar(Fig. 11,middle andright
panels,
slotc).Quantitative
analysis
of the BamHIrediges-tionof theHpaI-derivedDNA IIIrevealed that theinitial cut inthe naked DNA as wellas in thechromatin ismore
likely
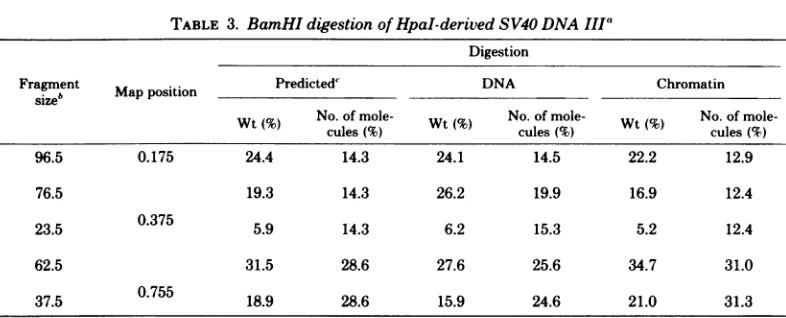
to occuratthe0.755 position thanatthe othertwosusceptible sites (Table 3). For theDNA, initialcutsoccurabout 1.5timesmorefrequentlyatthissite thanatthe othersites, whereasinthechromatin, initialcuts aremadeabout2.5timesmorefrequently than atthe other sites (Table 3). The 0.755position containstwoveryclosely spacedHpaI cleavage sites,sothat it is reasonable that the initialcuts should be morefrequent
in this region. For instance, ifit wereassumed thatcleavageoccurs withequalprobabilityateach ofthe four HpaI sites, there should have been twice as many DNA fragments generated by cleavage at the 0.755sites relativetoeachofthe othercleavage sites.However(Table 3), thefrequencyof cleav-age at the 0.755 site is significantly higher for cleavage of SV40 chromatin and significantly:'4a. Naked DNA
4 ?7C.ce o"i
.A I
J. VIROL.
less frequentfornaked SV40 DNA. These data indicate that theprobability ofcleavage is dif-ferent for each site and is different in naked DNAcompared with that in chromatin. Unfor-tunately, these experiments were done with a singlepreparation of SV40chromatin, and fur-ther work would be needed to determine the significance of these differences.
DISCUSSION
In thisstudy, thesusceptibility of SV40 chro-matintorestriction endonucleaseswasfoundto bedependentonthe ionic environment andon the topological configuration of the chromatin DNA. The SV40 chromatin employed in these experiments was devoid of Hi histones, so the observed
dependence
waspresumably
due to conformational changes between the nucleoso-mal histones and theSV40 DNA.Recent biochemical dataindicate that there are about 200DNA base pairs associated with eachSV40orpolyoma nucleosome andastretch of about50DNA basepairsconnecting adjacent nucleosomes (4, 6, 32,43). Consistent with this model isourfinding that only about 20% of the SV40 chromatin DNA was attacked by the
re-HpaIderived 'a de Naked DNAmA' : NA d e
I)NA
DNA11t DNA vit"sA
-_.
-VW
*ri. i.
i * tox
.)f!
FIG. 11. Cleavage ofHpaI-derivedDNAIII bysingle-site enzymes.32P-labeledSV40DNA III was removed
from preparativeagaroseslab gels after limit digestion ofSV40chromatin or after partialdigestion of naked
SV40 DNA. The DNAIII wasthendigestedbyeither EcoRI, HpaII, or BamHI in a buffer containing 70mM
NaCl,4mMKCI, 14mM Tris (pH7.5),3.5mM
MgCl,,
and 0. 7 mMII-mercaptoethanol.
DNA fragments wereseparatedon1.4% agarose slabgels.Left panel,SV40DNA digest products as markers: (a) untreatedDNA;
(b, c,andd) partial digestionbyHpaI; (e)HpaII digestion;
(t)
HpaI+ HpaII. Middle panel,HpaI-derivedDNAIIIfrom partial digestionof naked DNA: (a) untreated DNA; (b) EcoRIdigestion; (c)BamHIdigestion;
(d)HpaI digestion. Rightpanel,HpaI-derivedDNA IIIfromlimit digestion of chromatin: (a) HpaII digestion
(verylowactivity); (b) HpaIandHpaII(HpaII, very lowactivity); (c) BamHI digestion.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:11.507.121.406.346.565.2]TABLE 3. BamHIdigestion of HpaI-derivedSV40 DNA IIIa
Digestion
Fragment MapPOSitiOn Predicted DNA Chromatin
size' a oiinPeitd
Wt(%) No. of mole- Wt( NO.OfmOle Wt( No. of
mole-cules(% cules(%)ls %
96.5 0.175 24.4 14.3 24.1 14.5 22.2 12.9
76.5 19.3 14.3 26.2 19.9 16.9 12.4
23.5 0.375 5.9 14.3 6.2 15.3 5.2 12.4
62.5 31.5 28.6 27.6 25.6 34.7 31.0
37.5 0.755 18.9 28.6 15.9 24.6 21.0 31.3
aDatawerederived fromdensitometric tracings of gel patterns. Values represent averages for duplicate runs.
bFragment size represents percentage of the length of SV40 DNA. For each initial cut made by HpaI,
redigestion by BamHI will generatetwofragments. Thus, the 76.5 23.5 pair is generated by BamHI digestion
of DNA III terminatedatthe0.375mapposition, etc. A 3.5% fragment runs off this gel and is not included in
thisanalysis; thus, the96.5fragment is lacking its complement.
"Predictionswerebasedonthe assumption that each of the fourHpaI sites is cleaved with equal probability.
Since0.755represents adoublesite, cleavage at the site should be at twice the frequency of cleavage at the
othertwosites.
striction enzymes at 100 mM NaCl, suggesting that nucleosome-associated DNA was entirely resistant tothe enzymes atthis salt concentra-tion. However, ourkineticexperiments indicate that the susceptible DNA, presumably in the "spacer" regions between nucleosomes, didnot exist inastate
analogous
tofreeSV40 DNA.For example, inthe short timerequired to convert control naked SV40 DNA to its limit digest products,only about 4% ofthechromatinDNA wascleaved (Table1).Complete digestionof the susceptibleDNArequiredincubationtimes6 to 15timeslonger.Atlower salt concentrations, however,about 50% of the SV40 chromatin DNA wasdigested by the restriction enzymes. This value is in agreementwith values foundin studies in which chromatin was digested with nonrestricted en-donucleases (8, 9, 32, 41). Of this
susceptible
fraction, about half of the DNA was cleaved rapidly, i.e., duringatimeperiodcomparableto
that
required
for limitdigestion
ofequivalent
amounts of control naked SV40 DNA
(Fig.
3). From these experiments, we propose that the internucleosomal DNA has very little associa-tion with the histonesatlow saltconcentrations. In contrast, an incubation period at least 10 timesgreater wasrequiredtodigestthe remain-ingsusceptiblechromatin DNA.Thisfractionof thechromatin DNA may representsiteswithin nucleosomes whichareonly exposedatthelower saltconcentrations but nevertheless remainpar-tially
associated with histones. The fractionto-tally resistanttodigestionatthe lowersalt
con-centration probably contains sites within the
140-base-pair nucleosome core (1, 2, 10, 52, 53, 55, 57).
SV40 chromatin containing DNA in a topol-ogically relaxed configuration, i.e., with DNA having either a single- or a double-stranded break, was highly resistant tofurther cleavage by restriction enzymes. The resistance ofSV40 chromatincontaining nicked DNAwasrevealed in experiments with single-site enzymes, in which itwasfoundthat DNA IIinitiallypresent in chromatinfailedto be converted to DNA III (Fig. 3 and 4). Atthehigher salt concentration (100 mMNaCl) andatlowenzyme-to-chromatin ratios, the quantityof DNA IIactually increased during the course of the reactions (Fig. 2 and Table 1). In these experiments, the SV40 chro-matin used as asubstrate for the enzymes con-sisted
initially
of mixtures of chromatin with DNA I and II present, so that the possibility thatDNA II was anintermediatebeing formed and then converted to DNA III could not be ruledout.Nonetheless, kineticexperiments
suchas those shown in
Fig.
3 and 4 in which the fraction of DNA II remained essentially un-changedthroughoutprolongedincubationwhile DNA Iwasquantitatively convertedtoDNAIIIare notconsistent with suchamechanism.
Theconclusion thatasingledouble-stranded break can render SV40 chromatin resistant to
furtherdigestion by multiple-siteenzymes was
based onexperiments such astheoneshown in Fig. 5 in whichmostof the DNAwasconverted toDNA III and only 1% of the DNA could be detected atpositionscorrespondingtolimit di-gestfragments.Thesmallamountoflimit
digest
on November 10, 2019 by guest
http://jvi.asm.org/
730 LIGGINS, ENGLISH, AND GOLDSTEIN
products was not due to insufficient enzyme activity, sincenakedSV40DNA added to reac-tionmixtures before orduring thereactions was rapidly digested tolimit digestfragments (Fig. 8). In contrast to theseresults, Persico-DiLauro etal. (41)found that20 to25% ofthe chromatin DNA wasdigestedtolimitdigestfragments after exposure to HindIII. The discrepancy may be related todifferences in the purification of the chromatin and differencesinthe specificactivity andsource of enzymes.
Upon observing that most of the DNA in SV40 chromatin receivedonlyasinglecleavage after digestion with multiple-site enzymes, we attempted to determine whether this initial cleavage occurred at some preferred site(s) on the chromatin. Consequently, the initial cleav-age sites were mapped after digestion withHpaI. Subsequent experiments revealed that,innaked SV40DNAaswellas inchromatin, all potential siteswerecleaved,but notallsites were attacked withequalprobability (Table 3). For instance, it was shown that certain sites, specificallythose sites whichmap at position 0.755 on the SV40 genome, were cut at a much higher frequency thanthe other two sites.However,thefrequency wasnothighenough to justify a conclusion that these sites were devoid of histones. Had this been thecase,the ratio ofcutsat these sites to those atthe othersites should have been even greater, no DNA I ought to have remained un-digested, and limit digest fragments A and B should have beengenerated athigherlevels and in equal amounts (see Fig. 10
an4i
Table 3). Nonetheless,the results of ourexperimentwith HpaI demonstrate that thedistribution of his-tones in SV40 chromatin is not random. This conclusion is inagreementwith the data found for thedistribuition
ofhistones in polyoma chro-matin (44). It also raises the possibility that moreextensiveanalysisofchromatinby restric-tion enzymes might reveal certain sites which precludethebinding ofhistones.One type ofconformational change likely to render
internucleosomal
DNAresistant to diges-tion after relaxation of the SV40 DNA I in chromatin is one whichcauses the exposedDNA to associate with nucleosomes. Before relaxa-tion,SV40DNA I inchromatin is constrained in ahigh-energystate (64, 65), but upon relaxation the DNA canbecome free to rotate. In a model suggestedbyWorcel and Benyajati(63), succes-sive right-hand rotations of nucleosomes can causeinternucleosomal
DNA to coil bringing adjacent nucleosomes together. In this model, the spacer DNA forms a coil in the groove created by the contiguous nucleosomes. Alter-natively, it is conceivable that grooves exist orare formed upon rearrangement of the core structure. The extent ofcoiling andspacing of nucleosomes would of course depend on other factors,such as saltand divalent ion concentra-tion. An association between spacer DNA and nucleosomes similar to that proposed in this modelcanbe concluded fromrecent endonucle-ase studies which revealthat certain repeating elements found in nucleosomes extend intothe spacerregion (31, 44). Indeed, othershave sug-gestedthatnative chromatinordinarily existsin acompact structurewith nucleosomes almost in contact(15, 21,59).
The two proposed models ofchromatin, i.e., thecompactform and thebeaded-string config-urationappeartorepresent aconversionof one form to the other,dependingonthe conditions of observations. WithregardtoSV40chromatin, Griffith (21) observed thecompact form under physiological salt conditions and the beaded form atlow salt conditions. In
subsequent
ex-periments, Christiansen and Griffith(6) showed -thatmagnesium ions can condense SV40 chro-matinevenatlow salt concentrations intoaform which ressembles compact chromatin. This would suggest that, under the conditions em-ployed in our experiments (i.e., 1 to 10 mMMgCl2),
theSV40 chromatin became condensedwhen the DNA was relaxed, whereas, had the magnesium concentrations been reduced fur-ther, the internucleosomal DNAmayhave still remainedpartially extended. Inagreementwith this prediction, we noted above (see
Results)
that SV40 chromatinwas digestedto agreater extent atthe lower concentrations ofmagnesium (Fig. 6).
Additional data relate to the mechanism of action of the restrictionenzymes.Ruben and co-workers(47) and Modrich and Zabel (36) found that the reaction of EcoRI with DNA takes place in two distinct steps; a single-stranded break is made
rapidly
inonestrand,
followed by aslower reaction whichgeneratesabreakinthecomplementary
strand. When EcoRI wasre-acted withSV40 DNAat alow concentration of enzymerelativetoDNA,anicked-circular DNA IIintermediatewasgeneratedbeforeconversion to DNAIII (Fig. 2a). Incontrast, whenEcoRI was reacted with SV40 chromatin under the
same conditions, a significant amountof DNA II was generated, but itfailed to converted to DNA III. In agreement with this observation, Persico-DiLauroetal.(41)observed a 15 to 25% increase in DNAII
during
digestionwith EcoRI. In ourwork, however, this increase inDNA IIdidnotoccur athighconcentrationsofenzymes
(Fig. 3).If we are correct inassumingthat a nick inchromatin DNA can cause structuralchanges
J. VIROL.
on November 10, 2019 by guest
http://jvi.asm.org/
whichsequestercleavage sites,apossible
expla-nation for the failureto convertDNA IItoDNA
IIIatlowenzyme concentrations would be that
these changes occur before the enzyme can
cleave the complementary strand. We intendto
further investigate this possibility in more
de-tailed studies.
ACKNOWLEDGMENTS
We thank Michael Chenand Mary Gutai for theirgenerous
gifts of the Hindll+IIIenzymes,Abraham Worcel for his usefuldiscussions and hisinterest in this work, and Samuel Litwinfor his help with the analysisof the data.
This workwassupported by Public HealthServicegrant
no.CA11151from the National CancerInstitute. G.L.L. held Public Health Service fellowshipno. CA 05442, and D.A.G.
wasthe recipient of Public HealthService Research Career Development award no. CA 32425,both from the National Cancer Institute.
LITERATURE CITED
1. Axel, R.1975.Cleavage of DNA in nuclei and chromatin with staphylococcal nuclease. Biochemistry 14:2921-2925.
2. Axel, R.,W.Melchoir,B.Sollner-Webb, and G. Fel-senfeld.1974.Specificsites of interaction between
his-tones and DNA in chromatin. Proc. Natl. Acad. Sci. U.S.A.71:4101-4105.
3. Barban, S.,and R. S.Goor. 1971. Structuralproteinsof simian virus 40. J. Virol. 7:198-203.
4. Bellard, M., P. Oudet, J. E. Germond, and P. Cham-bon. 1976. Subunitstructureof simian virus-40 mini-chromosome. Eur. J. Biochem. 70:543-553.
5. Camerini-Otero,R.D.,B.Sollner-Webb,andG. Fel-senfeld. 1976. Theorganizationof histones and DNA in chromatin: evidence foranarginine-richhistone
ker-nel. Cell 8:333-347.
6. Christiansen, G.,and J. Griffith. 1977. Salt and divalent cation affect the flexiblenatureof the natural beaded chromatinstructure. Nucleic Acids Res. 4:1837-1851. 7. Christiansen, G.,T.Landers,J.Griffith,and P.Berg. 1977.Characterization ofcomponentsreleasedbyalkali disruptionof simian virus 40. J.Virol. 21:1079-1084.
8. Clark, R. J., and G. Felsenfeld. 1971. Structure of
chromatin. Nature (London) New Biol. 229:101-105. 9. Clark,R.J.,and G.Felsenfeld. 1974. Chemicalprobes
of chromatinstructure.Biochemistry13:3622-3628. 10. Compton, J.L., M.Bellard,and P. Chambon. 1976.
Biochemical evidence ofvariabilityin the DNArepeat
length in the chromatin of higher eukaryotes. Proc. Natl. Acad.Sci.U.S.A.73:4382-4386.
11. Cremisi, C., P. F. Pignatti, 0. Croissant, and M.
Yaniv.1976. Chromatin-likestructuresinpolyoma
vi-rusandsimianvirus40lytic cycle. J.Virol. 17:204-211. 12. Danna,K.J.,G. H.Sack, Jr.,and D. Nathans. 1973. Studies of simian virus 40 DNA. VII. Acleavagemapof theSV40genome.J. Mol. Biol. 78:363-376.
13. Estes, M. K.,E.-S. Huang, and J. S. Pagano. 1971. Structuralpolypeptidesof simian virus 40. J. Virol. 7: 635-641.
14. Fareed, G.,M.McKerlie,and N.Salzman. 1973. Char-acterization of simian virus 40 DNAcomponentII dur-ing viralDNAreplication.J. Mol. Biol. 74:95-111. 15. Finch,J.T.,M.Noll,and R. D.Kornberg.1975.
Elec-tronmicroscopyof definedlengthsof chromatin. Proc. Natl. Acad. Sci. U.S.A. 72:3320-3322.
16.Frearson, P.M.,andL. V. Crawford. 1972.Polyoma virusbasicproteins.J. Gen. Virol.14:141-155.
17. Friedmann,T., andD. David.1972.Structural rolesof
polyoma virusproteins.J.Virol. 10:776-782. 18. Germond, J.E.,B.Hirt, P. Oudet, M. Gross-Bellard,
andP.Chambon. 1975.Foldingofthe DNA double-helixinchromatin-likestructuresfrom simian virus40.
Proc. Natl.Acad. Sci. U.S.A. 72:1843-1847.
19. Gibson,W. 1974. Polyoma virus proteins:adescription ofthestructural proteins of the virions basedon poly-acrylamide gel electrophoresis and peptide analysis. Virology62:219-336.
20. Goldstein,D. A., M. R. Hall, and W. Meinke. 1973.
Properties of nucleoproteincomplexes containing
rep-licating polyoma DNA.J. Virol. 12:887-900.
21. Griffith, J. D.1975.Chromatinstructure:deduced from
aminichromosome. Science187:1202-1203.
22. Hall, M. R., W. Meinke, and D. A. Goldstein. 1973. Nucleoprotein complexescontaining replicating simian virus 40DNA:comparison with polyoma nucleoprotein complexes. J. Virol. 12:901-908.
23. Hedgpeth, J., H. M. Goodman,and H. W. Boyer.1972.
DNAnucelotidesequence restrictedby the RI endo-nuclease. Proc. Natl. Acad. Sci. U.S.A.69:3448-3452. 24. Hewish, D., and L. Burgoyne. 1973. Chromatin
sub-structure:thedigestion of chromatin DNAatregularly spaced sitesby nuclear deoxyribonuclease. Biochem. Biophys. Res. Commun.52:504-510.
25. Howe, C. C.,andK. B. Tan. 1977. Nucleoprotein
com-plexes from simian virus 40-infectedmonkey cells:
as-sociation withhistones and themajorviralstructural protein.Virology 78:45-56.
26. Huang, E.-S., M. K. Estes, and J. S.Pagano. 1972.
Structure and function ofthepolypeptidesin simian virus 40. I. Existence of subviral deoxynucleoprotein complexes. J. Virol.9:923-929.
27. Kornberg, R.D. 1974. Chromatinstructure:arepeating
unit of histones and DNA. Science 184:868-871. 28. Kornberg,R.D.,and J.0.Thomas. 1974. Chromatin
structure:oligomersofthe histones. Science 184:865-868.
29. Laemmli, U. K. 1970. Cleavage of structural proteins duringtheassemblyof the head ofbacteriophageT4. Nature(London)227:680-685.
30. Lake, R. S., S. Barban, and N. P. Salzman. 1973. Resolution and identification of thecore
deoxyribonu-cleoproteinsof the simian virus 40. Biochem.Biophys. Res. Commun. 54:640-647.
31. Lohr, D.,K.Tatchell,and K. E. Van Holde. 1977. On theoccurrence of nucleosome phasing in chromatin. Cell 12:829-836.
32. Louie,A. J. 1974. Theorganizationofproteinsinpolyoma and cellular chromatin. Cold Spring Harbor Symp. Quant.Biol. 38:259-265.
33. McMillen, J.,andR,A.Consigli.1974.Characterization ofpolyomaDNA-protein complexes.I.Electrophoretic identificationof theproteinsinnucleoprotein complex isolated frompolyoma-infectedcells. J. Virol. 14:1326-1336.
34. Meinke, W.,M. R.Hall, andD. A.Goldstein. 1975.
Proteins in intracellular simian virus 40nucleoprotein complexes: comparisonwith simian virus 40core
pro-teins. J. Virol. 15:439-448.
35. Mertz,J.E.,and R. W. Davis.1972.Cleavageof DNA by R,restriction endonucleasegeneratescohesive ends. Proc. Natl. Acad. Sci. U.S.A. 69:3370-3374.
36. Modrich, P.,and D. Zabel. 1976.EcoRIendonuclease. Physicaland chemicalpropertiesof thehomogeneous
enzyme.J. Biol. Chem. 251:5866-5874.
37. Mulder, C.,and H. Delius.1972.Specificityof the break
produced by restrictingendonucleaseR,in simianvirus
40DNA,asrevealedby partialdenaturationmapping. Proc. Natl. Acad. Sci. U.S.A. 69:3215-3219.
38. Noll,M. 1974. Subunit structure of chromatin. Nature
on November 10, 2019 by guest
http://jvi.asm.org/
732 LIGGINS, ENGLISH, AND GOLDSTEIN (London) 251:249-251.
39. Olins, A. L., and D. E. Olins. 1974.Sphericalchromatin units (vbodies).Science 183:330-332.
40. Oudet,P., M.Gross-Bellard,and P.Chambon. 1975. Electron microscopic and biochemical evidence that chromatinisarepeating unit. Cell 4:281-300.
41. Persico-Dilauro,M.,R. G. Martin, and D. M. Living-ston. 1977. Interaction of simian virus 40 chromatin with simian virus 40 T-antigen. J. Virol. 24:451-460. 42. Polisky,B., and B. McCarthy. 1975. Location of
his-tonesonSV40DNA. Proc.Natl.Acad.Sci.U.S.A. 72: 2895-2899.
43. Ponder, B. A. J., F. Crew, and L. V. Crawford. 1978. Comparisonof nucleasedigestionof polyoma virus
nu-cleoprotein complexandmousechromatin.J.Virol. 25:
175-186.
44. Ponder, B. A.J., and L. V. Crawford. 1977. The
ar-rangementofnucleosomesinnucleoproteincomplexes frompolvoma virus andSV40.Cell 11:25-49. 45. Qureshi,A.A., and P. Bourgaux. 1977. Distinct
non-structuralpolypeptidesinpolyomaandsimianvirus 40 DNA-protein complexes. Virology 77:418-420. 46. Roblin,R., E. Harle, and R. Dulbecco. 1971. Polyoma
virusproteins. I. Multiple virioncomponents.Virology 45:555-566.
47. Ruben, G.,P.Spielman,D. T.Chen-pei,E.Jay, B. Siegel,andR. Wu. 1977. RelaxedcircularSV40DNA
ascleavage intermediate oftworestriction
endonucle-ases.Nucleic Acids Res. 4:1803-1813.
48. Sahasrabuddhe, C.G., and K. E. VanHolde. 1974. The effect oftrypsinonnuclease-resistant chromatin
fragments. J. Biol. Chem. 249:152-156.
49. Seebeck, T.,and R. Weil. 1974. Polyoma viral DNA replicatedas anucleoprotein complexinclose
associa-tion with the hostcellchromatin. J. Virol. 13:567-576. 50. Sharp, P. A., W. Sugden, and J. Sambrook. 1973.
Detectionoftworestriction endonucleaseactivities in Hemophilusparainfluenza using analytical
agarose-ethidium bromide electrophoresis. Biochemistry 12: 3055-3063.
51. Shaw, B. R., J.L.Corden, C.G.Sahasrabuddhe, and K. E.Van Holde. 1974.Chromatographicseparation ofchromatinsub-units. Biochem. Biophys. Res.
Com-mun.61:1193-1198.
52. Shaw, B. R., T. M. Herman, R. T. Kovacic, G. S. Beaudreau, and K. E. VanHolde. 1976. Analysis of subunitorganizationin chicken erythrocytechromatin.
Proc.Natl.Acad.Sci.U.S.A. 73:505-509.
53. Simpson, R.,and J.Whitlock.1976.Chemicalevidence that chromatin DNA exists as 160 base pair beads interspersedwith 40 base pairbridges. Nucleic Acids Res.3:117-127.
54. Sollner-Webb,B.,R. D.Camerini-Otero,andG. Fel-senfeld. 1976. Chromatinstructure asprobedby
nu-cleases and proteases:evidence for thecentralrole of histones H3andH4.Cell9:179-193.
55. Sollner-Webb,B., and G. Felsenfeld.1975.A compar-isonof thedigestionofnuclei and chromatinby staph-ylococcal nuclease. Biochemistry14:2915-2920. 56. Tan, K. B.,and F. Sokol. 1972.Structuralproteinsof
simian virus40:phosphoproteins. J. Virol.10:985-994. 57. VanHolde,K. E.,C. G. Sahasrabuddhe, and B. R. Shaw.1974.Amodelforparticulatestructurein chro-matin. NucleicAcidsRes. 1:1579-1586.
58. Varshavsky, A. J.,V.V.Bakayev,P.M.Chumackov, and G.P.Georgiev.1976.Minichromosomeof simian virus40:presenceof histoneHI.Nucleic Acids Res. 3: 117-127.
59. Varshavsky, A. J., S. A. Nedospasov, V. V. Schmatchenko,V. V.Bakayev,P. M.Chumackov, and G.P.Georgiev.1977.Compactformof SV40viral minichromosome isresistanttonuclease:possible im-plicationsforchromatinstructure.Nucleic AcidsRes. 4:3303-3325.
60. Walter,G., R. Roblin, andR.Dulbecco.1972.Protein synthesisinsimianvirus40-infected monkey cells.Proc. Natl. Acad. Sci.U.S.A.69:921-924.
61. Wilson, G. A., and F. E. Young. 1975. Isolation of sequence-specific endonuclease (BamI) frombacillus amyloliquefaciens H. J. Mol. Biol.97:123-125. 62. Woodcock, C.L.F.1973.Ultrastructure ofinactive
chro-matin. J.Cell. Biol. 59:368a.
63. Worcel,A., andC. Benyajati.1977.Higher order coiling of DNAinchromatin. Cell12:83-100.
64. Vinograd, J., and J. Lebowitz.1966.Physical and to-pological propertiesofcircularDNA. J.Gen.Physiol. 49:103-124.
65. Vinograd, J.,J.Lebowitz, and R. Watson.1968.Early and late helix-coil transitions in closed circularDNA. The number ofsuperhelicalturnsinpolyoma DNA. J. Mol. Biol. 33:173-197.
66. Yang, R., K. Danna, A. Vande Voorde, and W. Fiers. 1975.Locationof thesmall restrictionfragments, Hind-L, Hind-M, andHpa-E, on the simian virus 40genome. Virology 68:260-265.
J. VIROL.