Copyright© 1977 American Society for Microbiology PrintedinU.S.A.
Encephalomyocarditis
Virus
RNA
II. Polyadenylic Acid Requirement
for
Efficient Translation
DENNIS E. HRUBY* AND WALDEN K. ROBERTS
Department of Microbiology and Immunology, University ofColorado Medical Center, Denver,
Colorado 80262
Receivedforpublication 7 March 1977
Differentiallypolyadenylated subpopulations ofencephalomyocarditis (EMC) viral RNA were isolated by affinity chromatography on oligodeoxythymidylic acid-cellulose. Translation of these RNA fractions in several in vitro protein-synthesizingsystems, isolated from Ehrlich ascites tumorcells, demonstrated thatpoly(A)+ EMC viral RNA wastranslatedtwo to three times moreefficiently than poly(A)- EMC viral RNA. Sodium dodecyl sulfate-polyacrylamide gel electrophoretic analysis ofthepolypeptidessynthesized by theinvitrosystemin response to the different RNAs showednodetectable differences in the size or relative amounts of the translational products. mRNA saturation curves indi-cated that the in vitro systems were stimulated maximally by equivalent amounts ofRNA, whether it be poly(A)- or poly(A)+ EMC viral RNA. Time course experiments showed that the differences in translatability were more pronounced late in the reaction when reinitiation was required, and that by eliminating reinitiation with high salt theapparenteffect ofpoly(A)on transla-tion wasdiminished. Together, these results suggest that poly(A) may be re-quired for efficient initiation and reinitiation ofprotein synthesis in the cell-freesystems. This interpretation is discussed relative to earlier data.
Many investigators have soughttoestablish acytoplasmic functions) forpolyadenylic acid [poly(A)], which is found covalently linked to the3'-terminusofmostRNAmolecules capable ofbeing translated byeucaryotic cells (4). Much ofthis work has been carried out in various eucaryotic cell-free protein-synthesizing sys-temsutilizing de-adenylatedmRNA as aprobe. These experiments have failed to discern any stringent requirement for
poly(A)
inthetrans-lation ofmRNA (2, 15, 27). This may, in part, be due to the
inefficiency
of the in vitro sys-tems being used or the conditions used for translation. Huez et al. (10) have utilized the Xenopus oocyte system,which has beenshown tocarry out translation of exogenously injected mRNA for long periods oftime, to show that poly(A) is important in maintaining the func-tional stability of globin mRNA. Also, recent careful analysis of ovalbumin message transla-tionhasshown that poly(A) facilitates the reu-tilization of this mRNA incell-free systems (6). Whether these effects are due to a protection of polyadenylated mRNA from nuclease degrada-tion, assuggestedby Hieter et al. (8), or rather a moredirect roleinproteinsynthesisis notyet clear.Wehave been interested in determining the role(s), ifany, of poly(A) in the replication of encephalomyocarditis (EMC) virus. To this
end, differentially polyadenylated fractions of EMC viral RNA have been isolatedand trans-lated in cell-freeextracts from Ehrlich ascites tumor cells. This approach has the advantage ofutilizing naturally occurring populations of homogeneous mRNAinwhich the only appar-ent differenceisthe lengthof the poly(A) moi-ety and translating these RNAs in a cell-free extract derived from cells that translate this message in vivo. The resultsreported here indi-catethat the presence of apoly(A)region stim-ulates the overall translation of EMC viral RNA two- tothreefold. Thisis inapparent con-trast toearlierwork onthe translation of polio-virus RNA (25)but is explicable dueto differ-ences inexperimental methods.
MATERIALS AND METHODS
Materials. [8-3H]adenosine (33 Ci/mmol),
[5-3H]uridine (28 Ci/mmol), and a 14C-labeled amino
acid mixture (algal profile) were obtained from
Schwarz/Mann; [32P]orthophosphate and
[35S]-methionine (525 Ci/mmol) were from Amersham/
Searle; oligodeoxythymidylic acid
[oligo(dT)]cellu-lose, T-3 grade, was from Collaborative Research,
Inc.; polyacrylamide gel electrophoresis materials
wereofelectrophoresis purity,purchasedfrom
Bio-Rad; Seakem agarose wasfromBauschand Lomb;
X-ray film and photochemicals were bought from
Eastman Kodak.
Virus growth. EMC viruswasgrown in
suspen-338
on November 10, 2019 by guest
http://jvi.asm.org/
sions of Ehrlichascitestumorcells andpurifiedas
previously described (9). The viral RNAwas
radio-actively labeled andextractedasbefore (9).
RNA analysis. Oligo(dT)-cellulose
chromatogra-phywasperformedessentiallyasdescribed by Aviv
and Leder(1), with certainmodifications(9).Intact
35S EMC viralRNAwasisolatedby centrifugation
through 11.5mlof5 to 20% sodiumdodecyl sulfate
(SDS)-sucrose gradients that were spun at 35,000
rpmfor4.75hat240CinaBeckmanSW41rotor(9).
Gel electrophoresis. (i) Agarose-acrylamide gels
(for RNA). Composite gelswere runaccordingtothe
methods ofLoening (14). Agarose(0.5%)-acrylamide
(1.8%)gelswerepolymerizedfor1h inacid-washed
glass tubes (15by 0.8 cm). Thegelsweresubjected
toelectrophoresis at5.5mA/gelfor10minin0.4 M
Tris-hydrochloride, 1 mM EDTA, 0.02 M sodium
acetate, 0.1% SDS (pH 5.5). RNA samples to be
analyzedweredissolvedin 251u of1mMEDTA,10
mM Tris-hydrochloride (pH 7.5), 0.2% SDS and
thendiluted1:1with50% sucrose-0.5%bromophenol
blue. Sampleswere heated at 850C for 2 min and
then applied immediatelytothe gel. Electrophoresis
was at 5.5 mA/gel for 3 to 5 h in running buffer.
After electrophoresis, gels were fractionated on a
Gilson gel slicerinto 2-mmslices. Sliceswere
incu-bated in0.5ml ofconcentrated ammonia overnight
andthencounted in10ml of Bray solution(5).
(ii) Polyacrylamide gel electrophoresis [for
poly(A)]. Polyacrylamide gels weresetupand run
accordingtoPeacock and Dingman(17). RNA
sam-plesweresubjectedtoelectrophoresison3-mm10%
polyacrylamide slab gelsat200Vat40C for4h inpH
8.3buffer. After electrophoresis, the gelwasfixedin
1 Macetic acid, stained with methylene blue, and
destained with water to locate nonradioactive
markers. The gelwasthenimpregnated with PPO
(2,5-diphenyloxazole) (3), driedundervacuum,and
putonKodakRoyal X-Omatfilm at-70°C for2to4
weeks. Theresulting autoradiogramwastraced
us-ingtheOrtec model 4310densitometer.
(iii) SDS-polyacrylamide gel electrophoresis (for
protein). Ten-microliter samples of cell-free protein
synthesis reactions were acetone-precipitated,
dried, dissolved in25
Al
of sample buffer, and boiledfor 5min. Sampleswerethenputon a10%, 1-mm,
0.1% SDS-polyacrylamide slab gel and subjectedto
electrophoresis at 100 V for 6 h (12). The gel was
thenstainedwith Coomassie blue anddestained in
methanol-acetic acid (5:7.5%,respectively). The gel
was then dried under vacuum and put on Kodak
Blue Brand film for 1to2weeks.
Cell-freeprotein synthesis. (i) Fractionated
sys-tem. An mRNA-dependentin vitroprotein
synthe-sizing system, consisting ofpurified ribosomes and
anammonium sulfatefraction, wasprepared from
Ehrlich ascites tumorcellsas previouslydescribed
by Sharma et al. (22). Reactions contained, in a
volume of 100 ,l: 20 mM Tris-hydrochloride (pH
7.5), 1mMATP, 0.2mMGTP,6mMcreatine
phos-phate (all adjusted to pH 7.5), 20 Mg of creatine
kinase,0.5 MiCiofa "C-labeledaminoacidmixture,
0.05mMeach of thesixnonradioactiveaminoacids
missing from the "4C-labeledaminoacidmixture, 6
mM /3-mercaptoethanol, 5 mM Mg(Ac)2, 120 mM
KCl,0.07opticaldensity(at260 nm)unitof
ammo-nium sulfate fraction, and 0.33 optical density (at
260 nm) unit ofpurified ribosomes. Followingthe
reaction, 0.2 mlof 0.1 N KOH was added, and the
tubes were incubated at370C for 20 min. A 1.0-ml
amountof cold 10% trichloroaceticacidwasadded,
and theprecipitate wascollectedonWhatmanGF/A
glassfiber filters.
(ii) S10-1 system (endogenous protein synthesis
reduced by preincubationof the cell-free extract).
Ehrlichascites cells from8-day tumors were washed
withphosphate-buffered salineto remove
erythro-cytes (20) andsuspendedto 5% inFischer medium
for leukemic cellsof mice (GrandIsland Biological
Co.),and the suspensionwasrotatedgentlyat370C
for 1 h to restore proteinsynthesis.The ascites cells
were then removed by centrifugation and used to
prepare an S10protein-synthesizing system, using
themethodpreviously described for L-cells (11).
Re-actions werecarriedout in atotal volume of 25 ul,
whichcontained:2.5 pulof S10 extract, 0.125
/Ci
ofthe amino acid mixtureplus the six nonradioactive
amino acids to 0.05 mM, 108 mM KCI, 5 mM
Mg(Ac)2, 20 mM HEPES(pH 7.6),7.2mM
,-mercap-toethanol, 4
,Ag
of creatine phosphokinase, 10mMATP, 1 mM GTP,6 mM CTP, and14mM creatine
phosphate. Following the reaction, 10-,ul samples
from each tube were spotted on Whatman 3 MM
filter squares andplunged into cold5%
trichloroace-ticacid. Filters were heated to 900C for 15 min and
washed twice with cold 5% trichloroacetic acid,
twice with cold ethanol, and twice with acetone.
Filters were dried and counted byliquid
scintilla-tion.
(iii) S10-2 system (endogenousproteinsynthesis
reduced by cellstarvation). We have found that by
incubating ascitescellsinnutrient-free mediumwe
canobtain an active S10 protein-synthesizing
sys-temwith low endogenous amino acid incorporation
without the usualpreincubation of the S10
superna-tant fraction. Ehrlich ascitescells were washed as
above andsuspendedto 3%in146mMNaCl-35 mM
Tris-hydrochloride (pH 7.5). The suspensionwas
ro-tated for 90 min at370C,and the cellswerecollected
by centrifugation. An S10 systemwas preparedas
above except that thepreincubation of theextract
was omitted. Assays were performed as with the
S10-1 system.
(iv) S10-3 system (endogenousproteinsynthesis
reduced bypreincubation withstaphylococcal
nu-clease). The S10-3 system wasprepared usingthe
S10-1 procedure, exceptthatinstead of
preincubat-ing the S10 extract at370C for45min, the extract
was preincubated briefly with staphylococcal
nu-cleaseaccording to the method of Pelham and
Jack-son(18).The S10-3 system wasassayedasabove and
showed low background incorporation and good
stimulation with EMC viral RNA.
RESULTS
Oligo(dT)-cellulose
chromatography.
We have previously shownthataffinity chromatog-raphyof EMC viral RNA onoligo(dT)-cellulose columns separates the RNA into three peak fractions, which differin bothpoly(A) content andbiologicalactivity (9). These threefractions 23, 1977on November 10, 2019 by guest
http://jvi.asm.org/
weredenotedaspeak1[poly(A)-] RNA, peak2
[oligo(A)] RNA, and peak 3 [poly(A)+] RNA andwereshowntocontainRNA molecules with
an averageof 16, 25, and 74nucleotides,
respec-tively, in their poly(A) tracts (9). In this
com-munication we will continue to use peaks 1
through3 asthedesignation for these
differen-tially polyadenylated subpopulations of EMC RNA.
The EMC viral RNA is notdegraded during fractionation on the oligo(dT)-cellulose
col-umns. The RNAs from peaks 1 through 3
mi-grate as a single band in agarose-acrylamide
gels (Fig. 1), thus corroborating the earlier SDS-sucrose analysis, which showed that the RNA from these fractions consisted of unde-gradedgenomic-length EMC RNA(9).
For this study we wanted to determine
whether peak 1EMC RNA contains auniform
population of RNAs with short or missing
poly(A) moieties,orwhether this RNA consists
ofpoly(A)- RNA together withaminor
contam-ination of poly(A)+ RNA sufficient to account
for the averagepoly(A) tractsizeof 16 nucleo-tides. Accordingly, [3H]adenosine-labeled peak
1 and3 EMC viral RNAswere treated witha
combination of RNases A and T1, and the
re-sistant fraction was isolated and run on 10%
polyacrylamide gel electrophoresis. Peak 3 EMC viral RNA poly(A)(Fig. 2B)ran as a
rela-tively homogeneous peak, with a mobility
slightly faster than that of 4S RNA. This is similartotheelectrophoretic mobility of polio-viruspoly(A) regions(24). Virtuallynopoly(A)
sequences weredetected in RNase-treated peak
1 EMC viral RNA (Fig. 2A). This eliminates thepossibility that peak1EMCviral RNA
con-mx0
DISTANCE(mm)
FIG. 1. Agarose-acrylamide gel electrophoresis of EMC viral RNA. Approximately100,000cpmofpeak
332P-labeled EMC viral RNA wassubjectedto elec-trophoresis through 0.5% agarose-1.8% acrylamide composite gels. Gels werefractionated, and the
ra-dioactivity ineach slicewasdetermined. The position
of the markerswassimilarly determinedon a
paral-lelgel using rRNA from Ehrlichascitestumorcells. Identicalpatterns were observed withpeak 1 and2
EMCviral RNA.
tainsafew longpoly(A) regions, whichaccount
for its residual infectivity, but does not prove
that this RNA is truly poly(A)-. One of the
stepsusedtoisolate the poly(A) regions is
bind-ingtobenzoylated cellulose (21), and it is possi-ble thatveryshortoligo(A)tractsfrom thepeak
1RNA donotbindtobenzoylated celluloseand
escape detection by this procedure.
Translation of oligo(dT)-cellulose-fraction-ated EMC viral RNA. Unlabeled EMC viral RNAwasfractionatedonoligo(dT)-celluloseas
usual. Peaks 1 through 3 were then run on
SDS-sucrose gradients, andtheRNA in the35S peakwasisolatedby ethanolprecipitation. The
three RNA fractions wereassayed by in vitro
protein-synthesizing systems prepared from Ehrlich ascitestumorcellsusing several
differ-entmethods. These different systemswere
uti-lized to investigate the possible effects of changes inpreincubation conditions, ions, and relative concentrations of translational
compo-nents on the translation of the EMC viral
RNAs.Reactionswerecarriedoutatsaturating
RNAconcentrations: 3.0 ,ug/assay for the frac-tionated system; 0.3 ,ug/assay for each ofthe
S10 systems. Regardless of the method of
sys-tempreincubation, reaction temperature, ionic conditions (Mg2+-spermidine), or presence of
hemin, peak 3 EMC viral RNA always stimu-latedtwo- tothreefoldmore radioactive amino
acid incorporation than peak 1 EMC viral RNA. Peak2EMCviral RNAwasusually 90to
E
c
0
0 v) LU u
z
0c
co
.5 .4 .3 .2
5S 4S a
.+ + a
I
20 4'0 6o 80 +
[image:3.499.290.437.405.560.2]DISTANCE (mm)
FIG. 2. Gel electrophoresis of EMC viral RNA poly(A) regions. [3H]adenosine-labeled poly(A)
re-gions were isolated from peak1 and 3 EMC viral
RNAby RNasetreatment,followed by chromatogra-phy on benzoylated cellulose (21). The poly(A)
re-gionswerethensubjectedtoelectrophoresison a10%
polyacrylamide gel along with 4S and 5S RNA markersfrom Ehrlich ascitestumorcells. Above, the densitometertracings of the autoradiogram are
dis-played. (a) Peak1EMC viral RNA; (b) peak 3 EMC
viralRNA.
48 28S 18S
4 4
36-
24-12
20 40 60 80 1
5 .4 .3 2 .I
IV 1.6
on November 10, 2019 by guest
http://jvi.asm.org/
[image:3.499.63.253.459.566.2]POLY(A) REQUIREMENT FOR EMC RNA TRANSLATION 341
100% as effective as peak 3 EMC viral RNA.
The data in Table 1 give the results ofseveral representative experiments. Anumber of other experiments were performed in which the
aforementioned variables were modulated,
in-cludingtranslation in the S10-2 and S10-3
sys-tems, and although the overall level of protein
synthesis was affected, the relative
translata-bilities of peak 1, 2, and 3 EMC viral RNA remained constant. This effect has been ob-served withEMC viral RNAs isolated from five
separate viruspreparations.
The nature of the polypeptides synthesized
by the S10 in vitrosysteminresponsetopeak 1, 2, and 3 EMC viral RNAwas investigated by
subjecting the 14C-labeled proteins to electro-phoresison a10%SDS-polyacrylamide gel. The
resultsof thisareshown in Fig.3.Little, ifany,
difference in the relative amounts or size of
products was detectable. This is in agreement
with earlier studies on the translation of
de-adenylated polioviral RNA (25).
Concentrations of RNA required for opti-mal translation. EMC viral RNA saturation experimentswerecarriedout toseeifthe
differ-encesintranslatabilities could beovercome by
simply addingmorepeak 1EMCviral RNAto
the cell-free system. As canbe seen in Fig. 4,
this was not the case. Radioactive amino acid
incorporation using 2.5 ,ul of S10 was
stimu-lated maximally by 0.3 ,ug of EMC viral RNA (12 ,ug/ml), regardless of whether itwaspeak1
orpeak 3 EMC viral RNA. Similar experiments werecarriedoutusing5 -lIof S10 (or the frac-tionated system), with identical results, the only difference being in the amount of RNA
requiredto reach saturation (5
Al
of S10 = 40pg/ml; fractionated system = 30 /Lg/ml).
At this point, it seemed possible that the lowered translatability of peak 1 EMC viral RNA wasdue tocontaminating host cell RNA
species, which alsosedimentedat35S.Tocheck this, EMC viruswasgrowninthepresenceof5
,tg
of actinomycin Dperml, which inhibitshostcell RNA synthesis, and labeled with [3H]uridine. The RNA was fractionated as
usual and thenrun on SDS-sucrose gradients.
Figure 5 shows that the radioactivity and ab-sorbancyat260nmprofiles of the 35S RNA in
peak 1, 2, and 3 EMC viral RNA were
coinci-dent. Theaveragespecific activities of the peak
fractions were virtually identical: peak 1 =
1,140cpm/gg,peak2= 1,170cpm/gg,and peak
3 = 1,190cpm/,g. Therefore, it appeared that
therewas little or no contamination of peak 1
EMC viral RNA by unlabeled host RNA
spe-cies.
Kinetics of translation. The incorporation of 14C-labeled amino acids into proteins in
re-sponse to peak 1 and 3 EMC viral RNA was
monitored with time (Fig. 6a). Translation of the two RNAswas foundto be similar during
the first 30 min of incubation; however, after this period the translation of peak 1 RNA slowedconsiderably, whereas the translationof
peak 3 RNA continued ata linear rate for an
additional 60 min, resulting in the eventual
two- to threefold difference in the extents of
translation. Similar kinetic patterns for the translation ofpoly(A)- ovalbumin RNA have been obtained by Doel and Carey (6). When
translation of the EMC viral RNAs was
initi-TABLE 1. Invitrotranslation ofoligo(dT)-cellulose-fractionatedEMC viral RNAa
In vitro Reaction conditions EMC viral RNA 14Cincorporated Stimulation
system
aC
Ions (mM) Fraction /19 (cpm) (fold)Fractionated 37 Mg2+, 5 0 5,100
K+, 120 Peak1 3 46,800 9.2
Peak2 3 128,700 25.4
Peak3 3 133,400 26.3
S10-1 30 Mg2+, 2 0 3,800
Spd, 0.3 Peak1 0.3 14,400 3.9
K+, 108 Peak2 0.3 26,500 7.2
Peak 3 0.3 29,100 7.9
S10-1 30 Mg2+, 5 0 400
K+,108 Peak 1 0.3 10,700 25.7
Peak2 0.3 22.800 54.6
Peak 3 0.3 23,400 55.9
a Oligo(dT)-cellulose-fractionated EMC viral RNAs were furtherpurified by zonal centrifugation and
translatedinseveraldifferentinvitro systemsprepared fromEhrlich ascitescells.Reactions wereallowed
toproceedfor 120 min, andthen14C-labeledaminoacidincorporation wasmeasuredasdescribedinthetext.
VOL. 23, 1977
on November 10, 2019 by guest
http://jvi.asm.org/
342 HRUBY AND
a
b
c
m
u
57-
45-EMC VIRAL RNA ADDED(Mg)
FIG. 4. RNA saturationcurves. Varyingamounts
ofpeak1 or 3EMC viral RNAwereaddedto25-piin
vitroprotein-synthesizingassays which contained 2.5
pIof S10-2. The reactionswerecarriedoutat5 mM
Mg2+-l08 mM K+ and30'C, for120 min. Symbols:
(0),peak3EMC viralRNA; (a), peak1EMC viral
RNA.
34-
23-FIG. 3. SDS-polyacrylamide gel electrophoresis
analysis of viral polypeptides synthesized in vitro. Peak 1 (a), peak 2 (b), or peak 3 (c) EMC viral RNA
(0.3pg) was added to a 25-
gl
protein synthesis assaywhich contained 2.5 pl of S10-1. After synthesis,
approximately 10,000 cpm of each reaction mixture
wasanalyzed by10%SDS-polyacrylamide gel
elec-trophoresis. Above is displayed the autoradiograph of the polypeptides synthesized in response to the added viral RNA. Numbers refer to the molecular
weights of marker proteins (xlO-3). Note that the
largest newly synthesized viral protein corresponds to
anapproximate molecular weight of 140,000.
ated asusual, followedbyshiftingto high-salt reaction conditionsto permit polypeptide elon-gation but prevent further initiation and
reini-tiation events (26), much ofthis difference in
theextentsoftranslation disappeared (Fig. 6b).
DISCUSSION
We have never failed to observe a two- to
threefold difference inefficiency of translation
25-
20-
15-
10-
5-
75-
60-01
C
45-X.
30-I
15-
50-
40-
30-
20-10.
.1.5
.1.0
.0.5
.0
*t
<
> z
.1.5 > a
.1.0 <
0
CLQ
.1.5
1.0
o.0
5 10 15 20 25
FRACTION NUMBER
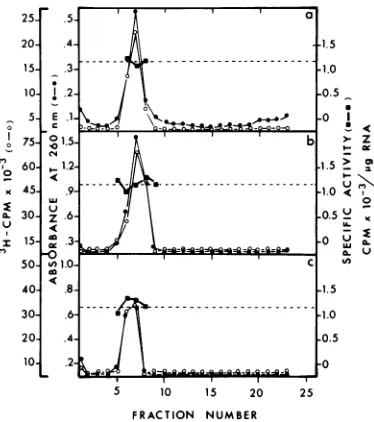
FIG. 5. Specific activity of [3H]uridine-labeled
EMC viral RNA oligo(dT)-cellulose fractions.
[3H]uridine-labeled EMC viral RNA was
fraction-atedon oligo(dT)-cellulose. Eachof the three
frac-tions wasthenanalyzed onSDS-sucrosegradients.
Each gradient was analyzed for absorbance at 260
nm
(A26&)
and trichloroaceticacid-precipitableradio-activity.Specificactivities werecalculatedassuming
1A260unitequals40pgofEMC viral RNA.(a)Peak
1 EMC viral RNA; (b) peak2EMCviralRNA; (c)
peak3EMC viral RNA.
betweenpoly(A)+ andpoly(A)- RNAs isolated from five different preparations of EMCvirus.
This differencewasobservedunderavarietyof ionic and temperature conditions used for
.-o
E .1_-%o_ f~
.8-C4
.2 -
--2-L
__
on November 10, 2019 by guest
http://jvi.asm.org/
[image:5.499.265.460.62.223.2] [image:5.499.83.231.65.433.2] [image:5.499.268.455.300.511.2]REQUIREMENT FOR EMC RNA TRANSLATION 343
0
x
I 0r
0.6
b)4
2
0-03
INCUBATION TIME (h) FIG. 6. Timecourse of EMC viral RNA
transla-tion. A 0.6-pgamountof peak1 (a)orpeak3 (0)
EMCviral RNA was addedto a50-p/X S10-1 assay
containing 5 piof S10. At theindicated times,
3-t1
samples were removed and processed to determine
hottrichloroacetic acid-precipitableradioactivity. (a)
Reactionrun at90 mMKCl; (b) reactionstartedat
90 mMKCl. At 15 min the KCl concentration was
shiftedto155 mM.
translation (Table 1). From the mRNA
satura-tioncurvesandspecific activity analysis itwas
evident that this difference could not be
ex-plained by thepresenceofcontaminatingRNA
species in the peak 1EMC viral RNA fraction (Fig.4and 5). Also,electrophoresis under dena-turing conditions showed the RNAstobe unde-graded (Fig. 1).
Thereare atleasttwopossible explanations
forthe reducedtranslatability ofpoly(A)-EMC
viral RNA. First, itmay be that
poly(A)-defi-cient RNAis moresusceptibletonuclease deg-radation (8) and therefore hasalowered
proba-bility of remaining intact and interacting
pro-ductively with the protein synthetic
machin-ery. Inpreliminary experiments,wehave been
unabletodetectanyincreasedsusceptibility of
thepoly(A)- RNAtoexonucleolytic or
endonu-cleolytic attack during the protein synthesis
incubation. However, a more subtle role of the poly(A) moiety, such asaffecting the degrada-tion ormodification of the5'initiation region of the viral RNA, cannotbe ruled out. A second possible explanation for the reduced translata-bility ofthe poly(A)- RNA is that poly(A) di-rectly influences one of the protein synthetic reactionsrequiredforinitiation, elongation, or termination. Since the patternsof polypeptides synthesizedinresponse topeak 1, 2, and 3 EMC viral RNA are identical (Fig. 3), this would indicate that the termination process is not af-fected.Also, thedecreased differentialbetween thetranslationsof poly(A)+and poly(A)-RNAs at early times or under high-salt conditions (Fig. 6) suggests that elongation is notinvolved inthe discrimination between the RNAs (13). It seems most likely that the reduced transla-tional capacityof poly(A)- EMC viral RNA is due to some impairmentwith the initiation or reinitiation process (6). Thisimpairmentcould resultinanincreasedfunctional inactivationof
the
poly(A)-
RNA wheneverit is not engagedwith ribosomesinproteinsynthesis, leadingto an accumulating inhibition of protein synthe-sis, which becomes increasingly apparent late intranslation.
There are several possible reasons why we have been able to demonstrate a reduction in the translational capacity of a poly(A)- mes-sagewhen similar analyses by others(2, 25, 27) have not. One possibility is that the Ehrlich ascites tumor cell-free system carries out pro-tein synthesis for long periods oftime in re-sponse to EMC viral RNA. Incorporation of radioactive amino acids continues for more than2h at 30'C (Fig. 6). This is in contrast to 30 to 60 min of linear incorporation that has beenmonitoredinother studies (2, 15). Huez et al. (10)have been themostsuccessfulin estab-lishinga role forpoly(A) inthe translation of mRNA,and thisisprobably duetothe fact that their studies have been done in the Xenopus oocyte system, which carries out manyrounds of translation.Thisis verydifferent frommany cell-free systems, such as the wheat germ em-bryosystem, which are veryinefficientat reini-tiating new rounds ofsynthesis (19). The Ehr-lichascites systemusedhere continues to reini-tiate newrounds ofsynthesis formorethan90 min (D. E. Hruby, unpublished data), which would facilitate detection ofaneffectofpoly(A) on the accumulating inactivation of mRNA. This is in contrast to themethodsofSpector et al. (25), who allowed initiation of poliovirus RNA toproceedforonly 15 min before shifting toahigher potassiumconcentration, which per-mits elongation but inhibits initiation. Using theirprocedure,we toocandetect little effect of
VOL. 23, 1977
on November 10, 2019 by guest
http://jvi.asm.org/
[image:6.499.63.216.53.367.2]poly(A) on the translation of viral RNA (Fig. 6b).
Inall of the reactions where it wasassayed, peak 2 EMC viral RNA was nearly as active as peak 3 EMC viral RNA. This was true even if peak2waschromatographed asecond time. It may be, as suggested for globin mRNA (16), that thereis a critical sizeforpoly(A)function. For globin mRNA this was estimated to be 32 nucleotides. This agrees fairly well with our data, aspeak 2 EMC viral RNA has previously been estimated to contain an average of 26 nucleotides in its poly(A) region (9).
Lowered infectivity of poly(A)- picornaviral RNA molecules has previously been observed by us (9) and others (7, 23). It is not known whether this effect of the poly(A) moiety on infectivity is at the level of penetration, protec-tion against nucleases, translation, or tran-scription. The loweredefficiency of translation ofpeak1EMC viral RNAmay, atleastinpart, explain its lowered infectivity. Currently, re-search is in progress to find conditions that potentiate the reducedtranslationalcapacityof peak1EMCviral RNAinordertobetterassess therole ofpoly(A) intranslation and infection.
ACKNOWLEDGMENTS
We thank0. K. Sharma for help in preparing the frac-tionated cell-free system. We also thank J. L. Brown for use ofthe Gilson gel crusher.
This work was aided by grant no NP-200 from the Ameri-canCancer Society.
LITERATURE CITED
1. Aviv, H.,and P. Leder. 1972.Purification ofbiologically activeglobin messenger RNAbychromatographyon
oligothymidylicacid-cellulose. Proc. Natl. Acad. Sci. U.S.A. 69:1408-1412.
2. Bard, E., D. Efron, A. Marcus, and R. P. Perry.1974. Translational capacity of deadenylated messenger RNA. Cell 1:101-106.
3. Bonner, W. M., andR. A.Laskey. 1974. A film detec-tionmethod for tritium-labeled proteinsand nucleic acidsinpolyacrylamide gels. Eur.J.Biochem. 46:83-88.
4. Brawerman, G.1976. Characteristics andsignificance ofthe polyadenylate sequence inmammalian mes-sengerRNA.Prog. Nucleic Acids Res.17:118-148. 5. Bray, G. A. 1960. A simple efficientliquidscintillator
forcounting aqueous solutions inaliquid scintilla-tioncounter. Anal.Biochem. 1:279-285.
6. Doel, M. T., and N. H. Carey. 1976.The translational capacityofdeadenylated ovalbumin messengerRNA. Cell 8:51-58.
7. Goldstein, N.O., I.U. Pardoe, andA.T.H. Burness. 1976.Requirement of aadenylic acid-rich segmentfor theinfectivityofencephalomyocarditisvirus RNA. J. Gen.Virol. 31:271-276.
8. Hieter, P.A., S. M.LeGendre, and C. C. Levy. 1976. Stabilization of an RNA molecule by 3'-terminal poly(A)-inducedinhibition ofRNaseactivity.J. Biol. Chem.251:3287-3293.
J. VIROL.
9. Hruby, D. E., and W. K. Roberts.1976.
Encephalomyo-carditisvirusRNA: variations inpolyadenylic acid contentand biologicalactivity. J. Virol. 19:325-330. 10. Huez, G., G. Marbaix, E. Hubert, M. Leclercq, M. Nudel, H. Soreq, R. Salomon, B.Lebleu,M.Revel, and M. Z. Littauer. 1974. Role of thepolyadenylate segmentinthe translation ofglobinmessengerRNA inXenopus oocytes. Proc. Natl. Acad. Sci. U.S.A. 71:3143-3146.
11. Kerr, I. M., R. E. Brown, M. J. Clemens, andC. S. Gilbert. 1976. Interferon-mediated inhibition of cell-free-protein synthesisinresponsetodouble-stranded RNA. Eur. J. Biochem. 69:551-561.
12. Laemmli, M. K. 1970. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature (London) 227:680-685.
13. Lodish, H. F. 1976. Translational control ofprotein synthesis. Annu. Rev. Biochem. 45:39-72.
14. Loening, U. K.1967.Thefractionationof high-molecu-lar-weight ribonucleic acid by polyacrylamide-gel electrophoresis. Biochem. J. 102:251-257.
15. Muller, W. E. G., G. Seibert, R. Steffen, and R. K. Zahn. 1976. Endoribonuclease IV.2.Further investi-gation on the specificity. Eur. J. Biochem. 70:249-258.
16. Nudel, U., H. Soreq, U. Z. Littauer, G. Marbaix, G. Huez, M. Leclercq, E. Hubert, and H. Chautrenne. 1976. Globin mRNA species containing poly(A) seg-ments of differentlengths:theirfunctional stability inXenopus oocytes. Eur. J. Biochem. 64:115-121. 17. Peacock, A. C., and C.W.Dingman.1967.Resolution
ofmultiple ribonucleicacidspeciesbypolyacrylamide gelelectrophoresis.Biochemistry 6:1818-1827. 18. Pelham, H. R. B., and R. J. Jackson.1976. Anefficient
mRNA-dependent translation system from reticulo-cytelysates. Eur. J. Biochem. 67:247-256.
19. Roberts, B. E., and B. M. Patterson. 1973. Efficient translationof tobacco mosaic virus RNA and rabbit globin 9SRNA in acell-freesystem fromcommercial wheat germ. Proc. Natl. Acad. Sci.U.S.A. 70:2330-2334.
20. Roberts, W. K.1965.StudiesonRNAsynthesisin Ehr-lich ascites cells: extraction and purificationof la-beled RNA. Biochim.Biophys.Acta108:474-487. 21. Roberts, W. K. 1974. Use ofbenzoylated cellulose
col-umns for theisolationofpoly(adenylic acid) contain-ingRNA and otherpolynucleotideswith little second-ary structure. Biochemistry 13:3677-3682.
22. Sharma, 0. K., W. K. Roberts, D.N.Beezley, andE. Borek.1975. AtransferRNA-dependent protein syn-thesizing system from Ehrlich ascites extracts. Biochim. Biophys. Acta 390:327-331.
23. Spector, D. H., and D. Baltimore. 1974. Requirement for3-terminalpoly(adenylicacid)for theinfectivity ofpoliovirus RNA. Proc. Natl. Acad. Sci. U.S.A. 71:2983-2987.
24. Spector, D. H., and D. Baltimore. 1975. Polyadenylic acid onpoliovirusRNA. II.Poly(A)onintracellular RNAs.J. Virol. 15:1418-1431.
25. Spector, D. H., L. Villa-Komaroff, and D. Baltimore. 1975. Studies on the function of polyadenylic acidon poliovirus RNA.Cell 6:41-44.
26. Villa-Komaroff, L., N. Guttman, D.Baltimore, and H. F. Lodish. 1975. Complete translationofpoliovirus RNA in a eukaryotic cell-free system. Proc. Natl. Acad. Sci. U.S.A. 72:4157-4161.
27. Williamson, R.,J. Crossley, and S. Humphries.1974. Translation ofmouse globin messengerribonucleic acid from whichthepoly(adenylicacid) sequencehas been removed.Biochemistry13:703-707.