JOURNAL OFVIROLOGY, Oct. 1971, p. 455-468
Copyright©1971 AmericanSocietyforMicrobiology
Vol.8, No. 4 Printed in U.S.A.
Heat
Induction
of Prophage 4105 in
Bacillus subtilis:
Replication of the Bacterial
and
Bacteriophage
Genomes
RICHARD W. ARMENTROUT AND LARS RUTBERG
DepartmentofBacteriology, Karolinska Institutet,S-10401Stockholm60,Sweden
Received for publication 14 June 1971
A temperature-inducible mutant oftemperate Bacillus bacteriophage 4105 was
isolated and used tolysogenize a thymine-requiring strain of Bacillus subtilis 168.
Synthesis of phage and bacterial deoxyribonucleic acid (DNA) was studied by
sucrose gradientcentrifugation and density equilibrium centrifugation of DNA
ex-tracted from induced bacteria. The distribution of DNA in thegradientswas
meas-ured by differential isotope and density labeling of DNA before and after induction
and by measuring thebiological activity of the DNA in genetic transformation, in
rescue ofphage markers, and in infectivityassays. At early times after induction,
but after at least oneround of replication, phage DNA remains associated with
high-molecular-weight DNA, whereas, later in the infection,phage DNA is
associ-ated with material ofdecreasing molecular weight. Genetic linkage between phage
and bacterial markerscanbedemonstrated inreplicated DNA from inducedcells.
Prophage induction is showntoaffectreplication of the bacterial chromosome. The
overallrateofreplication ofprelabeled bacterial DNA is identical in
temperature-induced lysogenics and in "mock-induced" wild-type 4105lysogenics. Therateof
replication of the bacterial marker phe-l (and also of nia-38), located closetothe
prophage in direction of the terminus of the bacterial chromosome, is increased in
induced cells, however, relativeto other bacterial markers tested. In
temperature-inducible lysogenics, where the prophage also carries a ts mutation which blocks
phage DNA synthesis, replication of both phage and bacterial DNA stops after
about 50% of the phage DNA has replicated once. The results of these
experi-ments suggestthat theprophage isnot initially excised in inducedcells,butrather
itisspecifically replicatedinsitutogether with adjacentpartsofthebacterial
chro-mosome.
The Campbell model (5) provides an elegant solutionto the
problem
ofintegration
andexci-sionoftemperate phage genomes.
Integration
isproposed
to occur through a reciprocal recom-binational event between aspecific region
on acircularized
phage chromosome and aspecific
attachment site on the bacterial chromosome. Prophage
excision
is thought to represent a reversalof
theintegration
event. There isstronggenetic
evidence forspecific integration
enzymes coded forby phagesP2(6),P22 (22),and lambda(8),aswellasanother enzyme, xis, in lambda (11)
which together with the int product promotes excision ofa resident prophage. Excision is
be-lievedto be theprimaryevent inprophage
induc-tion. This is supported by experiments which
show, forexample, that genetic linkage between
bacterial markers bracketing the prophage
in-creases after prophage induction
(13)
or thatprophage
deoxyribonucleic
acid(DNA)
can be recoveredunreplicated
in matureparticles
after heteroimmunesuperinfection
(15).
The primary DNA productafter induction is thoughtto bea circularized DNA moleculeequivalent
in size tomature
phage
DNA.Presumably,
excision
oflambda prophage
requires
protein
synthesis but notDNAsynthesis
(24).DNA extracted from Bacillus
bacteriophage
4105
particles is a homogenouscollection
ofmoleculesof lowinfectivity (3, 4, 18). Thepoor infectivity reflects some structural characteristic of the ends ofthe molecules
(17).
To study the structuralbasis
forthebiological
activityof4105
DNA, we sought to isolate excised prophage DNA fromlysogenic
bacteria carrying a tem-perature-inducible mutant of4105
as prophage. 455on November 11, 2019 by guest
http://jvi.asm.org/
The results of these experiments indicate, how-ever,that prophage replication rather than
pro-phage excision is the primary event associated
with heat induction of 4105 lysogenic bacteria.
The experiments alsosuggest a common
mecha-nism of control of replication of both bacterial
andphage DNA in induced cells.
MATERIALS AND METHODS
Bacteriaandphage.The bacterial strainsemployed
arelisted in Table 1. Someofthesestrainswereused
as recipients in transformation experiments, and the
map order of the markers usedonthebacterial chro-mosomeisorigin-purA16-leu-2, leu-S, -ilvAJ-prophage ¢105-phe-J-nia-38-metB5-terminus (7, 16).
Phage
0105
and the temperature-sensitive (ts) andsuppressor-sensitive (sus) mutants employed have
recentlybeen described (2). As aconvenientmarker
of low background, susl9 in gene C was used in
markerrescueexperiments.MutationtsN31ingeneK
(2) isreferredtohereasKts3l.Ithasbeenpreviously
shown that geneK is essential for phageDNA syn-thesis (2). The temperature-inducible mutant cts23
was isolated after treatment of infected cells with
N-methyl-N-nitroso-N'-nitroguanidine (2). The cts23
mutation mapsclose to susil ofgene J (unpublished
data). Lysogenicderivatives ofT bacteria were
iso-lated from the central growthin plaques formedon
these bacteria at 30 C. The growth rates of T- and
its lysogenic derivatives are identical in the media
employed; in Min-CH (see below) at 30C, the
dou-blingtime ofthese strains is about 60min.
Media and growthofbacteria andphage. Bacterial
strains were maintained on Tryptose Blood Agar
Base (TBAB, Difco) plates. Bacteria were grown in
Spizizen's minimal medium (23) supplemented with
0.05%'o
casein hydrolysate (Difco) and 20 MAg(per ml)ofany amino acid required and 10 ,ug ofthymidine
per ml when required. We refer to this medium as Min-CH. Phage assays and preparation of phage
stocksweredoneas described(2, 16, 18).
Assaysof the"biological activity"ofDNAsamples.
The relativeamounts of bacterialgenes, phagegenes,
andcomplete phage genomes present inDNA
sam-plesweremeasuredbytransformation, marker rescue,
ancd ttatisk;ction, respectively. For these
measure-metiEs, competentcells were obtainedby the method
ofAnagnostopoulosandSpizizen (1). Transformants
weteassayed byspreading0.1 ml of appropriate
dilu-tions of the competent culture on selective media
atterexposureofthe cellsto a dilutionofthe DNA
sample (2, 18). Inmarkerrescueexperiments,
compe-tentSR135 cellswereexposedtoaDNAsample,and,
20min after additionofDNA, thecells were
super-infected withan excessofmutantphage k105 CsusJ9.
Bacillus subtilis SR 135 carries a suppressor gene
which permits the superinfecting phage to grow and
recombine withphage DNA taken up bycompetent
cellsfromthe DNA sample. Priortolysis, the
super-infected cells were diluted and 0.1-ml samples were
plated with W168asindicator; W168doesnotpermit
growth of the superinfecting mutant phage. Details
ofthe methodhave recentlybeendescribed byus (2,
18). Assayofcomplete phagegenomes wasby
trans-fection as described (2, 18). Competent cultures of
SR135 were exposed to an appropriate dilution of
DNA sample, and prior to lysis the bacteria were
diluted and 0.1-ml samples were plated by using
SR135 as indicator bacteria. The number of plaques
that appearedontheplateswastakenas ameasureof
thenumber of whole "active" phageDNAmolecules
takenupbycompetentcells.
By use of these assays, the distribution of
trans-forming activity, marker rescue, and infectivity was
determinedoveragiven gradient.Thecompetenceof the different strains used in theseassays varied, and the results obtainedrepresentrelativeactivities rather
than specificactivities (e.g.,transformantsper
micro-gram of DNA corrected for constant competence).
The dataare presented onconvenient scalesto show
the distributions of activities overthe gradient, and
thescales neednotbe thesamefor allgraphs.
Inductionofheat-induciblelysogenicbacteria.
Lyso-genicbacteriaweregrownin20mlof Min-CHat30 C
toa density ofabout 5 X 107bacteria perml. For
prelabeling, 10,MCiof3H-thymidine (specific activity,
5Ci/mmole; The RadiochemicalCentre,Amersham)
wasaddedper20 ml inadditionto10,ugofcold
thy-midine per ml. The cells were centrifuged, washed
once with Min-CH, and suspended in0.5 volume of
fresh, thymineless Min-CH at 45 C. After 5 min at
TABLE 1. Bacterial strains
Strain Relevantproperties4 Source
W168 Prototrophic, suI J.
Spizizen
SR135 spoA9, trp-7, su+3 J. A. Hoch
BR26 trpC2, leu-2 J.
Spizizen
BR95 trpC2,phe-1, ilvAI J.
Spizizen
BR95 (l105
AtsJ5)
trpC2,phe-J,
ilvAl lysogenic for l105carrying
mu- Ourcollection tation tsNISBD25 purA16, leu-5, nia-38,metB5 A. Lindahl-Adams
168T- trpC2, thyAl, thyBI C.
Anagnostopoulos
aProperties:su+,carries thesu3suppressor gene; su-,
nonsuppressing,
spo,asporogenous; trp,trypto-phan; phe, phenylalanine; ilv, isoleucine-valine; pur, purine
(adenine);
leu,leucine,
nia, niacin, met,methionine, thy, thymine.
on November 11, 2019 by guest
http://jvi.asm.org/
[image:2.489.60.452.509.614.2]PROPHAGE
0105
IN B. SUBTILlSthistemperature, the culturewasshiftedto42 C and
an equal volume ofprewarmed Min-CH containing
20 ,ug of thymidine per ml was added. For density
labeling at42C,thymidine was omitted and instead
theMin-CHcontained20 ,ugof5-bromodeoxyuridine
(BUdR) perml(Sigma Chemical Co., St.Louis, Mo.)
and '4C-thymidine (specific activity, 58 mCi/mmole;
The Radioactive Centre, Amersham) to give afinal
concentration of 0.25 MCi per ml. Thebacteria were
incubated at 42 C with aeration, and samples were
withdrawn for phage and bacterial assays and for
extraction of DNA. T- bacteria lysogenic for
0105
wild type are not induced by temperature shifts.
WhenT-
(0105)
wastreatedasdescribed above, it isreferredto asmock-induced.
Extraction of DNA. Culture samples containing
2 X 108 to5 X 108 bacteria werepouredoverfrozen
Min-CH andcentrifuged.Thepelletsweresuspended
in2 ml of 0.2 M NaCl-0.001 M
ethylenediaminetetra-aceticacid(EDTA) pH8(saline), lysozymewasadded
to afinal concentration of 200
,g,/ml,
and thesus-pensionwasincubatedat37 Cfor5 to10min. When
lysis occurred,sodiumdodecylsulfatewasaddedto a
final concentration of about 0.5%. Afterstanding a
few minutes at room temperature, 2 ml of washed
[0.1 M tris(hydroxymethyl)aminomethane
(Tris)-0.01 M EDTA, pH 8] phenol was added. DNA was
extracted bygentle mechanicalrolling of the sample
at roomtemperature for 30 min(3). Thesamplewas
thencentrifugedandthe aqueousphasewas removed
with a "U"-shapedPasteur pipette. The DNA
solu-tionsweredialyzedat4Cfor 16 hragainstonechange
of about1,000 volumesofsaline. 14C-labeledT7 DNA
was prepared asdescribed (3); 32P-labeled P2 DNA
wasagiftfrom Jon Jonasson.
Sucrose gradients. All sucrose gradients were 5 to
20% sucrose in 0.05 MTris(pH 8.0)-0.001 M EDTA,
made up from boiled-stock solutions. Gradients
werecentrifugedat 14CinaSpinco modelL
centri-fuge. The SW50rotorwas run at 35,000rev/min for
150min, whereasthe SW25.1 rotorwas run at24,000
rev/min for 180 min.Fractionswerecollectedthrough
a needle inserted into the bottom of the tubes.
Ten-dropfractions werecollected from the SW50
gradi-ents, and 45-drop fractions were collected from the
SW25.1 gradients. In the case of SW50 gradients,
0.05 ml from eachfractionwasaddeddirectlyto5ml
ofdioxane-based scintillation fluid (3) and counted
forradioactivity. Samples (0.1 ml) from the SW25.1
gradientswereapplied tosquaresof Whatman no. 3
filter paper, placedin cold 10% trichloroaceticacid,
washedoncewithcoldethanol,and thendriedunder
vacuum. The radioactivity on the filter papers was
counted in 5 ml oftoluene-based scintillation fluid
(3).Inall cases, thescintillation vials were precounted
and the background radioactivity for each vial was
subtracted from the sample counts. The biological
activityof thefractionswasassayed.
Equilibrium densitycentrifugation.To 2mlof DNA
samplewasadded1ml ofsaline and 3.8 g of CsClto
givea
56%7
(w/w) solution. The samples werecentri-fugedat14 CinanSW50rotorat 31,000rev/min for
about 64 hr. Fractions ofeight drops were collected
asdescribed above. The refractive index of every fifth
fractionwasread on anZeissAbberefractometer. In
each case, alineardensity gradientwas obtained.To
each fraction was added 1 ml ofboiled saline, and
0.5mlwasadded to 2 ml ofcold 10% trichloroacetic
acid. These samples were filtered through HA filters
(Millipore Corp., Bedford, Mass.). The filters were
dried in air and counted forradioactivity byusinga
toluene-based scintillation fluid (3). The biological
activityof thesamples was assayed.
RESULTS
Heat induction of4105 cts23. When T-
bac-teria lysogenic for the 4105 mutant cts23 are
grown at 30 Candthen shiftedtoinducing
tem-perature, anincreased phage DNA synthesis is
seensome10minafter the shiftand afteran
addi-tional20 to 30 min the bacterialyse andliberate
aburstof phage (Fig. 1).
The 4105 cts23mutant wascrossed with4105
Kts3l and adouble mutant carrying both ofthe
ts mutations was isolated. This double mutant wasthenusedtolysogenizeT- bacteria to give the lysogen
T-(q605
cts23, Kts31). MutationKts3J
is located in gene K which has previ-ously been shown essential for phage DNA synthesis (2). After a shifttoinducingtemper-ature,
T(cts23,
Kts31) does not produce anyphageas longasit iskept at the high
temper-ature. Samples of DNA extractedfrom such a
culture afterthe temperature shift show no or very little increase in specific infectivity when assayed by transfection of competent SR135 cells (Fig. 1). The Kts31 mutation thus effect-ively blocks phage DNA synthesis also after
temperatureinduction of
lysogenic
bacteria.Irreversible induction of4105 cts23 lysogenic bacteria alsooccursintheabsence ofDNA syn-thesis;
T-(q505
cts23) is rapidly killed in the absence of thymine when shifted to inducingtemperature. The
viability
of T lysogenic forwild-type
4105
is not affected under the same conditions(Fig.
2). When cells from tempera-ture-induced andthymine-starved
cultures are plated with indicator bacteriain the presence ofthymine,
there is no decline in the number ofinfectious
centersformed duringa30-min incuba-tionperiod
without thymine (Fig. 2). Thus, the lack ofthymine aftertemperatureinduction does not damage the ability of the cells to producephage when returned to a thymine-containing
medium. On the other hand, there is evidence
that damageto theprophage does occur during incubation in the absence ofthymine. After 35 minwithoutthymine,thereis abouta 10-fold
de-creasein thespecific infectivity ofDNAextracted
from induced bacteria (Fig. 2). Marker rescue,
however,is not affected (not shown).
This damage to the prophage DNA must be 457
VOL.8, 1971
on November 11, 2019 by guest
http://jvi.asm.org/
in
z
0
.10
OA
z iC
VW
:3
10 20 30 40 50 60
[image:4.489.261.455.75.341.2]Minutes
FIG. 1. Temperature inductioni of T- (cts23) and
T(cts23, Kts31). The bacteria were grown to about
5 X107cellsperml,centrifuged,andsuspendedinfresh
medium at 42C. Samples were taken for infectious
centers andfor DNA extraction. The purified DNA
samples were assayedfor their infectivity by using
competenit SR135; the infectivity is expressed as
plaquesper microgram ofDNA. Symbols: 0-- -0,
7(cts23) infectivity of DNA; U---0, T-(cts23,
Kts31) infectivity of DNA; O--O, T-(cts23)
inifectious centers; O--EO, T(cts23, Kts3I)
in-fectiouscenters.
reversible because the bacteria willproducephage
when supplied with thymine. In mock-induced
wild-type lysogenics, no or avery slightdecrease
in prophage DNA infectivity is observed when
the cells are starved of thymine. Thus, during
induction in the absence of DNA synthesis,
prophage DNA suffers reversible damage which
decreases itsinfectivity.
Sucrose gradients of DNA from induced or
in-fected bacteria. To investigate whether the
dam-agethatprophage DNAsuffersafterinduction in
the absence of DNA synthesis (thymine
starva-tion) reflects some excision process, DNA from
induced lysogenswasexamined insucrose
gradi-ents. It was expectedthat high-molecular-weight
bacterial DNA could be separated from the
smaller phage-sized piece, and the excision
processmightbe examined.
T-(cts23)wasgrownat 30 C inthepresenceof
3H-thymidine. The prophage was induced by
heat, and the bacteria were incubated at 42C
with or without cold thymidine. Thirty minutes
after induction, DNA was extracted and
sedi-2
c
.n106
0
t
U
c
E4
O105
c
p
0
vc T-(cts
-2 0 5 10 15 20 25 30
M nutes 35
z
0
Qm
:k
Z
nL
z
c
FIG.2. Effects of temperature iniductionz oil
T-lysogenic bacteria in the absenice (f thymine. The
bacteriaweregrownto adenisityof about5 X 107cells
perml, centrifuged, andsuispenidedinfresh medium at
45C. Samples were takeniforviable counts, infectiouis
centers, andfor DNA extraction.Symbols: -- ,
T(p105) viable cells (v.c.); ----, T-(O105)
infectivityofDNA; O--Q ,7(cts23) v.c.;*--*,
T-(cts23)inifectiouscenters (i.c.); *---0, T-(cts23)
infectivity of DNA; A--i\ , T(cts23,KtsN31) v.c.;
A--A, T-(cts23, Kts3J) i.c.; A---A, T-(cts23,
Kts31)infectivity ofDNA.
mented in neutralsucrosegradients together with
"C-labeled T7 DNA as a size marker. After a complete run, fractions were collected and as-sayed forradioactivity, forphageDNA activity, and for phe-l transforming activity. A similar experiment was also performed with DNA
ex-tractedfromT-bacteria infected with 4105cts23
at inducing temperature. The results of these experiments areshowninFig. 3a-c. IntheDNA
sample extracted from lysogenic bacteria
in-duced in the absence ofthymine, tritium
radio-activity and phage and bacterial DNA
sedi-ment together as
high-molecular-weight
species(>200 x 106), indicatingthatprophageexcision
doesnot occurunder theseconditions (Fig. 3a).
When
T-(cts23)
lysogens are induced in thepresenceofthymidine, mostof the tritium
radio-activity is still associated with high-molecular-weightmaterial atthe time ofsampling,
on November 11, 2019 by guest
http://jvi.asm.org/
[image:4.489.62.257.77.302.2]PROPHAGE c105 IN B. SUBTILlS
3Ht
1500H
10ooo
[image:5.489.45.237.70.506.2]5 10 15 20 25 30 35 *0
FIG. 3. Sucrosegradientcentrifugation ofDNAfrom
induced cts23 lysogens or cts23-infected cells. The
bacteria were grown at 30C with prelabeling of the
bacterial chromosome with 3H-thymidine to about
5 X107cellsperml. Thebacteriawerewashedonceand
suspentdedin Min-CH (without any thymine) at45C for 5 min and then shifted to 42 C. After30 min at
inducing temperature, a sample was takenfor DNA
extraction. The purified DNA together with
14C-labeledT7DNAwascentrifugedinthe S W25.1rotor as
describedinMaterials and Methods. Thefractionswere
tested for 3H (0 ), 14C (X----X). phe-1+
transforming activity
(----
-0), rescue of phagemarker susl9 (0---0), and *for infectivity
(0--O). (a) T(cts23) with no thymine added
during the experiment. (b) T-(cts23) withz thymidine
ing no breakdown of preexisting DNA in the induced bacteria (Fig. 3b). Phage DNA active in the transfection or marker rescue assays is found
in three main peaks inthis sample. One peak of
infectivity and marker rescue cosediments with
the tritium peak. A second, more slowly
sedi-menting,
peak (fraction 29) corresponds to the infectious vegetative DNA previouslydemon-strated in pulse-labeled DNA from mitomycin
C-induced c105 wild-type lysogenics (19). A
third peak which exhibits marker rescue only is
mature DNA (18, 19). Most of the phe-J
trans-forming activity cosediments with the tritium radioactivity, but a distinct peak is found at fraction 29. This peak of phe-J activity is con-sistently reproducible and must represent newly synthesized bacterial DNA. Inthe DNAsample from infected bacteria, finally, tritium radioac-tivity andphe-1 transforming activity cosediment as high-molecular-weight material, whereasmost
of thephageDNAactivitysediments moreslowly
and corresponds tovegetativeand mature DNA
(Fig.
3c).
Thus, 15 min after infection of T- bacteria
with phage cts23, there is little phage DNA
present in thehigh-molecular-weight DNAfrom
the bacterial chromosome. Therefore, the
rap-idly sedimenting phage DNA observed after
prophage induction is not a very large form of
vegetativephage DNA. Thepersistence ofphage DNAactivityamonghigh-molecular-weight bac-terialDNA even 30 minafterprophage induction
suggests that excisionof prophageDNAhasnot
occurred under these circumstances.
Also,
the experiments give no evidence for the existence of a uniquely phage-sized DNA early after pro-phageinduction(see
alsoFig. 10a-c). Rather,the experiments suggest that excision may not be anecessary early eventin induction and thatafter
induction and subsequent replication, phage
DNA may stillbe associated with bacterialDNA.
Replication of phage and bacterial DNA after
prophage induction. Experiments were next
per-formedto
study
DNAreplication
in temperature-inducedT-(cts23).
The bacteria were grown at30 C in the presence of
3H-thymidine.
At aden-sity of about 5 x 10' bacteria per ml, the cells were induced at 45C in theabsence of thymine. After 5 min atthis temperature, theculture was
shifted to42Candanequalvolumeofprewarmed
medium containing BUdR and '4C-thymidine
was added.Samples were then taken at intervals
from the culture, and DNA was extracted. The
addedwheen cellsshiftedto42C. (c) T-infectedwith
cts23ataneffectivemultiplicity of about 2.2; burstsize
was48.Infectivitynotshown inthefigure. Thissample
wastakenat15 minafterinfection.
VOL.8, 1971 459
on November 11, 2019 by guest
http://jvi.asm.org/
DNA was
centrifuged
to equilibrium in CsCl, and the fractions were assayed for radioactivity and for biological activity. In the presence of excessBUdR, phage DNA synthesis will occur, but no burst of infectious particles is produced. InFig. 4 is shown an experiment where DNA was extracted from T(cts23) at 0 and 20 min afterinduction, respectively.Inthe 0-min sample, radioactivity andbiological
activity band atsimilar
densities,
about 1.703. After 20 min, about 11%of the prelabeled bacterial DNA hasreplicated
ascalculated from the shift of tritium toaposition of
increaseddensity.
Although
only
afraction
of thebacterial
DNA hasreplicated
at 20 min in the induced cells, about 80%of
the marker rescueactivity is found atahigherdensity
thanthemajor
tritium peakof
unreplicated DNA (the arrows in the figure
de-3H
2000- 200E
a.
E
(0 20 30 4
b)
2000 -200 200
!-i
100010MO100
notethe peaks of marker rescue in the gradient). In other words, phage DNA replicates more rapidly than the bulk of the bacterial DNAafter induction. The gradient fractions were assayed for their contents of the bacterial markersphe-J and leu-2 by transformation. Of these markers, about 45% of the phe-l transforming activity is associated with replicated material of increased
density,
whereas only about 10% of leu-2 trans-forming activity is found outside the main trit-ium peak of unreplicated DNA. The phe-l marker is linked to prophage 4405 by transforma-tion and transduction (14), whereas leu-2 is lo-catedfurther away from the prophage. The phe-l andthe leu-2 markers are replicated at the same rate in mock-induced wild-type lysogenic T or in temperature-induced T(cts23, Kts3l) (Fig. 5a-b). Thus, if the prophage is not induced by thetemperature shift employed or if the prophagez
c a
Fia.4. EffectoftemperatureinductionofT(cts23)
on the replication of the bacterial chromosome. The FIG. 5. Effectoftemperature-inducingtreatment on
cells were grown and induced as described in Materials replication of the bacterial chromosome in
mock-and Methods and in text. DNA was extracted as induced Th(105)andinT(cts23, Kts3). Cells were
described andcentrifugedtoequilibrium in CsCl. The grown and treated as in Fig. 4. The fractions
ob-fractionsobtained were testedfor 3H(0 O), 14C tainedfrom the CsCl gradient were assayed for 3H
(0---0), phe-l+ transformingactivity (0 0), (0 O), 14C (0---0), phe-l+ transforming
leu-2+transforming activity (a---0),andrescueof activity (--), and leu-2+ transforming activity
phage marker susl9; thepeaks ofmarker rescue are (---0). The DNA samples were taken 20 min
indicatedbyarrows.(a)DNA taken 0minafter shifting aftershiftingofthecells toinducingtemperature. (a),
the cells to inducing temperature. (b) DNA taken 20 T (j4105).(b), T(cts23, Kts3J).Data areexpressedas
minafter shiftingthecellstoinducingtemperature. per centof total recoveredfrom the gradients.
on November 11, 2019 by guest
http://jvi.asm.org/
[image:6.489.59.430.268.543.2]PROPHAGE
0105
TN B. SUBTILIScarries
amutation(Kts31),
whichpreventsphage
DNA
replication, in additionto thects23muta-tion,
there is nopreferential
replication of the phe-lregion
over the leu-2region on thebacterial
chromosome. Heat induction ofT(cts23) thus seems to be
associated
withpreferential
replica-tion of phage genes and withanalteration inthe pattern of replication of the bacterial chromo-some. At least one marker, phe-1, which is linked to the
prophage,
isreplicated
at an increased ratecompared
to anothermarker,
leu-2, after prophageinduction.
3H 10000
80001
6000)
1C
X00
20
The effects of prophage induction on
replica-tion
of phage and bacterial DNA were next in-vestigated in the followingexperiments.Tc(cts23)
andT-(cts23,
Kts3W)
were induced in thepres-ence
of BUdR, and DNA was extracted from each culture at various times after induction. Each DNA sample wascentrifuged
in a CsCl densitygradient.InFig. 6a-careshown the CsCl gradients for DNA fromTr(cts23),
and in Fig. 7a-c, the gradients for DNA fromT-(cts23,
Kts31).
The DNAsamples fromT-(cts23)
were10 15 20 25 30 35 40
FIG. 6. DNA wasextracted fromtemperature-induced Th(cts23) incubated with5-bromodeoxyuridine, as
de-scribed in Materials andMethods and centrifugedin CsCl. DNA wasextractedat 10min (a), 20 min (b),and
30min(c) after shiftingthecellstoinducingtemperature.Symbols:3H( );14C(0---0);unreplicated
DNA(LL);hybridDNA,replicatedonce(HL);heavyDNA,replicatedtwiceor more(HH).
|(a)
HL LL
I
_o--Cl-10 15 20 25 30 35 40
461
VOL.
8,
1971on November 11, 2019 by guest
http://jvi.asm.org/
[image:7.489.50.442.204.617.2]3H 1000
800
600 400 200
3H 1000
500
3H
1000
500
0o
20 30 40
FIG. 7. Same experiment as shown in Fig. 6, but
performedwith 7(cts23, Kts31). Symbolsidenticalto
thoseof Fig.6.Sample (c) wasextractedat40min.
also centrifuged in linear sucrose gradients (Fig.
lOa-c).
The fractions from the CsCl gradients were
pooled into fully light (LL), heavy-light (HL),
and heavy-heavy (HH) densitymaterial as
indi-catedinFig.6a-c and 7a-c. Thepooledfractions
were then assayed for radioactivity, for
trans-forming activity, for several bacterial markers,
and forphage markerrescueactivity.
Itshould be noted that the DNAwasextracted
so astominimizefragmentation ofthe material.
Asaresult of thelarge size of much oftheDNA,
e.g., Fig. lOa-c, some '4C-radioactivity appears
in the density gradients between LL and HL
density.
InFig. 8a-carepresented thedatafor the DNA
from induced T-(cts23). The curves represent
the relative ratesof replication of the particular
markersaswellastherateof replication of DNA
labeled with tritium before induction. For
in-stance, in Fig. 8a (per cent activity in LL
frac-tion), it is seen that phage marker rescue (the
lowest curve) leaves the unreplicated, LL, DNA
fraction ata muchfasterrate than, for example,
the terminal metBS transforming activity. By
scanning a fixed time such as the 10-min DNA
sample, it is seen that only 40% of phage DNA
activity (measured as marker rescue) remains
unreplicated (LL) at10 min after induction (Fig.
8a), and 60% of this activityappearsin the
once-replicated (HL) material (Fig. 8b), whereas no
activity is foundinthe HHmaterial(Fig. 8c). In
Fig. 8d-farepresented the results obtained with
DNA extracted from the induced Tr(cts23,
Kts31),toallowcomparison between thepatterns
of DNA replication observed after induction of
thetwo lysogens.
The results of theseexperiments,assummarized
inFig. 8a-f, show that prophage induction hasa
profound effect on DNA synthesis. Essentially
allphage DNA has replicatedatleastonce at30
min after induction of
T-(cts23),
and, of thebacterial markers, at least 80% of the phe-l
transforming activity has replicated atthis time.
About25% of the phe-J activityisfound in
ma-terialreplicated twiceormoreat30min (Fig. 8c).
However, about 50% of purA16, leu-2, and ilvAl
transforming activity is still in the LL material
at30min after induction (Fig. 8a), and only 10%
of the totalactivityof these markers is foundin
the HH material (Fig. 8c). The terminal metBS
marker replicates verylittle (about 10%) during
theexperiment, and noactivity ofthis marker is
recovered in the HH material. Thus, preferential
replication of phage DNA andthephe-1 marker
isconfirmed.
Itshould be noted that only 25% of the
pre-labeled DNA has replicatedat30min (Fig. 8b),
whereas somephage DNAandphe-l
transform-ing activity is found in the fraction replicated
twiceormore. Thiscould notoccuriftherewere
notpreferential local replication of the bacterial
chromosome. We havealso observedpreferential
replication of the bacterial marker nia-38,
lo-cated closeto phe-1 indirection of the
chromo-somal terminus (Table 2). Thus, preferential
replication of bacterial markers is not peculiar
tothephe-1 marker.
Induction ofTr(cts23, Kts3l) gives a totally
different pictureofDNAreplication (Fig. 8d-f).
About 50% of the phagemarker rescue activity
is found in the once-replicated (HL) fraction at
10to 20minafter inductioncompared toabout
20
Ib, Ic)
I
H '1
1
;,HL:JI
. L_LI0
on November 11, 2019 by guest
http://jvi.asm.org/
[image:8.489.54.245.70.448.2]PROPHAGE
0105
IN B. SUBTILIST-(cts 23) T-(cts 23, Kts31)
I .iI, I a. . *I I
10 20 30 L0 10 20 30 40
Minutes
FIG. 8. Rateof replication ofbacterialmarkers, phagemarkers, andtotal DNAaftertemperatureinduction of
T(cts23) and T(cts23, Kts3J). ThebracketedfractionsinFig.6 and 7werepooled, dialyzed overnight against
saline,andassayedforbiological activity.The dataaregivenaspercentoftotalactivity recoveredfromthegradient
inagiven timesample. Ontheleftisshown T(cts23) unreplicated (a),oncereplicated(b),andDNAreplicated
twiceor more(c).On therightisshown T(cts23, Kts31) unreplicated (d),oncereplicated(e),andDNAreplicated
twiceormore(f). Symbols:3H(O O),purA16+ transforming activity (O---0), leu-5+ activity (o D),
ilvA1+activity (X X),phe-l activity ( 0),metB5activity (0---0),rescueof phagemarkersusl9
(
---*)
15% for the bacterial markers purA16, leu-5,
ilvAl, phe-1. Thus,in the presence of the Kts3J
mutation, preferential replication of phage
DNA still occurs in the induced cells, although
at areduced rate. However, preferential
replica-tionof thephe-l marker isabolished, andphe-J
is replicated at the same rate as other bacterial
markers (Fig. 8d). Twenty minutes after
induc-tion, replication of both bacterial and phage
DNA virtually comes to a stop in T-(cts23,
Kts3J) and is not resumedduring theremaining
20minof theexperiment (Fig. 8e-f).
Prophage inductionhaslittleeffect ontherate
ofsynthesisof totalDNA, althoughit hasa
pro-100
80
60
40
20
4'0
4'
0-80
60
40
20
40
20
0
VOL. 8X l1971 463
on November 11, 2019 by guest
http://jvi.asm.org/
[image:9.489.97.384.70.496.2]TABLE2.Replicationofbacterial andbacteriophage
DNA in heat-inducedT(cts23)G
Percentmarkeractivityin DNAreplicatedonce
Time of or more
DNA
extraction
(min) purA16 leu-5 nia-38 metB5 sSUS19+105
10 20 23 42 7 59
20 37 30 74 14 81
30 41 43 71 14 97
aDNA extracted from induced
T(cts23)
anddescribed inFig.6,8,and 10wasusedtotransform
BD25.
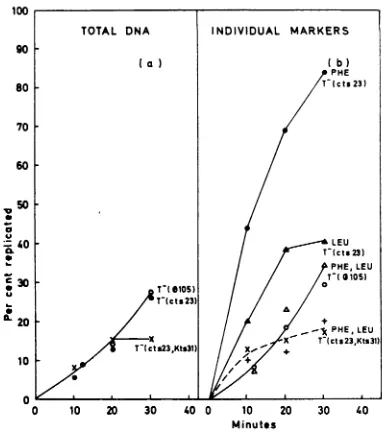
found effect on the rate of
replication
of indi-vidualportions
of the bacterial chromosome(Fig.
9). T-(105), T(cts23), and
T(cts23,
Kts31) wereinduced,
or mockinduced,
in the presence of BUdR.Samples
were taken at intervals for DNAextraction,
and the DNA was thencen-trifuged
in CsCl. Pooleddensity
fractions from
the
gradients
wereassayed
asdescribed above.
InFig. 9a is shown therateof
replication
ofDNAcontinuously
labeled with3H-thymidine
for severalgenerations
beforeinduction.
InFig.
9b is shown the rate ofreplication
(as
measuredby
shift to
higher
density)
of the bacterialmarkers
leu-2and
phe-J.
Theresultsoftheseexperiments
are in full agreement with those
presented
inFig.
8a-f.During
the first 20minafterinduction,
the rate of
replication
of theprelabeled
DNAis identical in the threelysogens.
After thistime,
however,
DNAreplication
stops inT-(cts23,
Kts3J). Only
inT(cts23)
is therepreferential
replication of
thephe-J
marker. The leu-2marker
is
replicated
at ahigher
rate ininduced
T-(cts23)
compared
tothewild-type
lysogen,
although
theirrates
of
replication
ofprelabeled
DNA areidenti-cal,
suggesting
thatreinitiation
ofDNAsynthe-sis may occur after
induction
notonly
in theprophage and the
phe-J
region
butalso,
lessfre-quently,
inotherplaces
onthebacterial chromo-some.The DNA
samples
takenfromtheinducedT(cts23) were
assayed
intransformation
forlink-age betweenthe bacterial
phe-1
marker and the phage AtslS marker. A lowdegree
oflinkage
could be detected evenin DNAreplicated twice or more
(Table 3).
Nolinkage
wasobserved
in DNA extractedfromT-cellsinfected withcts23 phage underconditions similartothose usedfor induction. SomephageDNAthusremains cova-lently bound to bacterial DNA after induction andreplication.
The DNA
samples
extracted fromT-(cts23)
and used in the
experiments
shown in Fig. 8a-c100
10so
a
u i0
c30
11I20
0 10 20 30 40 0 10 20 30 40
Minutes
FIG. 9. Rateofreplicationof the bacterial
chromo-some after temperature induction of li(cts23),
T-(cts23, Kts31), and mock induction of T(.105).
BacterialDNA wasprelabeled with 3H-thymidineawd
induced in the presence of BUdR as described in
Materials and Methods. DNA was extracted 10, 20,
and 30 min after induction and centrifuged in CsCl.
The gradientfractions were pooled as described for
Fig. 6 and 7. Eachfractionwasassayed for its content
of3H-prelabel (a) andforphe-J+andleu-2+
transform-ing activity (b). Results are presented as per cent
activityoftotal, whichisfoundinreplicated DNA. (a)
Per cent 3H-prelabel in replicated DNA: T(4d105),
O--O; T(cts23), * *;
T(cts23, Kts3J),
X--X. (b) Percent transformingactivity in
repli-catedDNA,phe-1+:
T-(105),
0 O; T(cts23),* *; T(cts23, Kts3l), X---X; leu-2+:
T(0105),
Ai A; 7(cts23), A A; 7T(cts23,Kts31), +---+.
were also
sedimented
in neutral sucrose with32P-labeled
P2 DNA asasize marker(Fig.lOa-c).
With
increasing
time afterinduction,
there is a gradual decrease in thesedimentation
rate of phage DNA, with thefirstappearance ofmature phageDNA at 30 minafter induction.Inno ex-perimenthave wefoundanabrupt, early change inthesedimentation
characteristics of prophage DNAafterheatinduction,
such asmight
be ex-pected from an excision event. The relativeamounts of rescue and
infectivity
(Fig. 10c) differfrom those shown in Fig.
3b, probably
due totheeffects of BUdR onthe
biological
activity ofthe DNA (9).
DISCUSSION
Twoconclusionsemergefromtheexperiments presented. First, we do not observe excision of
on November 11, 2019 by guest
http://jvi.asm.org/
[image:10.489.260.451.69.287.2] [image:10.489.56.250.72.198.2]PROPHAGE
0105
IN B. SUBTILlSTABLE 3. Linkagebetweenbacteriophage and bacterial markers in DNAfrom 4105lysogenicbacteriaa
Strainand treatment Fraction ts+/pkc-I+Transformnts
T-(cts23) heat induced (fractions shown in Fig.6) 10 minLL 9/104
HL 3/52
Unfractionated 9/156
Tr(cts23) heatinduced 20 min LL 0/70
HL
5/182
HH
0/80
Unfractionated
4/94
T-(cts23) heatinduced 30 min LL 0/52
HL 2/104
HH 2/80
Unfractionated 7/156
T-(40105)
mock induced 10 min 21/115Unfractionated
T(r0105)
mock induced 40 min 17/104Unfractionated
T infectedwith cts23 15min 0/298
Unfractionated
aBR95
(4105
AtslS)
wasgrown forcompetence as described.DNA extractedfrominducedT(cts23)
anddescribed inFig.6, 8, and10 wasused totransform the cellstophe-l+.The distributionof thets
and ts+alleles of the AtslS markerwasthendeterminedamongthetransformants (14).TenminutesLL,
HL, andHHreferstothefractions described inFig. 6. Unfractionated means thatthe DNAis used
priortoany centrifugation.
prophage DNA in heat-induced
4105
lysogenic
bacteria. That
is,
theexperiments give
noindica-tion
that both strandsof
theprophage
arepre-cisely
cut outof the bacterialchromosome
prior
to autonomous
replications
ofphage
DNA.Second
prophage
induction has aprofound,
butcovert,effectonthepattern of
replication
of thebacterial chromosome.
The first
conclusion,
thatprophage excision
need not be an early eventin
induction,
issup-ported
by
severalexperimental
findings.
Insu-crose
gradients
of DNAfrom induced
bacteria,
the bulkofphagegenesand wholephagegenomes cosediment with
high-molecular-weight
bacterial DNA, even when most of the phage DNA hasreplicated
at least once(see Fig. 8b, lOa).
Withincreasing
time afterinduction,
phage DNA is found in moreslowly
sedimenting material,
with a concomitant decline in the amount of
phage DNA associated withprelabeled bacterial
DNA. Ininduced E. coli (lambda), mostofthe
early
synthesized DNA sediments in neutralsucrose at a ratecharacteristic formaturephage
DNA (12, 20). Later in infection, this DNA is
converted to more rapidly
sedimenting
material,which most
likely
represents covalently closed, circularphageDNA (12, 25). ThelambdaDNAsedimentation pattern isthus the reverse ofthat
found for 4105. With thisphage,wehave never
observed DNA withsedimentation characteristics
of covalently closed circles either in induction, in infection, or in superinfected, immune, lyso-geniccells.
In DNAfrom4105lysogenics, genetic linkage
between phage and bacterial markers can be
demonstrated by transformation (14). The
de-gree oflinkagedecreases afterinduction of
lyso-genic bacteria, butsome
linkage
persistseven in DNAreplicatedtwo or more.When DNA from T bacteria infected with
cts23 phage, under conditions similar to those
used for induction, is sedimented in sucrose, phage DNA sediments more slowly than, and well separated
from,
bacterialDNA. Nogenetic linkage between phage and bacterial markers is found in such DNA.Phage genes do not leave the region of
bac-terial DNA inthe sucrosegradients when DNA
synthesis is blocked after induction (Fig. 3a). However, the lysogenic bacteria are fully
com-mitted to
induction;
whenthe cells are returnedto conditions permissive for DNA
synthesis,
morethan
90%
of themproduce
phage.Although the prophage state is maintained in
VOL. 8, 1971 465
on November 11, 2019 by guest
http://jvi.asm.org/
1s
11
3H
3H
125
100
75
50
25
2000 1600*' 1200 c
X
Boo-800
I100
Jn
[image:12.489.61.249.67.541.2]5 10 15 20 25 30
FIG. 10. Sucrose gradientcentrifugation ofthe DNA
samplesfrom 7(cts23), showninFig. 6,intheSW50
rotor. To the gradients wereadded0.15 ml of DNA
solution and 0.05 ml of appropriately diluted
32P-labeled P2 DNA. Symbols: 3H ( 0), 28p
(X---X),rescueofphagemarker susl9(O---0),
infectivity ofDNA (0 O). DNA samples were
takenat10min(a),20 min(b), and30 min(c) afterthe cellswereshiftedtoinducing conditions.
induced cells in the absence of DNA synthesis,
the infectivityofprophage DNA decreases. The
damage suffered by the prophageinthese
experi-ments is repairable, because in practice all
in-duced cells produce infectious phage when al-lowed to resume DNA synthesis. The thymine starvationdamage might be the result of abortive DNAsynthesis intheprophage region.
When the lysogen
T-(cts23,
Kts3J)
is inducedunder conditions nonpermissive for gene K ac-tivity, phage DNAis preferentially copied once (Fig. 8d, e), whereas the bulk of the bacterial DNA remains unreplicated even 40 min after induction. The preferential replication of phage DNAindicatesthatsomemechanism operates in the induced cells, which can specifically initiate replicationoftheprophage.
Noextensivebreakdown of bacterialDNA oc-curseither upon induction of lysogenic bacteria orafter infection of sensitive bacteria. After in-fection, replication ofthe bacterial chromosome
seems to proceed in a normal fashion at least
during thefirst 30min.
Preliminary
experiments indicate no preferential effect on replication of phe-J ininfection.Itshould bepointedoutthat therateof replica-tion of the bacterial
chromosome,
as measured by the transfer ofprelabeledDNAfrom lightto hybridmaterial,
is identical in induced and in mock-induced Tlysogenic bacteria,
whereas the rate ofreplication
ofphage
DNA, and at least the bacterial phe-J marker, ismore rapid in in-duced bacteria(Fig.
9).After induction of prophage cts23, bacterial
markers closer to the
origin
of the bacterial chromosomethan theprophage,
replicate
atsimi-larrates. Marker
phe-J,
situatedjust
distaltotheprophage,exhibitsadifferentpattern. Itis
initially
replicated at a faster rate than other bacterial
markers,
and it appears earlier infully
heavy
DNA. Induction of the
prophage
thus leads to preferentialreplication
notonly
ofphage
DNA but also of anadjacent
part of the bacterial chromosome. A new initiation site forreplica-tion mustthenbe
exposed
ininduced cellsat or closeto theprophage,
withreplication
proceed-ing
from this site in direction of theterminus,
but stopping short of the terminal metB.
Frag-mentsofasize
exceeding
thatofphage
DNAareproduced, as shown
by
sucrosegradient
sedi-mentationaswellasby
preserved linkage
between phage and bacterial markers inreplicated
DNA from inducedT-(cts23).
Theintegrated
state of theprophage,
initself,
generates animportant
element of the mechanism
controlling
replica-tion,
for induction alters the pattern of host DNAsynthesis,
whereas infection appears tohave no sucheffect.
Preferential
synthesis
ofadjacent
regions
of the bacterial chromosome afterprophage
inductioniH 32p
Xa)
50 300 i a
II It 200
II I
'K~~~~~~~
u
5 10 is 20 25 30
c
on November 11, 2019 by guest
http://jvi.asm.org/
PROPHAGE 4105 IN B. SUBTILIS
is not unique to the 4105 system. In lambda
lysogenic E. coli, there is evidence from DNA-DNA hybridization studies that replication of
the prophage may occur in situ after induction
andproceed into the gal operon (10). Such
repli-cation in induced lambda lysogenics has been
implied also in other types of experiments (13).
This regional replication appears to be under control of the 0andPcistrons.
Induction of integration-deficient (L) mutants
ofphageP22gives few infectiousparticles but a
substantial increase in proC andpurE transduc-ing particles (21). These markers are located
close to oneend of the prophage.Thedistribution
of phagegenes inthe defectiveparticlesisstrongly polar with genes distal to proC appearing with diminishing frequency. After induction of L
mu-tants of P22, replication thus seems to be
in-itiated at or close to the prophage and to
pro-ceed in the direction of proC. The mechanism
which ultimately produces phage-size DNA is unknown.
Ofparticular interest in the present context is
the defective phage PBSH carried by B. subtilis
168(9).PBSHisinducibleby mitomycin C. After induction, the relative amount of purA16 trans-forming activity increasesmanyfold due to pref-erentialreplication of this marker (9). However,
itcannot bedecided whetherpreferential
replica-tion of
purA16
isduetorepeated
reinitiations at thebacterialinitiation site atthe originorto theappearance of a new initiation site associated
with the phage. Other bacterial markers also replicate after PBSH induction albeit at a slow rate
compared
to purA16. Breakdown of the bacterial chromosome does not occur in the in-duced cells; even late after induction, purA16containing
DNA exists inpieces
atleasttwicethe size of the DNA found in the maturephage
particles (9).
Our
experiments
indicate that there is morethan one step involved in separating bacterial
and phage genes after heat induction of 4105
cts23 lysogens. After
induction, phage
genesreplicatealthough theyare still
genetically
linked to bacterialgenes and appear inlarge-size
DNA pieces.Apparently,
thereisagradual
decreaseinthe size of the DNA molecules which harbor
phage genes
during
thecourseof induction(Fig.
10a-c). Rather than an early excision event, the
processof
separating
phageandbacterialgenes ininduced bacteriaseems to be
gradual,
withfinal definitionoccurring
with the maturephage
DNA.
Insummary, excision doesnot seemto be the
only mechanism
by
which a prophage can enter the phase ofvegetative growth
after induction oflysogenic
bacteria. Whether inductionin-volves nucleases, recombinases, or polymerases, it requires recognition of the prophage region on the
bacterial
chromosome. Our dataindicate
thataDNAreplicationsystem mayrecognize the
4105
prophage after induction. Itwould appear,then, that in our system the initial events in
lysogenic induction arefundamentally similar to
the processof normal replication ofthebacterial
chromosome, in that some control apparatus directs a DNA replication system to a specific chromosomal site for initiation of
replication.
On thebasisofthedatapresented, this control
apparatus cannot be defined. Itcouldinvolve the
unveiling of an initiation site at the prophage, which is blockedin theuninduced lysogenic bac-teria, followed by binding ofthe
DNA-polymeriz-ingsystem alreadypresent in the
cell
tothissite.Or, the primaryevent ininduction may involve
some interaction between this
DNA-polymeriz-ing system and some (phage-coded) product
such thatthe specificity of the polymerizing
sys-tem is altered. Further
experiments
are now inprogress trying to resolve these
possibilities.
ACKNOWLEDGMENTS
Kerstin Nilsson provided excellent assistancethroughoutthis
work.
Theworkwassupported by grants from TheSwedishMedical
ResearchCouncil, Karolinska InstitutetsForskningsfonder, and
Emil ochWeraCornellsstiftelse. One of us (R.W.A.) holds a
postdoctoral fellowship from the American Cancer Society. LITERATURECITED
1. Anagnostopoulos, C., and J. Spizizen. 1962. Requirements
fortransformationinBacillus subtilis. J. Bacteriol.
81:741-746.
2. Armentrout, R. W., and L. Rutberg. 1970. Mapping of prophage and maturedeoxyribonucleicacid from temperate
Bacillus bacteriophage )105 by marker rescue. J. Virol. 6:760-767.
3.Armentrout, R. W., L. Skoog, and L. Rutberg.1971. Structure
andbiologicalactivity ofDNAfrom Bacillus bacteriophage
4105; effects of Escherichia coli exonucleases. J. Virol.
7:359-371.
4. Birdsell, D. C., G. M. Hathaway, and L. Rutberg. 1969.
Characterization oftemperate Bacillusbacteriophage4105. J.Virol. 4:264-270.
5. Campbell,A.1962.Episomes. Advan.Genet.11:101-145. 6. Choe,B. K.1969.Integration-defectivemutantsof
bacterio-phageP2.Molec. Gen.Genet. 105:275-284.
7. Dubnau,D., C.Goldthwaite,I.Smith, and J. Marmur. 1967. GeneticmappinginBacillus subtillis. J.Mol. Biol. 27:163-185.
8.Gingery,R., and H.Echols. 1967. Mutantsof bacteriophageX unabletointegrate into the host chromosome.Proc. Nat. Acad.Sci.U.S.A.58:1507-1514.
9. Haas,M., andH.Yoshikawa. 1969. Defective bacteriophage
PBSH in Bacillus subtilis. II. Intracellular development of theinducedprophage.J.Virol.3:248-260.
10.Imae, Y.,andFukasana, T. 1970.Regionalreplicationof the bacterial chromosomeinducedbyderepressionofprophage
lambda.J.Mol.Biol.54:585-597.
11. Kaiser,A.D., and T.Masuda. 1970.Evidencefor a prophage
excisiongeneinX. J.Mol.Biol.47:557-564.
12. Lipton,A., and A. Weissbach.1966.Theappearance of circu-467
VOL.8,]1971
on November 11, 2019 by guest
http://jvi.asm.org/
lar DNAafter lysogenic induction in Escherichia coli CR34 (X). J.Mol. Biol. 21:517-525.
13.Nishimune, Y. 1970. Prophage excision:onthestatusof host chromosome afterinduction.Virology41:541-548.
14.Peterson, A. M., and L. Rutberg. 1969. Linked transformation of bacterialandprophagemarkers inBacillus subtills 168
lysogenic for bacteriophage0105.J.Bacteriol. 98:874-877. 15. Ptashne,M. 1965. The detachmentandmaturationof
con-servedlambdaprophageDNA. J.Mol.Biol. 11:90-96.
16.Rutberg, L. 1969. Mapping ofa temperate bacteriophage activeonBacillus subtilis. J. Virol. 3:38-44.
17. Rutberg, L., and R. W. Armentrout. 1970. Low-frequency rescue ofa geneticmarker from Bacillus bacteriophage 0105bysuperinfecting bacteriophage. J. Virol. 6:768-771. 18.Rutberg, L., J.A.Hoch,andJ. Spizizen. 1969.Mechanism
oftransfection withdeoxyribonucleicacid fromthe tem-perateBacillusbacteriophage0105.J.Virol. 4:50-57.
19. Rutberg, L., and B. Rutberg. 1970. Characterization of
infectious deoxyribonucleic acidfromtemperateBacillus
subtilis bacteriophage4,105.J. Virol.5:604-608.
20. Salzman, L. A., and A. Weissbach. 1967. Formation of intermediates in the replication ofphage lambda DNA.
J. Mol. Biol. 28:53-70.
21.Smith, H.0.1968. Defective phage formation bylysogensof
integration deficientphageP22mutants. Virology
34:203-223.
22.Smith, H.O., and M. Levine. 1967.Aphage P22gene
con-trollingintegration of prophage. Virology 31:207-216. 23. Spizizen, J. 1958. Transformation of biochemically deficient
strains of Bacillus subtilis bydeoxyribonucleate. Proc. Nat. Acad. Sci. U.S.A.44:1072-1078.
24. Weisberg, R. A., and J. A. Gallant. 1967. Dual functionof
theXprophagerepressor.J.Mol.Biol. 25:537-544. 25. Young, E. T., III., and R. L. Sinsheimer. 1967. Vegetative
bacteriophage X DNA. II. Physical characterization and replication.J. Mol. Biol. 30:165-200.