0022-538X/91/073496-08$02.00/0
Copyright C 1991,AmericanSociety forMicrobiology
The
Yeast GAL4 Protein Transactivates
the
Polyomavirus Origin of
DNA
Replication
in
Mouse Cells
MOSHEBARU, MEIR SHLISSEL, AND HAIMMANOR*
Department of Biology, Technion-Israel Institute of Technology, Haifa32,000,Israel Received 19February 1991/Accepted5 April 1991
Wehavereplacedthepolyomavirus (Py)enhancer,which isanessentialcomponent of thePy originofDNA replication (ori), with five repeats ofa 17-bp oligonucleotide including the yeast GAL4 upstream activating sequence (5xGAL4sites). Plasmids containing this modified Py ori, designated test plasmids, and plasmids encoding either the GAL4 transcriptional activator protein or various derivatives of this protein were
cotransfected intomousecells whichconstitutively synthesizeatemperature-sensitive Py largetumorantigen (T-Ag). Replication ofthe testplasmids was monitored by Southern blotdeterminations of the amounts of plasmidDNA that becameresistanttocleavagebytheenzymeDpnI.Thesestudiesshowed that in thepresence ofafunctional T-Ag,the GAL4protein,andhybrid proteinsincludingthe GAL4DNA-bindingdomain andthe activating domainof theadenovirus Elaor herpesvirusVP16 proteintransactivated themodifiedPyori. A truncated proteinincluding just the GAL4DNA-bindingdomainwasinactive in these assays. The authentic GAL4proteinwasfoundtobea moreefficientreplicationtransactivator thanthehybrid proteins.Incontrast, chloramphenicol acetyltransferaseassaysshowed that thehybrid proteinsweremoreefficienttranscriptional activatorsthan theGAL4 protein.Theextentof theGAL4-dependent replicationofaplasmidinwhich thePy earlypromoterwasdeletedwas55 %lower than that ofaplasmidincludingthepromoter.However,the extents ofreplicationofplasmids includingtwotandem repeats of theremaining Pyorigincore and5xGAL4 sitesor twoorigincoresflankingasinglecluster of5xGAL4 siteswere4.8- and1.6-foldhigherthanthat of theplasmid includingasinglecopy ofeach element. The replication ofaplasmidincluding twoclustersof 5xGAL4 sites flankingasingle origincore wasbelow the limitof detection ofourassays.These resultsindicate that the GAL4 andhybridtransactivators donotactivate thePyoribyvirtue of their interactions withtranscriptionfactors thatbindpromoterelements.Rather,itappearsthat these activatorproteinsmayinteract with thereplication initiationcomplexes, therebyfacilitating orinhibitingtheinitiation ofreplication.
The polyomavirus (Py) origin of DNA replication (ori)
consists oftwoelements: the origin core and the enhancer (7). The origin core was defined genetically as an essential and specific part ofthe Py ori (22, 26, 44). Itextends from nucleotides 5268 to 42 in the Py A2 strainnumberingsystem (11). The core contains three main sequence elements: (i) a central palindrome(nucleotides 5281to22), which includes two repeats of thelargeT-antigen (T-Ag) recognition
penta-nucleotide sequence motif [5'-(G)(A/G)GGC-3'] on each
strand; (ii)anA/T-rich sequence (nucleotides 5268 to5283) that maps on the late side of the palindrome; and (iii) a
polypurine-polypyrimidine sequence (nucleotides 27 to 42) that maps onthe early side of the palindrome (7). The sites of initiation of bidirectional DNA replication have been mapped ontheearly sideof the core (15).
The Py replication enhancer overlaps with the
transcrip-tionalenhancerforthe early transcription unit (nucleotides 5021 to
5265)
and with the late promoter (4, 9, 14, 43, 45). Deletionof the entire enhancer inactivated the origin (9, 45).However, a functional origin could be reconstituted by
reinsertion ofvarious parts of the enhancer (3, 33, 39, 45). Moreover,functionalorigins were also obtained in which the Py enhancer was replaced with sequences from other viral andcellular enhancers (la, 9).
Previous work on eukaryotic enhancers focused on their role in transcription. It was found that enhancer sequences bind specific protein factors which act as transcriptional modulators (18, 20, 30, 38). Similar proteins bind to some
* Correspondingauthor.
upstream promoter elements, which are also found in en-hancers (28). A typical transcriptional modulator protein consists of a DNA-binding domain that specifically recog-nizes a short DNA sequence and a relatively nonspecific activating domain that interacts with general transcription factors and with other enhancer-binding proteins, thereby
enhancingorsilencingtheinitiation oftranscription (18,38). Anumber of mammalianand yeasttranscriptionalmodulator proteins were also found to regulate DNA replication, but
detailedknowledgeof the molecularinteractionsthat under-lie this regulationislacking (5, 19,24, 33, 34, 35).
Wereport here that the yeast GAL4transcriptional acti-vator protein and hybrid derivatives of this protein, which have been extensively used for studies of transcriptional
activation (38), also induce initiation of replication at a modified Py oriin which thePyenhancer hasbeenreplaced with aclusterof GAL4 upstreamactivatingsequences. We have begun a systematic study of this system and present here the dataobtained sofar, which indicate that transacti-vation of the Py ori by the GAL4 and hybrid proteins cannot be accounted for by their interactions with transcription
factors that bind promoter elements.
MATERIALS ANDMETHODS
Plasmids. Partial maps of the various plasmids used are shown inFig.1 to4.Plasmid
pPyAcat26.2'
(45) isapBR322 derivative which containsa PyDNAfragment (nucleotides 4632 to 152), in which the enhancer (nucleotides 5021 to 5262) has been replaced with two repeats ofa 26-bpoligo-nucleotideincludingthePy oxenhancerelement(nucleotides 3496
on November 10, 2019 by guest
http://jvi.asm.org/
5108 to 5130). Italsocontainsachloramphenicol
acetyltrans-ferase (CAT) gene attached to the Py early promoter. Plasmid pPyLT1 (49) contains a Py genome from which the large T-Ag intron has been deleted, which was cloned into
plasmid pAT153. Plasmid pGSE472cat (25) contains adeno-virus type 5 E4 promoter sequences (-240 to +32 from the transcription initiation site) inserted upstream of the CAT gene in apSP72 vector (Promega Biotec). In addition, this plasmid contains five repeats of a 17-mer oligonucleotide includingabindingsite forthe yeastGAL4 protein(SxGAL4 sites) insertedupstreamofthe E4 promoter. PlasmidspAG4
and pAG147 (21) include genes encoding the whole GAL4 protein and the truncated N-terminal GAL4 protein [GAL4
(1-147)protein], respectively, whichareexpressed under the control of the adenovirus major late promoter. Plasmid
pSGVP(40) includesageneencodingahybrid proteinwhich
consists of the N-terminal 147 amino acids of the GAL4
protein fused to thecarboxyl-terminal78amino acids ofthe herpesvirus protein VP16 (GAL4-VP16 protein). Plasmid pGAL4-Ela (25) includes a geneencoding a hybrid
protein
which consists of the N-terminal 147 amino acids of the GAL4 protein fused to amino acids 121 to 223 of the adenovirus Ela protein (GAL4-Ela protein). Plasmid
pSG147(25)includesthe geneencodingthetruncatedGAL4
(1-147) protein.The genesmentioned above in the last three
plasmidsareexpressed under thecontrol ofthesimianvirus 40 (SV40) earlypromoter.
Additional plasmids used were as follows. Plasmid pM96
was constructed by insertion of a fragment of Py DNA
extendingbetween the BamHI and EcoRI sites
(nucleotides
4632to1560) intothecorresponding restrictionenzymesites
in plasmid pUC119 (47). In this Py DNA fragment, the enhancer (nucleotides 5021 to 5264) was replaced with a
XhoI linker. PlasmidpMlOlwasconstructedby replacement ofthePy DNAfragment extending between the BamHIand
XhoI sites in plasmid pM96 with a fragment
extending
between the same two enzyme sitesin
plasmid pG5E472cat
(see above). The latter fragment includes the cluster of
5xGAL4 sites. Plasmid pM107 contains the cluster of
5xGAL4 sites fused to the late side of a Py DNA segment
(nucleotides5265to 90). Toconstruct this
plasmid,
plasmid
pMlOl
was digested withthe enzymes BamHI andBglI. Afragmentextendingbetween these twoenzyme
sites,
which includestheabove-mentioned sequences, waspurified
fromthis digest. This fragment was exposed to the enzyme T4 DNApolymerase firstat 12°C and thenat
37°C
todigest
the 3' extension formedby
theBglI cleavage
andtofillinthe 5'extension formed by the BamHI
cleavage (41)
and was inserted into the SmaI site ofplasmid
pUC119.
PlasmidpM108 contains two repeats of the above-mentioned BamHI-BglI fragmentinserted in tandem into theSmaI site of plasmid
pUC119.
Plasmid pMlll was constructedby
insertion ofaPy DNA segmentextending
between nucleo-tides 5265 and 90, which was obtainedby cleavage
ofplasmid
pM108 withBamHI andXhoI,
intoplasmid pM107
between the BamHI and Sall sites in the multiclonal seg-ment. To construct
plasmid
pM112, afragment
containing
the cluster of5xGAL4 sites was
prepared by
cleavage
of plasmid pM108 with the enzymes EcoRI and XhoI. Thisfragment waspurified and inserted between the
recognition
sites of the enzymes EcoRI and
KpnI
in the multiclonal segmentofplasmid
pM107.Theinsertionwasperformed by
first ligating thematching
EcoRIsites;
then T4 DNApoly-merase was used to
digest
the 3' extensionproduced
by
KpnI
and to fill in the 5' extensionproduced by
XhoI asdescribed
above;
finally,
the two blunt endsgenerated by
this
procedure
wereligated.
Transfection-replication
assays. Mouse WOP cells(23)
were
propagated
at37°C
in Dulbecco modifiedEagle's
medium
supplemented
with 5% fetal calf serum in aCO2
incubator as described
previously (29).
Twenty-four
hoursbefore the
transfection,
3 x105
cellswereseeded in60-mmplates
in 5 ml ofthesamemediumcontaining
20% fetalcalfserum. Each
plate
was transfected withplasmids by
thecalcium
phosphate
technique (10)
as follows. A240-pAl
vol-ume of a solution
containing
0.25 MCaCl2,
0.05 MN-2-hydroxyethylpiperazine-N'-2-ethanesulfonic
acid(HEPES;
pH 7.05),
and 10 ,ugofplasmid
DNAwasmixed with240IlI
of anothersolution
containing
0.05 MHEPES,
0.28 MNaCl,
0.75 mM
Na2HPO4,
and 0.75 mMNaH2PO4 (pH 7.05).
The mixturewasincubatedfor20minat25°C
andthenaddedto theplate.
The cultures were incubated 6 h at37°C,
after which time the medium was removed and the cells wereextensively
washed with fresh medium. Then 5 ml of me-diumcontaining
5%fetalcalfserum wasaddedtoeachplate,
and thecellswereincubatedfor 48 hateither33or
39°C,
asspecified
in thefigures.
The cells werethenharvested,
andsmall-molecular-weight
DNA wasselectively
extracted aspreviously
described(16,
31).
The DNA wasdigested
with the enzymeDpnI
andwithother enzymesasspecified
in thefigure legends.
Thedigests
wereanalyzed by
electrophoresis
in 1% agarose
gels
and thensubjected
to Southernblotting
and
hybridization
witha32P-labeled
plasmid probe
asprevi-ously
described(31).
Theprobe
used for these assays waspUC119
plasmid
DNA(47)
prepared by
oligonucleotide
labeling (17).
Forquantitative
analysis,
autoradiograms
ofthe blots were scanned with a Cliniscan 2 densitometer
(Helena
Laboratories).
Therelativeextentsofreplication
ofthe test
plasmids
were determinedby
estimations of thefollowing
ratio:intensity
ofabandcontaining
atestplasmid/
intensity
ofa bandcontaining
a referenceplasmid.
Theseratioswere normalized relativetothe smallest
ratio,
which wasgiven
the value 1.0.CAT assays. WOP cellswere transfected
by
the calciumphosphate
technique
as described above. After 48 h of incubation at33°C,
the medium was removed and each 60-mmplate
was washed twice with 5 ml of a solutioncontaining
0.13 MNaCl,
0.026 MKCl,
0.008 MNa2HPO4,
and 0.001 MKH2PO4.
Protein extracts wereprepared
at23°C
asdescribedby
Seed and Sheen(42).
Briefly,
thecellswere
scraped
with arubberpoliceman,
suspended
in 1.5 mlof the solution described
above,
and transferred to anEppendorf
tube. Thetubewas spunfor 10sat11,000
xgin aSorvallmicrofuge,
and thesupernatant
wasremoved.Theprecipitated
cells wereresuspended
in 1 ml ofa solutioncontaining
20mMTris-HCl(pH
7.5)
and 2 mMMgCI2
and were incubated for 5 min at23°C.
Then the cells wereprecipitated
again by
centrifugation
asdescribed above. Theprecipitated
cells wereresuspended
in 100,ul
ofa solutioncontaining
20 mMTris-HCl
(pH
7.5),
2 mMMgCl2,
and0.1%Triton andincubated
again
for5minat23°C.
Thistreatmentseparated
the nuclei from thecytoplasm.
The nuclei wereprecipitated by
spinning
thetube for 5 minat11,000
x g;thesupernatant
containing
thecytoplasmic
extract was heated for5 min at70°C
and used for the CAT assaysas follows.Fifty
microliters of theextractwas mixedin aplastic
vialat4°C
with 200[lA
ofa solutioncontaining
1.25 mMchloram-phenicol
and 100 mM Tris-HCl(pH
7.8).
To this mixture were added S,ul
of'4C-labeled
acetyl
coenzyme A(0.01
,uCi/,A;
specific
activity,
4Ci/mol)
and 5 mlofthescintilla-tion fluid Econofluor
(Dupont).
Themixture wasincubatedon November 10, 2019 by guest
http://jvi.asm.org/
at37°C and counted inaPackard scintillationcounterafter6 h. Theacetylatedchloramphenicol diffused into theorganic phase; hence, the recorded radioactivity represented the extentof the reactionandgavea measure of CATactivity. It should be noted that the radioactivity increased linearly in ourassaysforatleast 10 h.
-N'
5- 5. 5.
pftp
,
1AS' 'A ' e
(-.1> 1-7 4>4> lz
't
-B
BRESULTS
Experimental strategy. Two typesof shuttleplasmidswere used for intracellular replication assays. Plasmids of one type, designated test plasmids, contained modified Py
ori-gins, in which the yeast GAL4 upstream activating se-quences orPyenhancer elements were insertednext to the
origin cores. Plasmids of the second type, designated acti-vators, encoded the yeast GAL4proteinor various deriva-tives of this protein but did not contain active origins of
replication. These plasmids were transfected into Py-trans-formed mouse cells ofa line designated WOP (23), which continuously synthesize a temperature-sensitive Py large T-Ag, the viralproteinrequired forinitiation ofreplicationat the Py ori (7). All transfection-replication assays were
per-formedwith aconstant input ofplasmid DNA; i.e., plasmid pUC19 or pUC119 was added to the transfection mixtures such that the cells in each 60-mmplate were transfected with 10 ,Ig of plasmid DNA. Following transfection, the cells wereincubatedfor 48 h at either 33 or 39°C, the permissive
or nonpermissive temperature, respectively, for the large
T-Ag. Small-molecular-weight DNA was selectively ex-tracted from the cells at the end of these periods and was
digested withtherestrictionenzymeDpnI and with another enzymethat cutsthe testplasmidused for each experiment once. Thedigestswere analyzed by agarosegel electropho-resis, Southern blotting, and hybridization with a plasmid
probe. The amount of DpnI-resistant plasmid DNA, as
determined by blot hybridization, was a measure of the extentof replication of theseplasmids (36). In someassays,
referenceplasmids that had been propagated in Escherichia
coli dam were also cotransfected into the cells. These
plasmids were DpnI resistant even though they had not
replicatedin the mouse cells andwereused forquantitative
assessmentsofthe extentsofreplicationof thetestplasmids.
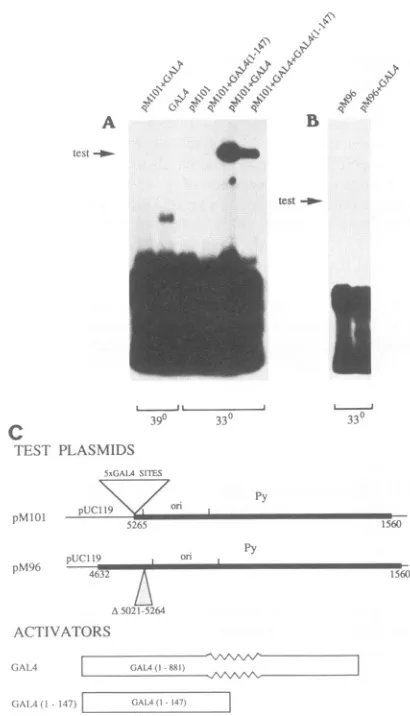
The yeastGAL4 proteincaninduce replication of aplasmid containing a Py origin core and GAL4 upstream activating sequences. The test plasmids designated pM1Ol and pM96 wereusedfor the firstseries of experiments shown in Fig. 1. AsillustratedinFig. 1C,plasmidpM1Olcontainsafragment
of Py DNAincluding the Pyorigincore,theearly promoter, andapartof the earlyregion encoding the small and middle Py T-Ags and a truncated large T-Ag. The Py enhancer was replaced in this plasmid with five repeats of a 17-bp DNA
oligomer containing a yeast GAL4 upstream activating se-quence (5xGAL4 sites; 21). Plasmid pM96 contains a frag-mentofPy DNAincluding the part of the early region which is also included in plasmid pM101 and a part of the late region. In this plasmid, the Py enhancer has been deleted. One of theactivatorplasmids used for these series of assays, pAG4, encodes the GAL4 protein, which includes a
DNA-binding domain and activating domains (21). A second activator plasmid, pAG147, encodes a polypeptide desig-nated GAL4 (1-147), which consists of the N-terminal 147 amino acids of the GAL4 protein; this polypeptide includes theDNA-bindingdomain of the protein (21).
The transfection-replication assays were performed in mouse WOP cells as described above. Autoradiograms of
theblots obtained inthese assays are presentedFig. 1A and
WSl-al
test _
396 35'
C
P1-t)SMIDES
i:( .4S I1-S
pUC(r
I
Pcv
5205
IIV
p'(.1t19 4f.1(2
.6 5e2 52(A1
ACTIIVATORS
AN IVA I 1KI
A1 (A. .1
FIG. 1. TransactivationofthePyoriby theyeastGAL4protein.
The plasmids whose structures are illustrated in panel C were
transfected into WOP cells as indicated above each lane. The transfection mixtures contained 10jigofDNA,including 1 ,ugofa testplasmid and/or2 ,ugofeachactivatorplasmid, the rest being pUC119 plasmid DNA, except for lane
pMlOl+GAL4+GAL4(1-147), in which themixture contained7 ,ugofplasmidpAG147, 1 ,ug
ofthe testplasmid, and2 ,ugof theactivatorplasmid. The cultures
wereincubated for48 h attheindicatedtemperatures.Then small-molecular-weightDNAwasselectively extracted and digested with DpnI and BamHI exceptforlaneGAL4, forwhichXhoIwasused instead of BamHI. The digests were analyzed by agarose gel electrophoresis, Southernblotting,andhybridizationwithapUC119 probe as described in Materials and Methods. Arrows indicate positions of the linear forms of thetestplasmids.
B.Thearrowspoint to the expectedpositionsofpMlOl (Fig.
1A)andpM96 (Fig. 1B) plasmidmolecules that have under-goneatleastoneround ofreplicationin themousecells,that
is, theDpnI-resistantlinear forms of thesetestplasmids. In theexperiments performed at33°C, the permissive temper-ature for the Py large T-Ag synthesized in WOP cells, no detectable replication ofplasmidpMlOl occurred whenthe cellsweretransfected with thisplasmid alone (Fig. 1A). This
plasmid did replicate in cells that had also been cotrans-fected with the nonreplicating plasmid pAG4, encoding the whole yeast GAL4 transactivator protein. In contrast, no detectable replication ofplasmidpM96 wasobserved when this plasmid was transfected into the cells either alone or with the plasmid encoding the GAL4 protein. Weconclude
A
p%1]10
I
on November 10, 2019 by guest
http://jvi.asm.org/
[image:3.612.329.534.74.432.2]A - I
1,; "I.
tes;t
_im
me mtest- _ m
ref.-_ *_ _
39) 33 )
B
pPyAcat26.2+
2xaENHANCER
ori CAT
4632
A5021-5264
Py
pPyl.T1
4632
A409-795
152
pAT153 4631
FIG. 2. Demonstrationthatawild-type large T-Agcanactivatea
Py ori in WOP cells incubated at
39°C.
Transfection-replicationassaysoftheindicatedplasmidswere carriedoutasdescribedin the
legendtoFig. 1exceptthat in lane pPyLT1 theDNAwascut with DpnI and XbaI. Each transfection mixture contained 1 p.gofatest plasmid, 0.5
,ug
of plasmidpUC19propagatedin E. colidam(ref.), and 8.5,ug
of plasmid pUC19propagatedin E.colidam'.TheDpnI cleavageproductsare not shown.that the GAL4 protein can transactivate the Py ori in a
plasmid that contains GAL4 upstreamactivating sequences
instead of the Pyenhancer.
In another assay shown in Fig. 1A, the cells had been
cotransfected with plasmid pMlOl and plasmid pAG147, encoding the truncated protein GAL4(1-147). It canbe seen
that this protein was unable to activate the Py ori. In
addition, when both the plasmid encoding the whole GAL4 protein andthe plasmid encoding the truncated GAL4
pro-tein were cotransfected into the cells with plasmid pMlOl,
replication induced by the whole GAL4 protein was
inhib-ited. This result is compatible with the notion that both proteinscompetefor bindingthe clusterof5xGAL4sitesand indicated that theinability of the truncated GAL4 proteinto
activate the Py ori is not due to a rapid turnover of this
protein. It appears, therefore, that the GAL4DNA-binding
domain alone cannotactivate the Py ori and that additional GAL4 protein sequences are required for this function.
Requirement for the Py large T-Ag. The experiments
described so far were carried out at
33°C,
the permissivetemperaturefor thePylargeT-Agsynthesized in WOPcells. As shown in Fig.1A, at
39°C,
the nonpermissive tempera-tureforthe large T-Ag, plasmidpMlOl didnotreplicateevenin the presence of the whole GAL4 protein transactivator.
This result, however,wasnotnecessarily duetoinactivation
of the T-Ag at
39°C.
It was possible, for example, thatreplication of this plasmid was inhibited at
39°C
becausecell-encoded protein components of the replication
appara-tus were damaged at the higher temperature. To examine
this question, we performed transfection-replication assays
of two plasmids including functional Py origins, whose structuresareillustratedin Fig. 2B. Inoneoftheseplasmids,
designated pPyAcat26.2', the complete Py enhancer was
replaced with two tandem repeats of a 26-bp sequence
including a segment of the Py enhancer designated a, which
provide the enhancer function needed for Py DNA
replica-tion (45). This plasmid does not encode any Py-specific proteins. The second plasmid, pPyLT1, contains a whole Py genome in which the large T-Ag intron has been deleted. Hence,this plasmid encodes the large T-Ag and the late viral proteins but not the middle and the small T-Ags. Each of these plasmids was transfected into WOP cells with a nonreplicatingDpnI-resistantreference plasmid (seeabove). The results of the transfection-replication assays per-formed with these plasmids arepresented in Fig. 2A, which showsjust the DpnI-resistant bands found in the gels. It can be seen that plasmid pPyAcat26.2', whichdoes notencode a Py large T-Ag, replicated at
33°C
but not at 39°C. In contrast, plasmid pPyLT1, whichencodes awild-type largeT-Ag,replicated atbothtemperatures. Theseresults showed that the cell-encoded proteins required for DNAreplication could functionat
39°C.
Hence, the inability of the WOPcellsto support the GAL4-induced replication of plasmid pMlOl at
39°C
wasduetoinactivation of the endogenous tempera-ture-sensitivelarge T-Ag.Transactivation ofthe Py ori by hybrid proteins including the yeast GAL4 DNA-binding domain and viral activating domains. Previous studies have indicated that hybrid pro-teins consisting of the GAL4 DNA-binding domain and activating domains derived from the viral transactivator protein adenovirus Ela (GAL4-Ela) or herpesvirus VP16 (GAL4-VP16) can enhance transcription from promoters
containing GAL4 upstream activating sequences in mamma-lian cells (25, 40). It was therefore of interest to determine whether the hybrid proteins can also transactivate the Py ori. For this purpose, plasmids pGAL4-Ela and pSGVP encod-ing these proteins were cotransfected into WOP cells with testplasmid pMlOl, and the extent of replication of the test plasmid was determined. Replication of plasmid pMlOl occurred at 33 but not
39°C
in the presence of either of the twohybrid transactivators (Fig. 3A). As shown previously, no replication was observed when plasmid pMlOl was transfected into the cells alone or with plasmid pSG147 encoding the truncated GAL4 (1-147) protein. Furthermore, in three plasmid cotransfection assays shown in Fig. 3A,the GAL4 (1-147) protein apparently competed with the hybrid transactivators for binding the 5xGAL4 sites and thereby reduced the extent of replication of plasmid pMlOl to an undetectable level.Relative efficiencies of transactivation of replication and transcription by the yeast GAL4 activating domains and the viral activating domains. A comparison ofthe intensities of the bands of test plasmid pMlOlin Fig.1A and 3A indicated that the authentic yeast GAL4 protein transactivated the Py ori more efficiently than did the hybrid proteinsincludingthe viral activating domains. Figure 3B shows an experiment that further substantiated this conclusion and allowed a quantitative assessment of the relative potencies of these proteins as replication transactivators. In this experiment, similar assays of the GAL4 and GAL4-Ela proteins were performed in the presenceof anonreplicating DpnI-resistant reference plasmid. In parallel, the extent of replication of plasmid pPyAcat26.2'was assayed again. Scanner readings of the intensities of the bands revealed that the whole yeast GAL4 protein was 10 times more efficient in inducing the replication of plasmid pM1Ol than was the hybrid protein GAL4-Ela. The scanner readings also revealed that the extent ofreplication of the GAL4-activated plasmidpMlOl was 1.7 times lower than that of plasmid
pPyAcat26.2',
which was activated by endogenous Py enhancer-binding
NIF
A
on November 10, 2019 by guest
http://jvi.asm.org/
[image:4.612.63.302.66.292.2]B
A
test-_
test-_ *
ref._- g
390 330
TEST PLASMIDS
5.GAL4SITES
pM101 PUC 5265
10 1 17 330
Py
2.0iENHANCER
oni pPyAcat26.27
ACTIVATORS
CAT
4632
A5021-5264
152
,A414ALA(1 881) GiAIA(1-147)L| GAIA(I- 147) i (GA1.4 -f.1a GALA (1-147)
GAIA VF'l6 GALA (1-147)
EIa (121-223)
[image:5.612.80.272.80.447.2]0iNT6(41041})
FIG. 3. Transactivation of the Py ori by hybrid proteins
includ-ing the yeast GAL4 DNA-binding domain and viral activating
domains. Transfection-replicationassaysofthe indicated plasmids
werecarriedoutasdescribed in the legendtoFig. 1. Thereference
plasmid (ref.)used intheassaysshowninpanelBwasthesame as
that used inFig.2. Numbersatthe bottom ofpanel Bindicatethe relative extents ofreplication of the test plasmids determinedas
described inMaterials and Methods andnormalizedwithrespect to theextentofreplication obtained inthe assayof plasmids pMlOl
plus pGAL4-Ela, whichwasgiven thevalue 1.0.
proteins. Similar estimations basedonthe data shown inFig.
2andonadditionalassaysof thesametyperevealed that the extentofreplication of plasmid
pPyAcat26.2'
was2.5to3.0 times lower than that of plasmid pPyLT1, which contains awholePyenhancer. Theseestimationswerecompatible with previous data of Veldman et al. (45), who compared the replication of plasmid
pPyAcat26.2'
with that of another plasmid containing awhole Py enhancer. Hence, under theconditions of our assays, the extent of GAL4-activated replication ofplasmid pMl0lwas4.0to5.0 times lower than thatof plasmids containingacomplete Pyori.
In view of the data described above on the relative
efficiencies of the GAL4 and hybrid proteins as replication
transactivators, it wasof interest todetermine the relative efficiencies of theseproteins astranscriptional activatorsin WOP cells under the conditions in which the replication
assays were performed. For these transcription activation assays, weused a reporterplasmid (pG5E472cat) in which the gene encoding the enzyme CAT was linked to an
adenovirus E4promoterincludingaclusterof 5xGAL4 sites of the type shown in Fig. 1 and 3 (25). This plasmid was cotransfected into WOP cells with theplasmids encodingthe activator proteins. CAT activities were determined in ex-tracts prepared from the transfected cells after 48 h of incubationat33°C. In contrast totheirrelative activities in transactivation of the Py ori, the hybrid proteins GAL4-VP16 and GAL4-Ela transactivated the modified E4 pro-motermuchmoreefficiently than did theyeastGAL4protein (Table 1). Also, the truncated GAL4(1-147) protein didnot activate thepromoter.These resultsarecompatible with the results obtained inother celltypes(25, 40).
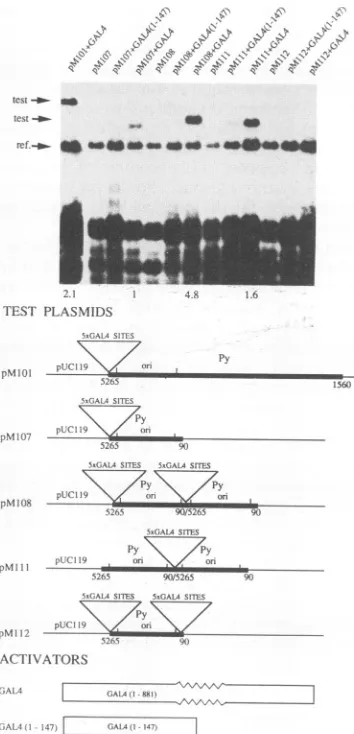
Activation ofthePyori occursintheabsenceofpromoter elements. Since plasmid pMlOl contains the Py early pro-moter,itwaspossible that the activation of the Py ori by the GAL4 and hybridproteinswascausedby their interactions
withtranscriptional elements. To examine this question,we
took advantage of the observation that in Py, unlike the related papovavirus SV40, thepromoterelementsmap out-side ori (7), and we deleted fromplasmid pM101 the early viral promoter elements, that is, the RNA start sites and about 60 bp upstream of these sites, including the TATA box.Replication of thenewplasmid, designated pM107,was then assayedas described above. As Fig. 4 shows, replica-tion of plasmid pM107 was induced by the intact GAL4 transactivator but not by the truncated protein GAL4 (1-147). Hence, the early promoter elementsare not required
TABLE 1. Transactivation of the adenovirus E4promoter by the yeast GAL4 protein and the hybrid proteins GAL4-Ela andGAL4-VP16 inmouseWOP cells
Radioactivity CAT
Transfected DNA' Transactivator
protein6
extractedinto activity RelativeCATorganic activity'
solvent(cpm)c (cpm)d
HerringDNA 1,436±43
pG5E472cat 1,809± 103 373
pG5E472cat+ pAG147 GAL4 (1-147) 1,557 ± 55 121
pG5E472cat+ pAG4 GALA 2,970 ± 390 1,534 1.0
pG5E472cat+ pSGVP GAL4-VP16 4,890± 790 3,454 2.25
pG5E472cat+ pGAL4-Ela GAL4-Ela 13,040± 580 11,604 7.6
a WOpcellsweretransfected withherringDNA or plasmid DNAs asdescribedinMaterialsandMethods.
bProteins encoded by the transfected activatorplasmids.
cCATassays wereperformed by thephaseextractionassay (42) as described in Materials and Methods. Each assay was carried out in duplicate. Numbers represent averagevalues andstandard deviations.
dValues in the firstrowofthe previous column weresubtractedfrom values in the other rows.
'TheCAT activitiesrecordedin thetwo rows atthebottomwereexpressedrelativeto theCATactivityinthe fourth row, whichwasgiven the value 1.0.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:5.612.66.558.580.679.2]'sV
n1 ax~~~4. 1.6 TFST PL.ASMID)S
0-iA s. 0-
0-rrl{)]~~~~~~~~~~~~~~~~~~~2
pM
i07
PMlIII
ilt'S 11I2
1 ',A'h TI77S. F.('( TA It rn
i'v
1561i
32: 0
. .,_ 15':AVi \//Pori
5265 111/5
115UAA11: 1G
4STl-l1, on
52659 9()/S :6 90
5vGAIA4SITI.S
;,III' on cr i
iS¢
S 11e0/5265
91)"AA[. SITES S5tiSLA4sLS-r- S
\7 917
ACTIVATORS
'IA
1.-,
i 41M|| vi(A1 1- 1t
FIG. 4. GAL4-dependent replication of a plasmid
promoterelementsandofplasmidscontainingduplications
origincoreand the 5xGAL4sites.Transfection-replication
the indicated plasmids werecarriedoutat33°C
legendstotheFig. 1and2exceptthat1.4instead pMlOl
plasmid DNAwas used. The reference plasmid
that had beenpropagatedinE. colidam. Numbers
indicatethe relativeextents of replicationofthe
for these assays, normalized relative to the extent
obtained in the assay ofplasmids pM107 plus GAL4,
giventhevalue 1.0.
forPy origin activation by the GAL4 transactivator.
ever, thedeletion didaffecttheefficiency oftransactivation
by GAL4, sincetheextent ofthe GAL4-induced of plasmid pMl07 was apparently less than one-half
plasmid pMlOl (Fig. 4). Thus, some involvement
promoterelements intheactivationofreplication
beruledout. Onthe otherhand, the reduction
of replication ofplasmid pMlO7couldbe due to
stronglarge T-Agbinding sites whichwere also
the deleted segment (6, 37).
Activationof replicationofplasmidscontainingduplications
of the origin core and of the cluster of 5xGAL4
determine whether deletion of the segment containing
early promoter elements and the T-Ag binding
plasmidpMlOl couldbe compensated byduplication
elements of the origin of replication, we constructed
assayed the replication of plasmid pM108, which contains two tandem repeats ofthe Py origin core and the cluster of
5xGAL4 sites but no promoter elements. As Fig. 4 shows, the extentof replication of plasmidpM108in thepresenceof the GAL4 transactivator was 4.8 times higher than that of
plasmid pM107. Apparently, the presence of two origin cores and two clusters of 5xGAL4 sites increased the probability of initiation of replication in a synergistic way
(see Discussion). Figure4 also shows that the extent of the GAL4-induced replication of plasmid pM108 was 2.3 times
higher than the extent of replication ofplasmid pMlOl and that replication of plasmid pM108 could not be induced by the truncated protein GAL4 (1-147).
To assess the contribution ofeach ofthe two duplicated elements to the synergistic effect on replication of plasmid
pM108,
we constructed and assayed the replication ofplas-mids pMll1 and pM112. Figure 4 shows the structures of these plasmids and the results of these replication assays. PlasmidpM1ll includes oneclusterof5xGAL4 sitesflanked by two origin cores. Theextentofreplicationof thisplasmid in the presence of the GAL4 transactivator was 1.6 times
higher than that of plasmid pM107. It should also be noted that some replication ofplasmid pMlllwas induced bythe
truncated protein GAL4 (1-147). Plasmid pM112 includes oneorigin core flanked bytwoclusters of 5xGAL4 sites. No detectable replication of this plasmid was observed even when it was cotransfected with the plasmid encoding the whole GAL4 protein or the plasmid encoding the GAL4 (1-147) protein. The origin core was not mutated in this plasmid, as revealed by DNA sequencing and by the ability of the same coretocausereplication of another plasmid into which it has beeninserted (1). Apparently, thepresence ofa second cluster of 5xGAL4 sitesnext tothe early boundaryof the core inhibited the initiation of replication of plasmid
pM112.
It should be noted that replication of anotherplas-mid includingjustonecluster of 5xGAL4 sites inserted next
to the early boundaryof thecore wasbarely detectableinthe
presence of the GAL4 protein (not shown). DISCUSSION
We have shown in this report that the yeast GAL4 transcriptional activator protein, which also enhances
tran-scription in mammalian cells (21,48), caninducereplication
of plasmids containing aPy ori in mouse cells. Replication
transactivation by the GAL4 protein wasfound to be abso-lutely dependentonthepresenceof GAL4upstream activat-ing DNA sequences next to the late boundary of the Py origin core and ofa functional Py large T-Ag. These data indicated that GAL4 protein molecules bound next to the origin core induced initiation of DNAreplication atthecore. It appears unlikely that the GAL4 protein also affected the replication indirectly by enhancement oftranscription from genes whose products are required for replication, because GAL4 recognition sequences are not found in mammalian promoters.
Since plasmid pMlOl, which was initially used in these
studies, contained the Py early promoter, we thought that
the GAL4 protein and its hybrid derivatives might activate the Py ori by virtueof their ability tostimulatetranscription from the promoter, or through interactions with general transcription factors, without actually causing transcription. Such interactions could, for example, affect the chromatin structure at the adjacent Py origin core and thus facilitate
assembly of thereplication initiation complex. The observa-tion that the truncated protein GAL4 (14147) could not
on November 10, 2019 by guest
http://jvi.asm.org/
[image:6.612.94.271.79.446.2]activate transcription and replication indicates that these two activities might be causally related. However, other results didnotsupport this hypothesis. First, wefound that theefficiencies ofactivation of the Py ori in plasmid pMlOl by the hybrid proteins GAL4-Ela and GAL4-VP16 were considerably lower than the efficiency of activation by the authentic yeast GAL4 protein. If activation of the Py ori were a consequence of transcriptional activation, then the relative efficiencies of these proteins as activators of
tran-scriptionand replication should have been correlated. How-ever, contrary tothis expectation, the hybrid proteins were foundtobemoreefficient transcriptional activators than the GAL4protein in thesame cells. Second, the GAL4 protein wasfoundtoefficiently induce replication of plasmidpM107, which contains thePyorigin coreand thecluster of 5xGAL4 sites but no Py promoter elements. Furthermore, even though the extentof theGAL4-dependent replication of the plasmid pM107wasless than half that ofplasmid pMlOl, the extentofreplication of plasmid pM108, which contains two repeats of the origin elements found in plasmid pM107 and also lackspromoterelements, was2.3 times that ofplasmid pMlOl (Fig. 4).
Other studies on the role that transcriptional activators playinreplication also supported the notion that replication transactivation by these factors may not result from the interactions that lead to transcriptional activation. In one study, it has been found that the CTF/NF-I transcriptional activators also stimulate initiation ofadenovirus DNA rep-lication (19). However, thereplication enhancement activity in this system required just the N-terminal portion of the CTF/NF-I proteins, whereas transcriptional activation re-quired, inaddition, aC-terminal domain (32). Similarly, the DNA-binding domain (POU domain) of the transcription factor Oct 1 (NFIII) suffices for stimulation ofadenovirus DNAreplication but doesnotactivate transcription(34, 46). Also, whereas the c-Jun/c-Fos heterodimer was found to activateboth transcription and replication, the cyclic AMP-responsive element-binding protein, CREB, stimulated
tran-scription but did not transactivate a Py ori containing a cyclic AMP-responsive element site (33). Thus, not all transcriptional activators alsostimulate replication. Finally, whereastransactivator proteins that bind enhancer elements stimulatetranscriptionevenwhenthese elementsareplaced atrelatively long distances from thepromoters oneither side of the transcription units, the same proteins could activate the Py ori only when the DNA sequences binding these proteins were placed at relatively short distances fromthe lateboundary of the core (14, 33).
It isinterestingtoconsiderourdataonthe GAL4-induced
replication of plasmids pM107, pM108, pMlll, and pM112 (Fig. 4)in relationtoother models that have been proposed
to account for the activation of SV40 and polyomavirus origins by transcriptional activators(8). One modelassumes
thatbinding of suchproteinstoaDNA-binding site inserted
next tothe origin core prevents chromatin assembly atthe originandthusfacilitates formation of the replication initia-tioncomplex. Suchamechanism appearsto accountforthe
invitro stimulation of SV40 DNAreplication byCTF/NF-I proteins (5). This model is compatible with the positive synergistic effectof the origin duplications onthe replication
ofplasmid pM108, for the repeat arrangement in this plas-mid, 5xGAL4 sites-core-5xGAL4 sites-core, could inhibit nucleosome assembly in a synergistic way and thus cause
theobserved4.8-fold increasein theextentof replication of this plasmid compared with thatof plasmid pM107, which contains the single-copy 5xGAL4 sites-core. However, to
accountfor the observation that the truncatedprotein GAL4 (1-147)did not activate the Pyorigins in these plasmids, one would have to assume that not only the DNA-binding domain but also the activating domains of the GAL4 or hybrid proteins were required for inhibition of chromatin assembly.Furthermore, it is difficult to reconcile this model with thecompleteinhibition of replicationof plasmid pM112
containing the repeat arrangement 5xGAL4 sites-core-5xGAL4 sites, because the presence of two clusters of 5xGAL4 sites in this plasmid should presumably prevent
chromatin assembly at the core at least as well as does a single cluster.
A second model assumes that, in analogy to the current models oftranscriptionalactivation, the DNA-binding
trans-activating proteins interact with proteins of the replication
initiationcomplex, e.g., the Py large T-Ag, thereby stabiliz-ing the complex and increasing the rate of initiation of
replication. Suchan interaction might also lead to enhance-mentofthelarge-T-Ag-dependentunwinding of the Py origin core, as suggested in studies of SV40 DNAreplication (12, 13). Thepositivesynergistic effects oftheduplicationsof the clusterof5xGAL4 sites and theorigincore onreplication of
plasmids pM108 and pM1ll could result from interactions between theproteinsthatbind theduplicatedelements. The inhibitory effect of the second cluster of 5xGAL4 sites inserted next to the early boundary of the origin core in plasmid pM112 could be accounted for if GAL4 protein
molecules bound to this element were to interact with the
replication initiation complex such that the initiation of
replication would be inhibited rather than stimulated.
Clearly, definitive experiments designed for testing these
models,whicharenotmutuallyexclusive,canbeperformed onlyin anin vitroreplication system.
ACKNOWLEDGMENTS
Wethank P. Clertant, M. R. Green,R. Kamen, D. Last, J. W.
Lillie,M. Ptashne,andG. M. Veldman for sendingusmanyofthe
plasmids used for this study. Wealso thank C. Basilico andF. G.
Kern for sending us the WOP mouse cells and A. Razin for providing theE. coli dam bacteria.
This workwassupportedbyagrantfrom theCouncil for Tobacco
Research (USA), bygrant 87-00267 from the UnitedStates-Israel Binational Science Foundation, and by a grant from the Israel Cancer ResearchFund.
REFERENCES 1. Baru,M. Unpublished data.
la.Bennet, E. R., M. Naujokas, and J. A. Hassel. 1989.
Require-ments for species-specific papovavirus DNA replication. J.
Virol.63:5371-5385.
2. Brand, A. H., G. Micklem, and K. Nasmyth. 1988. A yeast
silencer containssequencesthatcanpromote autonomous
plas-midreplication andtranscriptional activation. Cell51:709-719. 3. Campbell,B.A., and L. P. Villarreal. 1988.Functionalanalysis ofthe individual enhancer core sequences of polyomavirus: cell-specific uncoupling ofDNAreplication from transcription.
Mol. Cell. Biol. 8:1993-2004.
4. Cereghini, S., P. Herbomel, J. Jouanneau, M. Saragosti, B. Katinka, B. Bourachot, B. de Crombrugghe, and M. Yaniv. 1983.
Structure and function of the promoter-enhancer region of
polyomaand SV40. Cold Spring Harbor Symp. Quant. Biol. 47:935-944.
5. Cheng, L., and T. J. Kelly. 1989. Transcriptional activator
nuclear factorIstimulatesthereplication of SV40 minichromo-somes in vivoand in vitro. Cell59:541-551.
6. Cowie, A., and R. Kamen. 1984. Multiple binding sites for
polyomavirus large T antigen within regulatory sequences of
polyomavirusDNA.J. Virol.52:750-760.
on November 10, 2019 by guest
http://jvi.asm.org/
7. DePamphilis, M. L. 1987. Replication ofSV40and polyomavirus chromosomes, p. 1-40. In Y. Aloni (ed.), Molecular aspects of papovaviruses. Martinus Nijhoff, Boston.
8. DePamphilis, M. L. 1988. Transcriptional elements as compo-nents of eukaryotic origins of DNA replication. Cell52:635-638.
9. de Villier, J., W. Schaffner, C. Tyndall, S. Lupton, and R. Kamen. 1984. Polyomavirus DNA replication requires an en-hancer. Nature (London) 312:242-246.
10. Graham, F. L., and A. J. van der Eb. 1973. Transformation of rat cells by DNA of human adenovirus 5. Virology52:456-467.
11.
Griffin,
B. E., E. Soeda, B. G. Barrell, and R. Staden. 1981.Sequence and analysis of polyoma virus DNA, p. 843-910. InJ. Tooze (ed.), DNA tumor viruses, 2nd ed. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.
12. Guo, Z.-S., C. Gutierrez, U. Heine, J. M. Sogo, and M. L.
DePamphilis. 1989. Origin auxiliary sequences can facilitate initiation of simian virus 40 DNA replication in vitro as they do in vivo. Mol. Cell. Biol. 9:3593-3602.
13. Gutierrez, C., Z.-S. Guo, J. Roberts, and M. L. DePamphilis. 1990. Simian virus 40 origin auxiliary sequences weakly facili-tate T-antigen binding but strongly facilifacili-tate DNA unwinding. Mol. Cell. Biol. 10:1719-1728.
14. Hassel, J. A., W. J. Muller, and C. R. Mueller. 1986. The dual role of the polyomavirus enhancer in transcription and DNA replication. Cancer Cells 4:561-569.
15. Hendrickson, E. A., C. E. Fritze, W. R. Folk, and M. L. DePamphilis. 1987. The origin of bidirectional DNAreplication in polyoma virus. EMBO J. 6:2011-2018.
16. Hirt, B.
1967.
Selective extraction of polyoma DNA from infected mouse cell cultures. J. Mol. Biol. 26:365-369. 17. Hodgson, C. P., and R. Z. Fisk. 1987. Hybridization probe sizecontrol: optimized "oligolabelling." Nucleic Acids Res. 15: 6295.
18. Johnson, P. F., and S. L. McKnight. 1989. Eukaryotic transcrip-tional regulatory proteins. Annu. Rev. Biophem. 58:799-839. 19. Jones, K. A., J. T. Kadonaga, P. J.Rosenfeld,T. J.Kelly,and R.
Tjian. 1987. A cellular DNA-binding protein that activates eukaryotic transcription and DNA replication. Cell 48:79-89. 20. Jones, N. C., P. W. J. Rigby, and E. B. Ziff. 1988. Trans-acting
protein factors and the regulation of eukaryotic transcription: lessons from studies on DNA tumor viruses. Genes Dev. 2:267-281.
21. Kakidani, H., and M. Ptashne. 1988. GAL4 activates gene expression in mammalian cells. Cell 52:161-167.
22. Katinka, M., and M. Yaniv. 1983. DNA replication origin of polyomavirus early proximal boundary. J. Virol. 47:244-248. 23. Kern, F. G., and C. Basilico. 1986. An inducible eukaryotic
host-vector expression system: amplification of genes underthe control of the polyoma late promoter in a cell line producing a thermolabile large T antigen. Gene 43:237-245.
24. Kimmerly, W., A. Buchman, R. Kornberg, and J. Rine. 1988. Roles of two DNA-binding factors in replication, segregation and transcriptional repression mediated by a yeast silencer.
EMBO J. 7:2241-2253.
25.
Lillie,
J. W., and M. R. Green. 1989. Transcription activation bythe adenovirus Ela protein. Nature (London) 338:39-44. 26. Luthman, H., M. G.Nillson,and G. Magnusson. 1982.
Noncon-tiguous segments of the polyoma gene required in cis for DNA replication. J. Mol. Biol. 161:533-550.
27. Ma, J., and M. Ptashne. 1987. Deletion analysis of GAL4 defines two transcriptional activating segments. Cell 48:847-853.
28. Maniatis, T., S. Goodbourn, and A. J. Fischer. 1987. Regulation of inducible and tissue-specific gene expression. Science 236: 1237-1245.
29. Manor, H., and A. Neer. 1975. Effects of cycloheximide onvirus
DNAreplication inaninducible line of polyoma-transformed rat
cells. Cell 5:311-318.
30. McKnight, S.,and R.Tjian. 1986. Transcriptional selectivityof
viral genes in mammalian cells. Cell 46:795-805.
31. Mendelsohn, E., N. Baran, A. Neer, and H. Manor. 1982.
Integrationsiteofpolyoma virus DNA in the inducible LPTline ofpolyoma-transformed cells. J. Virol. 41:192-209.
32. Mermod, N., E. A. O'Neill, T. J.Kelly, and R. Tjian. 1989.The proline-rich transcriptional activator of CTF/NF-I is distinct fromthereplication and DNA binding domain. Cell 58:741-753. 33. Murakami, Y., M. Satake, Y. Yamaguchi-Iwai, M. Sakai, M. Muramatsu, and Y. Ito. Proc. Natl. Acad. Sci. USA, in press. 34. O'Neill, E. A., C. Fletcher, C. R. Burrow, N. Heinz, R. G. Roeder, and T. J. Kelly. 1988. Transcription factor OTF-1 is functionally identical to the DNA replication factor NF-III.
Science 241:1210-1213.
35. Passmore, S., G. T. Maine, R.Elble,C. Christ, and B.-K. Tye.
1988. Saccharomyces cerevisiae protein involved in plasmid maintenance is necessary for mating ofMATa cells. J. Mol.
Biol. 204:593-606.
36. Peden, K. W. C., J. M. Pipas, S. Pearson-White, and D. Nathans.
1980.Isolation ofmutantsofananimal virus inbacteria. Science
209:1392-1396.
37. Pomerantz, B. J., C. R. Mueller, and J. A. Hassel. 1983.
Polyomavirus large Tantigen binds independently tomultiple, unique regions onthe viral genome. J. Virol. 47:600-610.
38. Ptashne, M. 1988. How eukaryotic transcriptional activators work. Nature(London)335:683-689.
39. Rochford, R., C. T. Davis, K. K. Yoshimoto, and L. P. Villar-real. 1990. Minimal subenhancer requirements for high-level polyomavirusDNAreplication: acell-specificsynergyofPEA3
andPEAl sites. Mol. Cell. Biol. 10:4996-5001.
40. Sadowski, I., J. Ma, S. Triezenberg, and M. Ptashne. 1988.
GAL4-VP16 is an unusually potent transcriptional activator.
Nature(London) 335:563-564.
41. Sambrook, J., E. F. Fritsch, and T. Maniatis. 1989. Molecular
cloning: a laboratory manual. Cold Spring Harbor Laboratory Press,Cold Spring Harbor, N.Y.
42. Seed, B., and J.-Y. Sheen. 1988. A simple phase-extraction
assay forchloramphenicol acetyltransferase activity. Gene 67: 271-277.
43. Tang, W. J., S. L. Berger, S. J. Triezenberg, and W. R. Folk. 1987. Nucleotides in the polyomavirus enhancer that control viral transcription and DNA replication. Mol. Cell. Biol.
7:1681-1690.
44. Triezenberg, S. J., and W. R. Folk. 1984. Essential nucleotides
in the polyomavirus origin region. J. Virol. 51:437-444. 45. Veldman,G.M.,S. Lupton, and R. Kamen. 1985.Polyomavirus
enhancer contains multiple redundant sequence elements that activate both DNAreplication and geneexpression. Mol. Cell. Biol. 5:649-658.
46. Verrijzer, C. P., A. J. Kal,and P. C. vander Vliet. 1990. The
DNA binding domain (POU domain) of transcription factor oct-1 suffices for stimulation of DNA replication. EMBO J.
9:1883-1888.
47. Vieira, J., and J. Messing. 1987. Production ofsingle-stranded plasmid DNA. MethodsEnzymol. 153:3-11.
48. Webster, N., J. R. Jin, S. Green, M. Hollis, and P. Chambon.
1988. The yeast UASG is atranscriptional enhancerin human
HeLa cells in the presence of the GAL4 trans-activator. Cell
52:169-178.
49. Zhu, Z., G. M. Veldman, A. Cowie, A. Carr, B. Schaffhausen,
and R. Kamen. 1984. Construction and functional
characteriza-tion ofpolyomavirus genomes thatseparatelyencode thethree
earlyproteins. J. Virol. 51:170-180.