Copyrightq1997, American Society for Microbiology
Protective Cellular Immunity: Cytotoxic T-Lymphocyte Responses
against Dominant and Recessive Epitopes of Influenza Virus
Nucleoprotein Induced by DNA Immunization
TONG-MING FU, ARTHUR FRIEDMAN, JEFFREY B. ULMER, MARGARET A. LIU,*
ANDJOHN J. DONNELLY
Department of Virus and Cell Biology, Merck Research Laboratories, West Point, Pennsylvania 19486 Received 17 September 1996/Accepted 10 January 1997
DNA immunization offers a novel means to induce cellular immunity in a population with a heterogeneous genetic background. An immunorecessive cytotoxic T-lymphocyte (CTL) epitope in influenza virus nucleopro-tein (NP), residues 218 to 226, was identified when mice were immunized with a plasmid DNA encoding a full-length mutant NP in which the anchor residues for the immunodominant NP147-155 epitope were altered. Mice immunized with wild-type or mutant NP DNA were protected from lethal cross-strain virus challenge, and the protection could be adoptively transferred by immune splenocytes, indicating the role of cell-mediated immunity in the protection. DNA immunization is capable of eliciting protective cellular immunity against both immunodominant and immunorecessive CTL epitopes in the hierarchy seen with virus infection.
DNA vaccination provides a novel method to induce both humoral and cell-mediated immunities (8, 31). The simplicity of the technology, its ability to induce antibody (Ab) and cy-totoxic T-lymphocyte (CTL) responses in animals with differ-ent genetic backgrounds, and its preclinical efficacy in a variety of disease models (5, 7, 18, 31, 33) suggest that DNA vaccina-tion may be a useful approach for vaccine development. Ad-ditionally, the ability of DNA vaccines to induce immune re-sponses with nonreplicating materials offers an alternative to live vectors or pathogens for eliciting cell-mediated immunity for those instances where such agents may present potential safety concerns. In a previous study from our laboratory, we demonstrated that intramuscular injection of a plasmid DNA encoding influenza virus nucleoprotein (NP) induced a
vigor-ous CTL response specific for an H-2Kd-restricted epitope
147-155 in BALB/c mice (31). Furthermore, immunization with NP DNA, in which the sequence coding for NP was derived from influenza virus A/PR/8/34 (H1N1), conferred protection against a lethal dose challenge with the influenza virus A/HK/68 (H3N2), wherein much of the NP sequence
including theH-2Kd-restricted CTL epitope 147-155 was
con-served, despite a very significant antigenic difference in viral surface glycoproteins. In this report, we sought to address the following issues concerning the immunological and protective mechanisms of DNA vaccines: (i) whether a DNA vaccine permits epitope determinant selection to occur for the induc-tion of CTL responses restricted by different haplotypes; (ii) whether in vivo cross-strain protection is mediated by the cel-lular immunity induced by a DNA vaccine encoding a con-served viral internal antigen; (iii) whether the cell-mediated protective immunity is solely dependent on the CTL response against an immunodominant epitope; and (iv) whether DNA immunization can induce major histocompatibility complex (MHC) class I molecule-restricted CTL responses against epi-topes other than the immunodominant epitope.
CD81T cells recognize viral antigens in the form of short
peptides presented by MHC class I molecules on the surface of virus-infected cells (2, 30, 34). The recognition of these viral
peptides in the context of MHC molecules by CD81T cells can
trigger the specific lysis of virally infected cells (4, 19) or non-cytopathic intracellular inactivation of virus mediated by
cyto-kines such as gamma interferon and tumor necrosis factora
(15, 16). CD81CTL have thus been implicated as playing a
role in recovery or clearance of virus during a viral infection (6,
21, 22). Additionally, a role for CD41T cells in resistance to
influenza virus was indicated by experiments usingb
2-micro-globulin gene knockout (b2M2/2) mice which are defective in
expressing MHC class I molecules on the cell surface and lack
functional CD81T cells or using monoclonal antibodies to in
vivo deplete the CD81T-cell population. Several groups of
investigators reported that the b2M2/2 or CD81-depleted
mice could recover from influenza virus infection, though in a delayed fashion in some instances (1, 9, 10, 20). Thus, it ap-pears that cellular immune responses, mediated by either
CD81or CD41T cells, are important for recovery from
in-fluenza virus infection.
In this study, we generated a cDNA expression vector en-coding a mutant version of full-length NP in which the anchor residues for the NP147-155 CTL epitope were altered. By immunizing mice with the mutant version of NP DNA, we identified an immunorecessive CTL epitope in NP, residues 218 to 226, and showed that BALB/c mice immunized with the mutant NP DNA are resistant to cross-strain virus challenge. Furthermore, the resistance to cross-strain influenza virus challenge induced by the mutant form of NP DNA, like that induced by a cDNA immunization vector encoding wild-type NP, can be adoptively transferred to unprimed mice with the immune lymphocytes.
MATERIALS AND METHODS
Synthetic peptides and oligonucleotides.All synthetic peptides were custom synthesized by Chiron-Mimotopes, Inc., Raleigh, N.C., and their sequences are summarized in Table 1. The mutagenesis and sequencing oligonucleotides were synthesized by the Midland Certified Reagent Company, Midland, Tex.
DNA constructs.The expression vector used in the study, V1Ja, was derived from the vector V1J, which has been described previously (23). Briefly, it con-tains the human cytomegalovirus immediate-early gene enhancer and promoter, the intron A sequence, multiple restriction sites for cloning the gene of interest, and bovine growth hormone polyadenylation sequence. The NP gene from
in-* Corresponding author. Mailing address: Department of Virus and Cell Biology, Merck Research Laboratories, WP 16-101, West Point, PA 19486. Phone: (215) 652-3535. Fax: (215) 652-7320. E-mail: margaret _liu@merck.com.
2715
on November 9, 2019 by guest
http://jvi.asm.org/
fluenza virus A/PR/8/34 (H1N1) was cloned into theBglII andSalI sites. To generate the NPmut construct which contains two point mutations within the H-2Kd-restricted CTL epitope (residues 147 to 155), the NP DNA fragment was
cloned into pAlterSite vector from Promega Corp., Madison, Wis. The mutagen-esis was achieved by using an AlterSite kit purchased from Promega. The plasmid DNA used for immunization was purified fromEscherichia coliby a modified alkaline lysis procedure, and DNA was banded twice in a CsCl2gradient (29).
Animals.Four- to eight-week-old female BALB/c or C57BL/6 mice were purchased from Charles River Laboratories, Raleigh, N.C. The animals were maintained in the animal facility of Merck Research Laboratories at West Point, Pa. All experiments were performed in accordance with the procedures issued by International Animal Care and Use Committee.
Immunization, challenge, and adoptive transfer protocol.For immunization, mice under anesthesia induced by the administration of ketamine-xylazine were injected with 100mg of DNA dissolved in 100ml of sterile saline three times at 3-week intervals. Fifty microliters of DNA was injected bilaterally into the quad-riceps muscle each time. To prime mice with live influenza virus A/PR/8/34, mice were infected intranasally while awake (13). For challenge studies, mice were fully anesthetized and intranasally inoculated with 20ml containing a mouse-adapted strain, A/HK/68 (H3N2). This challenge dose has been characterized as 102.550% tissue culture infective doses, 300 50% mouse infective doses, or about 10 50% lethal doses. This routinely results in 90% death in unimmunized mice (13, 31). Survival and body mass of the challenged mice were monitored for up to 4 weeks.
The adoptive transfer protocol was modified from a method described by Wells et al. (32). The spleen cells harvested from the donor mice were cultured and restimulated for 6 or 7 days with syngeneic cells that had been either infected with A/PR/8/34 virus or pulsed with synthetic peptides. Interleukin-2 (IL-2; Cellular Products Inc., Buffalo, N.Y.) was added 10 U/ml on the second day of culture. Lymphocytes were washed three times with phosphate-buffered saline and resuspended at desired concentrations in phosphate-buffered saline. The recipient mice, 4 h after challenge infection with influenza virus A/HK/68, re-ceived 0.2-ml volume of lymphocytes (53107cells/mouse) through the tail veins, which had been dilated in hot water (about 508C). The survival and weight loss were monitored for up to 4 weeks.
Cells and CTL cultures.P815 (H-2d) and EL4 (H-2b) cells, routinely used as
target cells for the cytotoxicity assay, were maintained in Dulbecco’s modified Eagle medium supplemented with 10% heat-inactivated fetal bovine serum, 2 mML-glutamine, 100 U of penicillin per ml, 100mg of streptomycin per ml, 5 mM HEPES, and 0.0225% sodium bicarbonate. All cell culture reagents were purchased from GIBCO/BRL Life Technologies, Grand Island, N.Y.
Spleen cells were cultured to generate effector CTL as previously described (31). Spleen cell cultures were maintained in RPMI 1640 medium supplemented with 10% fetal bovine serum, 2 mML-glutamine, 531025Mb -mercaptoetha-nol, 25mg of pyruvic acid per ml, 100 U of penicillin per ml, 100mg of strepto-mycin per ml, 5 mM HEPES, and 0.0225% sodium bicarbonate. Irradiated syngeneic cells, either pulsed with peptide (10mM) or infected with A/PR/8/34 virus, were used as stimulator cells. Unless specified, no IL-2 was added in the first round of in vitro restimulation. Long-term CTL cultures were maintained in complete RPMI 1640 medium plus 10 U of recombinant human IL-2 per ml.
The CD41and CD81T-cell subset enrichments were performed by using T-cell subset purification columns (Biotex Laboratories, Inc., Edmonton, Al-berta, Canada). Briefly, splenocyte suspensions from mice immunized with DNA were washed and incubated with the monoclonal Abs supplied with the column. These Ab preparations consist of monoclonal Abs directed against surface marker antigens of B cells and of the T-cell subset which is intended to be depleted. The cells were then passed through a column of glass beads coated with anti-mouse immunoglobulin G which bound the cells coated with Abs. The unbound cells, the majority of which were the desired T-cell subset, were eluted
from the column and collected. The purity of CD41or CD81lymphocytes was confirmed by FACScan analysis.
Cytotoxicity assay.The cytotoxicity assays were performed as described pre-viously (31). Briefly, target cells labeled with Na51CrO4(Amersham Life Sci-ences, Arlington Heights, Ill.) were either infected with influenza virus A/Vic-toria (Vic)/73 (H3N2) or pulsed with synthetic peptides at the indicated concentration. The target cells were then mixed with CTLs at designated effec-tor/target cell ratios in 96-well plates and incubated at 378C for 4 h in the presence of 5% CO2. A 20-ml sample of supernatant from each cell mixture was counted to determine the amount of51Cr released from target cells. The per-centage of specific lysis was calculated by using the formula % specific lysis5
(E2S)/(M2S), whereErepresents the average counts per minute released from target cells in the presence of effector cells,Sis the spontaneous counts per minute released in the presence of medium only, andMis the maximum counts per minute released in the presence of 5% sodium dodecyl sulfate or 2% Triton X-100.
RESULTS
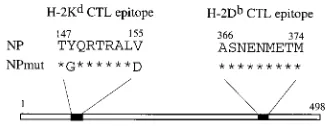
Generating the NPmut DNA construct.Two significant point mutations, in terms of both amino acid size and charge, were introduced into the NP147-155 CTL epitope sequence at the putative anchor positions (11, 28). The resultant DNA con-struct, named NPmut, encoded a full-length NP with these two
mutations. The H-2Db-restricted CTL epitope 366-374
re-mained intact in the NPmut construct (Fig. 1). A synthetic peptide corresponding to the mutant epitope sequence, NP147-155mut peptide, was prepared (peptide sequences are shown in Table 1). In a peptide competition assay, increasing concen-trations of competitor peptides, including NP147-155mut
pep-tide as well as H-2Db-restricted NP366-374 peptide and an
H-2Kd-restricted listeriolysin O CTL epitope peptide (17, 27),
were mixed with 1 nM NP147-155 peptide and used to pulse target cells. The cells were then incubated with CTL specific
for NP147-155 peptide. As shown in Fig. 2, the H-2Kd
-re-stricted listeriolysin O epitope peptide was able to effectively block the CTL recognition of NP147-155 peptide-pulsed target cells at high molar concentrations, whereas neither
NP147-155mut peptide norH-2Db-restricted NP366-374 peptide was
capable of inhibiting NP147-155-specific CTL recognition at any concentrations tested. This result indicated that the mu-tant peptide was not able to inhibit the CTL recognition of
NP147-155 peptide in the context ofH-2Kdmolecules,
presum-ably because NP147-155mut peptide did not bind to H-2Kd
molecules.
CTL responses in mice immunized with the NPmut DNA construct.(BALB/c3C57BL/6)F1mice (H-2dxb) were
immu-nized with NP DNA or NPmut DNA by intramuscular injec-tion, and their spleen cells were restimulated in vitro with
syngeneic spleen cells pulsed with either H-2Kd-restricted
NP147-155 peptide orH-2Db-restricted NP366-374 peptide for
7 days. The CTL responses were then examined in a standard cytotoxicity assay. As shown in Fig. 3, the spleen cells from the
F1 mice immunized with NP DNA lysed P815 cells (H-2d)
sensitized with NP147-155 peptide and EL4 cells (H-2b)
sen-sitized with NP366-374 peptide, respectively, but they did not
In single-letter code. Boldface letters indicate the potential anchor residues for the epitope.
bAbility of the peptide to bind toH-2Kdwas unknown.
cThe restricting MHC molecule for this peptide has not been reported.
on November 9, 2019 by guest
http://jvi.asm.org/
[image:2.612.353.516.70.133.2] [image:2.612.57.300.83.182.2]lyse cells pulsed with inappropriate peptides (P815 cells plus NP366-374 or EL4 cells plus NP147-155). Spleen cells from the NPmut DNA-immunized mice demonstrated cytolytic activity against EL4 cells pulsed with NP366-374 peptide, but killing against the NP147-155-sensitized P815 cells was absent in these splenocytes. This finding showed that NP DNA immunization could induce CTL responses of appropriate specificities ac-cording to the MHC haplotypes of the host. The results also indicated that the point mutations introduced in the NP se-quence functionally abolished the CTL response against the NP147-155 epitope, but that these mutations did not compro-mise the processing and presentation of the NP protein since
the response against the H-2Db-restricted epitope remained
intact.
Although they did not recognize the NP147-155 CTL epi-tope, virus-restimulated spleen cells from NPmut
DNA-immu-nized BALB/c mice (H-2d) still could lyse influenza
virus-in-fected target cells. As shown in Fig. 4, spleen cells from mice receiving vector DNA (plasmid not encoding any gene) did not recognize any target cells, whereas splenocytes from NP DNA-immunized mice, restimulated with virus or with NP147-155 peptide, demonstrated cytolytic activities toward target cells either infected with influenza virus A/Vic/73 or pulsed with NP147-155 peptide. Splenocytes of the NPmut DNA-immu-nized mice, when restimulated with NP147-155 peptide, did not lyse any target cells as expected, whereas restimulation of these splenocytes with A/PR/8/34 virus in vitro enabled them to recognize and lyse the A/Vic/73 influenza virus-infected target cells, even though the NP147-155-specific killing was absent. These results indicated that a cytotoxic immune response dif-ferent from the NP147-155 epitope response was induced by NPmut DNA immunization in BALB/c mice, and this CTL response could lyse virus-infected target cells.
Characterization of the cytotoxicity seen in NPmut DNA-immunized mice. To determine whether the cytolytic activity seen in the spleen cells from NPmut DNA-immunized mice
included a CD81CTL response, we enriched the T-cell
pop-ulation for either CD81or CD41T-cell subsets from the mice
immunized with NP DNA or NPmut DNA. The enriched T cells were restimulated in vitro with syngeneic cells infected with A/PR/8/34 influenza virus for 7 days and used as effector cells in the 4-h cytotoxicity experiment depicted in Fig. 4. The
enriched CD41T-cell cultures were included as a control.
As shown in Fig. 5, the CD81T cells derived from the mice
immunized with NP DNA recognized both virus-infected and
NP147-155 peptide-pulsed targets, and the CD81T cells from
the mice immunized with NPmut DNA lysed only the virus-infected target cells. As expected, no cytotoxicity against either virus-infected or NP147-155 peptipulsed targets was
de-tected in the control CD41-enriched T-cell cultures. The
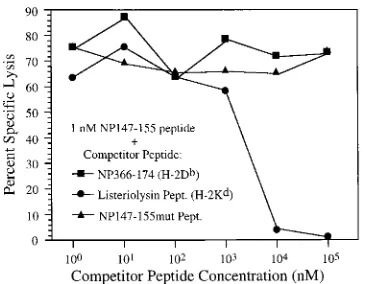
FIG. 2. NP147-155mut peptide does not inhibit the CTL recognition of NP147-155-pulsed cells. P815 cells (H-2d), labeled with51Cr and pulsed with mixture of 1 nM NP147-155 peptide and the competitor peptide at the indicated concentrations, were assayed with NP147-155-specific CTL for specific lysis at an effector to target cell ratio of 10 to 1, and the lysis of P815 cells pulsed with no peptide was less than 5%. The listeriolysin CTL epitope peptide (Pept.; residues 90 to 98) binds toH-2Kdmolecules (17, 27) and competitively inhibits the CTL
recognition of NP147-155 peptide. NP147-155mut peptide and NP366-374 pep-tide (H-2Db-binding peptide) fail to prevent the target cells from being sensitized
[image:3.612.88.271.69.211.2]by NP147-155 peptide. The results are representative of three experiments.
FIG. 3. Immunization with NPmut DNA still induces theH-2Db-restricted
CTL response but does not induce a CTL response specific for the H-2Kd
-restricted epitope NP147-155 inH-2dxbmice. (BALB/c3C57BL/6)F1mice were
immunized with 100mg of NP DNA or NPmut DNA intramuscularly three times at 3-week intervals. The spleen cells were collected and restimulated in vitro for 7 days with syngeneic splenocytes pulsed with NP147-155 or NP366-374 peptide (Pept.; 10mM) and used as effector cells in a 4-h cytolytic assay against P815 cells (H-2d) pulsed with NP147-155 or NP366-374 peptide or EL4 cells (H-2b) pulsed
with NP147-155 or NP366-374 peptide. The results are representative of three experiments.
FIG. 4. NPmut DNA immunization results in CTL which follows in vitro restimulation with virus can lyse influenza virus-infected target cells. BALB/c mice were intramuscularly immunized with 100mg of vector DNA (not encoding a protein), NP DNA, or NPmut DNA, and their spleen cells were restimulated in vitro for 7 days with syngeneic cells infected with influenza virus A/PR/8/34 or pulsed with NP147-155 peptide (Pept.). The CTL responses were measured in a cytotoxicity assay against P815 cells that were mock infected, infected with influenza virus A/Vic/73, or pulsed with NP147-155 peptide. The results are representative of two experiments.
on November 9, 2019 by guest
http://jvi.asm.org/
[image:3.612.339.534.70.239.2]CD81T cells from mice immunized with vector control DNA
likewise did not recognize any target cells. The CD81T cells
derived from the mice immunized with NP DNA recognized both virus-infected and NP147-155 peptide-pulsed targets, and
the CD81T cells from the mice immunized with NPmut DNA
lysed only the virus-infected target cells. These data demon-strated that the cytolytic activity induced in BALB/c mice im-munized with NPmut DNA, which was directed against a CTL epitope other than the NP147-155 epitope, included a
popu-lation of CD81T cells. Since our experimental conditions were
intentionally not optimized for detecting the lytic activity of
CD41cells, the existence of a CD41CTL component also is
possible.
Identification of an immunorecessive CTL epitope in NP.
MostH-2Kd-binding peptides identified to date contain a
ty-rosine at the second position and a hydrophobic residue at the C terminus to serve as two major anchor residues (11, 28). Therefore, we scanned the amino acid sequence of NP and
selected three peptides with this consensusH-2Kd-binding
mo-tif. Since the peptide corresponding to NP residues 39 to 47 was unable to be synthesized due to high hydrophobicity, we tested only the two synthetic peptides corresponding to NP110-119 and NP218-226 (Table 1). As shown in Table 2, virus-restimulated CTL from mice immunized with NP DNA or NPmut DNA recognized target cells infected with A/Vic/73 virus, whereas only the NP DNA-induced CTL could lyse the target cells pulsed with NP147-155 peptide. NP110-119 pep-tide-pulsed target cells were not recognized by either NP DNA- or NPmut DNA-induced CTL. However, target cells sensitized with NP218-226 peptide were specifically killed by NPmut DNA-induced CTL. In contrast, the NP DNA-induced CTL did not lyse the NP218-226 peptide-pulsed cells. Thus, NP218-226 behaves as a typical immunorecessive epitope in that the CTL response against this epitope can be detected only when the immunodominant CTL epitope, NP147-155, is
absent from the immunogens used to prime CTL precursors in vivo (24–26).
The NP218-226 CTL epitope overlaps with a Th epitope (residues 216 to 229) identified earlier in BALB/c mice (14). Therefore, to confirm that the epitope that we identified was
indeed an H-2Kd-binding peptide, we performed a peptide
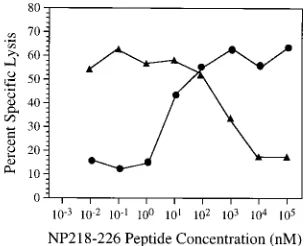
[image:4.612.89.266.73.239.2]competition assay. We used CTL cultures from NP DNA- or NPmut DNA-immunized mice to lyse the target cells sensitized by the two peptides, since the recognition of NP147-155 and that of NP218-226 epitopes by the NP DNA- and NPmut DNA-induced CTL were shown to be mutually exclusive (Ta-ble 2). NP147-155 peptide at a constant concentration (1 nM) was mixed with increasing concentrations of NP218-226 pep-tide. Target cells, pulsed with the mixture of peptides, were incubated with the CTL cultures described above. As shown in Fig. 6, at low molar concentrations of NP218-226 peptide, target cells were primarily lysed by NP147-155-specific CTL, whereas at high molar concentrations, NP218-226 peptide could abrogate the CTL recognition of NP147-155 and sensi-tize target cells for lysis by NPmut DNA-induced CTL. This finding is consistent with the hypothesis that, like NP147-155
peptide, NP218-226 peptide binds toH-2Kdmolecules.
Mice immunized with NPmut DNA were protected from influ-enza virus challenge.Experiments from our laboratory previously demonstrated that immunization of mice with a DNA plasmid
FIG. 5. The cytotoxicity of the splenocytes from both NP DNA- and NPmut DNA-immunized mice is mediated by CD81T cells. Spleen cells from mice immunized with vector DNA, NP DNA, or NPmut DNA were enriched for either CD41or CD81T cells by T-subset enrichment columns. The purities of the enriched T-cell subset were confirmed by FACScan analysis; the percentages of undesired T-cell subset in the enriched cell population are less than 5% for CD41T cells and less than 2% for CD81T cells. The enriched T-cell popula-tions were restimulated in vitro for 7 days with syngeneic cells infected with influenza virus A/PR/8/34 in the presence of 10 U of IL-2 per ml, and the cultures were diluted and used as effectors at the indicated effector-to-target cell ratios against P815 cells that were mock infected, infected with influenza virus A/Vic/ 73, or pulsed with NP147-155 peptide in a cytotoxicity assay.
FIG. 6. NP218-226 peptide competes with NP147-155 peptide for CTL rec-ognition. P815 cells pulsed with the mixture of 1 nM NP147-155 peptide and NP218-226 peptide at the indicated concentrations were incubated with NP147-155-specific CTL (å) or NP218-226-specific CTL (F) in the51Cr release assay. The effector-to-target cell ratio was 5 to 1, and the nonspecific lysis of P815 cells by both CTL cultures were less than 5%. The results are representative of three experiments.
2.5:1 32.7 7.2 2 18.2
aCTLs were cultured with syngeneic cells infected with A/PR/8 virus in the
presence of IL-2.
bTarget cells were pulsed with 10mM synthetic peptides. cThe data are representative of two separate experiments.
on November 9, 2019 by guest
http://jvi.asm.org/
[image:4.612.316.556.82.205.2] [image:4.612.360.512.541.664.2]encoding the wild-type NP sequence provided protection against death and morbidity following a lethal-dose influenza virus challenge (31). Since this method of immunization in-duced a robust CTL response against the NP147-155 epitope, we sought to determine whether the NPmut DNA immuniza-tion could still provide protecimmuniza-tion against the cross-strain chal-lenge with A/HK/68 (H3N2) in the absence of the CTL re-sponse to NP147-155 epitope. As shown in Fig. 7, like the mice immunized with NP DNA, all of the BALB/c mice intramus-cularly injected with NPmut DNA three times at 3-week inter-vals survived challenge, whereas the mice immunized with con-trol vector DNA were susceptible to the virus challenge.
To evaluate the role of cell-mediated immunity in the pro-tection that we observed with DNA immunization, we per-formed lymphocyte adoptive transfer experiments. Splenocytes from NP DNA- or NPmut DNA-immunized mice were acti-vated in vitro by restimulation with syngeneic spleen cells in-fected with A/PR/8/34 virus. Spleen cells from mice inin-fected with A/PR/8/34 virus were restimulated in vitro with syngeneic cells infected with A/PR/8/34 virus to serve as positive control, and spleen cells from mice injected with human
immunodefi-ciency virus (HIV) gag DNA were stimulated in vitro with
syngeneic spleen cells pulsed with Gag193-212 peptide to serve as negative control (12). The CTL cultures were functionally cytolytic, as indicated by their ability to specifically lyse P815 cells pulsed with NP147-155 peptide or Gag193-212 peptide, respectively (results not shown). The lymphocytes were
adop-tively transferred into age-matched naive mice (53107cells/
mouse) that had been challenged with influenza virus A/HK/68 4 h previously. As shown in Fig. 8, mice that received spleno-cytes from the mice immunized with NP DNA, NPmut DNA, or A/PR/8/34 virus were protected from death, whereas the majority of the mice receiving no treatment or spleen cells
from the mice immunized with HIVgag DNA succumbed to
challenge.
DISCUSSION
This study was undertaken to evaluate the CTL responses against NP epitopes presented in the context of MHC class I molecules after DNA immunization (31). We demonstrated
that DNA immunization generated CD81 CTL specific for
MHC class I-restricted epitope peptide. Determinant selection for the CTL epitope peptides was observed among different haplotypes, such that even if one haplotype-restricted epitope
was altered, an epitope peptide restricted by another allele remained functional (Fig. 3). By immunizing BALB/c mice with a mutant NP DNA construct in which the NP147-155 CTL epitope was functionally abolished (Fig. 2 and 3), no CTL response against the NP147-155 epitope was detected. How-ever, a CTL response was induced that was found to be
me-diated by CD81T cells and was directed against an
immuno-recessive CTL epitope (Fig. 4 and 5). The epitope, restricted by
H-2Kdmolecules, was identified and mapped to residues 218 to
226 of NP (Table 2). Although virus-infected cells presented
both peptides in the context ofH-2Kdmolecules, spleen cells
from BALB/c mice that had been infected with influenza virus in vivo and then restimulated with virus in vitro demonstrated the CTL response only against the immunodominant epitope, NP147-155 (results not shown). Like the immunorecessive CTL epitopes reported by others (24–26), the CTL response against NP218-226 could be primed in vivo only when the immunodominant epitope NP147-155 was altered due to site-directed mutagenesis in the immunizing NP DNA vector (NPmut DNA). Thus, in this aspect of antigen processing and presentation for the induction of CTL responses, DNA immu-nization appeared to mimic natural viral infection. Hence, DNA vaccines may provide a useful tool to dissect CTL epitopes within viral antigens and define their roles in viral immunity.
[image:5.612.94.262.69.195.2]The mechanisms underlying the immunorecessive properties of the NP218-226 epitope are yet to be understood. Since both NP147-155 and NP218-226 peptides were presented by influ-enza virus-infected cells (Table 2), it appeared that both pep-tides could be generated and presented by the MHC class I-dependent antigen processing pathway. Thus, it appears un-likely that a complete lack of binding of NP218-226 peptide to class I molecules in the presence of the NP147-155 epitope would account for the former being immunorecessive. How-ever, the relative amounts of the two peptides presented by virus-infected cells and the number of occupied MHC mole-cules required for cytolysis by the CTL for the two peptides are undetermined. Therefore, the preferential display of one
FIG. 7. Mice immunized with NPmut DNA are protected from lethal influ-enza virus challenge. BALB/c mice immunized intramuscularly three times at 3-week intervals with 100mg of the indicated DNA constructs (10 mice/group) were challenged under anesthesia intranasally with 300 50% tissue culture in-fective doses of influenza virus A/HK/68 (H3N2) (about 10 50% lethal doses). The proportion of surviving mice is shown. The results are representative of three experiments.
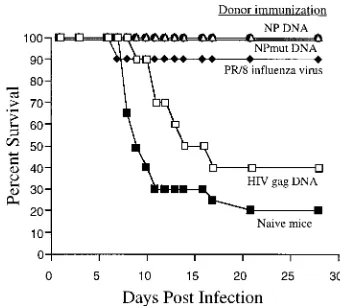
FIG. 8. NP DNA- and NPmut DNA-induced protection can be adoptively transferred by spleen cells. Spleen cells from mice primed with influenza virus A/PR/8/34 or immunized with NP DNA, NPmut DNA, or HIVgagDNA were harvested and restimulated in vitro, respectively, with syngeneic cells infected with A/PR/8/34 (for lymphocytes in groups denoted byF,Ç, and}) or syngeneic
cells pulsed with Gag193-212 peptide (for lymphocytes in the group denoted by
h) for 7 days in the presence of 10 U of IL-2 per ml. These lymphocytes were adoptively transferred into age-matched naive mice (53107cells/mouse, 10 mice/group) that had been intranasally challenged with A/HK/68 (H3N2) 4 h previously. The proportion of surviving mice is shown. The results are represen-tative of two separate experiments.
on November 9, 2019 by guest
http://jvi.asm.org/
[image:5.612.349.519.71.224.2]protection (31). It is of interest that NPmut DNA-induced
cellular immunity, including a CD81CTL response against the
NP218-226 epitope, can still be protective in the absence of the normally dominant NP147-155-specific CTL response (Fig. 7
and 8). Both CD81and CD41 T-cell populations are likely
important for the cross-strain protection with NP DNA immu-nization, as observed from in vivo T-cell subset depletion
stud-ies. In those studies, anti-CD81Ab treatment completely
ab-rogated NP DNA-induced protection, and anti-CD41 Ab
treatment diminished the protection (30a). Moreover, an
ad-ditional study showed that enriched CD81T cells from mice
immunized with NP DNA or NPmut DNA, after in vitro re-stimulation with syngeneic spleen cells infected with A/PR/8 virus, can confer full protection in naive mice when adoptively
transferred (13a). The importance of CD81T-cell response in
the protective immunity induced by NP DNA immunization
also was supported by adoptive transfer of theH-2Kd-restricted
NP147-155 peptide-restimulated splenocytes from mice immu-nized with NP DNA. These lymphocytes were shown to be fully protective for survival when adoptively transferred into virus-infected naive mice, compared to the 20% survival rate of untreated naive mice (results not shown), indicating that the
NP147-155 peptide-specific CD81CTL are protective against
cross-strain virus challenge. However, direct restimulation of splenocytes from NPmut DNA-immunized mice with NP218-226 peptide in the secondary culture did not result in detect-able NP218-226-specific CTL (results not shown), in contrast to restimulation with whole virus, which was effective at expanding NP218-226-specific CTL. This finding, although hindering our effort to demonstrate that the NP218-226 peptide-specific CTL are protective effectors in the lympho-cyte adoptive transfer experiment, may indicate the poten-tial mechanism underlying the immunorecessive nature of NP218-226, as demonstrated in a recent study that detection of the weak CTL response against a subdominant CTL epitope in influenza virus hemagglutinin, determined by the limited T-cell precursors, required in vitro restimulation of virus-infected splenocytes followed by restimulation with the synthetic pep-tide (3).
In summary, we generated a mutant version of NP which functionally abrogated the CTL response against epitope 147-155 in BALB/c mice. We showed that the CTL response against another epitope restricted by different allele remained intact. Moreover, we demonstrated that the cross-strain pro-tection by NP DNA and NPmut DNA immunization, with or without the induction of the dominant NP147-155-specific CTL response, respectively, was mediated through cell-medi-ated immunity. Furthermore, DNA immunization with the
NPmut construct in mice uncovered an H-2Kd-restricted
im-munorecessive epitope (NP218-226). Our studies show that DNA immunization can effectively induce CTL responses in accordance with a hierarchical rank in CTL epitope immuno-genicities. This observation also suggests that the pathways of antigen processing and presentation for MHC class I-restricted
2.Bjorkman, P. J., M. A. Saper, B. Samraoui, W. S. Bennett, J. L. Strominger, and D. C. Wiley.1987. The foreign antigen binding site and T cell recognition regions of class I histocompatibility antigens. Nature (London)329:512–518. 3.Cao, W., B. A. Myers-Powell, and T. J. Braciale.1996. The weak CD81CTL response to an influenza hemagglutinin epitope reflects limited T cell avail-ability. J. Immunol.157:505–511.
4.Christinck, E. R., M. A. Luscher, B. H. Barber, and D. B. Williams.1991. Peptide binding to class I MHC on living cells and quantitation of complexes required for CTL lysis. Nature (London)352:67–70.
5.Cox, G. J. M., T. J. Zamb, and L. A. Babiuk.1993. Bovine herpesvirus 1: immune responses in mice and cattle injected with plasmid DNA. J. Virol.
67:5664–5667.
6.Doherty, P. C., W. Allan, and M. Eichelberger.1992. Roles ofabandgdT cell subsets in viral immunity. Annu. Rev. Immunol.10:123–151. 7.Donnelly, J. J., A. Friedman, D. Martinez, D. L. Montgemery, J. W. Shiver,
S. L. Motzel, J. B. Ulmer, and M. A. Liu.1995. Preclinical efficacy of a prototype DNA vaccine: enhanced protection against antigenic drift in in-fluenza virus. Nat. Med.1:583–587.
8.Donnelly, J. J., J. B. Ulmer, J. W. Shiver, and M. A. Liu.1997. DNA vaccines. Annu. Rev. Immunol.60:163–172.
9.Eichelberger, M., W. Allan, M. Zijlstra, R. Jaenisch, and P. C. Doherty.
1991. Clearance of influenza virus respiratory infection in mice lacking class I major histocompatibility complex-restricted CD81T cells. J. Exp. Med. 174:875–880.
10. Epstein, S. L., J. A. Misplon, C. M. Lawson, E. K. Subbarao, M. Connors, and B. R. Murphy.1993.b2-Microglobulin-deficient mice can be protected against influenza A infection by vaccination with vaccinia-influenza recom-binants expressing hemagglutinin and neuraminidase. J. Immunol.150:5484– 5493.
11. Falk, K., O. Ro¨tzschke, S. Stevanovicˇ, G. Jung, and H. G. Rammensee.1991. Allele-specific motifs revealed by sequencing of self-peptides eluted from MHC molecules. Nature (London)351:290–296.
12. Frankel, F. R., S. Hegde, J. Lieberman, and Y. Paterson.1995. Induction of cell-mediated immune responses to human immunodeficiency virus type 1 Gag protein by usingListeria monocytogenesas a live vaccine vector. J. Im-munol.155:4775–4782.
13. Friedman, A., J. J. Donnelly, R. R. Deck, M. A. Liu, and D. Martinez.1993. Modulation of the virulence of an H3N2 influenza virus in mice by method of infection.InH. S. Ginsberg, F. Brown, R. M. Chanock, and R. A. Lerner (ed.), Modern approaches to new vaccines including prevention of AIDS. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y. 13a.Fu, T.-M., et al.Unpublished data.
14. Gao, X.-M., F. Y. Liew, and J. P. Tite.1989. Identification and characteriza-tion of T helper epitopes in the nucleoprotein of influenza A virus. J. Im-munol.143:3007–3014.
15. Guidotti, L. G., P. Borrow, M. V. Hobbs, B. Matzke, I. Gresser, M. B. A. Oldstone, and F. V. Chisari.1996. Viral cross talk: intracellular inactivation of the hepatitis B virus during an unrelated viral infection of the liver. Proc. Natl. Acad. Sci. USA93:4589–4594.
16. Guidotti, L. G., T. Ishikawa, M. V. Hobbs, B. Matzke, R. Schreiber, and F. V. Chisari.1996. Intracellular inactivation of the hepatitis B virus by cytotoxic T lymphocytes. Immunity4:25–36.
17. Harty, J. T., and M. J. Bevan.1992. CD81T cells specific for a single nonamer epitope of Listeria monocytogenes are protective in vivo. J. Exp. Med.175:1531–1538.
18. Huygen, K., J. Content, O. Denis, D. L. Montgomery, A. M. Yawman, R. R. Deck, C. M. DeWitt, I. M. Orme, S. Baldwin, C. D’Souza, A. Drowart, E. Lozes, P. Vandenbussche, J. Van Vooren, M. A. Liu, and J. B. Ulmer.1996. Immunogenicity and protective efficacy of a tuberculosis DNA vaccine. Nat. Med.2:893–898.
19. Ka¨gi, D., B. Ledermann, K. Bu¨rki, P. Seiler, B. Odermatt, J. Olsen, E. R. Podack, R. Zinkernagel, and H. Hengartner.1994. Cytotoxicity mediated by T cells and natural killer cells is greatly impaired in perforin-deficient mice. Nature (London)369:31–37.
20. Lightman, S., S. Cobbold, H. Waldmann, and B. A. Askonas.1987. Do L3T41T cells act as effector cells in protection against influenza virus infection. Immunology62:139–144.
on November 9, 2019 by guest
http://jvi.asm.org/
21. McMichael, A.1994. Cytotoxic T lymphocytes specific for influenza virus. Curr. Top. Microbiol. Immunol.189:75–91.
22. McMichael, A. J., F. M. Gotch, G. R. Noble, and P. A. S. Beare.1983. Cytotoxic T-cell immunity to influenza. N. Engl. J. Med.309:13–17. 23. Montgomery, D. L., J. W. Shiver, K. R. Keander, H. C. Perry, A. Friedman,
D. Martinez, J. B. Ulmer, J. J. Donnelly, and M. A. Liu.1993. Heterologous and homologous protection against influenza virus A by DNA vaccination: optimization of DNA vectors. DNA Cell Biol.12:777–783.
24. Mylin, L. M., R. H. Bonneau, J. D. Lippolis, and S. S. Tevethia.1995. Hierarchy among multipleH-2b-restricted cytotoxic T-lymphocyte epitopes
within simian virus 40 T antigen. J. Virol.69:6665–6677.
25. Oldstone, M. B. A., H. Lewicki, P. Borrow, D. Hudrisier, and J. E. Gairin.
1995. Discriminated selection among viral peptides with the appropriate anchor residues: implications for the size of the cytotoxic T-lymphocyte repertoire and control of viral infection. J. Virol.69:7423–7429.
26. Oukka, M., N. Riche´, and K. Kosmatopoulos.1994. A nonimmunodominant nucleoprotein-derived peptide is presented by influenza A virus-infected H-2bcells. J. Immunol.152:4843–4851.
27. Pamer, E. G., J. T. Harty, and M. J. Bevan.1991. Precise prediction of a dominant class I MHC-restricted epitope of Listeria monocytogenes. Nature (London)353:852–855.
28. Rammensee, H.-G., K. Falk, and O. Ro¨tzschke.1993. Peptides naturally presented by MHC class I molecules. Annu. Rev. Immunol.11:213–244.
29. Sambrook, J., E. F. Fritsch, and T. Maniatis.1989. Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
30. Townsend, A. R. M., J. Rothbard, F. M. Gotch, G. Bahadur, D. Wraith, and A. J. McMichael.1986a. The epitopes of influenza nucleoprotein recognized by cytotoxic T lymphocytes can be defined with short synthetic peptides. Cell
44:959–968.
30a.Ulmer, J. B., et al.Unpublished data.
31. Ulmer, J. B., J. J. Donnelly, S. E. Parker, G. H. Rhodes, P. L. Felgner, V. J. Dwarki, S. J. Gromkwoski, R. R. Deck, C. M. DeWitt, A. Friedman, L. A. Hawe, K. R. Leander, D. Martinez, H. C. Perry, J. W. Shiver, D. L. Mont-gomery, and M. A. Liu.1993. Heterologous protection against influenza by injection of DNA encoding a viral protein. Science259:1745–1749. 32. Wells, M. A., F. A. Ennis, and P. Albrecht.1981. Recovery from a viral
respiratory infection. II. Passive transfer of immune spleen cells to mice with influenza pneumonia. J. Immunol.126:1042–1046.
33. Xiang, X., and H. C. J. Ertl.1995. Manipulation of the immune response to a plasmid-encoded viral antigen by coinoculation with plasmids expressing cytokines. Immunity2:129–135.
34. Zinkernagel, R. M., and P. C. Doherty.1979. MHC-restricted cytotoxic T cells: studies on the biological role of polymorphic major transplantation antigens determining T-cell restriction-specificity, function and responsive-ness. Adv. Immunol.27:52–177.