Vol.63, No. 5 JOURNALOFVIROLOGY, May1989,P. 2233-2243
0022-538X/89/052233-11$02.00/0
Copyright © 1989,American SocietyforMicrobiology
The
Sendai Virus
Nucleocapsid Exists
in
at
Least
Four
Different Helical States
E. H. EGELMAN,L* S.-S. WU,1 M. AMREIN,2 A. PORTNER,3 AND G.
MURTI3
Department of Molecular Biophysics and Biochemistry, Yale University, Box6666, 260 Whitney Avenue, NewHaven, Connecticut065111;Institutefor Cell Biology, E.T.H.-Honggerberg, Zurich, Switzerland2; and Laboratoryof Virology,
St. Jude Children's ResearchHospital, Memphis, Tennessee38101-03183 Received 15 December1988/Accepted 27 January 1989
Sendai virusnucleocapsidshave been observed by electron microscopy tocoexist inthree differenthelical pitch conformations, 5.3, 6.8, and37.5 nm. The5.3- and 6.8-nm conformations arepresentboth in uranyl acetatenegatively stained preparations andin tantalum-tungstenmetal-shadowed preparations, whereas the 37.5-nm conformation, whichhasnotbeenpreviously reported, ispresentonlyinthe shadowed preparations. The5.3-nm pitch conformationappearstobeamixtureoftwodiscretestructuralstates,withasmall difference inthe twistof thestructurebetween thetwo. We haveusedimagereconstruction techniques on anaveraged datasetfrom eight negatively stained nucleocapsidstoproduceathree-dimensionalreconstruction at2.4-nm resolutionofthestructureinoneofthe5.3-nm pitchstates.Thereare13.07nucleocapsid protein(NP) subunits
ineachturnofthe helix inthisstate.Thehelicalrepeatis79.5nm,containing 196 subunits in15turnsof the
left-handed 5.3-nmhelix. Thearrangementof subunits producesa5.0-nm-diameter hollowcorewhichforms aninternal helicalgroove.The RNAaccountsforabout 3%ofthemassofthenucleocapsid, andsoitslocation
is notconspicuous in the reconstruction. Becausethe RNAremains associatedwith the NPsubunits during mRNAtranscription and genome replication, structural transitions in the nucleocapsid may determine the accessibility ofthegenometopolymerases. Alternatively, thelarge hollowcoreand internal helicalgroove we
have reconstructedmayallow accesstothe RNAevenin thetightlycoiled5.3-nm pitch conformation.
Sendai virus has been studiedextensively as aprototype
of the Paramyxoviridae, a family of enveloped
single-stranded RNA-containing animal viruses whose site of
rep-licationis thecell cytoplasm. Progenyparamyxovirus parti-cles areliberated from infected cellsby budding. Thus, the
paramyxovirus envelope is derived from the cell surface membrane, The lipid composition in thisenvelope is similar tothatinthe surface membranes of uninfected cells, but the
proteins are virus specified (23). The intact virus particles areusuallyrounded but appearirregularin shape. They are of variable size;diametersrangingfrom 100 to 800 nm have beenreported (11, 16, 17, 20, 22).
A single nucleocapsid, consisting of a strand of RNA
associatedwithalargenumber ofnucleocapsid protein (NP) subunits in arod-shaped helical structure, isnormally
con-tained within theenvelope. Fromthe nucleotide sequence, thepredicted
molecular
weight of the NPprotein is 56,543 (27). The nucleocapsid rod is about 1.0 ,um long (11, 18, 20) and has adiameter ofabout 20.0 nm (11). A hollow core of about 5.0-nmdiameter runsalongitslength (11, 17,21). Theone-start helix of the nucleocapsid has been characterized
bothasleft-handed (31) andright-handed (18) and has been
previously suggested tocontaineither 11 or 13 NPsubunits in each turn(11), 13 subunits (19), or 15 subunits (18). This helix has a pitch ofabout 5.0nm in its most tightly coiled
state(1, 11, 16, 17, 20, 21). However, extended
conforma-tions ofthenucleocapsid have been previously observed for Sendaivirus(14) and for otherparamyxoviruses (4, 30).
TheRNAof theparamyxovirusgenome iscomplementary in sequence to the viral mRNA. As a result,
paramyxovi-ruses are described as negative-strand viruses. mRNA is
produced by transcription of the genome into a number of
partial replicas (2), and transcription and replication of the
* Correspondingauthor.
viral RNA occur while it
remnains
associated with the NPsubunits(23).This characteristicsetsparamyxoviruses apart from most other typesof viruses. Theassociation oftheviral RNA with the nucleocapsid is so strong that the only
unencapsidated virus-specific RNAmolecules found in cells infected byparamyxoviruses arethe viral mRNAs (3, 25).
There are two additional proteins present in the intact
nucleocapsid at alower
stoichiometry,
and these areimpli-cated in theenzymaticprocesses ofRNAtranscriptionand
replication. Thesearethepolymerase-associated(P)protein
(79,000Mr) and the Lprotein(>200,000 Mr), and themolar ratio of NP-P-L in the virionis approximately 100:10:1(26). These proteins act in concert; neither one is capable of
catalyzingRNAsynthesiswhen addedalonetonucleocapsid templatesin the highlyhomologous Newcastle disease virus system, buttogethertheyformanactivetranscriptive com-plex (13). Both the Pprotein(32) and the Lprotein(33) have been visualized by immunogold labeling and electron mi-croscopy as associated with the nucleocapsid at clusters along its length. Thus, the NP, P, and L proteins together form a complex which is the enzymatic machinery for
transcriptionandreplication.Thisnotion issupportedby the
factthatinvitro, thefully encapsidatedRNAis thetemplate
for RNA synthesis in cell-free systems (35).
The mechanismby which apolymerase can interactwith theviralRNAwhile it is stillencapsidated is not known.On
the basis ofobservation of extendedandcompressed states of the nucleocapsid, it has been proposed that extension of the nucleocapsid, perhaps onlylocally, may. allow the
poly-meraseto obtainaccess to the encapsidated RNA (14).
Wereport heretheobservationof fourdistinctlydifferent structural states of the nucleocapsid, with all statespresent under the same conditions. We have generated a three-dimensionalreconstruction from electronmicrographsof the mostorderedstate of the nucleocapsid.The
nucleocapsid
is 2233on November 10, 2019 by guest
http://jvi.asm.org/
FIG. 1. ThecomputedFouriertransform ofanegatively stainednucleocapsidin the 5.3-nmpitchstate(left) and inthe6.8-nmpitchstate
(right).Thelayer linelabeled1 isat1/(79.5nm),the layer linelabeled14isat1/(5.7nm), thelayer line labeled 15isat1/(5.3nm), and the
layer line labeled 16isat 1/(5.0nm). On theright, the layer line labeled 1isat1/(6.8nm).
thus a dynamic structure which can maketransitions among at least four different discrete states. It is believed that a
knowledge of thethree-dimensional structure of the Sendai virus nucleocapsid in various conformations and the
mech-anisms governing the transitions between these states will provide some insights into the mechanisms of mRNA
tran-scription and genome replication.
MATERIALSAND METHODS
Virus preparation. Samplesof Sendaivirusnucleocapsids (isolated from infected cells, rather than fromvirions)were prepared as described by Portner and Murti (32) and were stored in an aqueous buffercontaining 0.01 M Tris
hydro-chloride, 0.1 M KCl, 0.015 M MgCl2 (pH 7.4), and 0.3 M
NaCl. Each of the samples was then further purified by
running it with sample buffer through aBio-Gel500column
(Bio-Rad Laboratories).
Electron microscopy. Sendai virus nucleocapsids were taken onto electron microscope gridsand negatively stained with uranyl acetate by the following procedure. A carbon-coated collodion grid which had been made hydrophilic by glow discharge was touched in sequence to each of the
drops, 10 sto the nucleocapsid sample, 10 s to the buffer,
and 1min toeachof2 drops of uranyl acetate (2%wt/vol). Excessfluid was removed with filter paper at each transfer. The specimens were examined at an instrumental magnifi-cation of 49,000 on aPhillips EM-420 electron microscope
equipped witha low-dosekit.
Thenucleocapsids havesome sensitivity to electron dose.
Higher-dose micrographs provided fewer filaments with a well-determined helical symmetry than did minimal-dose
micrographs, inwhich the specimen received no exposure to theelectron beamathighmagnification beforetheimagewas recorded on film. Helical symmetry was assessed from the computed Fourier transforms of images which had been
densitometered (see below). Nucleocapsids were also
ob-served by using a technique of heavy metal shadowing. Samples were adsorbed outofa drop of 5 mM magnesium acetate onto acarbon-film grid which had been previously exposed to glow discharge. Itwas then washed in distilled water for 2 min andfrozen by dipping it in liquidnitrogen. Freeze-drying was carried out in a BAF400 freeze-etch unit (Balzers Union, Fla.) for 3 h at 193 K. The specimen was shadowedunidirectionally with atantalum-tungsten alloyto
give an average metal thickness of0.5 nm. Film thickness
wasmeasured with aquartzcrystal film monitor (QSG201; Balzers). The shadowingwascarriedout at193 K and atan elevation angle of 300. A carbon backing was then applied
from above, with an average thickness of 5 nm. This shadowing technique has been described by Gross et al.(12).
The shadowed nucleocapsids were observed on a Phillips EM-420 electron microscope at a magnification of49,000. Tobacco mosaic virus and RecA filaments were imaged as controls to determine the absolute handedness from shad-owing.
Image analysis. Relatively straight images of nucleocap-sidswereselected from theelectronmicrographs. Theywere digitized byusing anOptronics P-1000scanning densitome-teranddisplayedon anAED 767raster-graphicsdevice.The films were typically recorded at 50,000 magnification and scanned at 25
VLm
per pixel, resulting in a sampling ofthe images of 0.5nm perpixel. Curvature of the flexible nucle-ocapsids in theimageswascorrectedby usinganassumptionof a normal mode ofbending (9). Since this method is only
valid for relatively small amounts of curvature, only
fila-mentswere used whichrequiredarelativelyminimal amount
of correction. The smallest radius of curvature which was
corrected wasgreaterthan 500 nm.Fouriertransformsofthe
straightened images were then computed. Typical filament
lengths which were transformed were 250 nm (approxi-mately 500pixels),andthetransformsweretypically128by 1,024points.The imagesofnegativelystainedobjectsinthe transmission electronmicroscope correspond tothe
on November 10, 2019 by guest
http://jvi.asm.org/
3333
ages
ssag33.
I.
3ss
aa~~~ ~ ~ ~ ~~~~3
aa 33333333333za..
j
~L=.0
N=-2L:
33 33333;
3 3
333 3
.
,.L16,N12
.s' ...'
,,,,,.'
33$s'~~~~~~3 ,
3333.3...
333333333 ,3La,,
L=1 N=13
h[3 L=O N=O
tions of these three-dimensional objects onto two dimen-sions (the recording film). The Fourier transform of the projection of a helical filament is non-zero only on lines called layer lines (6, 24). The spacing of these layer lines is determined by the pitch and symmetry (number of units per turn)of the helix.
Figure 1 shows acomputed Fourier transform ofa nega-tively stained Sendai virus nucleocapsid in the 5.3-nm pitch state. The strongest layer lines are the equatorial and the 1/(5.3 nm) layer line, which is labeled as layer line 15. The 1/(5.3 nm) layer line arises from the 1-start 5.3-nm pitch helix, and a weak second order of this layer line can usually
beseen at 1/(2.65 nm). Mosttransforms will only show these
features. About 10 nucleocapsid sections were found where otherlayer lines were present. In Fig. 1, a near-equatorial layer line lying at an axial spacing of about 1/(79.5 nm) can be identified, labeled as layer line 1. On the basis of this position, otherlayer linescanbepredicted to fallatspacings of 1/(5.7 nm)(layerline 14) and 1/(5.0 nm) (layer line 16). However, the intensities of these predicted layer lines are not in general above the level of the noise in transforms of individual nucleocapsids which possess a1/(79.5 nm) layer line.
Layer lines at 1/(79.5 nm), 1/(5.7 nm), 1/(5.3 nm), 1/(5.0 nm), and 1/(2.65 nm) were extracted from the eight best specimens which gave a significant 1/(79.5 nm) layer line. The 1/(79.5nm)layer line wasdetermined to contain an odd Bessel order, due to the 1800 phase difference between the left and right sides of the transform. On the basis of the
positionof the maximum onthe 1/(79.5 nm) layer line, this layerline would have to be of either Bessel order 11 or 13. Given that these twopossibilities could each correspond to either a right-handed or left-handed set of helices, there would be fourpossible sets of indices for all the layer lines
[the
indexing
of the1/(79.5 nm) layer
lineuniquely
defines the1/(5.7 nm) and1/(5.0 nm) layerlines, given that the 1/(5.3 nm)layer line is of Bessel order 1]. In all four possibilities, however, the1/(5.7nm)and 1/(5.0 nm) layer lines are even. Wethereforewereabletoaveragetheeight sets of layer line data before establishing the indexing for these layer lines.An individual helix gives rise to two redundant sets of layerlines, arising from the near and far sides of the helix. Averaged layer lines (near plus far sides) were extracted from each of the transforms. Alignment of the individual layer line sets was achieved by searching for the rotation,
translation, andup-downorientationof each, which gave the minimumamplitude-weighted phase residualagainst a refer-ence nucleocapsid. An average wasconstructed with these parameters,andthis average was used as the new reference data set. This procedure was repeated until further cycles producednochange in the average. The averaged layer line data is shown in Fig. 2.
Thequality oftheaveraging can be discerned in Fig. 3, in which the phase residuals are displayed between the individ-ual layer line datasetsand the averaged layerline set of Fig. 2.
The mean radial density distribution of the nucleocapsid
(given by the Fourier-Bessel transform of the equatorial layer line [Fig. 4]) demonstrates that the structure in cross-section is annular, with a hollow core about 5.0 nm in diameter. The outer boundary of the filament determined
FIG. 2. The layer lines from an averageddata setofeight
nu-cleocapsids, labeled with the indexingshown inFig. 1.The
ampli-tudes areinarbitrary units, and theamplitude scaleonthe equator
(L = 0,N = 0)is 10 times greater thanthat for the otherlayerlines.
2235
L=14 N=-14
1
4
1
I
I
.01
.02
.03
.04
on November 10, 2019 by guest
http://jvi.asm.org/
[image:3.612.91.252.45.743.2]o /
90 75 60 45 30 15
BETTER PHASE RESIDUAL
FIG. 3. The total amplitude-weighted phase residual between
eachoftheeight layerlinesetsused in theaverageand theaveraged layer line set itself (Fig. 2), as a function ofthe polarity ofthe
individualsetagainsttheaverageddataset.Thisplotcan serve as a
measure of the intrinsic polarity of the structure. For a nonpolar structure(or forhighly noisy data),therewouldnotbeasignificant
difference betweenthetwodifferentorientations ofeachindividual
data set, andallpointswould fallnearthediagonalline. Thefarther
points are from the diagonal, the greaterthe measure ofpolarity.
The centroid of the distribution is49.60for thebetter-phaseresidual
and71.30for theopposite orientations.
0.020
0.0 0
0.000
--0.020
from the equator is in very
good
agreement with the outerboundary
determinedby
the radial extent ofhelical modu-lationarisingfrom thevery strong1/(5.3
nm) layerline(layer
line
15),
asshown inFig.
4.We have used thisinformationto determine whether layer line 1 contains n = +11 orn = ±13. Ifn were to be equal to 11, one would have a
helical structure consisting of discrete subunits with no modulation of the stainattheouterperiphery (Fig. 4)arising
from the discretesubunits, but instead with strong modula-tion of the stain in the center of the annulus of
density
formed
by
the subunits. This seems rather unlikely and would be different from any other helical tubular structure looked at in negative stain (e.g., microtubules, tobacco mosaic virus, bacteriophage tails, etc.). In the absence of other information, we can only conclude that n = ±11 ishighly unlikely, but not impossible. Once we accept that
n = ±13,one mustdetermine whethern = +13or n = -13,
given that n = -1 on layer line 15 (see Results). This information issuppliedbythe 1/(5.7 nm)and11(5.0nm)
layer
lines. Bessel orders mustbe chosentogive helical modula-tion at radii within the annulus ofdensity. The only possi-bilitywhichgavethiswas n = +13onthe1/(79.5nm), which
generatesn = -14onthe1/(5.7nm) andn = 12onthe11(5.0
nm) layerlines.
The layer line positions give very precise information about the helical twist, and although there was variation
among specimens, the mean ratio of the spacing of the
0.100
-\
LL15,
N=-1
0.070
\V\
--LL1,
N=13
0.040
----LL1,
N=
11
0.010
-0.020
-0.050
-0.080
*.
LL
0,
N=0
-0.110
-0.140
0
2
4
6
8
10
12
14
Radius
(nm)
FIG. 4. The meanradialdensity distribution,thefunction
go(,(r)
(24) from the equator of thetransform(LL 0, N =0), isshown with thescaleontheright.The modulus of the
g.,(r)
functionforthe very strong layer line LL 15, N = -1, is shown with the scale on the left. Onecanseehowconsistentboth are withrespect to the outer boundary of thefilament,between 9 and 10 nm. Theagreementis good, since this
data isanaverage fromninefilaments.The datafromlayer line 1 hasbeentransformedwithtwo possibleoptions,that n = 11 orn = 13, and
the modulus of theresultingg131(r) is shown, as is thecorrespondingplot forg11 1(r). One can see theassumptionthatn = 13 on the first layer
lineyieldsaradialdensity ofmodulation in excellent agreement with the outer boundary of the filament as determined by the equator and
the15thlayerline, whereas theassumptionthatn = 11yields no helical modulationat theouterperipheryof thetubular structure.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:4.612.98.278.66.248.2] [image:4.612.71.564.366.657.2]SENDAI VIRUS NUCLEOCAPSID 2237
-4
. 49 *.* *N
.~~
:ZI'-
-
J00-
~
-WS't5
~~I
FIG. 5. Electron micrographs of negatively stainednucleocapsids. The space bar in panel Ais100 nm.ThearrowinpanelApointstoa
region of6.8-nmpitch,whichis characterizedbyincreasedflexibility. The straight segment at the bottomis inthe 5.3-nmpitchstate, as is
the relativelystraight region above the arrow. A similar transition can be observed in panel C. The nucleocapsid in panel B isentirelyin the
5.3-nmstate. In panel D, mostof the nucleocapsidsareinthe6.8-nm state. One mayfindfairly straightsegments of 6.8-nm pitch nucleocapsid,
asindicatedby the arrow at the topin panelD. Thetransform ofthissegment is shown on the right in Fig. 1. The arrow at the bottompoints
toaregionof5.3-nmpitch, indicating thatastructural transition has occurred within thenucleocapsid.
1/(79.5 nm)layerline to the1/(5.3nm)layerline was close to 1:15. Theindexing which hasbeen used corresponds to 196
subunits in 15 turns of the 5.3-nm pitch helix.
Theintensities and phases along the layer lines contain the
informationabout the shapeof the subunit within the helix, which is theasymmetric unit in the structure. Since the layer
line intensitiesand phasescontainall theinformationneeded
toreconstruct ahelix,athree-dimensional reconstructionof
the nucleocapsid structure was obtained by performing a
Fourier-Bessel transform of the averaged layer line data. Thesurfacesofthereconstructionsweredisplayed by using an algorithm described byEgelman and Stasiak(10).
RESULTS
In both the negatively stained and the heavy metal-shadowed preparations,the majorityof nucleocapsids were
ina statecharacterizedbyameanpitch ofabout 5.3 nm(Fig.
5and 6). A smaller number ofindividualnucleocapsids and
sections of nucleocapsids were observed with a more ex-tendedpitch ofabout 6.8 nm, and these were seenin both
negatively stained and shadowed specimens. Large
aggre-gated masses ofnucleocapsids, however, could be seen in
shadowingtobe almostentirelyin the 6.8-nm pitch
confor-mation,whereasinnegative stainsuch structures would not have discernible detail. Nucleocapsids inahighlyextended 37.5-nmpitch conformationwerefrequentlyobserved in the
shadowed preparations but were never seen in negative
stain.
Because thehelical stripesgenerated by the 5.3-nm pitch helix are nearly horizontal (Fig. 6A and C), the perceived handedness ofthis helix can actually depend on the
shad-owingdirection. It is thuspossible to find segments in this
conformationwhich appear left-handed and other segments which appear right-handed. To avoid this problem, we
computedtheFourier transforms of nine segments of shad-owed nucleocapsidsin the 5.3-nm pitch state andaveraged
the intensities of the transforms. The resulting averaged transform is shown in Fig. 6G. This transform displays intensitytotheright ofthevertical lineshownbythearrows
(called the meridian) in the top half and to the left of the
meridian in the bottom half. This pair of reflections is
generatedby the stripes correspondingtothe topsurface of aleft-handed helix.Aright-handed helixwouldgiveriseto a
pairof reflections on the opposite sides of the meridian. If the6.8- and 37.5-nm nucleocapsid conformations ariseby a
simpleextension of the 5.3-nmpitchhelix,then these helices will also beleft-handed. Shadowed images (Fig. 6) show that
they areindeed left-handed.
Reconstruction ofthe5.3-nmpitch nucleocapsid. The
com-puted Fouriertransforms(Fig. 1, left) ofimages of
nucleo-capsids in the 5.3-nmpitch state yieldthe symmetry of the
structure toaratherhigh precision, if theambiguities
inher-ent in the indexing of the transform can be overcome (see
MaterialsandMethods).Our best estimate of this symmetry,
given the potential for alternate indexing schemes, is 196
copies ofthe NPproteinin 15turnsofthe 5.3-nmpitchhelix or 13.07 NP molecules per turn of this helix. There is therefore an axial rise persubunit in thisstate of 0.41 nm.
Ahelixconsistsofasingleasymmetricunitrepeated along
thelength of the structureby asymmetry operation
involv-ing rotation and translation. We thus havechemically iden-tical subunits inchemically identicalenvironments,ignoring, ofcourse, those subunitsatthe ends ofthestructure. If the helix isviewed in projection, asit is in a negatively stained electronmicrograph, manydifferentprojectionsofthe same subunitaregenerated alongthelengthof the helix. This set ofprojections contains all theinformation needed to
recon-struct the nucleocapsid in three dimensions (6), and sucha
reconstruction is shown inFig. 7and8.The data whichhas been used for the reconstruction (Fig. 2) extends out to about 2.4 nm. Figure 7 displays a surface view of the
nucleocapsid, looking perpendicular to the filament axis,
I
VOL.63, 1989
on November 10, 2019 by guest
http://jvi.asm.org/
4W~~~~~~~~~~~~4'
°) C0*
,r
2
CU
';ca = U
C 4) C . .4
0,CU
.0 0
>a
- 8=.u.
Q4-en C
co
,a CUJ
CU*
rA
c:^to'A0,
-Q x 2cX0* °C, UOE-sJ
CU's co
J*N.- .CU
C
.C0
<'-° CUoV
CX ,C.
on November 10, 2019 by guest
http://jvi.asm.org/
SENDAI VIRUS NUCLEOCAPSID 2239
FIG. 7. Asurface view ofathree-dimensional reconstruction ofanucleocapsid in the 5.3-nm pitch state, computed from the data shown
inFig. 2.On the right, half of the nucleocapsid has been cut away to expose the hollow core.
whereas Fig. 8 shows the internal density distribution in an
orthogonal view,looking alongthefilamentaxis. The central hole in thehorizontal sliceshowninFig.8 has adiameterof about 7.5 nm. However, this hole is precessing about the
filament axis as one travels along the axis, generating an internal helicalgroovevisiblein the cutaway surface shown ontherightinFig. 7. Asaresult, the diameter of the largest
continuous cylinder which could be placed in the hollow core is about 5.0 nm, but the hole can accommodate a
less-extended objectabout 7.5 nm across. The outer diame-ter ofthe nucleocapsid is about 20.0 nm. These values are
similar to those that have been reported previously forthe
Sendaivirus nucleocapsid (11, 17, 21).
Asecond5.3-nmpitchstate. Both thepitchandtwist ofa
helixdetermine theposition of layer linesseenin theFourier
transform ofahelical specimen (Fig. 1). If the pitchof the Sendai virus helixwere tobekept approximately fixedat5.3 nmand the numberofsubunits (NPproteins) per turn were to be changed from 13.07, then the weaker layer lines
(labeled as 1, 14, and 16 in Fig. 1) would shift in their
spacings. Because smallchanges in twist ofthe elementary subunit accumulate over manycopies of the subunit, small
changesintwistcangeneratelargechanges in the transform
ofa helical object, even at low resolution. We have found several segments ofnucleocapsids in which the first layer line is moved to about 1/(23.8 nm) from 1/(79.5 nm). This
position for a near-equatorial layer line, 1/(23.8 nm), is similar to that shown by Finch and Gibbs (11) for one segment ofa Sendai virus nucleocapsid. Since the pitch of the helix is apparently unchanged in these segments, as determined by the relative constancy of the spacing of the strong (1/(5.3 nm) layer line, this change inposition
corre-spondsto achangein thetwist of thestructure. We havenot been able to characterize this state any further than by determiningthatthislayerline arises fromanoddnumber of helices. Sincethe peak ofthis layerlineoccurs at asimilar
distance fromthe meridian as that on the 1/(79.5 nm) layer line, it is most likely that it also arises froma13-starthelix. However, the handedness of this 13-start helix could be either left orright, and the twist of this secondstate would
therefore be either 12.78 or 13.22 units per turn,
respec-tively. The smallest transition between these two discrete 5.3-nm pitch states would therefore be from 13.07 to 13.22 units perturn. While this transition appears striking at low
resolution,itwouldamount to achange inrotation between twoadjacentsubunits ofabout0.32°or aradial shift inmass
at8-nm radius of about 0.05nm (about half the diameter of ahydrogen atom).
Comparisonof the 5.3- and6.8-nmpitchstates.Negatively
stained images of segments ofnucleocapsid in the 6.8-nm
pitch state give rise to transforms with only an equator, a 1/(6.8nm) layer line, and a 1/(3.4nm) layerline
(Fig.
1). InVOL.63, 1989
on November 10, 2019 by guest
http://jvi.asm.org/
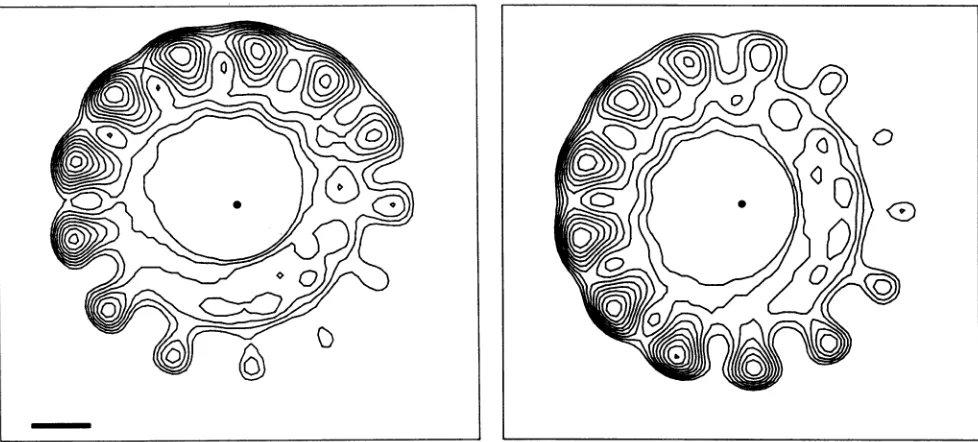
FIG. 8. Two slices,perpendiculartothefilament axis,ofthethree-dimensionalreconstructionshown inFig.7,shownwith contourplots.
Theslicesareseparatedby 1.0 nm. The space bar on the left is 3.0 nm, and the dot in the center of each slice marks thelocation ofthefilament
axis.One can see that the hollow core precesses about the filament axis as one travelsalongtheaxis,and this creates thehelicalgroove in
thecore, which is seen in thecutaway view in Fig. 7.
contrast, we have been able to find six layer lines in the transforms of filaments in the 5.3-nm pitch 13.07 units per turn state. A three-dimensional reconstruction was made with the three layer lines in the 6.8-nm pitch state. This reconstruction, shown in Fig. 9, contains only the continu-ous helical density along the 6.8-nm pitch one-start helix, with the modulation of the density dueto discrete subunits
absent. For purposes ofcomparison, we have generated a
similarreconstruction in Fig. 9 ofthe 5.3-nm stateshownin
Fig. 7 and 8, but using only theequator, the1/(5.3 nm), and the 1/(2.65 nm) layer lines. This averages out the subunits
seeninFig. 7for thecomparison of Fig.9. Onecan seethat
the 6.8-nm pitch state is significantly narrower than the
5.3-nmpitchstate,withachangein diameterfrom20.0nmto
about 16.5 nm. The 6.8-nm state is also significantly more
polar,consistent with thechevronlikeimage seen in
projec-tion in negative stain (Fig. SD). The narrowing and the
increasing polarity ofthe 6.8-nm pitch state could both be generated byalargeincrease in the tilt of subunits from their position inthe5.3-nm pitchstate. Alternatively, thechange
could be due to a large conformational change within the
subunitsor toacombination of both of these events.
DISCUSSION
Changesin saltconcentration(14)andtrypsin cleavageof
the NP subunits (28, 29) have previously been reported to
cause conformational changesinparamyxovirus
nucleocap-sids. In this study, different states have been observed to
occurinthe samepreparation, often in the same
nucleocap-sid (Fig.4and5). Aheterogeneity ofNPsubunits,resulting from proteolytic cleavage, cannot be the source of the
structuralheterogeneity, since theNPproteinalwaysrunsas
asingle bandon sodium dodecyl sulfate-polyacrylamide gel electrophoresis (7, 8, 32) at the same time that structural heterogeneity canbe seenin theelectron microscope.
Fur-ther, trypsin-cleaved nucleocapsids appear to be in the
tightly coiled 5.3-nm state (14), so the transitions between
thedifferent helical formscannotsimplybeaccounted forby proteolysis. Asaresult, theabove-mentioned factors may be able to influence thedistribution ofstatesbutarenotneeded toproduce nucleocapsids in5.3-, 6.8-, and 37.5-nm
confor-mations.
Under the sample preparation conditions used in this
study, the majority of nucleocapsids were in the 5.3-nm
state. Within this state, we have observed two substates
which correspond to a small variation in the twist of the structure, from 13.07 to either 13.22 or 12.78 subunits per turnof the helix. Since wehavenot found anyintermediate
positions, there are probablytwo discrete statesof twist in the 5.3-nmpitchstate. Similarly,because formsintermediate
between the three different pitches (two5.3-nmpitch states and a6.8- anda37.5-nmstate) havenotbeenobserved, it is likely that the different conformations represent discrete states.
The 37.5-nm pitch conformation is never observed in negative stain and is onlyseenin shadowedpreparations. It is possible that this state is an artifact of the particular specimen preparation techniques used. There are several
reasons why this isparticularly unlikely. First, this stateis
alwaysseenwithareproducible helical pitch. Ifthiswere an artifact of preparation, it is hard to imagine that it would always adopt the same helical conformation. Second, one canobserve transitions from the 5.3-nmpitch stateinto the
37.5-nm pitch state within a nucleocapsid (Fig. 6B). Third,
fast-freezing and etching, which reveals this state, is more likely to preserve a fragile structure than the drying em-ployed in negative staining. Because previous studies of Sendai virusnucleocapsidshaveallemployednegativestain,
it isnotsurprisingthatthishighlyextendedstructure has not been previouslyobserved.
Thehandedness of the helixof Sendai virusnucleocapsids inthe 5.3-nm pitchstate has beenpreviouslydetermined by
tilting negatively stained specimens (31) and by observing
asymmetry innegative staining(18). Thetiltingexperiments
on November 10, 2019 by guest
http://jvi.asm.org/
[image:8.612.69.558.74.295.2]SENDAI VIRUS NUCLEOCAPSID 2241
FIG. 9. Asurface representation oft4e6.8-nmpitchstateisshown ontheleft. This reconstructionwasmadewithonlythe equator, the
1/(6.8nm)layer line, andthe 1/(3.4nm)layerline and so contains no modulation of thecontinuous helical density duetodiscrete subunits.
For purposesof comparison, the5.3-nmpitch reconstruction shown inFig. 7 has beendisplayedbyusingonlytheequator, the 1/(5.3nm)
layer line,andthe1/(2.65nm)layer line, so that thediscretesubunits are averaged out and thecontinuoushelicaldensityisdisplayed. One
canseethe narrowing ofthenucleocapsidinthe6.8-nmstate, as well as theincreaseinpolarity,withbothmostlikely dueto alargeincrease
in the tilt angleof the subunits.
concluded that the helix isleft-handed,whereas thestaining
asymmetry observations indicated that the helix is right-handed. Our shadowed observations arein agreement with thenegative staintiltresults. Compansetal. (5) usedtilting ofnegatively stained specimenstoexamine "stretched-out" segmentsofnucleocapsids fromtwootherparamyxoviruses,
simian virus 5 and mumpsvirus, and concluded that both are
left-handed. Theysurmisedthat theSendai virus nucleocap-sid would also be left-handed, due to the structural
similar-ity. It is possible in principle forthe compact stateofall of these nucleocapsids to have one-handedness and for the stretched out states to have a different handedness. This could occur through a switching ofdominant contacts
be-tween two different one-start helices. A helix
handedness-reversing transition occurs in bacterial flagella (36) between twodifferent conformations. It is also possible in principle (5) thatenantiomorphicforms mayoccurwithinasingle viral group. We have directly shown in Sendai virus that the 5.3-nm conformation has the same handedness as the stretched 6.8- and 37.5-nmstatesandthat this handedness is in agreement with that determined by Nonomura and Ko-hama (31).
The presence ofapproximately 13subunits in eachturnof
the helix has been previously determined by the use of rotational harmonics (19), but a similar study led to the conclusion that there were 15 subunits per turn (18). Finch and Gibbs (11) used a method similar to that used in this report, ofcalculating radii ofhelical modulation, which led themtoconclude that thereareeither 11 or13 subunitsina turn. We have found that arelativelyunambiguous indexing canonly be achieved byusinglayerlines [at1/(5.7 nm) and 1/(5.0nm)] which are onlyslightly above the noise level in individual transforms but which can be averaged to yield reliable data.
An estimate for the RNA-protein stoichiometry may be obtained fromaknowledgeof the number ofproteinsubunits
perturn. If all ofa nucleocapsid were tobe in the state we
have characterized, there would be approximately210 heli-cal turns in the 1.1-,um modallength (11). There would then be about2,745 NP subunits inanucleocapsid (again,
assum-ing that all of the nucleocapsid were to be in the same conformational state). The RNA genome consists of15,383 bases (34) which would suggest an NP-RNA ratio of 5.6
basesperNP subunit.Helicalsymmetry suggestsaninteger
stoichiometry between bases and NP subunits. Since there areprobably specificcontactsbetween the RNA and theNP
VOL.63, 1989
on November 10, 2019 by guest
http://jvi.asm.org/
[image:9.612.58.556.74.417.2]protein, a nonintegral ratio would break the helical symme-try relating every NP subunit to every other one. For example, if there were 5.5 bases perprotein, the asymmetric unit in the helix would be two NP proteins, and not one. While it is possible that this symmetry breaking would not be seen at low resolution, many examples exist in which small perturbations from helical symmetry can be seen dramati-cally at low resolution. Alternatively, one might argue that helical symmetry within the nucleocapsid is obeyed, but that there is a nonspecific association between the RNA and the NPsubunits. Thus,
one
could have anyrelationship in this scheme. We think that this is even less likely on physical grounds, given that highly extended forms of the nucleocap-sid have been observed in which the RNA appears to hold together disassembled NP subunits as "beads on a string"(14). Therefore, the factors affecting this ratio must be
considered. The most likely explanation is that parts of the
nucleocapsid are extended, and the total number of NP
copies is therefore less than 2,745. A 6:1 stoichiometry
wouldimply 2,564copies of the NPprotein. Thealternative, that there are actually greater than 2,745 copies of the protein and a 5:1 stoichiometry, seems less plausible.
The reconstruction of Fig. 7 and 8 shows a single asym-metric unit which has a volume (approximately 70 nm3, given areasonable choice of cutoff density) consistent with theknownmolecular weight of the NP protein, 56,543. The additional Sendai virusproteins, P and L, have been shown to beassociated with thenucleocapsid (32, 33) but located in clusters along the filament and not with a helical symmetry. Thus, these proteins would not contribute to the density seen in thereconstructions, in which the helical symmetry of the NP protein has been imposed onlengths of filaments for the purposes of averaging.
The physiological role of the different states we have described here is not known. Shadowing has shown that highly coiled masses ofnucleocapsids are in a 6.8-nm pitch state, consistent with the observations from negative stain thatthe transition from the 5.3- to the 6.8-nm pitch ismarked by alargeincrease in the flexibility of the nucleocapsid. It is most likely that the highly coiled nucleocapsid within the intact virion is in the 6.8-nm pitch state.
Transcription occurs without dissociation of the NP pro-teinsubunits from the RNA. It has been suggested (14) that an uncoiling of thenucleocapsid may be necessary for viral transcription and RNA replication. One might expect that an RNA which is completely ensconced within the nucleocap-sid helix would become exposed when the nucleocapsid makes a transition to an extended state. However, the transition from the 5.3-nm state to the 6.8-nm pitch state does not appear to fully open the helical grooves, as one would expect in such a model. This structural observation is consistent with the biochemical report (15) that the RNA within the nucleocapsid is inaccessible to digestion by ribo-nuclease regardless of the unwinding of the nucleocapsid helix. On the other hand, both the extended 6.8-nm state and thecompact 5.3-nm state possess a large central corewhich
could allow a polymerase to reach the RNA while the
nucleocapsid protein is bound to the RNA. The observation ofthe 37.5-nm conformation is particularly interesting be-cause the RNA must undoubtedly be exposed in such a highly extended state.
ACKNOWLEDGMENT
This research was supported by Public Health Service grant GM35269 from theNational Institutes ofHealth to E.H.E.
LITERATURE CITED
1. Amano, Y., T. Takano,K.Takahashi,and N. Ishida. 1971. Fine
structure of Sendai virus nucleocapsid. Jpn. J. Microbiol. 15:
549-551.
2. Baltimore, D. 1971. Expression of animal virus genomes.
Bac-teriol.Rev. 35:235-241.
3. Blair, C. D., and W. S. Robinson. 1970. Replicationof Sendai
virus. II. Steps in virus assembly. J. Virol.5:639-650.
4. Choppin, P. W., and W.Stoeckenius. 1964. Themorphology of SV5virus. Virology 23:195-202.
5. Compans, R. W., W. E. Mountcastle, and P.W.Choppin.1972.
Thesenseof the helix ofparamyxovirus nucleocapsids. J.Mol.
Biol. 65:167-169.
6. DeRosier, D. J., and A. Klug. 1968. Reconstructions of
three-dimensional structures from electron micrographs. Nature
(London) 217:130-134.
7. Deshpande, K. L., and A. Portner. 1984. Structural and
func-tional analysis ofSendai virus nucleocapsid protein NP with
monoclonal antibodies. Virology 139:32-42.
8. Deshpande, K. L., and A. Portner. 1985. Monoclonalantibodies
to the P protein of Sendaivirusdefine itsstructure androle in
transcription. Virology 140:125-134.
9. Egelman, E. H. 1986. An algorithmforstraighteningimages of
curved filaments. Ultramicroscopy 19:367-374.
10. Egelman, E. H., and A. Stasiak. 1988. Structure of helical RecA-DNA complexes. Il. Localconformational changes visu-alized in bundles of RecA-ATP--y-S filaments. J. Mol. Biol.
200:329-350.
11. Finch, J. T., and A. J. Gibbs. 1970. Observations on the
structure of the nucleocapsids of some paramyxoviruses. J.
Gen. Virol. 6:141-150.
12. Gross, H., T.Mueller, I.Wildhaber,and H. Winkler.1985. High
resolutionmetalreplication, quamtified byimageprocessing of
periodic test specimens. Ultramicroscopy 16:287-305. 13. Hamaguchi, M., T. Yoshida, K. Nishikawa, H. Naruse, and Y.
Nagai. 1983.Transcriptive complex of Newcastle disease virus. I. Both L and P proteins are required toconstitute an active complex. Virology 128:105-117.
14. Heggeness, M. H., A.Scheid,and P. W.Choppin. 1980. Confor-mation of the helical nucleocapsids of paramyxoviruses and vesicular stomatatis virus: reversible coiling inducedbychanges
insaltconcentration. Proc. Natl.Acad. Sci. USA77:2631-2635.
15. Heggeness, M. H., A. Scheid, and P. W. Choppin. 1981. The relationship of conformational changes in the Sendai virus nucleocapsid to proteolytic cleavage of the NP polypeptide.
Virology 114:555-562.
16. Horne, R. W., and A. P. Waterson. 1960. Ahelical structure in mumps, Newcastle disease, and Sendai viruses. J. Mol. Biol. 2:75-77.
17. Horne,R. W., A. P. Waterson, P.Wildy, and A. E. Farnham.
1960. The structure and composition of the myxoviruses. I.
Electron microscope studies of the structure of myxovirus
particles bynegative stainingtechniques. Virology 11:79-98. 18. Hosaka, Y. 1968. Isolation andstructureofthenucleocapsidof
HVJ. Virology35:445-457.
19. Hosaka, Y., and J. Hosoi. 1983. Study of negatively stained
images of Sendai virus nucleocapsids using minimum-dose
system. J. Ultrastruct. Res.84:140-150.
20. Hosaka, Y., H. Kitano, and S. Ikeguchi. 1966. Studies on the
pleomorphismof HVJ virions. Virology29:205-221.
21. Hosaka, Y., Y. Nishi, and K. Fukai. 1961. Thestructureof HVJ.
lI. The fine structure ofsubunits. Biken J. 4:243-254. 22. Ishida, N., and M. Homma. 1978. Sendai virus. Adv. Virus Res.
23:349-383.
23. Kingsbury, D.W. 1977. Paramyxoviruses, p. 349-382. In D. P.
Nayak (ed.), The molecularbiology ofanimal viruses, vol. 1. Marcel Dekker, Inc., New York.
24. Klug,A., F. H. C. Crick, and H. W. Wyckoff. 1958. Diffraction
byhelical structures. ActaCryst. 11:199-213.
25. Kolakofsky, D., and A. Bruschi. 1975. Antigenomes in Sendai virions and Sendaivirus-infectedcells. Virology 90:226-234. 26. Lamb, R. A., B. W. J. Mahy, and P. W. Choppin. 1976. The
synthesis of Sendai virus polypeptides in infected cells.
on November 10, 2019 by guest
http://jvi.asm.org/
SENDAI VIRUS NUCLEOCAPSID
ogy69:116-131.
27. Morgan, E. M., G. G. Re, and D. W. Kingsbury. 1984. Complete
sequence of the Sendai virus NPgene from a cloned insert.
Virology 135:279-287.
28. Mountcastle, W. E., R. W. Compans,L. A.Caliguiri,andP.W.
Choppin. 1970.Nucleocapsid protein subunits of simian virus 5, Newcastle disease virus, and Sendai virus. J. Virol. 6:677-684. 29. Mountcastle, W. E., R. W.Compans, H. Lackland, and P. W.
Choppin. 1974. Proteolytic cleavage of subunits of the nucleo-capsid of the paramyxovirus simian virus 5. J. Virol. 14: 1253-1261.
30. Nakai,T., F.L.Shand, and A. F.Howatson.1969.Development ofmeasles virus in vitro. Virology 38:50-67.
31. Nonomura, Y., and K. Kohama. 1974. Determination of the absolute hand of the helix in the nucleocapsid of hemagglu-tinating virus (Japan). J. Mol. Biol. 86:621-626.
32. Portner, A., and K. G. Murti. 1986.Localization ofP,NP,and
M proteins on Sendai virus nucleocapsid using immunogold
labeling. Virology 150:469-478.
33. Portner, A., K. G. Murti, E. M. Morgan, and D. W. Kingsbury. 1988. Antibodies against Sendai virus L protein: distribution of the protein in nucleocapsids revealed by immunoelectron
mi-croscopy.Virology 163:236-239.
34. Shioda, T.,K.Iwasaki, andH.Shibuta. 1985. Determination of thecomplete nucleotide sequence of the Sendai virus genome
RNA and thepredicted amino acidsequencesof the F, HN and
Lproteins. Nucleic Acids Res. 14:1545-1563.
35. Stone, H. O., D. W. Kingsbury, and R. W. Darlington. 1972.
Sendai virus-inducedtranscriptase from infected cells: polypep-tides inthe transcriptive complex. J. Virol. 10:1037-1043. 36. Trachtenberg,S., and D. J. DeRosier. 1987. Three-dimensional
structure of the frozen-hydrated flagellar filament. The left-handed filament of Salmonella typhimurium. J. Mol. Biol. 195:581-601.
VOL. 63, 1989 2243