Copyright0 1974 American Society forMicrobiology Printed in U.S.A.Vol.14,
Properties and Location
of
Poly(A)
in
Rous
Sarcoma
Virus
RNA
LU-HAI WANG AND PETER DUESBERG
Department of Molecular Biology and Virus Laboratory, UniversityofCalifornia,Berkeley,California 94720 Received forpublication28 August 1974
The poly(A) sequence of 30 to 40S Rous sarcoma virus RNA, prepared by
digestion oftheRNA with RNase T, showed a ratherhomogenous
electropho-reticdistributioninformamide-polyacrylamide gels. Its size was estimated tobe
about 200 AMP residues. The poly(A) appears to be located at or near
the 3' end ofthe 30 to 40S RNAbecause: (i) it containedone adenosineper 180
AMP residues, and because (ii) incubation of 30 to 40S RNA with bacterial
RNase H inthe presence ofpoly(dT) removed its poly(A) without significantly
affecting itshydrodynamic or electrophoretic properties in denaturing solvents.
Theviral 60 to70SRNAcomplexwas found to consist of 30 to40S subunitsboth
with (65%) and without (approximately 30%) poly(A). The heteropolymeric
sequences of these two species of 30 to 40S subunits have the same RNase
T,-resistant
oligonucleotide composition. Some,perhapsall, RNaseT,-resistant
oligonucleotidesof 30 to40S Roussarcomavirus RNAappeartohavea
unique
lo-cationrelativeto the
poly(A)
sequence, becausethecomplexity
ofpoly(A)-tagged
fragments of30 to40S RNA decreasedwithdecreasingsize ofthe fragment.Two
RNase
T,-resistant
oligonucleotides which distinguish sarcoma virus Prague BRNA from that of atransformation-defective deletion mutant ofthe samevirus
appear tobe associated with an 11Spoly(A)-tagged fragmentofPragueB RNA.
Thus RNA sequences concerned with cell transformation seem to be located
within 5 to 10% ofthe3'terminus ofPragueBRNA.
Poly(A) stretches have been found in the
RNA of tumor viruses and many cytocidal viruses as well as in many cellular messenger
and
heterogenous
nuclear RNAs(1, 3, 7, 13, 14,16-18, 21-23, 26, 27,33, 35, 37, 45,47).Although the function of poly(A) in these RNAs is
un-known, it has been suggested that poly(A) is
added tomRNAs
post-transcriptionally (7, 34).
Consistent with this notion, the
poly(A)
stretches have been found at the 3' end of a
number ofmRNAs (31, 32, 40).
Based on endgroup-labeling techniques tumor virus RNA wasreported toterminate at
the 3' end with a
poly(A)
segment about 30residues longin the case of an avian virus (43)
and with a
poly(A)
segment of 190 residues inthecase ofmurinesarcoma-leukemiavirus(38). In this report we present evidence in agree-ment with earlier studies (23) that the poly(A) sequence ofRous sarcoma virus (RSV) isabout 180 nucleotideslongand confirm its location at or near the 3' end
by
two methods:(i)
poly(A)
prepared enzymatically from RSV RNA
con-tainedoneadenosineper180AMPresidues. (ii) Removal of thepoly(A) stretch fromRSV RNA
by digestion with bacterial RNase H in the
presence ofpoly(dT) resulted in 30 to40S RNA
which was only marginally smaller than 30 to
40SRNA containingpoly(A). Further, no
inter-nal poly(A) sequences were found in 30 to40S RSV RNA.
In agreement with earlier observations on RSV RNA (23) and with a recent report on murine tumor virus RNA (20), we found that
about30%ofthe30 to40S RNAs ofRSV hadno
poly(A).Finger printanalysesof 30to40S RSV RNAs with and without poly(A) indicated that the two species are indistinguishable with re-gard to their heteropolymericsequences.
Moreover, fingerprint
analyses
ofpoly(A)-tagged fragmentsof 30to40S RSV RNA showed
a decreasing
complexity
withdecreasing
size.This indicates that locations of some, perhaps
all, RNase
T,-resistant
oligonucleotides
and ofpolv(A) are the same on all 30to 40S RNAs. MATERIALS AND METHODS
Reagents.Thefollowingreagentswerepurchased.
[3H]uridine (40Ci/mmol) wasfrom NewEngland Nu-clear Corp.; [3H]adenosine (10 Ci/mmol or 16 Ci/ mmol was from New England Nuclear Corp. or Schwarz/Mann Research; Carrier-free 32p was from ICN; [3H]poly(A) (73
gCi/gumol
of phosphate) and poly(dT) were from Miles Laboratories; oligo(dT)-1515on November 10, 2019 by guest
http://jvi.asm.org/
cellulose was from Collaborative ResearchInc.; RNase A, RNase
T,,
DNase I, and E. coli alkaline phospha-tase were from Worthington Biochemical Corp.; and RNase T2 was from Calbiochem. E. coli RNase H was prepared asdescribed previously (24).Virus. Prague RSV ofsubgroup B (PR RSV-B) was used in all of these studies. It was propagated and purified according to published procedures (10, 11).
RNA preparations.Tobacco mosaic virus (TMV) RNA, [3H]uridine, [3Hladenosine,or32P-labeled 60to 70S RSV RNAs as well as30 to 40S subunitswere prepared according to published procedures (9, 11). All RSV RNAs used in these studies were isolated from radioactive virus harvested from infected cul-turesat3-h intervals.
[3HJuridineor32P-labeledchick cell 4, 18, and 28S RNAs were prepared as follows. Primary chick em-bryo fibroblastcells wereseededat aconcentrationof 4 x 101cells per 10-cmpetridish andweregrown in medium 199 supplemented with 2% tryptose
phos-phate broth (TPB), 1%calf serum, 1%chick serum, 1%dimethylsulfoxide;0.05%glucose,100unitsper ml ofpenicillin, 50Agperml ofstreptomycin, and 0.5 gg per ml offungizone. Twelve hourslater, mediumwas changedto8-ml per dish of Dulbecco modifiedEagle
medium with the same supplements as described aboveexcept that TPBwasomitted anddialyzed calf and chick sera were used. A 200-uCi amount of [3HJuridine or 1 mCi of 32P was added per dish; generally, fourdisheswerelabeledat onetime. After 10to12 h ofincubationat 42 C,disheswereremoved from the incubator and placed on an ice bath. Mediumwasremoved and the cellswerewashed twice with Tris saline; then, 1.5 ml of hypotonic buffer
containing0.01MTris-hydrochloride, pH7.4,0.01M NaCl, 2mM EDTA, and 0.05% Triton X-100 was addedtoeach dish. Aftersittingfor5minonice,cells wereharvested witharubberpolicemanandpipetted into aDouncehomogenizer.Cellswerebroken up with sixto sevenstrokes and under theseconditions,very few nuclei were broken. Nuclei were pelleted by
centrifugationat 600xg for5minin aSorvall centri-fuge. Thesupernatantwasthendiluted with standard buffer (0.01 M Tris-hydrochloride, pH 7.4, 0.1 M NaCl, and 1 mMEDTA), and the RNA extractedas described above for viral RNA. Afterprecipitationthe RNAwaspelleted and washed twice with 75% ethanol andredissolved in 0.3 ml of standard buffer contain-ing0.2%sodiumdodecylsulfate (SDS).Thesolution was heated at 100 C for 1 min, quicklychilled, and wasthen layered on a5-ml 10 to 25%linear sucrose gradient containing standard buffer plus 0.1% SDS. Sedimentation was in aSpincoSW65rotor at65,000 rpm for2 h at 20C. Fractions(0.3 ml) were collected and asmallportionofeachfractionwascountedin 3 ml oftoluene-based scintillation fluidcontaining 10% NCS (Nuclear Chicago). The sedimentation profile showed three distinct peaks representing 4, 18, and 28S RNAs.Peak fractions ofeach RNA were pooled, ethanol precipitated, and redissolved in buffer con-taining 0.01 M Tris-hydrochloride, pH 7.4, 10 mM NaCl, and1 mM EDTA. ThepurifiedRNAsamples were storedat -70C.
Conditions for enzyme reactions. DNase I:0.01M Tris-hydrochloride, pH 7.2, 4 mM MgCl2 and 20,ig
per ml of DNase, incubated at 38C for 30 min. Combined digestion with RNase A and RNase T,: 0.01MTris-hydrochloride, pH 7.2, 0.3 M NaCl, 1 mM
EDTA, 20ug perml of RNaseA, and150units perml ofRNaseT, incubatedat38C for 30 min. RNaseT,
alone: 0.01 M Tris-hydrochloride, pH 7.2, 0.15 M NaCl, 1 mMEDTA, and 150unitsperml ofenzyme incubatedat38C for60min. RNase T2 alone:0.04M ammonium acetate, pH 4.4, 1 mM EDTA, 5 units per ml of enzyme, incubated at 38C for2h. Combined digestion with RNase A,T,andT2: same conditions asforRNaseT2 alone plus20flgpermlof RNaseA, and 150 units per ml of RNaseT,,incubationat38C for2h. E. coli RNase H (24): reaction mixture (100 or 150uliters) contained 0.02 MTris-hydrochloride, pH 8.0, 10 mM MgCl2,6 mMdithiothreitol,25units per mlof enzyme and 350 pmol (- 1.85 x 104counts/min) of[3H]poly(A) plus175pmol ofpoly(dT). When RSV RNA (-0.4 Mg) was used as substrate, 0.25 gg of poly(dT) was added, whereas [3H]poly(A) poly(dT) wasomitted. The reaction was carried out at 38 C for 1h.Dephosphorylation and fractionation of commer-cial [3H]poly(A): 25 gtg (3.7 x 106 counts/min) of [3Hlpoly(A) (Miles) was treated with E. colialkaline phosphatase in a 1-ml solution containing 0.05 M Tris-hydrochloride, pH 8.2, and 2 units of enzyme. Afterincubationat38Cfor30min,itwasdiluted to4 ml with standard buffer containing 0.1% SDS and phenol-extracted as described for viral RNA. After ethanol precipitation, the poly(A) was pelleted and washed three times with 75% ethanol. It was redis-solved in 0.5 ml 0.01 M Tris-hydrochloride, pH 7.2. Forfurther fractionation aportionofpurifiedpoly(A) was heat denatured (100 C 1 min) in 0.3 ml of standard buffercontaining 0.2% SDS and was sedi-mentedthrough a5-ml10 to25%sucrosegradient as describedabove. Sedimentation wascarried out in a Spinco SW50.1 rotor at49,000 rpm for12 hat 20C. Peak fractions ofpoly(A)werepooled,ethanol precip-itated, and resedimentedsimilarly. Three subsequent sedimentations were performed to prepare poly(A) with a uniform sedimentation profile and a peak at about8S.
Isolation of[3H]adenosineor32P-labeled poly(A) from RSV RNA. Purified 60-70S RSV RNAin(<0.2 ml) 0.01 M Tris, pH 7.2, 1 mM EDTA was heat denaturedat100 Cfor 1minanddigestedeither with RNase A plus RNase T, or RNase
T,
alone as described above. Afterdigestion,thereactionmixture wasdiluted with standard buffer to 4 to5 ml after addition of 20Mg of carrier TMVRNA and SDS to 0.1% thesolution was phenol-extracted three times. The RNA was ethanol precipitated twice to remove all soluble nucleotides. Such poly(A) was either used directly forend-group analysisorfurtherpurified by heating (100 C, 1 min) followed by sucrose gradient sedimentationin anSW50.1rotor at49,000 rpmfor10 hat 20C.Peakfractionsrepresenting more than 80% of thetotalradioactivitywerepooled, andpoly(A)was then ethanol precipitated twice and used for end-group analysis.End-group analysis. Purified commercial (Miles, Inc.) 7.8S, [3H]poly(A) (0.7to 0.9Mg) or [3H]adeno-sinelabeledpoly(A)from(0.5-1.5Mg)60 to70S RSV RNAwascompletelydigestedtomonoculeotideswith J. VIROL.
on November 10, 2019 by guest
http://jvi.asm.org/
RNase T2 in a volume of 0.2 ml (see above). The reaction mixture wasthen lyophilizedand redissolved in 30Mlitersof water. A200-ggamount of an equimo-lar mixture of each of the fourribonucleoside mono-phosphates and adenosine in 12 hliters buffer was added to it as internal standards. The mixture was subjected to electrophoresis on a Whatman 3 MM paper (12 by 56 cm) according to published proce-dures (23) except thatelectrophoresis was at 1,500V for 2.5 h. Afterthe paper had been air dried the spots representing adenosine and thefournucleotides were located by exposing the paper to UV light in a dark room. Each spot as well as regions between spots were cut outseparately. The nucleoside or nucleotides were elutedfromthepaper with0.4 mlofwater.Theeluted radioactivitywasmeasuredin15 mloftoluene-based scintillation fluid containing20%NCS andcounted in a Packard liquid scintillation counter. Afterelution, the paperwastested foranyremainingradioactivity
by the same method.Sincenoradioactivity could be detected on theeluted paper, the recoveryappeared
tobecomplete.
Oligo(dT)-cellulose column fractionation of
poly(A)-containing RNA. The procedures were a modification of the method of Aviv and Leder (4). One-half gram (dry weight)ofoligo(dT)-cellulose was washed and soaked in 0.01MTris-hydrochloride (pH
7.4) buffer for several hours at room temperature. Afterdecanting the fineparticles severaltimes,itwas packed intoacolumn (0.7by3cm). Before each run, the column was washed with 2mlof 0.1MKOH and subsequently washed with ice-cold buffer I (0.5 M LiCl, 0.01 MTris, pH 7.4, 0.05% SDS)until thepH wasneutral.AnRNAsamplein 0.2ml ofcoldbufferI wasapplied to the top of the column andallowedto enterthe column. Fractionation was carried out by
stepwise elution with 2-ml batches ofbufferI,buffer II containing0.1 M LiCl, but otherwise thesame as bufferI,8andbufferIIIcontaining0.01M
Tris-hydro-chloride and 0.05% SDS. Fractions (0.5 ml) were collected. The RNA ineach fractionwasdetermined
byitsradioactivity.
Milliporebindingand elution. Theprocedureisa slight modification of that described byMendeckiet al. (31).Briefly,theRNAwasdissolvedin/ordiluted atleast20-fold with buffercontaining0.5MKCI,0.01 MTris-hydrochloride (pH7.4), and1mMMgCl2and filtered slowly by gravity (approximately 6 ml/h)
through a Millipore filter previously soaked in the same buffer for 30 min. After washing three times with1-mlportionsofthesamebuffer,the RNAbound wasrecoveredby elutingwith three0.5-mlportionsof buffer containing0.1M Tris-hydrochloride (pH9.0) and0.5% SDS.
RESULTS
Sizeofpoly(A)inRSVRNA.Earlier studies
inthis (23) and other laboratories (16, 17) have estimated the size of the
poly(A)
stretch of tumorvirus RNA to be around 60,000daltons.However, more
recently
thepoly(A)
stretch ofavianmyeloblastosisvirusRNAwasreportedto
be only 9,000 daltons (43). Since both the size
and the absolute amount of poly(A) in viral RNA must be known to distinguish between several relatively small or one relatively large poly(A) stretch, we have redetermined the size of thepoly(A) in RSV RNA.
Previous size estimates were done with poly(A) prepared from avian tumor virus RNA by RNase A and RNase
T,
digestion at highionic strength (see abovereferences). Although
poly(A) is known to be relatively resistant to RNase A athigh ionic strength (5), it is conceiv-able that the enzyme mayintroducesome nicks
inthis condition. This was indicated by
experi-ments in which poly(A) prepared from viral
RNA by digestion with RNase T1, which is
specificforG,andwithacombinationofRNase
T,
and RNase A were compared. It was foundthat after electrophoresis in formamide-polyac-rylamide gels the RNase T1-prepared poly(A)
wasmuchmorehomogenous than thatprepared
earlier byRNase A (Fig. 1, ref. 23). If
commer-cialpoly(A) wastreated under the same
condi-tions as used to prepare viral poly(A) with RNase T1, its electrophoretic distribution was the same as that of an untreated control.
However, commercial poly(A) was rendered
heterogenous if treated with RNase A and
RNase
T,
(notshown). We conclude that RNaseA at high ionic strength nicks viral and other
poly(A) sequences. Nicking during isolation by
RNase A may be the reason forthe size
differ-ence between the
poly(A) prepared by
us andthat described
by Stephenson
et al. (43) andmay also account for the heterogeneity of
A-rich fragments rangingfrom
poly(A)
tooligo(A)describedby Horstetal.(19).
The molecular
weights
of commercial 7.8Spoly(A) and of the poly(A) of RSV RNA
pre-pared by RNase
T,
were estimated from theirelectrophoretic mobilitiesin
formamide-polyac-rylamide gels to be 116,000and70,000,
respec-tively (Fig.1 insert), using18S rRNA (0.7 x 106
[42]) and tRNA (2.5 x 104 [42]) as standards
and assuming a linear inverse relationship
be-tweenthe
logarithm
ofthemolecularweightof apolynucleotide
and itselectrophoretic
mobility
(42).However, this sizeestimateof
poly(A)
maybe subject to 10 to 20% error since deviations
from a linear log molecular weight-mobility
relationship have been observed under similar
conditions (12).
Further it is conceivable that viral poly(A)
prepared by RNase
T,
contains at its ends astretch of nucleotides containing U, A, and C which would notbe digested byRNase
T,.
This may affect slightly the molecular weightofthepoly(A)
prepared by
RNaseT,.
However,
base analyses of viral poly(A) prepared with
on November 10, 2019 by guest
http://jvi.asm.org/
I
'-4
x
0s
0
=L
3
-0
x
0
0-CM
#LzL4~ <4L2:
0
20
40
60
80
100
[image:4.499.118.402.61.368.2]DISTANCE MOVED
(mm)
FIG. 1. Determinationofthe sizeof marker 7.8Spoly(A)andRSVpoly(A) by formamidepolyacrylamide gel electrophoresis. RSVpoly(A)wasisolatedfrom30 to40S [32P]RNA subunitsby RNase
T,
digestion. Thedigest wasphenol-extractedthreetimes and ethanol-precipitatedin thepresence of 30,g
of TMVRNA. The RNA recovered was then passed through anoligo(dT) -cellulose column (Fig. 3). The poly(A) peak eluted with buffer IIIand wasdivided into twoportions. One portion was ethanol-precipitated for analysis of its base composition asdescribed for Table 1. The other portion was mixed with 3H-labeled 4S, 18S RNAs, and 7.8S poly(A). After ethanolprecipitation the mixturewasdissolved in buffered formamide and subjected to electrophoresis in a 5% polyacrylamidegelcontaining98%formamideat 100Vfor 7 hasdescribedpreviously (12). The insert shows a log molecularweight/electrophoreticmobility plot (42) used to estimate the molecular weight of RSVpoly(A).RNase
T,
and a combination of RNase Aand RNase
T,
werevery similar andsuggestedthat very few (<3%) bases other than A were
present inour
poly(A)
(not shown). Weassumethat some ofthose bases come from the5 end
of RNase
T,-prepared
viral poly(A). Thus itappears that our size estimate of viral poly(A)
was little, at most 3%, affected by bases other
than A. Moreover, we may conclude that, in contrast to anearlier report(19), the poly(A) of RSV RNA is free of interspersed G-residues sensitive to digestion by RNase
T,,
since it isnotdegraded bythis enzyme.
Locationof thepoly(A)at or near the 3' end
ofRSVRNA.There issuggestive evidence from end group analyses that the poly(A) of tumor virus RNA, like the poly(A) of other viral or
cellular mRNAs is located at the 3' end of the
RNA molecule (2, 38, 43, and J. Keith, M.
Gleason, and H. Fraenkel-Conrat, Proc. Nat.
Acad. Sci. U.S.A., in press). One method used
todetermine the3' terminallocation ofpoly(A)
was based on oxidation and reduction of the
vicinal 2' and3' OH groups at the 3' terminus
ofthe RNA (38, 43). However this method has
ledto contradictory resultswhen appliedto
tu-morvirusRNA (15, 25, 30).
Another method was based on the ratio of
nucleosides to nucleotides after complete
hy-drolysis of radioactive RNA or poly(A) (2, 38,
and J. Keith, M. Gleason, and H.
Fraenkel-Conrat, Proc. Nat. Acad..Sci. U.S.A., inpress).
However, the accuracy of this method suffers
from the low
nucleoside/nucleotide
ratios ofRSV RNA (1:200 for poly(A), 1:10,000 for 30
to 40S RNA). We describe below two
ap-proaches to locate the poly(A) stretch in RSV
RNA.
End-group analysis ofviral poly(A). First
we determined the end-group of biologically
labeled poly(A) from 30 to 40S RSV RNA. If
poly(A) is at the 3' terminus of the RNA it
1518
on November 10, 2019 by guest
http://jvi.asm.org/
should, after complete digestionto mononucleo-tides, containoneadenosineperabout 200AMP residues, assuming our size estimate was
cor-rect. We have chosen enzymatic digestion of
poly(A) instead ofdigestion by alkali, because
pilot experiments had shown that approxi-mately 50% ofthe 3H of poly(A) was rendered
volatile during lyophylization after KOH hy-drolysis, presumable due to isotope exchange with water (39, 46). Exhaustive digestion with RNase A also proved unsatisfactory, as the
digestion productwasmostly ApAprather than
AMP (not shown). Complete digestion of viral,
as well as commercial poly(A) to mononucleo-tideswas obtained with RNase T2 (Table 1).
It is shown in Table 1 that the poly(A) from
RSV RNA contained one adenosine per 180
AMP residues and that commercial poly(A) containedone per320. This is compatible with our size estimate of about 200 nucleotides for
viral poly(A) and with the size estimate of 116,000for commercial 7.8S poly(A).
Nosignificant radioactivitywasdetectableat the origin of the pherogram usedto resolve the digest (Table 1), neitherwasthereradioactivity
between the spots of the four UV-absorbing monophosphate ribonucleotide markers usedto monitor the electrophoresis. However, there
weresmallamountsofradioactivity in the CMP spot if viral or commercial poly(A) were
ana-lyzed (experiments 1 and2, Table 1). Thiswas
most likely due to slight trailing of AMP,
because afterprolongedelectrophoresis
(experi-ment 3, Table 1) no radioactivity was seen in
the CMP position. Significant amounts of
ra-dioactivity migrated with the GMP spot, when
the poly(A) ofRSV was analyzed, especially in
experiment 3 (Table 1). This could be due to
residual heteropolymeric sequences associated
with this preparation of viral poly(A) or less
likely (see above; 38) due to RNase
TI-resis-tant G residues interspersed into poly(A). The
radioactivity inGMPmayhave originated from
cellular conversion of [3H ladenosine into
[3H ]GMPorfrom [3H ]guanosine contaminating
the [3H]adenosine used by us (from New
Eng-land Nuclear Corp., experiments 1 and 2,
Ta-ble 1; from Schwarz-Mann, experiment 3, Table 1). The amount of 3H electrophoresing with GMPvaried relativetothat found inAMP
in the experiments described in Table 1. By contrast, the radioactivity associated with the
adenosine spot.relative to that in the AMP
spot remained constant (.55%) in all experi-mentswithpoly(A) fromRSV(Table 1).
Control experiments using 28S [32P]rRNA (1.25 x 105 counts/min) showed that <0.04%
(<50 counts/min) of the mononucleotides had
beendephosphorylated underour conditionsof
enzymatic digestion using RNase T2, RNase
TI,
and RNaseA (Fig. 2). Thismaybeaproblemto be considered if alkali is used to digest RNA (28). Therefore, we believe that the
adeno-sine residues detected by complete enzymatic digestion of viral poly(A) werederived from the
3' end of the RNA, and consequently that
TABLE1. End-group analysis of marker and RSV poly(A)
Marker[3H]poly(A)" [3H]poly(A)from 60to70SRSV RNAb
Poly(A) tested I II III I II III
experiments
Counts/ % Counts/
%,
Counts/! Counts/! Counts/! Counts/min' min min min min min
Adenosine 400 0.31 328 0.32 400 0.31 314 0.59 144 0.55 430 0.52
CMP 246 0.18 145 0.14 0 0 162 0.30 46 0.19 0 0
AMP 128,780 99.43 103,410 99.44 129,900 99.66 52,890 98.65 24.190 98.58 80,000 97.29
GMP 72 0.06 70 0.07 36 0.03 218 0.41 144 0.55 1,800 2.19
UMP 32 0.04 36 0.04 0 0 31 0.06 13 0.05 0 0
Total 129,530 100 103,990 100 130,340 100 53,165 100 24,540 82,230 100
AMP/adenosine 322:1 315:1 325:1 170:1 168:1 186:1
Avg ofAMP/ 320±5 178+8
adenosine
aComplete digestion of poly(A) by RNase T2 was described in Materials and Methods. In experiments I and II, dephosphorylatedandethanol-precipitated Miles poly(A)was used: inexperimentIIIpolv(A) hadbeen furtherpurified by
threesubsequentsedimentations. l
[3H]adenosine-labeledpoly(A) fromRSV RNAwasisolatedby digestionwith RNasesT,and Aasdescribed in Materials and Methods. In experiments I and II, poly(A) isolated from same RNA preparation was used directly without further
purificationbysedimentation. InexperimentIII, poly(A) isolated fromadifferent RNApreparationwasfurtherpurified by
sedimentation to remove the oligonucleotides that might have co-precipitated with poly(A). Paper electrophoresis of
experimentIIIwasrunfor 3h instead of2.5h.
cThirtycountsper minute ofbackground eluted frominterspot regionhas beensubtracted.
1519
on November 10, 2019 by guest
http://jvi.asm.org/
O
L
20 23
DISTANCE MOVED (Cm)
FIG. 2. Electrophoresis of the 32P-mononucleotides produced by complete enzymatic digestion of 28S chick rRNA. Approximately I 1sgof [32PV28S rRNA (1.2 x 105 counts/mm) was digested with RNases A,
T,,and T2 in a total volume of 0.2 ml at 38 C for 2 h. The digest was analyzed by paper electrophoresis. A
20-Muliter amount of [3H]32P04 (2 x 104counts/mmn)
was spotted on the same paper and run in parallel as a p043- marker. The32p nucleotides and pQ43 marker were detected by determining the radioactivity of 1-cm stripsof the pherogram in toluene-based scintil-lation fluid.
poly(A) appears to be located at the 3' end of viral RNA. However, our experiment does not rule out the possibility te the penultimate base(s) of viral poly(A) are not A but a few U, C, G residues resistant todigestion with RNasesTA
and A used to prepare our poly(A). Further, it may be argued that our evidence for the 3'
location of poly(A) is not compelling, since it is
based entirely on the end-group of the viral
poly(A) which represents only 0.5% ofthe
radio-activity associated with the poly(A) molecule. 30 to 40S RSVbefore and after removal of
poly(A)
with RNase H of E. coli. Therefore,independent
evidence was sought to locate theposition of
poly(A)
on viral 30 to 40S RNA.Using RNase H in the presence ofpoly(dT) to
digest specifically the
poly(A),
but not theheteropolymeric sequences of viral RNA, it should be possible to
determine
thelocation of poly(A) on the RNA by analysis of the RNase H-resistantheteropolymeric
sequences of theRNA.
An RNase H which is free of othernucleolytic
activities
is essential for thisexperi-ment. Starting material for
these
experiments was RSV RNA,[3Hsadenosine
or 32P-labeled, selectedfor an intactpoly(A)
stretch by binding and subsequent elution fromoligo(dT)-cellulose
at lowionic
strength (fraction III, Fig. 3). This RNA was incubatedwith E. coli RNase H in the presence ofpoly(dT) (24)
(see Materials andMethods). The RNase H-treated RNAwasthen
compared to untreated RNA with regard to its
Millipore filter binding capacity, its poly(A)
content andsize.
Analysis of 30 to 40S RSV RNA before and
after digestion with RNase H and in the
pres-ence of poly(dT) indicated that 80% of the
RNaseA-and RNase
T,-resistant,
viralpoly(A)can be digested by RNase H (Table 2). It
appears then thatmost orall poly(A) (about15
to 20 pmol) ofabout 1,200 pmol ofviral RNA
(specific activity -105
counts/min/,ug)
wasdi-gested in our conditions. Control experiments
using the same amount of enzyme, indicated
that 110 to 140 pmol of
[3Hlpoly(A)
.poly(dT)were digested in the same conditions. Thus,
digestion of the viral poly(A) was at enzyme
excess (Table 2).
Moreover
Millipore
binding capacity
of 30to40S
RSV RNA was reduced from about 90 to10%bytreatmentwith RNaseH(Table 2). The
enzyme/substrate
ratios in these experimentswere about the same as those used above
Isee
[image:6.499.64.253.60.239.2]control experiments with [3H]poly(A).poly(dT),
Table 21.
The size ofthe RNase H-treated 30 to
40S
RNA was
indistinguishable
from that ofun-treated RNA on the basis ofsedimentation in
formaldehyde-sucrosegradients (Fig. 4) butwas
found to be slightly smaller on the basis of its
0
x 0 0L
cI
cL
30
20
10
0
I
II
'III
.5M
.1M Tris
K
1k1
I
L
0
4
8
12
16
FRACTION NUMBER
FIG. 3. Oligo(dT)-cellulose chromatography of 30-40SRSVRNA.Purified 30 to40S [32P]RNAwas dissolvedin 0.2 mlof bufferIandheatedat100 C for1 min. Fractionationwas asdescribedinMaterials and Methods. Arrows indicate where the elution with different buffers (I, II,andIII) started. Fractions (0.5 ml) were collected and
5-gsliter
portions of each fractionwerecountedin3mloftoluene-based scintil-lationfluid containing10%NCS.1520
on November 10, 2019 by guest
http://jvi.asm.org/
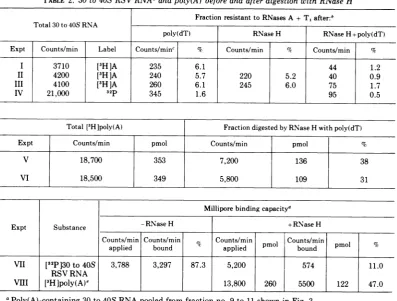
[image:6.499.267.453.389.587.2]TABLE 2. 30 to40S RSVRNAaandpoly(A) before and after digestion with RNase H Fraction resistant to RNases A +T,after:' Total 30 to 40S RNA
poly(dT) RNaseH RNaseH+poly(dT)
Expt Counts/min Label Counts/minc % Counts/min % Counts/min %
I 3710 [3H]A 235 6.1 44 1.2
II 4200 [3H]A 240 5.7 220 5.2 40 0.9
III 4100 [3H]A 260 6.1 245 6.0 75 1.7
IV 21,000 32p 345 1.6 95 0.5
Total [3H]poly(A) Fraction digested by RNase H with poly(dT)
Expt Counts/mmn pmol Counts/min pmol %
V 18,700 353 7,200 136 38
VI 18,500 349 5,800 109 31
Milliporebindingcapacity'
Expt Substance -RNaseH +RNase H
Counts/min Counts/min Counts/min pmol Counts/min pmol %
applied bound applied p bound
VII [32P130 to40S 3,788 3,297 87.3 5,200 574 11.0
RSV RNA
VilI [3HIpoly(A)e 13,800 260 5500 122 47.0
aPoly(A)-containing 30to40SRNApooledfrom fractionno.9 to11showninFig.3.
RNase H digestionwascarriedout asdescribed in Materials and Methodsusing approximately 0.4ug of
[3H]adenosine (1.5 x 104counts/min) or 32P (7 x 104 counts/min) labeled RSV RNAineach assay. Control tubes eitherminusRNaseH orminuspoly(dT) (0.25,g) but with RNase H were incubated in parallel. At the end of reaction, duplicate portions of reaction mixture from each tube were taken separately. They were incubated with DNasetoremovepoly(dT)afteradjustingtheir volumeto100
pliters
and the saltconditionsto those describedinMaterials and Methods for digestion with DNase. Subsequently they were adjusted to 0.5 ml pertube andsubjectedtoRNaseAplus RNaseT1digestionforthe assay of the remaining poly(A) sequences. At the endofdigestionby RNases, 100,gofyeasttRNAwasadded andtrichloroacetic acid precipitable material wasdetermined (44). Each number represents the averageofduplicate assays. Fifteen counts per minute of background had been subtracted.cTheamountofRNA resistant toRNases.
dRNase Hdigestionwascarriedout asdescribed in ". A control without RNase H was incubatedinparallel. Each tube contained 1.5 x 104counts/minof32P-labeledpoly(A)-containing 30to40S RNA. After incubation eachreaction mixturewasadjustedto0.1 MNaCl and1 mMEDTA anda 100-fold excess ofcold poly(A) (25 Mg)wasaddedtopreventpoly(dT) from complexingtoRSV RNA. They were thenheat-denatured at 100 C for 40 sand chilled. Each reaction mixturewasadjustedto atotal volumeof 0.3ml withSDS standard buffer and sedimented through a5 ml 10 to25%linearSDS sucrosegradient afteradding 18S [3H]rRNA as an internal marker(Materials andMethods). Aftersedimentation, 30to40SRSV RNAswerepooledseparately, ethanol-precipitated andsubjectedtoMilliporebinding(MaterialsandMethods).
eStandard assaysofRNaseHwith [3H]poly(A) andpoly(dT) wereperformedinparallel and carried outas described in Materials and Methods. Eachnumber represents theaverage of duplicate assays.
electrophoretic mobility in
formamide-polya-crylamide gels (Fig. 5). This suggests that the
poly(A) segment is locatedatornearaterminus
ofthe RNA, since its removal did not
signifi-cantly affect the size of the RNA. Given a
terminal location of the
poly(A),
we would expect at most a small increase in electropho-retic mobility after removal ofpoly(A),
sincethepoly(A) segment represents onlyabout 1.5% ofthe 30to40S RSV RNA.Further,therather
homogenous distribution ofRNaseH-treated30 to40S RSV RNA informaldehyde gradients or formamide gels indicates that the RNase H usedby us wasfree ofdetectableendonuclease.
The second minor peakofRNA observedboth in the RNase H-treated and untreated RNA
1521
on November 10, 2019 by guest
http://jvi.asm.org/
I 28S - NsH 28S + 2S
RNaseH I +RNase H Complete
cl 4 o poly(dT) poly(dT) ( an
RNase H(C)asdsrbdi al .Sbeuety ahsml a iue it tnadbfe o2m fe
X 3 -x
addingtoeah1C-)ro f02MDAan 0y fTVN.Atrpeoletato 8 SPr a
21-0 I 0
016e04 8 12 4 8 12 160 4 8 12 16
FRACTION NUMBER
FIG. 4. Formaldehydesucrosegradientsedimentationof30 to40SRS V RNAbeforeandaftertreatmentwith RNaseH.Poly(A)-containing [3H]adenosine-labeled30 to40SRSVRNAwaspreparedasdescribedfor Fig. 3. Three identicalportions of30 to 40S RNA were incubated with poly(dT) (A), RNase H(B), poly(dT), and RNase H(C)asdescribed in Table2.Subsequently, eachsamplewasdiluted with standardbufferto2mlafter addingto each 10 litershof 0.2MEDTAand 30 ugofTMVRNA.After phenolextraction28S [3eP]rRNAWas
added toeachsample; the RNAs werethen ethanolprecipitated. After pelletingandwashingtwice with 75% ethanol,the RNA ineachtubewasredissolved in0.3mlofbuffer containing0.1MNaCl,2mMEDTA, pH 7.2, 0.1%SDS,and1.1Mformaldehyde. Afterincubationat65 Cfor15mineach RNAsamplewaslayeredonthe top ofa 5-mI10 to20%sucrosegradient containingthesamesalts andformaldehydeas the bufferdescribed above. SedimentationwasinparallelinaSpincoSW65rotor at65,000rpmfor3hat20 C.Fractions(0.3 ml)
were collected through the bottom of the tube, and
100-Mgliter
portions of each fraction were measuredfor radioactivity.control seen in the
formamide-polyacrylamide
gels (Fig. 5) is
thought
to be 30 to 40S RNAofsize class
b,
derivedfromtransformation-defec-tive virus which segregates
spontaneously
from stocks ofnondefective aviansarcomavirus(29).We conclude thatRNase H treatment
effec-tivelyremovesallor mostofthe
poly(A)
associ-ated with viral RNA without
significantly
re-ducing its size. It follows that the
poly(A)
islocated near a terminus and that there are no
largepoly(A) stretches located within the30to
40SRNA chain.
Notall30 to40S RNA subunits of60to70S
RSV RNA contain
poly(A). During
this andearlier (23) studies, it was repeatedly observed
thatheat-denatured 60 to 70S RNAorpurified
30 to
40S
RNA had a much lowerbinding
capacitytoeither
Millipore
filtersoroligo(dT)-cellulosethan intact60 to70S RNA. This raised
the question whether all subunits in 60to 70S RNA contain a poly(A) sequence. It seemed
plausible that if there exist some 30 to 40S
subunits which do not contain poly(A) se-quences, they could bind tooligo(dT)-cellulose
only ifcomplexed toother, poly(A)-containing
subunits witnin the same 60 to 70S RNA
complex. Therefore, the
oligo(dT)-cellulose
binding capacityofpurified60 to70S and 30 to
40S RNAs were compared at different ionic
strengths. The results, summarized inTable3,
indicate that 90% of the60 to70S butonly65%
ofthe30 to40S RNA bindtooligo(dT)-cellulose athighionicstrength, implying that about1 out of3 30 to40S RNAspecies isdevoidofapoly(A)
stretch that confers binding capacity for
oli-go(dT)-cellulose. This is in agreement with a
recent report by Ihle et al. (20) that only
two-thirds ofpurified 30 to40SRNA of mouse
leukemia virus RNA binds to
poly(U)-sepharose. As expected, most of an 18S
ribo-somal RNA used as internal control in the
binding
studies wasincapableofbindingto anoligo(dT)-cellulose (Table3).
Direct analyses of the poly(A) content of
fractions of 60 to 70S and 30 to 40S RNA,
distinguished by
their differential bindingca-pacities to oligo(dT)-cellulose, confirmed the
above conclusions. It was found that after
complete enzymatic
digestion
ofheteropolym-eric sequences with RNase A and RNase
T,
(Materials and
Methods),
30 to 40S and 60to70S RNA eluting from oligo(dT)-cellulose at
high ionicstrength (fraction I, Fig. 3, Table 3) were devoid of enzyme-resistant RNA and therefore did not contain poly(A) (not shown). However, 60 to 70S and 30 to 40S RNA eluting atlowionicstrength (fraction
III,
Fig. 3, Table 3)contained1.5%resistantRNA,iflabeled with 32P, and 6% if labeled with [3H]adenosine asexpected for a poly(A) containingRNA (Table
2).The RNAspecieswhichelutedat intermedi-ateionic strength contained 1to 1.5%resistant
1522
on November 10, 2019 by guest
http://jvi.asm.org/
[image:8.499.113.396.81.222.2]0 20 40 60
DISTANCE MOVED
(mm)FIG. 5. Formamide polyacrylamide gel electrophoresis of 30 to 40S RSV RNA before (A) and after (B)
treatmentwith RNase H.Poly(A)-containing 32P-labeled30to 40S RSV RNAselutingat low ionicstrength from oligo(dT)-cellulose(fractionIIIshown inFig.3)wasincubated withRNase HasdescribedforFig.4.After phenolextraction, [3H]adenosine-labeledpoly(A) containing 30 to 40S RNA and 18SrRNA wereaddedas
internalmarkers. The RNAswereethanolprecipitated.Electrophoresiswasinpolyacrylamidegelscontaining
98%formamideat 100 Vfor 12hasdescribed (12).
RNA, if labeled with [3H ]adenosine (not shown). The degree ofRNase resistance in this RNAspeciesvaried with different preparations,
sothisfraction of RNA islikelytoinclude RNA
species carrying poly(A) segments which are
much shorter than those of the RNA fraction
eluting from oligo(dT)-cellulose only at low ionicstrength (fractionIII, Fig. 3).Weconclude that 60 to 70S viral RNA contains 30 to 40S
RNAsubunits withpoly(A) (-65%), some
with-out poly(A) ("30O%) and a few subunits (5 to 10%) with short segments ofpoly(A).
The sizes of 30 to 40S viral RNA subunits which lack poly(A) (fractionI, Table3, Fig. 5)
were compared to the 30to 40SRNAsubunits which contain poly(A) (fraction III, Table 3, Fig. 3) by simultaneous electrophoresis in form-amide-polyacrylamide gels. It can be seen in
Fig. 6A and B that 30to40S [3H]-or [32P]RNA
withpoly(A) migrated alittle slowerthan 30to
CV
0
V'-4 x
Q-(O4
8
4
0
x
0 CL 0
Q r)
15'):3
on November 10, 2019 by guest
http://jvi.asm.org/
[image:9.499.102.388.64.482.2]TABLE 3. Oligo(dT)-cellulose column binding of RSV RNAa
60to70S RNA 30 to40S RNA 18SrRNA5 Experimentsand RNA Labeling %Eluted %Eluted %Eluted
preparations
IP [3HlU 5.7 4.3 90.0 18.0 24.0 58.0 86.0 5.6 8.4
32p 5.0 4.0 91.0 14.0 18.0 68.0 87.0 5.8 7.2
II [3HJA 5.4 3.6 91.0 17.6 15.7 67.0
III [3H
]A
29.5 8.5 62.0IV 32P 25.0 13.0 62.0
Vd 32P 21.8 5.2 73.0
Average 5.4 4.0 90.7 21.0 14.1 j 65.3 86.5 5.7 7.8
Purified RSV60 to70S or 30 to 40SRNAsweresubjectedtooligo(dT)-cellulosecolumn fractionationas de-scribedin Materials and Methods andforFig.3.Anelutionprofileof 30to40S is shown inFig. 3.Percentages ofradioactivityelutingathigh,intermediate, and low ionicstrengthweretermed I,II,and III,respectively, and correspondtotheelutionprofileshown inFig. 3.
h [3H]uridineor32P-labeled 18SrRNAswereusedasinternalcontrols in the elution of 60to70S RSV RNAin Exp. 1.
[3H]uridineand 32P were addedtodifferent dishesofthe samevirus-infected culture and the RNAswere isolated inparallel.Thesedif'ferentially labeled RNAswereusedforcomparisonoftheir size(Fig.4and 5).
dFractionated30 to40S RNAsofthisexperimentwereusedforfingerprintanalysis(Fig. 7).
40S [3H]-or [32P]RNA withoutpoly(A). An18S
rRNAmarker isincluded in these experiments;
in addition, a shoulder is seen at the leading edgeof eachpeak,thisisthoughttobe30to40S RNA of size class b (see above). A control
experiment showsthatpoly(A) containing30to
40S [3H]- and [32P]RNA coincide upon
electro-phoresis in the same conditions (Fig. 6C). It
appearsthen,that in agreement withthe exper-iments described above in which poly(A) was
removed enzymatically from 30 to 40S RNA,
the presence of poly(A) reduces slightly the
electrophoretic mobilityof 30 to 40SRNA.
Theexistence of 30to40SRNAsubunitswith and without poly(A) in 60 to 70S tumor virus
RNA observed here and by others (20, 23),
raises the question ofwhether
poly(A)-contain-ing andnonpoly(A)-containingRNAsrepresent
genetically different fractions of the RNA. To
test this the fingerprint patterns of RNase
T,-resistant
oligonucleotidesofthetwotypes of30to40SRNAwerecompared. Itcanbeseen in
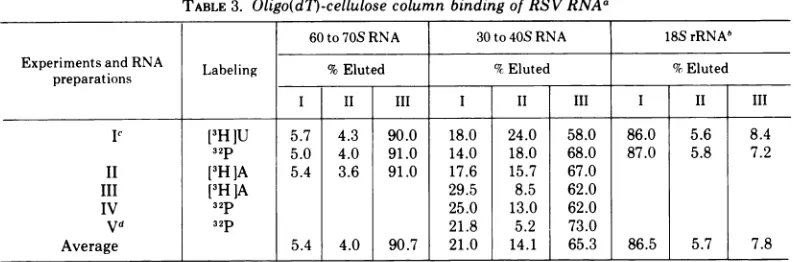
Fig. 7 that, except for the absence of the
poly(A) spot (lower right corner in Fig. 7A) in
thepoly(A) lacking30 to40S RNA, the
finger-prints ofpoly(A)-containing and poly(A)-lack-ing30to40SRNAspeciesareidentical. There-fore, we conclude that the 30 to 40S RNA species with and without poly(A) stretch are
genetically identical. This conclusion is
com-patible with thefinding that the complexity of
aviantumorvirus RNA is onthe order of 3.5 x
106daltons, whichistheapproximate
complex-ity ofone 30to 40Ssubunit only (K. Beemon,
P. Duesberg, and P. Vogt, Proc. Nat. Acad.
Sci.U.S.A., inpress).
Locations of some RNase
T,-resistant
oli-gonucleotides and of poly(A) appear to be the same in all 30 to 40S RNAs. The above
experiments and those of others (43) indicate
thatpoly(A)is locatedat or nearthe 3' end of 30
to 40S RSV RNA. If the heteropolymeric se-quences of 30 to 40S RNA all had the same location on the
polynucleotide
relative topoly(A), instead ofbeing circularly
permuted,
poly(A)-tagged natural or artifically
produced
fragments would be very useful for sequence
analysesofviral RNAs. It would beexpected,for
example, that ifthe
heteropolymeric
sequencesof all 30 to 40S RNAs are located
identically
relativetopoly(A), the
complexity
ofapoly(A)-tagged fragment of 30 to 40S RNA should be
directly
proportional
to the size of thatfrag-ment. However, if the RNA sequences are
extensively
permuted
thecomplexity
of apoly(A)-tagged
fragment must notbe less thanthat of the total RNA. To
distinguish
between thesealternativeswehavealkali-degraded 30 to40S
[32P]RNA
into fragments sedimentingbroadly with apeakof11S (36), corresponding
to anaverage size ofabout240,000daltons(41).
The poly(A) containing fragments of the
de-graded RNA were isolated by binding and
subsequentelution fromMilliporefilters
(Mate-rials and Methods). The recoveries ranged
be-tween 5 to 10%indifferent experiments.This is
expectedifone considers that an 11S fragment
of240,000 daltons corresponds
roughly
to 5 toDUESBERG 1524)
on November 10, 2019 by guest
http://jvi.asm.org/
~~~~~~~1-.-v,-\,.. .. .,
15 18 (^) Fraction Il-1
04~ ~~ab
o0
4 20phrei. 30t 0 N ihutpl()wspe
0 ~~~~~~~~~~~~~~~0~
C 30-40S Peaka
15
-l(a)FractionIII 15
18S
pl( Fraction III
10 b 10
5 -5
0l
00 20 40 60
[image:11.499.51.237.71.547.2]DISTANCE MOVED(mm)
FIG. 6. Comparison of the size of poly(A)- and
non-poly(A)-containing30 to 40S RSV RNA isolated
from 60 to 70S viral RNA byformamide gel electro-phoresis. 30 to 40S RNA without poly(A) was pre-pared byelutionfromoligo(dT)-celluloseathighionic strength (fraction I, Fig.3) and30to40Swithpoly(A)
was thatwhich elutedatlow ionic strength (fraction III,Fig. 3).Portionsof30to40SRNAfrom
oligo(dT)-cellulosefractions IandIIIweremixedtogetherwith
18S3rRNAandanalyzed by formamide gel
electropho-resisas describedforFig.5. (A) 30to40S RSVRNA withoutpoly(A) (fraction 1)and 30to40S RNA with
10%ofa 30 to
40S
RNAof 3.5 x 101daltons.We have compared the complexity of these
poly(A)-tagged fragments of 30 to40SRNA to
that of undegraded 30 to 40S RNA by
finger-printing. It can be seen in Fig. 7C that the
numberofRNaseT1-resistant oligonucleotides,
and thus the complexity ofpoly(A) containing
11S fragments of PR-B RNA, is significantly
lower than that of 30 to 40S RNA (Fig. 7A).
Many large oligonucleotidesseen in the
finger-print of 30 to 40S RNA are completely absent
from the pattern of poly(A)-tagged 11S RNA
fragments. These arethoughttobe
oligonucleo-tides located most distantly from the poly(A)
stretch on the 30 to
40S
RNA. Someoligonu-cleotides marked
by
letters inFig.
7AandCarepresent athigh and some atintermediate con-centrations relative to the total counts per
minute on the fingerprint or to the counts per
minute ofthepoly(A) spot(Table4).Theseare
thought to be located at or near the poly(A)
stretch of 30 to 40S RNA. Varying relative
concentrations ofoligonucleotides are expected from the heterogenous size distribution of the
poly(A)-tagged fragments used to prepare the
fingerprint pattern. Since fragments of larger
size would contributeoligonucleotidesnot
pres-ent in smaller fragments. We conclude that
some, perhaps all, RNase
T,-resistant
se-quences of tumor virus RNA have the same
location relative tothe poly(A).
Itshould then be
possible
tomapthe positionof large oligonucleotides relative to that of
poly(A) on the 30 to 40S
polynucleotide
bycomparing the mass ratios ofa given
oligonu-cleotide tothe total RNAor tothatofpoly(A)in
the fingerprint patterns derived from intact
poly(A)-tagged 30 to 40S RNA and
poly(A)-tagged fragmentsofRNA. Forexample, it can
bededucedfrom the datainTable4thatspota
and b must be more distant from the poly(A)
stretchthan spot c.
DISCUSSION
The experiments described here confirm and
extend earlier studieson
poly(A)
in tumorvirusRNA (16, 17, 23) andsuggestthattwo-thirdsof
the 30 to 40S subunits of 60 to70SRSV RNA contain a poly(A) stretch consisting of about
200nucleotidesat or nearthe 3' endofthe RNA.
The finding that some 30 to 40S RNA
sub-poly(A) (fraction III). (B) Different preparations of thesame twospecies of30to40SRSVRNAasin(A).
(C) Two preparations of30 to 40S RSVRNA with poly(A). Peakarepresents 30to40S RSV RNA. Peak b is the18S[3H]rRNAaddedas aninternal standard.
on November 10, 2019 by guest
http://jvi.asm.org/
-.
--0
-i be *
A
*
o
1~ f it
_
i~~
A. %
a,
ID0
[image:12.499.57.452.77.298.2]B.0
Cl_
D
FIG. 7. (A-O) Fingerprint analyses of30to40SRSVRNA containingpoly(A), (A); of30to40SRSVRNA withoutpoly(A), (B);andofpoly(A)-tagged 11Sfragments of 30to40SRSVRNA, (C); and of the 30to40SRNA ofatransformation-defectivePRRSV-B(D).30to40S RSVRNA without poly(A)wasprepared by elution from
oligo(d7)-celluloseathighionicstrengthand 30to40SRSV RNA withpoly(A) byelutionatlowionic strength (see Fig. 3). The RNAs were exhaustively digested with RNase T1 and the digests were analyzed by two-dimensionalfingerprint analysisas described (K.Beemon, P. Duesberg, and P. Vogt, Proc.Nat. Acad.Sci. LS.A.,inpress).AnI S,poly(A)-containing fragment of30to40SRNAwasprepared byincubating 30to40S RSVRNA in50 Mliters 0.05 MNa2CO,(pH 10)at50 Cfor 6min (36). The solutionwas thenmixed with300
Aliters ofstandardbuffercontaining 0.2%oSDS andsedimentedinthepresenceofan18S
[9H]rRNA
standard.Asymmetrical peak sedimentingat11Swasobtainedandethanolprecipitated. The pelletwasredissolvedand
passed throughaMilliporefilter.Millipore-bound 11SRNA(5 to10%of the 11S RNAapplied) waselutedas
described inMaterialsandMethods andfingerprintedasdescribedfor (A).
TABLE 4. Radioactivitya of certainoligonucleotidesand of the total RNA from which itwasderivedafter fingerprintingasshowninFig. 7aandc
60 to70SRNA Poly(A) fragment Poly(A) fragment
(Fig. 7a) (1OS) (11S) (Fig.7c)
Partofdigestanalyzed
Counts/min Counts/min % Counts/min %
Total 1.17x 106 100 48,750 100 70,400 100
Poly(A) 15,560 1.33 8,140 16.7 9,190 13.1
Spota 1,470 0.13 31 0.06 79 0.11
Spot b 2,900 0.25 48 0.10 108 0.15
Spotc 1,730 0.15 391 0.8 570 0.81
Spot d° 1,550 0.13
aQuantitations were doneaccordingtothe procedures described elsewhere (K. Beemon,P. Duesberg, and P.Vogt, Proc.Nat.A-cad. Sci.U.S.A., inpress).
b Anotherpossible spottocorresponding spotcofFig. 7c.
units existwhich contain poly(A) while others
lack poly(A), but appear otherwise identical, could reflect either heterogeneity of the 60 to 70S RNA population extracted from our virus
preparationorsubunit differences within the60
to70S complex. Since binding of RNA, bothto
Millipore filters and to oligo(dT)-cellulose, is thoughttoinvolvepoly(A)sequences,and since over 90% of the 60to 70S RNA binds to these substrates, itappearslikely thatmostofthe30 to40S RNA species without poly(A) arepartof
a 60 to 70S complex which includes other
h
W.
1526
on November 10, 2019 by guest
http://jvi.asm.org/
subunits containing poly(A). If correct, this
would be an independent argument in favor of
the subunitstructure of 60 to70SRNA; it would
imply that 60 to 70S complexes exist between
poly(A)-containing and poly(A)-free 30 to 40S
subunits. The phenomenon could not be
ex-plained interms of a hypothesis which assumes
that the 60 to 70S RNA is a conformational isomer of 30 to 40SRNA.
Results of three different experiments now
suggest that the poly(A) of tumor virus RNA is at or near the 3' end of the RNA. It was shown
bytwodifferent end-group methodsthat
adeno-sine is the 3' terminal nucleotide of poly(A) as wellas ofviralRNA (2, 38, 43, and J. Keith, M.
Gleason, and H. Fraenkel-Conrat, Proc. Nat.
Acad. Sci. U.S.A., in press). Nevertheless,
end-group analyses of macromolecules are subject to error, because only a very small fraction of
the molecule is analyzed. However, these
re-sultsare complementedby our observation that
RNaseHremoves poly(A) from viral RNA
with-out significantly reducing its size. The sum of
these experiments suggests strongly that the
poly(A)of 30 to40Stumor virus RNA is at the 3'
endofthepolynucleotide,aswith other
poly(A)-containing RNAs (31, 32, 40). There is no
evi-dencefor internal poly(A), because (i) one
poly-(A) stretch ofthe size found in this study can
account forallpoly(A) associated withan aver-age 30 to 40S RNA, (ii) 30 to 40S RNAs with
and without poly(A) (eithernaturally occurring
orafter treatment with RNase H) have
approxi-mately the same size, (iii) the poly(A) of 30 to
40S tumor virus RNA was found to reside at a
unique location relative to other
heteropoly-mericsequences of the RNA. However, prelim-inary experiments suggest that A-rich runs,
other than the poly(A) of 200 nucleotides
de-scribed above, exist in the RNA. This
sugges-tion isbasedonthe observation that 20 to 25%
of all 11S fragments of 30 to 40S RNA were
found to bind to oligo(dT)-cellulose, whereas
only 5 to 10% of such fragments would be
expected to bind ifbinding were solely due to
poly(A)-tagged fragments of an RNA with an
originalsize ofaround3.5 x
106
(12).Some,
perhaps
all, individual RNaseT,-re-sistant oligonucleotides appear to have the
samerelative location onall the 30to40SRNA
molecules with respect to-the
position
ofpoly(A), suggesting that individual 30 to 40S
RNAs are not
permuted
withregard
to se-quences of the RNA. This iscompatible
withgeneticexperimentswhich indicate that certain
markers of different avian tumor virus strains
recombine at stable
frequencies
(J.
Wyke,
J. Bell, and J. Beamand, Cold Spring Harbor
Symp.
Quant. Biol., in press; P. K. Vogt,per-sonal communication).Thus, thepoly(A) should prove a useful marker for mapping oligonucleo-tides on the RNA.
Given acomplexityof only 3.5 x 106daltons
forRSVRNA (K. Beemon, P. Duesberg, and P. Vogt, Proc. Nat. Acad. Sci. U.S.A., in press) and several (two to three) 30 to 40S subunits
(H. Delius, P.
Duesberg,
and W.Mangel,
ColdSpringHarbor
Symp.
Quant.
Biol.,
inpress)
for60to70S RNA it maybe
argued
that thediffer-ent sequences are
distributed
unevenly
overseveral subunits. In this case one subunit may
contain onlyone-halfofall different sequences,
but all ofthesesequenceswould berepresented
twice per 30 to 40S polynucleotide. The
com-plexityofpoly(A)-tagged fragmentsof 30 to40S
RNAwould then notonly bea function of their
size but also of the number of subunits over
whichthe different sequences of RSV RNA are distributed. The lowcomplexityofthe 10 to11S
fragments analyzed here (containing one to
three largeoligonucleotidesout ofabout20 to 30
per 30 to40SRNA) argues against the
possibil-ity that the differentsequences ofRSV RNAare
distributed repetitively over many subunits.
Although our evidence does not exclude the
possibility that sequences of RSV RNA are
unequally distributedovertwo subunits, thisis
considered unlikelyforotherbiologicalreasons.
Nondefective sarcoma viruses containing 30 to
40S RNA of size class a segregate
transforma-tion-defective viruses which contain
only
30to40S RNAof sizeclass b atratherhigh
frequen-cies(11,29). If the sequences ofRSVRNA were
distributed over several different 30 to 40S
subunits transformation defectives would be
expected which contain both class aandclass b
RNA. This was neverobservedin many
analy-ses of
transformation-defective
viruses (11; P.Duesberg and P. Vogt,
unpublished
observa-tions).
ACKNOWLEDGMENTS
We thank J. Hurwitz, J. Leis, M. Jacquet, and I. Berkowerforseveralgiftsof RNase Hessentialforthisstudy.
We aregratefultoSunYung Kim and MarieStanleyfor
as-sistance with these experiments, and Karen Beemon, Jan Maisel, andEliCanaaniforreview of themanuscript.
This work was supported by Public Health Service
re-search grant CA-11426 from the National CancerInstitute
andbythe CancerProgram-NationalCancerInstitute,under Contractno.N01CP 43212.
ADDENDUM IN PROOF
It wasshown recentlythat the fingerprintpattern of the transformation-defective deletion mutant(11, 12, 29) of PR RSV-B lacks the two oligonucleotide
on November 10, 2019 by guest
http://jvi.asm.org/
spotstermedaand b in Fig. 7but is otherwise
indis-tinguishable from thatofPR RSV-B RNA(Fig. 7;P. Duesberg and P. Vogt, unpublished observations). Moreover, theRNA ofthis deletionmutant, like that ofother transformation-defective mutants(11, 12, 29) isabout 12 to 15%shorter than that of thewild-type PR RSV-B (P. Duesberg and P. Vogt, unpublished observations). Thus, it appears that RNAsequences
concerned with cell transformation and lacking in this transformation-defective mutant are located
within5to10%ofthe3'terminusof PR RSV-BRNA. LITERATURE CITED
1. Adesnik, M., M.Salditt,W.Thomas, and J. E. Darnell.
1972. Evidence that all messenger RNA molecules (except histone messenger RNA) contain poly(A) se-quencesand that thepoly(A)hasanuclearfunction. J.
Mol. Biol. 71:21-30.
2. Ahmad, M. S.,P. D.Markham, andD. G.Glitz. 1972.
Terminal nucleotides of avian myeloblastosis virus RNA and ofribosomal RNA from chicken leukemic myeloblasts. Biochim. Biophys.Acta281:554-563.
3. Armstrong, J. A., M. Edmonds, H. Nakazoto, B. A.
Phillips, and M. H. Vaughan.1972.Polyadenylic acid
sequencesinthevirion RNA ofpoliovirus andeastern
equine encephalitis virus. Science176:526-528. 4. Aviv, H., and P. Leder.1972.Purificationofbiologically
active globin messenger RNA by chromatography on
oligothymidylic acid-cellulose. Proc. Nat. Acad. Sci. U.S.A.69:1408-1412.
5. Beers, R. F., Jr.1960.Hydrolysisofpolyadenylic acidby pancreaticribonuciease. J. Biol.Chem.235:2393-2398. 6. Canaani, E., and P. Duesberg. 1972. Role ofsubunitsof
60 to 70S avian tumor virus ribonucleic acid in its
template activity for the viral deoxyribonucleic acid polymerase. J. Virol. 10:23-31.
7. Darnell, J. E., R. Wall, and R. J. Tushinski. 1971. An
adenylicacid-richsequenceinmessengerRNAofHeLa
cells and itspossiblerelationshiptoreiteratedsitesin
DNA.Proc. Nat. Acad. Sci. U.S.A.68:1321-1325. 8. Darnell, J. E., L. Philipson, R. Wall, and M. Adesnik.
1971. Polyadenylicacidsequences:rolein conversion of nuclear RNA into messenger RNA. Science
174:507-510.
9. Duesberg, P. H. 1968. Physical propertiesofRous
sar-coma virus RNA. Proc. Nat. Acad. Sci. U.S.A.
60:1511-1518.
10. Duesberg, P. H.,H. L.Robinson, W. S. Robinson,R.J. Huebner, and H. C. Turner. 1968. Proteins ofRous
sarcomavirus.Virology36:73-86.
11. Duesberg, P. H., and P. K. Vogt. 1973. RNA species obtained fromclonallines of avian sarcomaandfrom
avian leukosisvirus.Virology54:207-219.
12. Duesberg, P. H., and P. K. Vogt.1973.Gel electrophore-sis ofavianleukosisandsarcomaviralRNAin
formam-ide: comparison with other viral and cellular RNA species.J. Virol. 12:594-599.
13. Edmonds, M.,and M.G. Caramela. 1969.Theisolation andcharacterizationofadenosine monophosphate-rich polynucleotides synthesized byEhrlich ascites cells. J. Biol.Chem.244:1314-1324.
14. Ehrenfeld, E.,and D. F. Summers.1972.Adenylate-rich
sequencesinvesicular stomatitis virusmessenger ribo-nucleicacid. J.Virol. 10:683-688.
15. Erikson,R.L., E.Erikson,and T. A. Walker. 1971.The
identificationofthe 3-hydroxylnucleoside terminus of avianmyeloblastotisvirusRNA.Virology45:527-528.
16. Gillespie, D., S. Marshall,and R.C. Gallo.1972.RNAof
RNAtumourviruses containspoly(A).Nature N. Biol.
236:227-231.
17. Green, M., and M. Cartas. 1972. Thegenomeof RNA
tumor viruses contains polyadenylic acid sequences.
Proc. Nat.Acad. Sci.U.S.A. 69:791-794.
18. Greenberg, J. R., and R. P. Perry. 1972. Relative occur-renceofpolyadenylic acidsequencesinmessengerand heterogeneous nuclear RNA of L cellsasdeterminedby poly(U)-hydroxylapatite chromatography. J. Mol. Biol.
72:91-98.
19. Horst, J., J. Keith, and H. Fraenkel-Conrat. 1972. Characteristic two-dimensionalpatternsofenzymatic digests ofoncorna and other viral RNAs. Nature N.
Biol. 240:105-109.
20.Ihle, J. N., K-L. Lee,and F. T.Kenney.1974. Fractiona-tion of 34S ribonucleic acid subunits from
oncor-navirusesonpolyuridylate-Sepharose columns. J. Biol.
Chem. 249:38-42.
21. Johnston, R. E., and H. R. Bose. 1972. Anadenylate-rich
segmentin the virionRNA of Sindbis virus. Biochem. Biophys. Res. Commun.46:712-718.
22. Kates, J. 1970. Transcription of the vaccinia virus
ge-nome and the occurrence of polyriboadenylic acid
sequences in messenger RNA. Cold Spring Harbor Symp. Quant. Biol. 35:743-752.
23. Lai, M. M. C., and P. H. Duesberg. 1972. Adenylic acid-richsequencein RNAs of Roussarcomavirusand Rauscher mouse leukaemia virus. Nature (London) 235:383-386.
24. Leis, J. P.,I.Berkower,andJ. Hurwitz.1973.Mechanism
of action ofribonuclease Hisolatedfromavian
myelo-blastosis virus and Escherichia coli. Proc. Nat.Acad. Sci.U.S.A. 70:466-470.
25. Lewandowski, L. J., and S.H.Leppla.1972.Comparison of the3' termini ofdiscretesegmentsof the double-stranded ribonucleicacidgenomesofcytoplasmic poly-hedrosis virus, wound tumor virus, and reovirus. J. Virol. 10:965-968.
26. Lim, L., andE. S.Canellakis. 1970.Adenine-rich
poly-mer associated with rabbit reticulocyte messenger
RNA.Nature (London) 227:710-712.
27. Lindberg, U., andT.Persson. 1972.Isolation of mRNA from KB-cells by affinity chromatography on
polyu-ridylic acid covalently linked to Sepharose. Eur. J.
Biochem.31:246-254.
28. Maitra, U.,Y.Nakata,and J. Hurwitz. 1967.Therole of
deoxyribonucleic acid in ribonucleic acid synthesis. XIV. A study of the initiation of ribonucleic acid synthesis.J. Biol.Chem.242:4908-4918.
29. Martin, G. S., and P. H. Duesberg. 1972.The a-subunit
in the RNA oftransforming avian tumor viruses: I.
Occurrence in differentvirus strains. II. Spontaneous
loss resulting in nontransforming variants. Virology
47:494-497.
30. Maruyama, H. B., M. Hatanaka, and R.V.Gilden.1971. The 3-terminal nucleosides of the high molecular weight RNA of C-type viruses. Proc. Nat. Acad. Sci. U.S.A.68:1999-2001.
31. Mendecki, J., S. Y. Lee, and G. Brawerman. 1972. Characteristics of thepolyadenylicacidsegment associ-ated withmessengerribonucleic acidinmousesarcoma
180ascitescells.Biochemistry11:792-798.
32. Molloy, G. R., M. B. Sporn, D. E. Kelley, and R. P.
Perry. 1972. Localization of polyadenylic acid
se-quences in messengerribonucleic acidofmammalian
cells.Biochemistry11:3256-3260.
33. Pemberton,R.E., and C. Baglioni.1972.Duck hemoglo-binmessengerRNA containsapolynucleotidesequence
rich inadenylicacid. J. Mol. Biol.65:531-535. 34. Penman, S.,M.Rosbash,and M.Penman.1970.
Messen-ger and heterogeneous nuclear RNA in HeLa cells:
differential inhibitionby cordycepin. Proc.Nat. Acad.
1528
on November 10, 2019 by guest
http://jvi.asm.org/
LOCATION OF POLY(A)IN RSV RNA Sci.U.S.A. 67:1878-1885.
35. Philipson, L., R. Wall, G. Glickman, and J. E. Darnell. 1971. Addition of polyadenylate sequences to virus-specificRNA duringadenovirus replication. Proc. Nat. Acad.Sci. U.S.A. 68:2806-2809.
36. Pollet,R.,P.Knolle, andC. Weissmann. 1967. Replica-tion ofviral RNA. XV. Purification and properties of
Q, minus strands. Proc. Nat. Acad. Sci. U.S.A.
58:766-773.
37. Pridgen, C.,and D. W. Kingsbury. 1972.Adenylate-rich
sequences in Sendai virus transcripts from infected
cells.J. Virol. 10:314-317.
38. Rho, H. M., and M. Green.1974.Thehomopolyadenylate andadjacent nucleotidesatthe3-terminus of30-40S RNA subunits in the genome of murine
sarcoma-leukemia virus. Proc. Nat. Acad. Sci. U.S.A.
71:2386-2390.
39. Shelton, K. R., and J. M. Clark, Jr. 1967. A proton
exchangebetweenpurines andwaterand its
applica-tiontobiochemistry. Biochemistry6:2735-2739. 40. Sheldon, R., J. Kates. D.E.Kelley,and R. P. Perry. 1972.
Polyadenylic acidsequences on3' termini of vaccinia
messenger ribonucleic acid and mammalian nuclear
and messenger ribonucleic acid. Biochemistry 11:3829-3834.
41. Spirin,A.S.1963.Someproblems concerning the
macro-molecularstructureofribonucleic acids. Progr. Nucleic Acid Res. 1:301-345.
42. Staynov,D. Z., J. C.Pinder, and W. B. Gratzer. 1972.
Molecularweight determinationof nucleicacids bygel electrophoresis in non-aqueous solution. Nature N.
Biol.235:108-110
43. Stephenson,M.L.,J. F.Scott,and P.C. Zamecnik. 1973.
Evidence thatthe polyadenylic acidof"35S"RNA of avian myeloblastosis virus is located at the 3-OH terminus. Biochem.Biophys. Res.Commun.55:8-16.
44. Wang,L-H., andP. H.Duesberg.1973. DNApolymerase ofmurine sarcoma-leukemia virus: lackofdetectable
RNaseHandlowactivitywith viral RNA and natural
DNAtemplates. J.Virol. 12:1512-1521.
45. Weinberg, R. A.,Z. Ben-Ishai,and J. E. Newbold. 1972.
Poly(A) associatedwithSV40messengerRNA. Nature N.Biol.238:111-113.
46. Wilt,F. H.1969. An artifact in the alkalinehydrolysisof
RNA labeled with guanosine-8-T. Anal. Biochem.
27:186-189.
47. Yogo,Y., andE. Wimmer. 1972.Polyadenylicacidatthe
3-terminus ofpoliovirusRNA. Proc. Nat. Acad.Sci.
U.S.A. 69:1877-1882. VOL.14,
![FIG. 1.waselectrophoresis. Determination of the size of marker 7.8S poly(A) and RSVpoly(A) by formamide polyacrylamide gel RSVpoly(A) was isolated from 30 to 40S [32P]RNA subunits by RNase T, digestion](https://thumb-us.123doks.com/thumbv2/123dok_us/1578183.110411/4.499.118.402.61.368/waselectrophoresis-determination-rsvpoly-formamide-polyacrylamide-isolated-subunits-digestion.webp)