Copyright e 1993, AmericanSocietyforMicrobiology
Gibbon Ape Leukemia Virus and the
Amphotropic Murine
Leukemia Virus
4070A
Exhibit
an
Unusual
Interference
Pattern
on
E36
Chinese
Hamster
Cells
MARTIN A. EGLITIS,* MARIBETH V. EIDEN,ANDCAROLYN A. WILSON Laboratory of Cell Biology, National Institute of Mental Health, Bethesda, Maryland 20892
Received 27 May1993/Accepted 21 June 1993
Thegibbonapeleukemia virus (GaLV), the amphotropic mouseleukemiavirus (A-MLV) 4070A,and the
xenotropic mouseleukemiavirus (X-MLV) exhibit widebutnotidentical species hostranges.However,most Chinese hamster cells resist infectionbyallthree viruses. We havenowdetermined that the Chinese hamster
cell lineE36 differs from other Chinesehamstercelllines inthatit issusceptibletoinfectionbywild-type GaLV, A-MLV, and X-MLV. Surprisingly, analysis of theinterference patternof GaLV andA-MLV in E36 cells indicated that GaLV and A-MLV interfere in a nonreciprocal fashion. E36 cells productively infected with
GaLVwereresistant tosuperinfection byboth GaLVandamphotropicallypackagedrecombinant retroviral
vectors. In contrast, E36 cells infected with A-MLVwereresistant to superinfection with an amphotropic vectorbut could still be infected byaGaLVvector.These resultsimplytheexistence ofareceptoronE36 cells that interacts with both GaLVand A-MLV.
While thegibbonapeleukemia virus(GaLV), the
ampho-tropicmouseleukemia virus (A-MLV), and the xenotropic
mouseleukemiavirus (X-MLV) possessbroadspecies host
range,theyareinseparate interference classes (26) andare
believedto use differentreceptors to infectcells. Thegene
for the human GaLV receptor has been cloned (22) and mapped to chromosome 2 (12). The gene for the A-MLV receptorhasnotyetbeencloned, butit isknowntomapto chromosome 8 in both humans(5)andmice(6).Thegenefor the X-MLV receptor has been mapped to mouse chromo-some 1 (13). None of these viruses hasbeen described as
capableofefficiently infecting Chinesehamster cells(8, 29). Wild-type GaLV cannot infect Chinese hamster ovary
(CHO-K1)cells. However,whenanMLVcore ispackaged in the GaLV envelope, CHO-Kl cells are efficiently in-fected, demonstrating that CHO-Kl cells do express a
functional GaLV receptor on their surface (29). Thus, the block to GaLV infection appears related to-events subse-quent to the virus-receptor interaction (29). In contrast, neitherCHO-KlnorDon Chinese hamstercellsareinfected
when the sameMLV coreand genome are packaged inan
A-MLV envelope. Therefore, the block to infection by wild-typeA-MLV inChinese hamster cells isatthe level of receptor. This block can be overcome by treatment with tunicamycin,aninhibitorof N-linkedglycosylation.
Presum-ably, thistreatmentaffectsglycosylatedmoieties that inter-ferewith theinteractionbetween the A-MLVenvelope and theCHO-KlA-MLVreceptor(21).
We have now found a Chinese hamster cell line, E36,
which is efficiently infected by both GaLV and A-MLV. Since differentmechanisms accountforCHO-Kl cell resis-tance toinfectionby GaLV and A-MLV, it is likely that both changesinthereceptorandrelief ofapostpenetration block
account for the altered infection pattern inE36 cells. The acquisition of susceptibility to infection by two different retroviruses suggests that a common factor could be in-volved inthischange. We have used recombinant retroviral
*Correspondingauthor.
vectorsdiffering onlyin theirenvelopetodeterminewhether thefactor(s) accountingfor alteredsusceptibilitytoinfection functions atthe level of viral entry.
MATERIALS AND METHODS
Cells and viruses. E36 cellsarederived from thelungofa
male Chinese hamster (7). CHO-Klcells, derived from an
adult Chinese hamster ovary (23),were obtained from the AmericanTypeCulture Collection(ATCC) (CCL 61). Rat-2 embryonic rat fibroblasts (27) and NIH 3T3 embryonic
mouse cells (10)wereobtained from the ATCC (CRL1764
and CRL 1658, respectively). TblLu batlung and MvlLu mink lung fibroblasts were also obtained from the ATCC (CCL88 and CCL64, respectively). HOS cellsare ahuman osteogenicsarcoma(15) (ATCCCRL1543).MDBK cellsare
derived from normal bovinekidney (14) (CCL 22). Cultured tail fibroblasts derived from the feral mouse Mus dunni
(MDTF) (1)wereprovided byJ.Hartley,National Institute ofAllergyand Infectious Diseases. All cellswerecultured in Dulbecco's modified Eagle's medium supplemented with 10% fetal bovine serum, except for CHO-Kl cells, which
werecultured inalphaminimumessential medium. GaLV of the SEATO subtype was collected from infected TblLu
cells. A-MLV of the 4070A subtype was collected from infected MvlLu cells. X-MLVof theAKR40-Xsubtype (4, 9) was collected from infected MvlLu cells. The recombi-nantretroviral vectorGlBgSvN (17) carryingthe bacterial lacZ andneomycinresistancegeneswasused in interference
assays. The vector was collected from either PG13/
GlBgSvNorPA317/GlBgSvNcells. PG13 isahybrid retro-viral packaging cell line, expressing Moloney ecotropic MLV core proteins and the GaLV envelope (19) (ATCC
CRL10686).PA317isahybridretroviralpackagingcell line expressingMoloney ecotropic MLVcore proteins and the envelope from the 4070A A-MLV(18) (ATCC CRL9078). Titer of thevector produced byboth packaging cellswas
approximately 105 G418-resistant CFU/ml, as assayed on
HeLa cells.
Wild-typevirus infections. Medium containing wild-type
5472
on November 9, 2019 by guest
http://jvi.asm.org/
virus was prepared by changing medium on the producer
cellswhenthey hadreached approximately 50% confluence. Virus-containing mediumwas collected 24to 48 h after the medium change, passed through a0.22-,um-pore-size filter,
and storedat-70°C untiluse. Targetcells were seededat2
x 104to3 x 104cellsperwell in 24-well dishes. Twenty-four
hoursafter seeding,cellswereinfected with virus in medium
containing3 ,ug of Polybreneperml.Twenty-four hours after
exposure to virus, the cells were fed with fresh medium.
After an additional 24 h of culture, the cellswere passaged
1:10. Sixdays aftersplitting, the mediumwascollected and
assayed for reverse transcriptase. Reverse transcriptase
assays wereperformed aspreviously described (29).
Interference analysis. For the interference assay, a series
ofvirally infected cell linesweregenerated by infecting with GaLV-SEATO, 4070A, orAKR 40-X as described above. CellstestedwereE36, HOS, and MDTFcells expressing the
cloned GaLV receptor (22). Productive infection with a
particular virus was confirmed by detection of significant
levels ofreversetranscriptase in the mediumatlevels100to 1,000 times abovebackground. Toassaythecells'
suscepti-bilitytosuperinfection witharecombinant hybrid retroviral vector, targetcellswereseeded in three individual wells ofa
12-wellcultureplateataninitialdensity of 104 cellsperwell.
The next day, the cells were exposed to the vector by replacing theirmediumwith medium harvested fromeither PG13/GlBgSvN or PA317/GlBgSvN producers.
Vector-containing medium was prepared as described above for wild-type virus. To facilitate infection with retroviral vec-tors,Polybreneatafinalconcentrationof 8 ag/mlwasadded to the medium. Target cells were exposed to retroviral vector overnight, and then vector-containing medium was
removedandreplacedwithfreshmedium. Afterafurther 24
hof culture,thecellswerefixed andstained for thepresence
of,B-galactosidase (3-gal) asdescribed
previously
(25). Construction of MDTF cells expressing the human GaLV receptorcDNA.ThepOJ9 plasmid containing the cDNA for thehumanGaLVreceptorwasobtained fromBrian O'Hara(Lederle Laboratories, PearlRiver, N.Y.). The 2.3-kb Hin-dIII to EcoRV restriction fragment from pOJ9 was
intro-duced into thepLNSXretroviral expressionplasmid (20) at theHindIIItoClaIsites (the Clalsitewasfilledinwith T4
DNApolymerase)to makethe pLNS-OJ9 plasmid. PA317 cellsweretransfected with 20 ,ugofpLNS-OJ9 plasmid (2). Transfected cellswere selected with 400 pgofactive G418 per ml. Supernatant medium from the stably transfected PA317 cells was used to infect MDTF cells. MDTF cells expressingthe LNS-OJ9vectorwere selected in 400 ,ug of activeG418perml.
RESULTS
Weusedthewild-type GaLV, A-MLV,orX-MLV viruses to confirm their species host range and simultaneously determine whether E36 cells are susceptible to infection (Table 1). Productive infection of different target cells
ex-posedtoeitherGaLV,A-MLV,orX-MLVwasassessedby
a reverse transcriptase assay of the cell supernatant
per-formed 8daysafter the cellswereexposedtofiltered virus. Thethreeretroviruseswereabletoinfecthuman,mink,bat, and rat cells but differed in their ability to infect bovine MDBK, murine MDTF, and NIH 3T3 cells. Bovine cells
weresusceptibletoinfectionbyGaLV and X-MLV butwere
resistanttoA-MLV. Themurinecell line MDTFwas
capa-bleofbeing infectedby bothmurine retroviruses (A-MLV and X-MLV)but nottheprimate retrovirusGaLV. Murine
TABLE 1. Susceptibility of cells to infection with GaLV, A-MLV,and X-MLV
Susceptibility'with Targetcell Ongin challenge virus
GaLV A-MLV X-MLV
HOS Human + + +
MvlLu Mink + + +
TblLu Bat + + +
Rat-2 Rat + + +
MDBK Bovine + - +
MDTF Musdunni - + +
NIH3T3 Musmusculus - +
CHO-Kl Chinese hamster - - +
E36 Chinesehamster + + +
aSusceptibilityto infection was determined by assay for reverse
tran-scriptasein the medium. For reverse transcriptase assays: +, >2x back-ground reverse transcriptase activity; -, <2x backback-ground reverse tran-scriptaseactivity.
NIH3T3cellswereonlysusceptible to infection by A-MLV. The CHO-Kl Chinese hamster cell line was resistant to infection by GaLV and A-MLV but was susceptible to infection by X-MLV. Surprisingly, a second Chinese ham-stercell line,E36,supportedproductive infectionby GaLV, A-MLV, and X-MLV retroviruses. Because of the novel infectionproperties ofE36cells comparedtoCHO-Kl cells, the identity of E36 cells was confirmed to be Chinese hamster by isozyme analysis (assay performed by the ATCC; datanotshown).
We then useda
3-gal
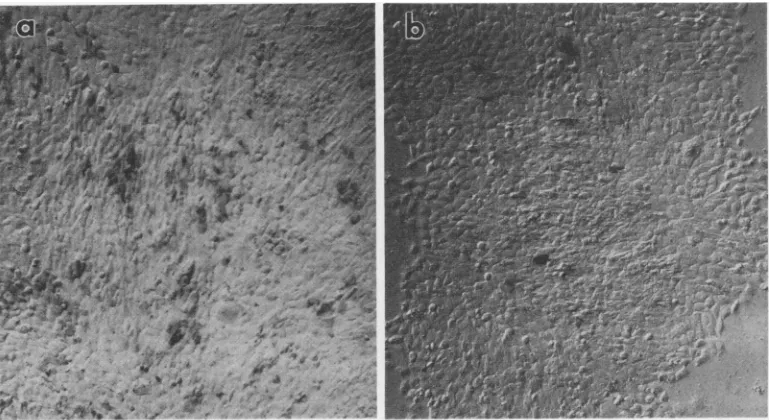
retroviralvectorpackagedineither PG13 cells (bearing the GaLV envelope) or PA317 cells (bearing a A-MLV envelope), to determine whether E36 cells productively infected with either GaLV or A-MLV wereresistant tosuperinfection.Both vectors infected 20 to 40% of E36 cells (Fig. la and b). E36 cells infected with GaLV-SEATOvirus exhibited resistance to superinfection with PG13/GlBgSvN (Fig. lc). Unexpectedly, GaLV-SEATO-infected E36cells alsoexhibitedareduction in their susceptibility to superinfection by amphotropically pack-aged PA317/GlBgSvN recombinant vector compared with that ofuninfectedE36 cells(Fig. ld). In contrast,E36 cells infectedwith the 4070A strain of A-MLV could be infected by the GaLV-enveloped PG13/GlBgSvN vector (Fig. 2a), whereassuperinfectionwith amphotropicPA317/GlBgSvN wasblocked(Fig. 2b). Thepatternof interference was then evaluated quantitatively by measuring the effective,3-gal
titer on uninfected E36 cells and E36 cells infected with various wild-type viruses (as described in Materials and Methods). The amphotropicvectorPA317/GlBgSvN had a titer of 2.2x 105bluecoloniespermlonE36cells,compared with a titerof only 1 blue colony per ml on E36 cells infected by GaLV-SEATO.This representsagreaterthan
1,000-fold
reduction in susceptibility toinfection(Table 2). Whilethe titer of the GaLVvectorPG13/GlBgSvNwas alsoreduced bymorethan
1,000-fold
in E36cells infectedwithSEATO, the titer of this vector remained essentially unchanged on E36 cells infected withwild-type 4070Aamphotropicvirus. GaLV-infected E36 cells remained susceptible to infection by X-MLVpseudotypedretroviralvectors(datanotshown). It ispossible that other retroviruses that infect E36 cells through a common entry pathway can also cause anonre-ciprocal
pattern
of infectioninterference when E36 cellsarechallenged with an amphotropic vector. Since
GaLV,
A-MLV, and X-MLV all enter cells by a
pH-independent
on November 9, 2019 by guest
http://jvi.asm.org/
. o,f'''J
~;;'z - ' {t 5 ;; $ _ t
9trt' A.%% ' . i
4-~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~J.
FIG. 1. Interferencepatternof PG13/GlBgSvN and PA317/GlBgSvN on E36 cells infected with GaLV-SEATO. Uninfected E36cells or E36 cellsproductively infected with wild-type GaLV-SEATO were tested for their ability to be transduced by retroviralvectorsbearing different viral envelopes and containing theEscherichia coli lacZ gene. All cells were stained foracquired ,B-gal activity as described in Materialsand Methods.(a)E36 cellstransduced with aPG13/GlBgSvN vector bearing a GaLV envelope. (b) E36 cellstransduced with a PA317/GlBgSvN vector bearing an amphotropic envelope. (c) E36 cells productively infected with GaLV-SEATO, transduced with PG13/GlBgSvNvector. (d) E36 cells productively infected with GaLV-SEATO, transduced withPA317/GlBgSvN vector.
mechanism (16, 30), we determined the susceptibility of E36 an over 1,000-fold reduction in p-gal titer when compared cellsinfected with X-MLV to superinfection with GaLV- or with uninfected HOS. GaLV-infected HOS cells exhibited A-MLV-hybrid vectors. As indicated in Table 2, X-MLV did essentiallyno change in effective a-gal titer when exposed to notinterfere with either GaLV or A-MLV superinfection. amphotropicPA317/GlBgSvNvector (data not shown).
In contrast to the results obtained with E36 cells, no MDTF cells are resistantto GaLV infection because of the cross-interference between GaLV and A-MLV was seen in absence of a functional receptor (19). Expression of the HOS cells. HOS cells exhibited a conventional pattern of human GaLVreceptor in MDTF cells confers susceptibility
interferenceinthat GaLV-infected HOS cells were resistant to infectionby bothPG13/GlBgSvN(Table 3) andwild-type
onlytosuperinfection by GaLV-packaged vector, exhibiting GaLV (data not shown). In contrast to E36 cells
on November 9, 2019 by guest
http://jvi.asm.org/
[image:3.612.101.507.70.552.2]
4-FIG. 2. Interference pattern of PG13/GlBgSvN and PA317/GlBgSvN on E36 cells infected with wild-type A-MLV. (a) E36 cells
productively infected withA-MLV, transduced with PG13/GlBgSvN. (b) E36 cells productively infected with A-MLV, transduced with PA317/GlBgSvN.
tively infected with GaLV,MDTF cellsinfected withGaLV remain susceptible to the amphotropic vector
PA317/
GlBgSvN(Table
3).
These results suggestthat expressionof thehuman GaLVreceptorinMDTFcellsisnot sufficient to induce nonreciprocal interference between GaLV and A-MLVanalogous tothepattern
observed inE36 cells.DISCUSSION
We have identified a Chinese hamster-derived cell line, E36, that issusceptibletoinfection by GaLV, A-MLV, and X-MLV.Although Chinese hamster cells, suchasCHO-Kl, express a functional receptor for GaLV on their surface, they exhibitapostbinding blocktoinfection withGaLV(29). Wedemonstrate that E36 cells are susceptible to infectionby bothwild-type GaLV andGaLV-enveloped MLVvectors. Since GaLV infection of other Chinese hamster cells is normally blocked by cellular factors operating at a point postpenetration, E36 cells must lack these restrictive fac-tors.Such factorsmayaffect the
ability
totransport a GaLV coreorgenome tothenucleus, improve the abilityto reverse transcribe the GaLV genome, improve the ability of the GaLV genome tointegrate intothechromosomes,oroperate through other as-yet-undefined mechanisms. The specificTABLE 2. Summary of interference analysis of productively infectedE36cells
Transductionefficiency(104) of E36 cells
Vector infectedwithb:
envelopea
Novirus GaLVc A-MLVd X-MLVe
GaLV 1.4 0.0006 0.56 1.0
Amphotropic 22.0 0.0001 0.01 5.2
aGaLV=PG13/GlBgSvN; Amphotropic =PA317/GlBgSvN.
bTransduction efficiencyexpressed in number of
(3-gal-positive
coloniesperml.
IWild-typeGaLV-SEATO.
dWild-type 4070Aamphotropic MLV.
e
Wild-type
AKR 40-Xxenotropic MLV.change required to relieve the postpenetration block to GaLV infection in other Chinese hamster cells such as CHO-Klremains to be elucidated.
Incontrast to
GaLV,
A-MLVinfectionof CHO-Kl cells is restrictedby the absence of afunctionalreceptor. CHO-Kl cells exhibit veTy low or no binding of A-MLV (11) and becomesusceptible to infection by A-MLV only after treat-mentwith tunicamycin (21). E36 cells express a functional receptor for A-MLV on theirsurface, lending them suscep-tibility toinfection bybothwild-type A-MLV and ampho-tropicvectors.Toevaluate whether a common receptor serves for both A-MLVand GaLVinE36cells,we performedan interfer-enceassay. The type of assay weused has a major advantage over conventional assays in that it employs viral vectors insteadofreplication-competent virusfor superinfection of infected cells. Staining for expression of a histochemical markerreplaces syncytium formation(26)ortheproduction ofviralparticles (3)astheendpoint inthis assay. Therefore thisassay can be usedwithtargetcellsthat do notefficiently form foci or syncytia orwhich do not support productive infection of the test retrovirus. Furthermore, by using re-combinantvectorsdiffering onlyintheirenvelope, theassay focuses on interference at the level of receptor-envelope interaction.
TABLE 3. Interference analysis of GaLV and A-MLVonMDTF cellsexpressing the human GaLV receptor
Transductionefficiency (104)ofMDTF/huGaLVW
Vector cellsinfectedwithe:
envelopea
No virus GaLVc A-MLV
GaLV 0.5 0.0007 0.11
Amphotropic 1.0 1.0 <0.0001
a GaLV= PG13/GlBgSvN; Amphotropic= PA317/GlBgSvN.
bTransductionefficiency expressed in number of
(3-gal-positive
coloniesperml.
cWild-type GaLV-SEATO.
on November 9, 2019 by guest
http://jvi.asm.org/
[image:4.567.108.493.65.275.2]We found that GaLV and A-MLV exhibited a nonrecip-rocal interference pattern in E36 cells. E36 cells produc-tively infected with A-MLV were exclusively resistant to infection with amphotropically packaged recombinant vec-tors. On theother hand, E36 cells productively infected with GaLV were resistant to infection by recombinant vectors packaged with either GaLV or amphotropic envelopes but remained susceptible to infection with vectors packaged in xenotropic envelope. Similarly, E36 cells infected with X-MLV remained susceptible to infection by either GaLV or A-MLV, demonstrating that in E36 cells only GaLV exhib-itedthe ability to block the entry of a virus from a different receptor class. Furthermore, the nonreciprocal interference between GaLV and A-MLV is unique to E36 cells and is not found in other cells. HOS cells do not exhibit any interfer-ence between these viruses. Our observations of MDTF cells expressing the human GaLV receptor further support that theinterference we observe is an inherent quality of the receptor-envelope interaction. Such cells acquire suscepti-bility to GaLV infection and exhibit the same interference pattern seen in human cells. Therefore, the interference patternweobserve in HOS cells is attributable to the human GaLV receptor and does not involve other cellular factors specific to these cells.
Our results suggest that E36 cells infected with GaLV interfere with A-MLV infection at the level of virus entry. We postulate two mechanisms that may account for this interference. The first is that E36 cells express a cell surface receptor capable of interacting with both the A-MLV and GaLV envelopes. Such a mechanism has been proposed for the 1504A A-MLV and the 1OAl MLV, which exhibit nonreciprocal interference properties in NIH 3T3 cells (24). In NIH 3T3 cells productively infected with 10A, superin-fection with both A-MLV and 1OAl isinhibited; however, NIH 3T3 cells infected with A-MLV remain susceptible to infection with 1OAl. Rein and Schultz (24) account for this pattern by postulating that two receptors exist, one that exclusively binds 1OAl and a second that binds both 1OAl and A-MLV. Byanalogy, E36cells would also express two receptors. GaLV would interact with both receptors and thereby block both GaLV and A-MLV superinfection, while A-MLVwould interact with and inactivate only one of the receptors, blocking further A-MLV infection but permitting GaLV infection.
Asecond, alternative mechanism is that a factor associ-atedwithGaLV replication in E36 cells blocks the A-MLV receptor. This
mechanism
would be analogous to that re-cently described in cells infected with the human immuno-deficiency virus (HIV) (28). Willey et al. report that the HIV-encodedintegral membrane protein Vpu contributes to the inhibition of HIV superinfection in HIV-infected cells by rapidly degrading the HIV receptor, CD4, when it is com-plexed to the HIV envelope within the endoplasmic reticu-lum. Thus, subsequent infection by challenge virus is blockedby anonenvelope-mediated depletion of functional receptor.Should A-MLV and GaLV share a cellular receptor on E36 cells, the E36 receptor for A-MLV might bear some level of homology to the authentic GaLV receptor. The cDNAforthehuman GaLV receptor has been cloned (22). If similarities exist between theauthentic GaLV receptor and theputative dually active A-MLV/GaLV receptor expressed in E36 Chinese hamster lung cells, it may be possible to isolate the E36 cell cDNA that encodes the A-MLV receptor by screeningforgenes similar to the authentic GaLV recep-torcDNA.
ACKNOWLEDGMENTS
We aregratefultoMaryCatherine Walshe for excellenttechnical assistance. C. KozakprovidedE36 cells and MvlLu cells produc-tivelyinfected with theAKR40-X virus. J.Hartleyprovided MvlLu cellsproductivelyinfectedwith the 4070Avirus.B.O'Hara kindly provided the human GaLV receptor expression plasmid,pOJ9.
REFERENCES
1. Chattopadhyay, S. K., M. R. Lander, S.Gupta, E. Rands,and D. R. Lowy. 1981. Origin of mink cytopathic focus-forming (MCF) viruses:comparisonwithecotropic and xenotropic mu-rine leukemia virusgenomes.Virology 113:465-483.
2. Chen, C., and H. Okayama. 1987. High-efficiency transforma-tion ofmammalian cells by plasmid DNA. Mol. Cell. Biol. 7:2745-2752.
3. Chesebro,B., and K.Wehrly.1985. Different murine celllines manifestuniquepatterns of superinfectionby murineleukemia viruses.Virology141:119-129.
4. Cloyd, M. W., J. W. Hartley, and W. P. Rowe. 1979. Cell surface antigens associatedwith recombinant mink cell focus inducingmurine leukemiaviruses. J. Exp. Med. 149:702-712. 5. Garcia, J. V., C. Jones, and A. D. Miller.1991.Localization of
the amphotropic murine leukemiavirus receptor to the pericen-tromeric regionof human chromosome 8. J. Virol. 65:6316-6319.
6. Gazdar, A. F., H.Oie,P.Lalley,W. W.Moss, and J. D. Minna. 1977.Identificationof mousechromosomesrequired for murine leukemia virus replication.Cell 11:949-956.
7. Gillin, F. D., D. J. Roufa, A. L. Beaudet, and C. T. Caskey.1972. 8-Azaguanine resistance in mammalian cells. I. Hypoxanthine-guaninephosphoribosyltransferase. Genetics 72:239-252. 8. Hartley, J. W., and W. P. Rowe. 1976. Naturally occurring
murine leukemiaviruses in wild mice:characterizationof a new "amphotropic" class.J.Virol. 19:19-25.
9. Hartley, J. W., N. K. Wolford, L. J. Old, and W. P. Rowe. 1977. A new class of murine leukemia virus associated with the development of spontaneous lymphomas. Proc. Natl. Acad. Sci.USA74:789-792.
10. Jainchill,J. L., S. A. Aranson, and G. J. Todaro. 1969. Murine sarcoma and leukemia viruses: assay using clonal lines of contact-inhibitedmousecells.J.Virol. 4:549-553.
11. Kadan, M. J., S. Sturm, W. F. Anderson, and M. A. Eglitis. 1992. Detection of receptor-specific murine leukemia virus binding to cells by immunofluorescence analysis. J. Virol. 66:2281-2287.
12. Kaelbing, M., R. Eddy, T. B. Shows, N. G. Copeland, D. J. Gilbert, N. A. Jenldns, H. P. Klinger, and B. O'Hara. 1991. Localizationof the humangeneallowing infection by gibbonape leukemia virusto human chromosome region2q11-q14and to the homologous region on mouse chromosome 2. J. Virol. 65:1743-1747.
13. Kozak, C.1985.Susceptibility of wildmousecellstoexogenous infectionwithxenotropic leukemiaviruses: control by asingle dominantlocusonchromosome 1. J.Virol. 55:690-695. 14. Madin, S. H., and N. B. Darby. 1958. Established kidney cell
lines ofnormal adult bovine and ovine origin. Proc.Soc. Exp. Biol. Med. 98:574-576.
15. McAllister, R. M., M. B. Gardner, A. E. Greene, C. Bradt, W. W.Nichols, and B. H.Landing.1971.Cultivation in vitro of cells derived from a human osteosarcoma. Cancer 27:397-402. 16. McClure, M. O., M. A. Sommerfelt, M.Marsh, and R. A. Weiss. 1990. ThepH independence of mammalianretrovirus infection. J. Gen. Virol.71:767-773.
17. McLachlin, J. R., N. Mittereder, M. B. Daucher, and M. A. Eglitis. 1993. Factors affecting retroviral vector function and structuralintegrity. Virology195:1-5.
18. Miller, A. D., and C. Buttimore. 1986. Redesign ofretrovirus packaging linestoavoid recombination leadingtohelpervirus production.Mol. Cell. Biol.6:2895-2902.
19. Miller, A. D., J. V. Garcia, N. von Suhr, C. Lynch, C. A. Wilson, and M. V. Eiden. 1991.Construction and properties of retrovi-rus packaging cells based on gibbon ape leukemia virus. J. Virol.65:2220-2224.
on November 9, 2019 by guest
http://jvi.asm.org/
20. Miller, A. D., and G. J. Rosman. 1989. Improved retroviral
vectorsforgenetransfer andexpression.BioTechniques
7:982-989.
21. Miller, D.G., and A. D. Miller.1992.Tunicamycintreatmentof CHO cellsabrogates multipleblockstoretrovirusinfection,one
ofwhichis duetoasecreted inhibitor.J. Virol. 66:78-84.
22. O'Hara, B. S., S. V. Johann, H. P. Klinger, D. G. Blair, H. Rubinson, K. J. Dunne, P. Sass, S. M. Vitek, and T. Robins. 1990.Characterizationofahumangeneconferring sensitivityto infection by gibbon ape leukemia virus. Cell Growth Differ. 1:119-127.
23. Puck, T.T.,S.J. Cieriura,and A. Robinson. 1958. Genetics of somatic mammalian cells. III. Long-termcultivationofeuploid cells from human and animal subjects. J. Exp.Med. 108:945-957.
24. Rein, A., and A. Schultz. 1984. Different recombinant murine leukemia viruses usedifferent cell surfacereceptors. Virology 136:144-152.
25. Sanes, J. R., L. Rubenstein,and J.-F.Nicholas. 1986.Useofa
recombinantretrovirustostudypost-implantation cell lineage in
mouseembryos. EMBO J. 5:3133-3142.
26. Sommerfelt, M. A., and R. A. Weiss. 1990. Receptor
interfer-encegroupsof20retroviruses platingonhumancells. Virology
176:58-69.
27. Topp, W. C. 1981. Normalrat cell lines deficient in nuclear thymidinekinase.Virology 113:408-411.
28. Willey,R.L., F. Maldarelli, M. A. Martin, and K. Strebel. 1992. Human immunodeficiency virus type I Vpu protein induces rapid degradation ofCD4.J.Virol.66:7193-7200.
29. Wilson,C.A., and M. V. Eiden.1991. Viral andcellular factors governing hamster cell infection by murine and gibbon ape
leukemiaviruses. J. Virol.65:5975-5982.
30. Wilson, C. A., J. W. Marsh, and M. V. Eiden. 1992. The requirements forviralentrydiffer from thoseforvirally induced syncytiumformation in NIH3T3/DTrascellsexposedto
Molo-neymurineleukemia virus.J.Virol. 66:7262-7269.