organic papers
o2746
Ma and Xian C18H14N6S2 doi:10.1107/S1600536805023779 Acta Cryst.(2005). E61, o2746–o2747 Acta Crystallographica Section E
Structure Reports
Online
ISSN 1600-5368
2,3-Bis[(pyrimidin-2-ylsulfanyl)methyl]quinoxaline
Zi-Chuan Ma* and Hong-Shi Xian
College of Chemistry, Hebei Normal University, Shijiazhuang 050016, People’s Republic of China
Correspondence e-mail: mazichuan@eyou.com
Key indicators
Single-crystal X-ray study T= 293 K
Mean(C–C) = 0.001 A˚ Rfactor = 0.035 wRfactor = 0.096
Data-to-parameter ratio = 17.6
For details of how these key indicators were automatically derived from the article, see http://journals.iucr.org/e.
#2005 International Union of Crystallography Printed in Great Britain – all rights reserved
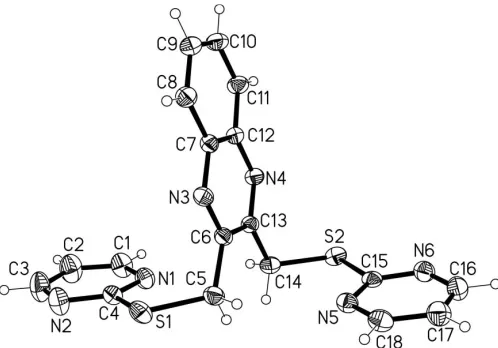
In the title compound, C18H14N6S2, the two terminal
pyrimidine groups attached to the central quinoxaline system adopt an anti conformation, with the thiopyrimidine groups on opposite sides of the quinoxaline plane. The dihedral angles formed by each of the two terminal pyrimidine and the central quinoxaline planes are 81.1 (3) and 82.0 (3), while the two
pyrimidine planes are almost parallel to each other, forming a dihedral angle of 6.0 (3).
Comment
The type of coordination architecture in complex compounds is mainly determined by a combination of two factors, namely the coordination preference of the metal ions and the nature of the ligands (Sun et al., 2002). The purposeful design of specific ligands capable of controlling the assembly of supra-molecular architectures has recently evolved as a popular and rapidly growing discipline (Zaworotko, 2000). In particular, multithioether ligands have been shown to exhibit high potential for structure control in inorganic chemistry, and many unusual crystal structures of complexes with multithio-ether ligands have been reported (Hartshorn & Steel, 1998). In this paper, we describe the crystal structure of a dithioether ligand, namely 2,3-bis[(pyrimidin-2-ylsulfanyl)methyl]quin-oxaline, (I), which has already been used in the construction of silver(I) complexes (Wanget al., 2001).
The molecular structure of (I) is shown in Fig. 1. Two terminal pyrimidine groups lie on the opposite sides of the planar quinoxaline system and their planes are almost ortho-gonal to the quinoxaline plane; the dihedral angles formed by the N3/N4/C6–C13 plane with the N1/N2/C1–C4 and N5/N6/ C15–C18 planes are 81.1 (3) and 82.0 (3), respectively. The
two pyrimidine planes are almost parallel to each other, the dihedral angle being 6.0 (3).
Experimental
2,3-Bis[(pyrimidin-2-ylsulfanyl)methyl]quinoxaline, (I), was prepared by a procedure similar to that used for the synthesis of
(phenylsulfanyl)butane (Hartley et al., 1979). A simple synthesis procedure has also been described by Wanget al.(2001), even though these authors did not isolate the free ligand. 2-Mercaptopyrimidine (1.12 g, 10 mmol) was added to a stirred solution of KOH (0.56 g, 10 mmol) in ethanol (20 ml). The mixture was heated to reflux, then an ethanol solution of 2,3-bis(bromomethyl)quinoxaline (1.57 g, 5 mmol) was added dropwise and the mixture was further refluxed for about 6 h. After adding more water (30 ml), the mixture was left to stand overnight. The precipitate was filtered off and washed with water, giving a fine white powder in 85% yield.1H NMR (CDCl3):
5.00 (s, 4H, SCH2), 6.95–8.37 (m, 10H, Ar-H); elemental analysis
found: C 57.52, H 3.45, N 22.01%, calculated for C18H14N6S2: C 57.12,
H 3.73, N 22.21%. Colorless single crystals of (I) were obtained by recrystallization from EtOH.
Crystal data
C18H14N6S2 Mr= 378.47
Orthorhombic,Pccn a= 22.485 (5) A˚ b= 12.361 (3) A˚ c= 12.560 (3) A˚ V= 3490.9 (13) A˚3 Z= 8
Dx= 1.440 Mg m 3
MoKradiation Cell parameters from 4783
reflections = 2.5–25.6
= 0.32 mm1 T= 293 (2) K Block, colorless 0.220.200.20 mm
Data collection
Bruker SMART 1000 CCD area-detector diffractometer ’and!scans
Absorption correction: multi-scan (SADABS; Bruker, 1998) Tmin= 0.933,Tmax= 0.939 21857 measured reflections
4126 independent reflections 3069 reflections withI> 2(I) Rint= 0.027
max= 27.9
h=29!29 k=15!16 l=16!14
Refinement
Refinement onF2 R[F2> 2(F2)] = 0.035 wR(F2) = 0.096 S= 1.04 4126 reflections 235 parameters
H-atom parameters constrained
w= 1/[2(F
o2) + (0.0485P)2 + 0.5889P]
whereP= (Fo2+ 2Fc2)/3 (/)max= 0.004
max= 0.24 e A˚3
min=0.27 e A˚3
Table 1
Selected geometric parameters (A˚ ,).
S1—C4 1.7465 (7) S1—C5 1.8052 (7)
S2—C14 1.8060 (7) S2—C15 1.7500 (6)
C4—S1—C5 101.88 (3) C15—S2—C14 102.69 (3)
All H atoms were included in calculated positions and refined as riding atoms [C—H = 0.93 or 0.97 A˚ andUiso(H) = 1.2Ueqof the
carrier atom].
Data collection:SMART(Bruker, 1998); cell refinement:SAINT
(Bruker, 1998); data reduction: SAINT and SHELXTL (Bruker, 1998); program(s) used to solve structure: SHELXS97 (Sheldrick, 1997); program(s) used to refine structure:SHELXL97(Sheldrick, 1997); molecular graphics: SHELXTL; software used to prepare material for publication:SHELXTL.
This work was supported by the NNSF of China (No. 20477009).
References
Bruker (1998).SMART(Version 5.051), SAINT(Version 5.01),SADABS (Version 2.03) andSHELXTL(Version 6.1). Bruker AXS Inc., Madison, Wisconsin, USA.
Hartley, F. R., Murray, S. G., Levason, W., Soutter, H. E. & McAuliffe, C. A. (1979).Inorg. Chim. Acta,35, 265–277.
Hartshorn, C. M. & Steel, P. T. (1998).J. Chem. Soc. Dalton Trans.pp. 3935– 3940.
Sheldrick, G. M. (1997). SHELXL97 and SHELXS97. University of Go¨ttingen, Germany.
Sun, W. Y., Kusukawa, T. & Fujita, M. (2002).J. Am. Chem. Soc.124, 11570– 11571.
Wang, R. H., Hong, M. C., Su, W. P., Liang, Y. C., Cao, R., Zhao, Y. J. & Weng, J. B. (2001).Polyhedron,20, 3165–3170.
[image:2.610.315.564.70.244.2]Zaworotko, M. J. (2000).Angew. Chem. Int. Ed.39, 3052–3054. Figure 1
supporting information
sup-1 Acta Cryst. (2005). E61, o2746–o2747
supporting information
Acta Cryst. (2005). E61, o2746–o2747 [https://doi.org/10.1107/S1600536805023779]
2,3-Bis[(pyrimidin-2-ylsulfanyl)methyl]quinoxaline
Zi-Chuan Ma and Hong-Shi Xian
2,3-bis[(pyrimidin-2-ylsulfanyl)methyl]quinoxaline
Crystal data
C18H14N6S2 Mr = 378.47 Orthorhombic, Pccn
Hall symbol: -P 2ab 2ac
a = 22.485 (5) Å
b = 12.361 (3) Å
c = 12.560 (3) Å
V = 3490.9 (13) Å3 Z = 8
F(000) = 1568
Dx = 1.440 Mg m−3
Mo Kα radiation, λ = 0.71073 Å Cell parameters from 4783 reflections
θ = 2.5–25.6°
µ = 0.32 mm−1 T = 293 K Block, colorless 0.22 × 0.20 × 0.20 mm
Data collection
Bruker SMART 1000 CCD area-detector diffractometer
Radiation source: fine-focus sealed tube Graphite monochromator
φ and ω scans
Absorption correction: multi-scan (SADABS; Bruker, 1998)
Tmin = 0.933, Tmax = 0.939
21857 measured reflections 4126 independent reflections 3069 reflections with I > 2σ(I)
Rint = 0.027
θmax = 27.9°, θmin = 1.9° h = −29→29
k = −15→16
l = −16→14
Refinement
Refinement on F2
Least-squares matrix: full
R[F2 > 2σ(F2)] = 0.035 wR(F2) = 0.096 S = 1.04 4126 reflections 235 parameters 0 restraints
Primary atom site location: structure-invariant direct methods
Secondary atom site location: difference Fourier map
Hydrogen site location: inferred from neighbouring sites
H-atom parameters constrained
w = 1/[σ2(F
o2) + (0.0485P)2 + 0.5889P]
where P = (Fo2 + 2Fc2)/3
(Δ/σ)max = 0.004
Δρmax = 0.24 e Å−3
Δρmin = −0.27 e Å−3
Special details
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic)
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2,
conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used
only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2
are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger.
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
x y z Uiso*/Ueq
S1 0.612609 (8) −0.080394 (13) −0.256292 (12) 0.04873 (4)
S2 0.565136 (7) 0.096057 (14) 0.191282 (11) 0.04578 (4)
N1 0.63995 (2) −0.18181 (4) −0.07659 (4) 0.04599 (14)
N2 0.65941 (3) −0.27153 (5) −0.24066 (5) 0.06522 (19)
N3 0.69335 (2) 0.08019 (4) −0.13370 (4) 0.03752 (12)
N4 0.69035 (2) 0.06352 (4) 0.08821 (4) 0.03748 (12)
N5 0.52223 (2) 0.18037 (4) 0.01111 (4) 0.04305 (13)
N6 0.51128 (2) 0.28060 (4) 0.17222 (4) 0.05007 (15)
C1 0.66457 (3) −0.26523 (6) −0.02506 (6) 0.0585 (2)
H1A 0.6664 −0.2634 0.0489 0.070*
C2 0.68713 (4) −0.35282 (6) −0.07703 (7) 0.0757 (3)
H2A 0.7044 −0.4101 −0.0403 0.091*
C3 0.68315 (4) −0.35277 (6) −0.18593 (7) 0.0765 (3)
H3A 0.6977 −0.4122 −0.2231 0.092*
C4 0.63976 (3) −0.18897 (5) −0.18177 (5) 0.04302 (16)
C5 0.59366 (3) 0.01581 (5) −0.15362 (5) 0.04304 (16)
H5A 0.5777 0.0805 −0.1868 0.052*
H5B 0.5626 −0.0149 −0.1093 0.052*
C6 0.64551 (2) 0.04731 (4) −0.08376 (4) 0.03484 (14)
C7 0.74216 (2) 0.10546 (4) −0.07436 (4) 0.03490 (14)
C8 0.79506 (3) 0.13769 (5) −0.12524 (5) 0.04394 (16)
H8A 0.7964 0.1443 −0.1990 0.053*
C9 0.84414 (3) 0.15913 (5) −0.06589 (5) 0.04777 (17)
H9A 0.8791 0.1804 −0.0995 0.057*
C10 0.84283 (3) 0.14964 (6) 0.04482 (5) 0.05076 (18)
H10A 0.8770 0.1647 0.0839 0.061*
C11 0.79237 (3) 0.11874 (5) 0.09640 (5) 0.04649 (17)
H11A 0.7920 0.1125 0.1702 0.056*
C12 0.74077 (2) 0.09632 (4) 0.03687 (4) 0.03486 (14)
C13 0.64375 (2) 0.04045 (4) 0.02997 (4) 0.03434 (14)
C14 0.59015 (3) 0.00126 (5) 0.09145 (5) 0.04161 (15)
H14A 0.5999 −0.0667 0.1258 0.050*
H14B 0.5579 −0.0123 0.0419 0.050*
C15 0.52895 (2) 0.19489 (5) 0.11541 (5) 0.03884 (14)
C16 0.48498 (3) 0.35857 (6) 0.11685 (6) 0.0576 (2)
H16A 0.4716 0.4196 0.1529 0.069*
C17 0.47664 (3) 0.35317 (6) 0.00876 (6) 0.0572 (2)
H17A 0.4587 0.4093 −0.0287 0.069*
C18 0.49594 (3) 0.26131 (6) −0.04127 (5) 0.05132 (18)
supporting information
sup-3 Acta Cryst. (2005). E61, o2746–o2747
Atomic displacement parameters (Å2)
U11 U22 U33 U12 U13 U23
S1 0.06156 (9) 0.04730 (8) 0.03733 (7) −0.00420 (7) −0.01123 (7) −0.00311 (6)
S2 0.03942 (7) 0.06442 (9) 0.03351 (6) −0.00177 (7) 0.00118 (6) 0.00452 (6)
N1 0.0539 (3) 0.0407 (2) 0.0434 (3) −0.0004 (2) 0.0000 (2) 0.0003 (2)
N2 0.0839 (4) 0.0557 (3) 0.0561 (3) 0.0099 (3) 0.0006 (3) −0.0122 (3)
N3 0.0402 (2) 0.0390 (2) 0.0333 (2) −0.0026 (2) −0.00357 (19) −0.00025 (19)
N4 0.0329 (2) 0.0446 (2) 0.0350 (2) −0.00230 (19) −0.00072 (18) 0.00038 (19)
N5 0.0452 (2) 0.0488 (3) 0.0351 (2) −0.0047 (2) −0.0006 (2) −0.0013 (2)
N6 0.0484 (3) 0.0568 (3) 0.0450 (3) −0.0056 (2) 0.0068 (2) −0.0089 (2)
C1 0.0768 (4) 0.0490 (3) 0.0498 (3) 0.0045 (3) −0.0034 (3) 0.0057 (3)
C2 0.0982 (5) 0.0532 (4) 0.0758 (5) 0.0238 (4) −0.0098 (4) 0.0031 (4)
C3 0.0995 (6) 0.0542 (4) 0.0759 (5) 0.0230 (4) 0.0012 (5) −0.0129 (4)
C4 0.0428 (3) 0.0403 (3) 0.0459 (3) −0.0040 (2) −0.0011 (3) −0.0051 (3)
C5 0.0415 (3) 0.0428 (3) 0.0448 (3) −0.0013 (2) −0.0097 (2) −0.0029 (3)
C6 0.0358 (3) 0.0307 (2) 0.0380 (3) −0.0003 (2) −0.0047 (2) −0.0013 (2)
C7 0.0352 (3) 0.0320 (2) 0.0375 (3) −0.0012 (2) −0.0007 (2) −0.0009 (2)
C8 0.0432 (3) 0.0469 (3) 0.0417 (3) −0.0050 (3) 0.0060 (3) 0.0000 (3)
C9 0.0375 (3) 0.0509 (3) 0.0548 (3) −0.0076 (3) 0.0074 (3) −0.0027 (3)
C10 0.0332 (3) 0.0631 (4) 0.0560 (4) −0.0065 (3) −0.0048 (3) −0.0072 (3)
C11 0.0365 (3) 0.0628 (3) 0.0401 (3) −0.0030 (3) −0.0049 (2) −0.0049 (3)
C12 0.0309 (2) 0.0368 (2) 0.0368 (3) 0.0001 (2) −0.0010 (2) −0.0021 (2)
C13 0.0323 (2) 0.0319 (2) 0.0388 (3) 0.0004 (2) −0.0030 (2) 0.0007 (2)
C14 0.0362 (3) 0.0440 (3) 0.0446 (3) −0.0064 (2) −0.0011 (2) 0.0053 (2)
C15 0.0307 (2) 0.0478 (3) 0.0380 (3) −0.0101 (2) 0.0043 (2) −0.0021 (2)
C16 0.0569 (4) 0.0491 (3) 0.0669 (4) −0.0003 (3) 0.0066 (3) −0.0097 (3)
C17 0.0613 (4) 0.0466 (3) 0.0637 (4) −0.0013 (3) −0.0046 (3) 0.0074 (3)
C18 0.0557 (3) 0.0554 (3) 0.0429 (3) −0.0077 (3) −0.0056 (3) 0.0039 (3)
Geometric parameters (Å, º)
S1—C4 1.7465 (7) C5—H5A 0.9700
S1—C5 1.8052 (7) C5—H5B 0.9700
S2—C14 1.8060 (7) C6—C13 1.4316 (8)
S2—C15 1.7500 (6) C7—C12 1.4019 (8)
N1—C4 1.3240 (8) C7—C8 1.4078 (8)
N1—C1 1.3375 (9) C8—C9 1.3578 (9)
N2—C3 1.3289 (10) C8—H8A 0.9300
N2—C4 1.3356 (9) C9—C10 1.3958 (10)
N3—C6 1.3099 (7) C9—H9A 0.9300
N3—C7 1.3630 (7) C10—C11 1.3611 (9)
N4—C13 1.3093 (7) C10—H10A 0.9300
N4—C12 1.3659 (7) C11—C12 1.4078 (8)
N5—C15 1.3308 (8) C11—H11A 0.9300
N5—C18 1.3355 (9) C13—C14 1.5110 (8)
N6—C16 1.3275 (9) C14—H14A 0.9700
C1—C2 1.3622 (11) C16—C17 1.3721 (11)
C1—H1A 0.9300 C16—H16A 0.9300
C2—C3 1.3707 (13) C17—C18 1.3683 (10)
C2—H2A 0.9300 C17—H17A 0.9300
C3—H3A 0.9300 C18—H18A 0.9300
C5—C6 1.5102 (8)
C4—S1—C5 101.88 (3) C7—C8—H8A 120.2
C15—S2—C14 102.69 (3) C8—C9—C10 120.89 (6)
C4—N1—C1 115.63 (6) C8—C9—H9A 119.6
C3—N2—C4 115.07 (7) C10—C9—H9A 119.6
C6—N3—C7 118.07 (5) C11—C10—C9 121.02 (6)
C13—N4—C12 117.73 (5) C11—C10—H10A 119.5
C15—N5—C18 115.73 (5) C9—C10—H10A 119.5
C16—N6—C15 115.34 (6) C10—C11—C12 119.30 (6)
N1—C1—C2 122.35 (7) C10—C11—H11A 120.3
N1—C1—H1A 118.8 C12—C11—H11A 120.3
C2—C1—H1A 118.8 N4—C12—C7 120.85 (5)
C1—C2—C3 116.96 (8) N4—C12—C11 119.46 (5)
C1—C2—H2A 121.5 C7—C12—C11 119.67 (5)
C3—C2—H2A 121.5 N4—C13—C6 121.50 (5)
N2—C3—C2 122.88 (7) N4—C13—C14 115.00 (5)
N2—C3—H3A 118.6 C6—C13—C14 123.43 (5)
C2—C3—H3A 118.6 C13—C14—S2 113.28 (4)
N1—C4—N2 127.06 (6) C13—C14—H14A 108.9
N1—C4—S1 118.98 (5) S2—C14—H14A 108.9
N2—C4—S1 113.95 (5) C13—C14—H14B 108.9
C6—C5—S1 113.74 (4) S2—C14—H14B 108.9
C6—C5—H5A 108.8 H14A—C14—H14B 107.7
S1—C5—H5A 108.8 N5—C15—N6 126.74 (5)
C6—C5—H5B 108.8 N5—C15—S2 119.65 (4)
S1—C5—H5B 108.8 N6—C15—S2 113.61 (5)
H5A—C5—H5B 107.7 N6—C16—C17 122.94 (7)
N3—C6—C13 121.25 (5) N6—C16—H16A 118.5
N3—C6—C5 115.84 (5) C17—C16—H16A 118.5
C13—C6—C5 122.90 (5) C18—C17—C16 116.84 (7)
N3—C7—C12 120.57 (5) C18—C17—H17A 121.6
N3—C7—C8 119.80 (5) C16—C17—H17A 121.6
C12—C7—C8 119.60 (5) N5—C18—C17 122.40 (6)
C9—C8—C7 119.51 (6) N5—C18—H18A 118.8
C9—C8—H8A 120.2 C17—C18—H18A 118.8
C4—N1—C1—C2 −1.05 (10) C8—C7—C12—N4 −178.92 (5)
N1—C1—C2—C3 −0.56 (12) N3—C7—C12—C11 177.52 (5)
C4—N2—C3—C2 0.28 (12) C8—C7—C12—C11 −0.43 (8)
C1—C2—C3—N2 0.97 (14) C10—C11—C12—N4 178.90 (6)
C1—N1—C4—N2 2.58 (10) C10—C11—C12—C7 0.38 (9)
supporting information
sup-5 Acta Cryst. (2005). E61, o2746–o2747
C3—N2—C4—N1 −2.20 (11) C12—N4—C13—C14 178.11 (5)
C3—N2—C4—S1 176.64 (6) N3—C6—C13—N4 −2.12 (8)
C5—S1—C4—N1 1.38 (6) C5—C6—C13—N4 176.75 (5)
C5—S1—C4—N2 −177.56 (5) N3—C6—C13—C14 −178.90 (5)
C4—S1—C5—C6 58.65 (5) C5—C6—C13—C14 −0.03 (8)
C7—N3—C6—C13 1.50 (7) N4—C13—C14—S2 57.53 (6)
C7—N3—C6—C5 −177.45 (5) C6—C13—C14—S2 −125.50 (5)
S1—C5—C6—N3 52.02 (6) C15—S2—C14—C13 75.91 (5)
S1—C5—C6—C13 −126.91 (5) C18—N5—C15—N6 1.23 (9)
C6—N3—C7—C12 −0.03 (7) C18—N5—C15—S2 −177.82 (4)
C6—N3—C7—C8 177.91 (5) C16—N6—C15—N5 −0.74 (9)
N3—C7—C8—C9 −177.72 (5) C16—N6—C15—S2 178.36 (5)
C12—C7—C8—C9 0.25 (8) C14—S2—C15—N5 5.34 (5)
C7—C8—C9—C10 −0.01 (9) C14—S2—C15—N6 −173.83 (4)
C8—C9—C10—C11 −0.04 (10) C15—N6—C16—C17 −0.52 (10)
C9—C10—C11—C12 −0.15 (10) N6—C16—C17—C18 1.15 (11)
C13—N4—C12—C7 0.40 (8) C15—N5—C18—C17 −0.48 (9)
C13—N4—C12—C11 −178.09 (5) C16—C17—C18—N5 −0.61 (10)