metal-organic papers
m72
Caiet al. [Cu(C2H6NO3S)2(C10H8N2)]C10H8N2H2O doi:10.1107/S1600536805040900 Acta Cryst.(2006). E62, m72–m73 Acta Crystallographica Section E
Structure Reports Online
ISSN 1600-5368
catena
-Poly[[[bis(2-aminoethanesulfonato-
j
2N
,
O
)-copper(II)]-
l
-4,4
000-bipyridine-j
2N
:
N
000] 4,4
000-bipyridine
monohydrate]
Jin-Hua Cai,a,bYi-Min Jiang,a* Xiu-Ju Yin,bXu-Hui Liuband Xiu-Jian Wanga
aCollege of Chemistry and Chemical
Engineering, Guangxi Normal University, Guilin, Guangxi 541004, People’s Republic of China, andbDepartment of Chemistry and Life Sciences, Hechi Normal College, Yizhou, Guangxi 546300, People’s Republic of China
Correspondence e-mail: cjhzse@163.com
Key indicators
Single-crystal X-ray study
T= 293 K
Mean(C–C) = 0.004 A˚ Disorder in main residue
Rfactor = 0.047
wRfactor = 0.099
Data-to-parameter ratio = 15.3
For details of how these key indicators were automatically derived from the article, see http://journals.iucr.org/e.
#2006 International Union of Crystallography Printed in Great Britain – all rights reserved
The title compound, {[Cu(C2H6NO3S)2(C10H8N2)]C10H8N2 -H2O}n, was obtained by the hydrothermal reaction of Cu(CH3COO)22H2O, 4,40-bipyridne and taurine at 380 K. The 4,40-bipyridine ligands bridge CuII
ions, forming polymeric chains, in which CuII ions and the bridging ligands lie on twofold axes. Two taurine anions chelate to the CuIIion by the terminal N and O atoms, completing the distorted octahedral coordination. The uncoordinated bipyridine molecule is located on an inversion center.
Comment
Several Schiff bases derived from taurine have recently been prepared in our laboratory (Zhang & Jiang, 2002; Zenget al., 2003; Jianget al., 2004) because of the important physiological properties of taurine. As part of our ongoing investigation on taurine derivatives, we report here the synthesis and crystal structure of the title taurine–copper(II) complex, (I).
The crystal structure of (I) consists of one-dimensional complex chains, uncoordinated 4,4-bipyridine (bipy) and water molecules. The bipy ligands bridge neighboring CuII ions, forming infinite polymeric chains along the
[image:1.610.208.459.569.717.2]crystal-Received 7 November 2005 Accepted 6 December 2005 Online 10 December 2005
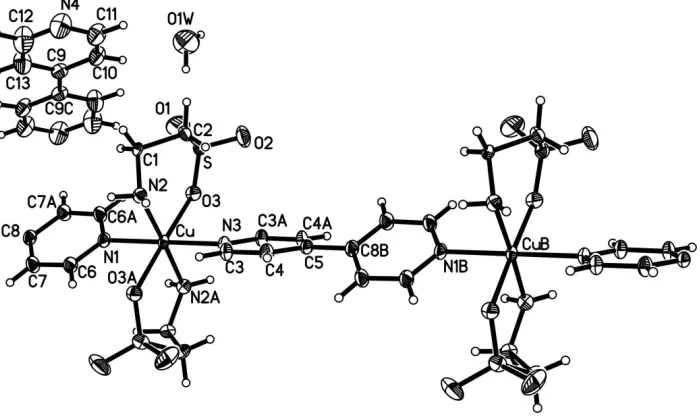
Figure 1
A segment of the polymeric structure of (I) with 30% probability displacement ellipsoids (arbitrary spheres for H atoms) [symmetry codes: (A)x,y,1
lographicbaxis (Fig. 1). The CuIIions and bridging ligands lie on twofold axes. Two taurine anions chelate to the CuIIion by terminal N and O atoms to complete the distorted octahedral coordination. The longer Cu—O3 bond distance (Table 1) shows the typical Jahn–Teller distortion effect for the CuIIion. Within the bridging bipy ligand, the two pyridine rings are twisted with respect to each other; the dihedral angle is 53.11 (4).
The uncoordinated bipy is located on an inversion center
and displays a planar configuration. Weak C—H O
hydrogen bonding occurs between the bipy and the water molecules. Classical O—H O hydrogen bonding is also observed between the water and the complex chain (Table 2).
Experimental
A methanol solution (10 ml) of taurine (1.5 mmol) and KOH (1.5 mmol) was mixed with another methanol solution (10 ml) of Cu(CH3COO)22H2O (0.5 mmol). After stirring for 10 min, 4,40
-bipyridine (1 mmol) was added slowly to the mixture, which was then dropped into a 25 ml Teflon-stainless steel reactor and heated at 380 K for 5 d. After cooling the reactor to room temperature, single crystals of (I) were obtained (yield 65%). Analysis found (%): C 44.87, H 4.66, N 13.05, S 9.98; calculated (%): C 44.84, H 4.67, N 13.08, S 9.97. IR (KBr, cm1): 1033.3, 1168.1, 1237.4 (—SO3), 3258.4,
3304.3 (N—H).
Crystal data
[Cu(C2H6NO3S)2(C10H8N2)]
-C10H8N2H2O Mr= 642.2
Monoclinic,P2=c a= 9.8647 (11) A˚
b= 11.3846 (9) A˚
c= 12.9516 (14) A˚ = 107.446 (5)
V= 1387.6 (2) A˚3
Z= 2
Dx= 1.537 Mg m
3
MoKradiation Cell parameters from 2887
reflections = 3.3–27.5 = 0.99 mm1 T= 293 (2) K Prism, blue
0.250.200.15 mm
Data collection
Rigaku Mercury CCD diffractometer ’and!scans
Absorption correction: multi-scan (SADABS; Sheldrick, 1996)
Tmin= 0.775,Tmax= 0.860
10598 measured reflections
3176 independent reflections 2681 reflections withI> 2(I)
Rint= 0.039
max= 27.5 h=11!12
k=14!14
l=16!15 Refinement
Refinement onF2 R[F2> 2(F2)] = 0.047
wR(F2) = 0.099 S= 1.08 3176 reflections 208 parameters
H-atom parameters constrained
w= 1/[2(F
o2) + (0.031P)2
+ 1.3917P]
whereP= (Fo2+ 2Fc2)/3
(/)max= 0.001
max= 0.42 e A˚
3
min=0.40 e A˚
[image:2.610.312.569.93.137.2]3
Table 1
Selected geometric parameters (A˚ ,).
Cu—N1 2.166 (3)
Cu—N2 1.990 (2)
Cu—N3 2.127 (3)
Cu—O3 2.3067 (19)
N2—Cu—N2i
177.17 (13) N2—Cu—N3 91.42 (6)
[image:2.610.310.566.207.263.2]Symmetry code: (i)x;y;zþ1 2.
Table 2
Hydrogen-bond geometry (A˚ ,).
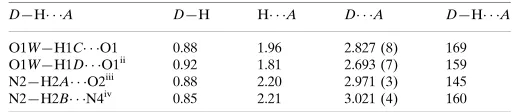
D—H A D—H H A D A D—H A
O1W—H1C O1 0.88 1.96 2.827 (8) 169 O1W—H1D O1ii
0.92 1.81 2.693 (7) 159 N2—H2A O2iii
0.88 2.20 2.971 (3) 145 N2—H2B N4iv
0.85 2.21 3.021 (4) 160
Symmetry codes: (ii)xþ1;y;zþ3
2; (iii)x;yþ1;z 1
2; (iv)xþ1;y;zþ 1 2.
The methylene group of aminomethylenesulfonate is disordered over two sites with 0.5 site-occupancy factors. The water molecule is located close to the twofold axis and has a 0.5 site-occupancy factor. H atoms on N and O atoms were located in a difference Fourier map and refined as riding in their as-found relative positions, with
Uiso(H) = 1.2Ueq(carrier). Other H atoms were placed in calculated
positions, with C—H = 0.93 (aromatic) or 0.97 A˚ (methylene), and refined using a riding-model approximation, with Uiso(H) =
1.2Ueq(C).
Data collection: CrystalClear (Rigaku, 2000); cell refinement:
CrustalClear; data reduction:CrystalClear; program(s) used to solve structure:SHELXS97(Sheldrick, 1997); program(s) used to refine structure: SHELXL97 (Sheldrick, 1997); molecular graphics:
SHELXTL (Bruker, 1997); software used to prepare material for publication:SHELXTL.
This work was supported by the Natural Science Founda-tion of Guangxi Chuang Autonomous Region of China (grant No. 0339034) and the Science Research Foundation of Guangxi University.
References
Bruker (1997).SHELXTL. Bruker AXS Inc., Madison, Wisconsin, USA. Jiang, Y.-M., Zeng, J.-L. & Yu, K.-B. (2004).Acta Cryst.C60, m543–m545. Rigaku (2000).CrystalClear. Rigaku Corporation, Tokyo, Japan. Sheldrick, G. M. (1996).SADABS. University of Go¨ttingen, Germany. Sheldrick, G. M. (1997). SHELXS97 and SHELXL97. University of
Go¨ttingen, Germany.
supporting information
sup-1
Acta Cryst. (2006). E62, m72–m73supporting information
Acta Cryst. (2006). E62, m72–m73 [doi:10.1107/S1600536805040900]
catena
-Poly[[[bis(2-aminoethanesulfonato-
κ
2N
,
O
)copper(II)]-
µ
-4,4
′
-bipyridine-κ
2N
:
N
′
] 4,4
′
-bipyridine monohydrate]
Jin-Hua Cai, Yi-Min Jiang, Xiu-Ju Yin, Xu-Hui Liu and Xiu-Jian Wang
S1. Comment
Several Schiff bases derived from taurine have recently been prepared in our laboratory (Zhang & Jiang, 2002; Zeng et
al., 2003; Jiang et al., 2004) because of the important physiological properties of taurine. As part of our ongoing
investigation on taurine derivatives, we report here the synthesis and crystal structure of the title taurine–copper(II)
complex, (I).
The crystal structure of (I) consists of one-dimensional complex chains, coordinated 4,4-bipyridine (bipy) and water
molecules. The bipy ligands bridge neighboring CuII ions, forming infinite polymeric chains along the crystallographic b
axis (Fig. 1). The polymeric chain is located on a twofold axis. Two taurine anions chelate to the CuII ion by terminal N
and O atoms to complete the distorted octahedral coordination. The longer Cu—O3 bond distance (Table 1) shows the
typical Jahn–Teller distortion effect for the CuII ion. Within the bridging bipy ligand, the two pyridine rings are twisted to
each other with a dihedral angle of 53.11 (4)°.
The bipy is located on an inversion center and displays a planar configuration. Weak C—H···O hydrogen bonding
occurs between the bipy and the water molecules (Table 2). Classic O—H···O hydrogen bonding is also observed between
the water and the complex chain.
S2. Experimental
A methanol solution (10 ml) of taurine (1.5 mmol) and KOH (1.5 mmol) was mixed with another methanol solution (10
ml) of Cu(CH3COO)2·2H2O (0.5 mmol). After stirring for 10 min, 4,4′-bipyridne (1 mmol) was added slowly to the
mixture. It was then dropped into a 25 ml Teflon-stainless steel reactor and heated at 380 K for 5 d. After cooling the
reactor to room temperature, single crystals of (I) were obtained (yield 65%). Analysis found (%): C 44.87, H 4.66, N
13.05, S 9.98; calculated (%): C 44.84, H 4.67, N 13.08, S 9.97. IR (KBr, ν cm−1): 1033.3, 1168.1, 1237.4 (–SO
3), 3258.4,
3304.3 (ν N—H).
S3. Refinement
The methylene group of aminomethylenesulfonate is disordered over two sites with 0.5 site-occupancy factors. The water
molecule is located close to the twofold axis and has a 0.5 site-occupancy factor. H atoms on N and O atoms were located
in a difference Fourier map and refined as riding in their as-found relative positions, with Uiso(H) = 1.2Ueq(carrier). Other
H atoms were placed in calculated positions, with C—H = 0.93 (aromatic) or 0.97 Å (methylene), and refined using a
Figure 1
A segment of the polymeric structure of (I) with 30% probability displacement ellipsoids (arbitrary spheres for H atoms)
[symmetry codes: (A) −x, y, 1/2 − z; (B) x, 1 + y, z; (C) 1 − x, −y, 1 − z].
catena-Poly[[[bis(2-aminoethanesulfonato-κ2N,O)copper(II)]-µ-4,4′- bipyridine-κ2N:N′] 4,4′-bipyridine
monohydrate]]
Crystal data
[Cu(C2H6NO3S)2(C10H8N2)]·C10H8N2·H2O
Mr = 642.2 Monoclinic, P2/c
Hall symbol: -P 2yc
a = 9.8647 (11) Å
b = 11.3846 (9) Å
c = 12.9516 (14) Å
β = 107.446 (5)°
V = 1387.6 (2) Å3
Z = 2
F(000) = 666
Dx = 1.537 Mg m−3
Mo Kα radiation, λ = 0.71073 Å Cell parameters from 2887 reflections
θ = 3.3–27.5°
µ = 0.99 mm−1
T = 293 K Prism, blue
0.25 × 0.20 × 0.15 mm
Data collection
Rigaku Mercury CCD diffractometer
Radiation source: fine-focus sealed tube Graphite monochromator
φ and ω scans
Absorption correction: multi-scan (SADABS; Sheldrick, 1996)
Tmin = 0.775, Tmax = 0.860
10598 measured reflections 3176 independent reflections 2681 reflections with I > 2σ(I)
Rint = 0.039
θmax = 27.5°, θmin = 3.3°
h = −11→12
k = −14→14
l = −16→15
Refinement
Refinement on F2 Least-squares matrix: full
R[F2 > 2σ(F2)] = 0.047
wR(F2) = 0.099
3176 reflections 208 parameters 1 restraint
supporting information
sup-3
Acta Cryst. (2006). E62, m72–m73Secondary atom site location: difference Fourier map
Hydrogen site location: inferred from neighbouring sites
H-atom parameters constrained
w = 1/[σ2(F
o2) + (0.031P)2 + 1.3917P] where P = (Fo2 + 2Fc2)/3
(Δ/σ)max = 0.001 Δρmax = 0.42 e Å−3 Δρmin = −0.40 e Å−3
Special details
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full
covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes.
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2,
conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2 are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger.
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
x y z Uiso*/Ueq Occ. (<1)
Cu 0.0000 0.37558 (3) 0.2500 0.02478 (14)
S 0.25875 (8) 0.39859 (6) 0.49274 (6) 0.03598 (18)
N1 0.0000 0.1853 (2) 0.2500 0.0332 (7)
N2 0.1844 (2) 0.37126 (19) 0.21701 (18) 0.0325 (5)
H2A 0.1791 0.4239 0.1658 0.039*
H2B 0.1885 0.3042 0.1896 0.039*
N3 0.0000 0.5624 (2) 0.2500 0.0280 (7)
N4 0.7300 (4) 0.1558 (3) 0.3845 (3) 0.0790 (11)
O1 0.3188 (3) 0.3010 (2) 0.5593 (2) 0.0904 (10)
O2 0.2828 (3) 0.5085 (2) 0.5489 (2) 0.0729 (8)
O3 0.1092 (2) 0.37865 (16) 0.43436 (15) 0.0370 (4)
O1W 0.5829 (8) 0.2895 (8) 0.7232 (5) 0.134 (3) 0.50
H1C 0.4958 0.2944 0.6791 0.161* 0.50
H1D 0.5936 0.2934 0.7961 0.161* 0.50
C1A 0.3188 (5) 0.3414 (5) 0.3031 (4) 0.0278 (11) 0.50
H1A1 0.3974 0.3384 0.2727 0.033* 0.50
H1A2 0.3101 0.2652 0.3340 0.033* 0.50
C2A 0.3459 (7) 0.4380 (6) 0.3919 (6) 0.0350 (15) 0.50
H2A1 0.3093 0.5126 0.3591 0.042* 0.50
H2A2 0.4472 0.4464 0.4265 0.042* 0.50
C1B 0.3085 (7) 0.4331 (8) 0.2996 (6) 0.0589 (19) 0.50
H1B1 0.3880 0.4387 0.2703 0.071* 0.50
H1B2 0.2799 0.5123 0.3113 0.071* 0.50
C2B 0.3536 (8) 0.3728 (8) 0.4012 (7) 0.053 (2) 0.50
H2B1 0.4521 0.3932 0.4364 0.063* 0.50
H2B2 0.3505 0.2891 0.3866 0.063* 0.50
C3 0.0111 (3) 0.6237 (2) 0.1647 (2) 0.0328 (6)
H3 0.0175 0.5827 0.1043 0.039*
C4 0.0134 (3) 0.7442 (2) 0.1622 (2) 0.0359 (6)
C5 0.0000 0.8084 (3) 0.2500 0.0302 (8)
C6 −0.0586 (3) 0.1244 (2) 0.1593 (2) 0.0395 (7)
H6 −0.0997 0.1655 0.0956 0.047*
C7 −0.0606 (3) 0.0027 (2) 0.1561 (3) 0.0424 (7)
H7 −0.1025 −0.0361 0.0912 0.051*
C8 0.0000 −0.0610 (3) 0.2500 0.0358 (9)
C9 0.5482 (3) 0.0322 (2) 0.4758 (3) 0.0414 (7)
C10 0.5921 (4) 0.1457 (3) 0.5069 (3) 0.0540 (9)
H10 0.5626 0.1830 0.5602 0.065*
C11 0.6802 (4) 0.2032 (3) 0.4582 (4) 0.0678 (12)
H11 0.7058 0.2802 0.4790 0.081*
C12 0.6910 (5) 0.0463 (4) 0.3565 (4) 0.0864 (15)
H12 0.7262 0.0102 0.3054 0.104*
C13 0.6007 (4) −0.0176 (3) 0.3989 (3) 0.0671 (11)
H13 0.5756 −0.0939 0.3754 0.081*
Atomic displacement parameters (Å2)
U11 U22 U33 U12 U13 U23
Cu 0.0296 (2) 0.0165 (2) 0.0310 (3) 0.000 0.01326 (19) 0.000
S 0.0381 (4) 0.0362 (4) 0.0315 (4) 0.0053 (3) 0.0072 (3) −0.0069 (3)
N1 0.0352 (18) 0.0186 (14) 0.046 (2) 0.000 0.0124 (15) 0.000
N2 0.0344 (12) 0.0353 (11) 0.0311 (12) 0.0008 (9) 0.0149 (10) −0.0025 (10)
N3 0.0365 (17) 0.0196 (13) 0.0311 (16) 0.000 0.0150 (14) 0.000
N4 0.086 (2) 0.0542 (19) 0.122 (3) −0.0166 (17) 0.070 (2) 0.006 (2)
O1 0.084 (2) 0.0679 (18) 0.093 (2) 0.0237 (15) −0.0139 (17) 0.0292 (16)
O2 0.0695 (17) 0.0516 (14) 0.106 (2) −0.0207 (12) 0.0397 (16) −0.0400 (14)
O3 0.0366 (11) 0.0424 (11) 0.0327 (11) 0.0007 (8) 0.0114 (9) 0.0010 (8)
O1W 0.126 (6) 0.192 (8) 0.057 (4) 0.038 (6) −0.015 (4) −0.033 (5)
C1A 0.026 (3) 0.031 (3) 0.030 (3) 0.005 (2) 0.013 (2) −0.010 (2)
C2A 0.034 (3) 0.036 (3) 0.037 (4) −0.009 (3) 0.014 (3) −0.013 (3)
C1B 0.037 (4) 0.080 (6) 0.068 (5) −0.009 (4) 0.029 (4) −0.010 (4)
C2B 0.030 (3) 0.067 (5) 0.058 (5) 0.007 (4) 0.009 (3) −0.008 (5)
C3 0.0472 (16) 0.0235 (12) 0.0312 (14) −0.0006 (11) 0.0170 (13) −0.0008 (10)
C4 0.0530 (18) 0.0242 (12) 0.0330 (15) −0.0001 (11) 0.0167 (13) 0.0047 (11)
C5 0.0309 (19) 0.0196 (16) 0.039 (2) 0.000 0.0092 (17) 0.000
C6 0.0476 (17) 0.0238 (12) 0.0429 (17) 0.0017 (12) 0.0073 (14) 0.0014 (12)
C7 0.0531 (19) 0.0247 (13) 0.0440 (17) −0.0009 (12) 0.0062 (15) −0.0031 (12)
C8 0.038 (2) 0.0234 (18) 0.047 (2) 0.000 0.0141 (19) 0.000
C9 0.0352 (15) 0.0318 (14) 0.059 (2) −0.0020 (12) 0.0173 (14) 0.0101 (14)
C10 0.053 (2) 0.0329 (16) 0.088 (3) −0.0010 (14) 0.0392 (19) 0.0029 (16)
C11 0.067 (2) 0.0331 (16) 0.119 (4) −0.0088 (16) 0.051 (3) 0.006 (2)
C12 0.109 (4) 0.069 (3) 0.112 (4) −0.022 (3) 0.080 (3) −0.012 (3)
supporting information
sup-5
Acta Cryst. (2006). E62, m72–m73Geometric parameters (Å, º)
Cu—N1 2.166 (3) C2A—H2A2 0.9700
Cu—N2 1.990 (2) C1B—C2B 1.432 (11)
Cu—N2i 1.990 (2) C1B—H1B1 0.9700
Cu—N3 2.127 (3) C1B—H1B2 0.9700
Cu—O3i 2.3067 (19) C2B—H2B1 0.9700
Cu—O3 2.3067 (19) C2B—H2B2 0.9700
S—O1 1.422 (3) C3—C4 1.373 (3)
S—O2 1.431 (2) C3—H3 0.9300
S—O3 1.460 (2) C4—C5 1.391 (3)
S—C2B 1.741 (8) C4—H4 0.9300
S—C2A 1.822 (7) C5—C4i 1.391 (3)
N1—C6 1.336 (3) C5—C8ii 1.487 (5)
N1—C6i 1.336 (3) C6—C7 1.386 (4)
N2—C1A 1.493 (6) C6—H6 0.9300
N2—C1B 1.534 (8) C7—C8 1.388 (3)
N2—H2A 0.8838 C7—H7 0.9300
N2—H2B 0.8482 C8—C7i 1.388 (3)
N3—C3i 1.338 (3) C8—C5iii 1.487 (5)
N3—C3 1.338 (3) C9—C13 1.375 (5)
N4—C11 1.313 (5) C9—C10 1.384 (4)
N4—C12 1.323 (5) C9—C9iv 1.482 (6)
O1W—H1C 0.8790 C10—C11 1.381 (5)
O1W—H1D 0.9192 C10—H10 0.9300
C1A—C2A 1.556 (8) C11—H11 0.9300
C1A—H1A1 0.9700 C12—C13 1.384 (5)
C1A—H1A2 0.9700 C12—H12 0.9300
C2A—H2A1 0.9700 C13—H13 0.9300
N2—Cu—N2i 177.17 (13) S—C2A—H2A1 109.7
N2—Cu—N3 91.42 (6) C1A—C2A—H2A2 109.7
N2i—Cu—N3 91.42 (6) S—C2A—H2A2 109.7
N2—Cu—N1 88.58 (6) H2A1—C2A—H2A2 108.2
N2i—Cu—N1 88.58 (6) C2B—C1B—N2 112.6 (6)
N3—Cu—N1 180.0 C2B—C1B—H1B1 109.1
N2—Cu—O3i 87.22 (8) N2—C1B—H1B1 109.1
N2i—Cu—O3i 92.83 (8) C2B—C1B—H1B2 109.1
N3—Cu—O3i 89.13 (5) N2—C1B—H1B2 109.1
N1—Cu—O3i 90.87 (5) H1B1—C1B—H1B2 107.8
N2—Cu—O3 92.83 (8) C1B—C2B—S 117.5 (6)
N2i—Cu—O3 87.22 (8) C1B—C2B—H2B1 107.9
N3—Cu—O3 89.13 (5) S—C2B—H2B1 107.9
N1—Cu—O3 90.87 (5) C1B—C2B—H2B2 107.9
O3i—Cu—O3 178.27 (9) S—C2B—H2B2 107.9
O1—S—O2 113.56 (19) H2B1—C2B—H2B2 107.2
O1—S—O3 111.42 (15) N3—C3—C4 123.1 (3)
O1—S—C2B 94.0 (3) C4—C3—H3 118.4
O2—S—C2B 116.9 (3) C3—C4—C5 119.9 (3)
O3—S—C2B 106.7 (3) C3—C4—H4 120.0
O1—S—C2A 115.3 (3) C5—C4—H4 120.0
O2—S—C2A 96.1 (2) C4—C5—C4i 116.7 (3)
O3—S—C2A 106.7 (2) C4—C5—C8ii 121.67 (16)
C6—N1—C6i 117.5 (3) C4i—C5—C8ii 121.67 (16)
C6—N1—Cu 121.25 (16) N1—C6—C7 122.9 (3)
C6i—N1—Cu 121.25 (16) N1—C6—H6 118.5
C1A—N2—Cu 120.6 (2) C7—C6—H6 118.5
C1B—N2—Cu 115.4 (3) C6—C7—C8 119.8 (3)
C1A—N2—H2A 123.7 C6—C7—H7 120.1
C1B—N2—H2A 93.9 C8—C7—H7 120.1
Cu—N2—H2A 107.0 C7i—C8—C7 117.0 (3)
C1A—N2—H2B 88.0 C7i—C8—C5iii 121.51 (17)
C1B—N2—H2B 125.2 C7—C8—C5iii 121.51 (17)
Cu—N2—H2B 105.9 C13—C9—C10 116.3 (3)
H2A—N2—H2B 107.2 C13—C9—C9iv 122.1 (3)
C3i—N3—C3 117.2 (3) C10—C9—C9iv 121.6 (4)
C3i—N3—Cu 121.42 (15) C11—C10—C9 119.6 (3)
C3—N3—Cu 121.42 (15) C11—C10—H10 120.2
C11—N4—C12 116.4 (3) C9—C10—H10 120.2
S—O3—Cu 128.64 (12) N4—C11—C10 124.1 (3)
H1C—O1W—H1D 117.0 N4—C11—H11 118.0
N2—C1A—C2A 107.9 (4) C10—C11—H11 118.0
N2—C1A—H1A1 110.1 N4—C12—C13 123.7 (4)
C2A—C1A—H1A1 110.1 N4—C12—H12 118.2
N2—C1A—H1A2 110.1 C13—C12—H12 118.2
C2A—C1A—H1A2 110.1 C9—C13—C12 119.8 (3)
H1A1—C1A—H1A2 108.4 C9—C13—H13 120.1
C1A—C2A—S 110.0 (4) C12—C13—H13 120.1
C1A—C2A—H2A1 109.7
Symmetry codes: (i) −x, y, −z+1/2; (ii) x, y+1, z; (iii) x, y−1, z; (iv) −x+1, −y, −z+1.
Hydrogen-bond geometry (Å, º)
D—H···A D—H H···A D···A D—H···A
O1W—H1C···O1 0.88 1.96 2.827 (8) 169
O1W—H1D···O1v 0.92 1.81 2.693 (7) 159
N2—H2A···O2vi 0.88 2.20 2.971 (3) 145
N2—H2B···N4vii 0.85 2.21 3.021 (4) 160
![Figure 1A segment of the polymeric structure of (I) with 30% probabilitydisplacement ellipsoids (arbitrary spheres for H atoms) [symmetry codes:(A) �x, y, 12 � z; (B) x, 1 + y, z; (C) 1 � x, �y, 1 � z].](https://thumb-us.123doks.com/thumbv2/123dok_us/684763.571399/1.610.208.459.569.717/figure-segment-polymeric-structure-probabilitydisplacement-ellipsoids-arbitrary-symmetry.webp)