Acta Crystallographica Section E
Structure Reports Online
ISSN 1600-5368
2-(9-Hydroxy-3a,5a,5b,8,8,11a-hexamethylicosa-hydro-1
H
-cyclopenta[
a
]chrysen-1-yl)propanoic acid
(3b
-hydroxylupan-29-oic acid)
Pakakrong Thongdeeying,a Suchada Chantrapromma,a‡ Hoong-Kun Fun,b* Shazia Anjum,cShamsher Alicand Chanita Ponglimanonta
aDepartment of Chemistry, Faculty of Science,
Prince of Songkla University, Hat-Yai, Songkhla 90112, Thailand,bX-ray Crystallography Unit, School of Physics, Universiti Sains Malaysia, 11800 USM, Penang, Malaysia, andcHEJ Research Institute of Chemistry, University of Karachi, Karachi 75270, Pakistan
‡ Additional correspondence author, email: suchada.c@psu.ac.th.
Correspondence e-mail: hkfun@usm.my
Key indicators Single-crystal X-ray study T= 293 K
Mean(C–C) = 0.006 A˚ Rfactor = 0.053 wRfactor = 0.117 Data-to-parameter ratio = 8.4
For details of how these key indicators were automatically derived from the article, see http://journals.iucr.org/e.
The title compound, C30H50O3, a lupane triterpene, was
isolated from the leaves of Ceriops decandra (Griff.) Ding
Hou. There are two crystallographically independent mol-ecules in the asymmetric unit. In both molmol-ecules, the cyclopentane ring adopts an envelope conformation. In the crystal structure, molecules are interconnected into a
two-dimensional network by intermolecular O—H O hydrogen
bonds.
Comment
Ceriops decandra (Griff.) Ding Hou (Rhizophoraceae) is a mangrove plant widely distributed from East Africa and Madagascar throughout tropical Asia and Queensland to
Melanesia and Micronesia (Tomlinson, 1986).C. decandrahas
many local Thai names,e.g.Prong Khao and Prong Nu, and
also a synonym of Ceriops roxburghiana Arn (Smitinand &
Larsen, 1970). The bark of this plant has been used as a folk medicine for the treatment of diarrhoea, vomiting, amoebiasis and ulcers (Bamroongrugsa, 1999). An ethanol extract of the leaves has shown antinociceptive activity (Uddinet al., 2005).
The title compound, 3-hydroxylupan-29-oic acid, (I), was
previously isolated from Gymnosporia wallichiana
(Kulsh-reshtha, 1977). As part of our research on bioactive
compounds from Thai medicinal plants (Chantrapromma et
al., 2003, 2004; Waratchareeyakulet al., 2004; Boonnaket al., 2005), compound (I) was isolated for the first time from
Ceriops decandra (Griff.) Ding Hou, collected from Phang-Nga province in the southern part of Thailand. We have undertaken the X-ray crystal structure analysis of (I) in order to establish its molecular structure and relative stereo-chemistry.
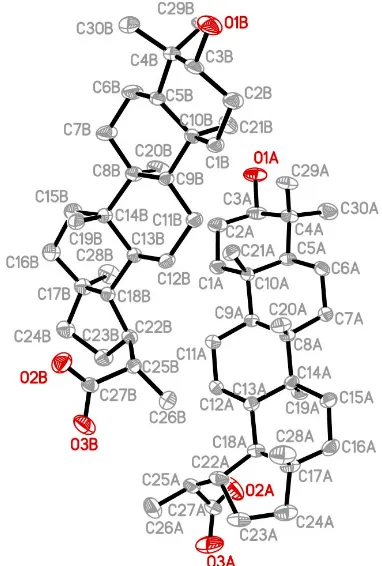
The asymmetric unit of (I) contains two crystallographically
independent molecules, A andB, which have similar
chiral-ities, bond lengths and angles (Fig. 1). The molecules are approximately related by a local twofold rotation axis. The bond lengths in (I) show normal values (Allenet al., 1987). All
the ring junctions in the lupane nucleus are trans-fused. In
with atom C17 displaced from the C18/C22/C23/C24 plane by
0.692 (7) A˚ in moleculeA and 0.674 (7) A˚ in molecule B. The hydroxyl group is equatorially attached at atom C3. The
C23—C22—C25—C27 torsion angle of55.1 (5)[45.7 (5)
in B] describes the orientation of the propanoic acid group
with respect to the lupane nucleus. The C18—C22—C25—C26 torsion angle is170.1 (4) [161.8 (3)inB].
The molecular structure is stabilized by C—H O
hydrogen bonds (Table 2). O—H O intermolecular
hydrogen bonds link the molecules into a two-dimensional network parallel to theacplane (Fig. 2).
Experimental
Air-dried leaves ofCeriops decandra(Griff.) Ding Hou (3.9 kg) were ground and extracted with hexane (220 l) at room temperature. The mixture was filtered and concentrated under reduced pressure to give a crude hexane extract (66.4 g). The crude extract was separated by quick column chromatography (QCC) on silica gel and eluted initially with hexane, followed by ethyl acetate and finally with methanol to give nine fractions (B1–B9). Fraction B7 (2.7 g) was subjected to column chromatography with MeOH–CH2Cl2(0.2:9.8v/
v) and further purified by recrystallization from CHCl3–EtOAc– MeOH (4:4:1) for a few days to obtain colourless single crystals of compound (I) [m.p. 518–520 K, []28
D 42.6
(c= 0.125, MeOH)].
Crystal data
C30H50O3 Mr= 458.70
Monoclinic, P21
a= 8.1669 (8) A˚
b= 24.110 (2) A˚
c= 14.4666 (14) A˚
= 104.057 (2)
V= 2763.2 (4) A˚3 Z= 4
Dx= 1.103 Mg m
3
MoKradiation
Cell parameters from 14 091 reflections
= 1.5–25.0
= 0.07 mm1
T= 293 (2) K Plate, colourless 0.230.150.08 mm
Data collection
Siemens SMART CCD area-detector diffractometer
!scans
Absorption correction: multi-scan (SADABS; Sheldrick, 1996)
Tmin= 0.988,Tmax= 0.994
14 091 measured reflections
4986 independent reflections 3296 reflections withI> 2(I)
Rint= 0.050 max= 25.0
h=9!9
k=23!28
l=14!17
Refinement
Refinement onF2 R[F2> 2(F2)] = 0.053 wR(F2) = 0.117 S= 1.02 4986 reflections 596 parameters
H-atom parameters constrained
w= 1/[2(F
o2) + (0.0486P)2
+ 0.1104P]
whereP= (Fo2+ 2Fc2)/3
(/)max< 0.001 max= 0.16 e A˚
3 min=0.14 e A˚
3
Table 1
Selected geometric parameters (A˚ ,).
O1A—C3A 1.452 (5) O2A—C27A 1.234 (6) O3A—C27A 1.264 (5)
O1B—C3B 1.445 (5) O2B—C27B 1.236 (5) O3B—C27B 1.284 (5)
O2A—C27A—O3A 122.9 (5) O3A—C27A—C25A 118.1 (5)
[image:2.610.339.530.72.355.2]O2B—C27B—O3B 123.8 (5) O3B—C27B—C25B 116.0 (4)
Table 2
Hydrogen-bond geometry (A˚ ,).
D—H A D—H H A D A D—H A
O1A—H1A O2Bi
0.82 1.91 2.705 (6) 161 O1B—H1D O2Aii 0.82 1.86 2.662 (6) 165 O3A—H3A O1Aiii
0.82 1.98 2.724 (6) 151 O3B—H3C O1Biv
0.82 1.82 2.625 (5) 163 C18A—H18A O2Av
0.98 2.52 3.262 (6) 132 C18B—H18B O2Bv 0.98 2.46 3.206 (5) 133 C23A—H23B O3Av
0.97 2.49 3.222 (7) 132 C23B—H23D O3Bv
0.97 2.49 3.237 (6) 134 C30A—H30A O1Av 0.96 2.56 2.957 (7) 105 C30B—H30D O1Bv
0.96 2.51 2.917 (7) 105
Symmetry codes: (i) xþ1;yþ1
2;z; (ii) x;y 1
2;z; (iii) x;y;zþ1; (iv) xþ1;y;zþ1; (v)x;y;z.
H atoms were placed in calculated positions, with O—H distances of 0.82 A˚ and C—H distances in the range 0.93–0.98 A˚. TheUiso values were constrained to be 1.5Ueqof the carrier atom for hydroxyl and methyl H atoms, and 1.2Ueqfor the remaining H atoms. In the absence of significant anomalous dispersion effects, Friedel pairs were merged before the final refinement and the absolute config-uration was assigned arbitrarily.
Data collection:SMART(Siemens, 1996); cell refinement:SAINT
(Siemens, 1996); data reduction:SAINT; program(s) used to solve structure: SHELXTL (Sheldrick, 1997); program(s) used to refine structure:SHELXTL; molecular graphics:SHELXTL; software used to prepare material for publication:SHELXTLandPLATON(Spek, 2003).
Figure 1
[image:2.610.313.565.435.543.2]PT thanks the Higher Education Development Project: Postgraduate Education and Research Program in Chemistry for financial support. The authors thank Prince of Songkla
University and the Pakistan Government, and also the Malaysian Government and Universiti Sains Malaysia for the Scientific Advancement Grant Allocation (SAGA) grant No. 304/PFIZIK/653003/A118.
References
Allen, F. H., Kennard, O., Watson, D. G., Brammer, L., Orpen, A. G. & Taylor, R. (1987).J. Chem. Soc. Perkin Trans. 2, pp. S1–19.
Bamroongrugsa, N. (1999).Songklanakarin J. Sci. Technol.21, 377–386. Boonnak, N., Chantrapromma, S., Fun, H.-K., Anjum Ali, S., Rahman, A. &
Karalai, C. (2005).Acta Cryst.E61, o410–o412.
Chantrapromma, S., Fun, H.-K., Ibrahim, A. R., Laphookhieo, S. & Karalai, C. (2003).Acta Cryst.E59, o1864–o1866.
Chantrapromma, K., Saewan, N., Fun, H.-K., Chantrapromma, S. & Rahman, A. A. (2004).Acta Cryst.E60, o312–o314.
Kulshreshtha, D. K. (1977).Phytochemistry,16, 1783–1785.
Sheldrick, G. M. (1996).SADABS. University of Go¨ttingen, Germany. Sheldrick, G. M. (1997).SHELXTL. Version 5.1. Bruker AXS Inc., Madison,
Wisconsin, USA.
Siemens (1996).SMARTandSAINT. Siemens Analytical X-ray Instruments Inc., Madison, Wisconsin, USA.
Smitinand, T. & Larsen, K. (1970).Flora of Thailand, Vol. 2, Part 1, pp. 10–12. Bangkok: The ASRCT Press.
Spek, A. L. (2003).J. Appl. Cryst.36, 7–13.
Tomlinson, P. B. (1986).The Botany of Mangroves, pp. 352–357. Cambridge University Press.
Uddin, S. J., Shilpi, J. A., Barua, J. & Rouf, R. (2005).Fitoterapia,76, 261– 263.
[image:3.610.45.295.72.287.2]Waratchareeyakul, W., Chantrapromma, S., Fun, H.-K., Razak, I. A., Karalai, C. & Ponglimanont C. (2004).Acta Cryst.E60, o1964–o1966.
Figure 2
supporting information
Acta Cryst. (2005). E61, o1861–o1863 [https://doi.org/10.1107/S1600536805015606]
2-(9-Hydroxy-3a,5a,5b,8,8,11a-hexamethylicosahydro-1
H
-cyclo-penta[
a
]chrysen-1-yl)propanoic acid (3
β
-hydroxylupan-29-oic acid)
Pakakrong Thongdeeying, Suchada Chantrapromma, Hoong-Kun Fun, Shazia Anjum, Shamsher
Ali and Chanita Ponglimanont
2-(9-hydroxy-3a,5a,5 b,8,8,11a-(hexamethylicosahydro-1H-cyclopenta[a]chrysen- 1-yl)propanoic acid
Crystal data C30H50O3
Mr = 458.70
Monoclinic, P21
Hall symbol: P 2yb a = 8.1669 (8) Å b = 24.110 (2) Å c = 14.4666 (14) Å β = 104.057 (2)° V = 2763.2 (4) Å3
Z = 4
F(000) = 1016 Dx = 1.103 Mg m−3
Melting point = 518–520 K Mo Kα radiation, λ = 0.71073 Å Cell parameters from 14091 reflections θ = 1.5–25.0°
µ = 0.07 mm−1
T = 293 K Plate, colourless 0.23 × 0.15 × 0.08 mm
Data collection
Siemens SMART CCD area-detector diffractometer
Radiation source: fine-focus sealed tube Graphite monochromator
Detector resolution: 8.33 pixels mm-1
ω scans
Absorption correction: multi-scan (SADABS; Sheldrick, 1996) Tmin = 0.988, Tmax = 0.994
14091 measured reflections 4986 independent reflections 3296 reflections with I > 2σ(I) Rint = 0.050
θmax = 25.0°, θmin = 1.5°
h = −9→9 k = −23→28 l = −14→17
Refinement Refinement on F2
Least-squares matrix: full R[F2 > 2σ(F2)] = 0.053
wR(F2) = 0.117
S = 1.02 4986 reflections 596 parameters 1 restraint
Primary atom site location: structure-invariant direct methods
Secondary atom site location: difference Fourier map
Hydrogen site location: inferred from neighbouring sites
H-atom parameters constrained w = 1/[σ2(F
o2) + (0.0486P)2 + 0.1104P]
where P = (Fo2 + 2Fc2)/3
(Δ/σ)max < 0.001
Δρmax = 0.16 e Å−3
Special details
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes.
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2,
conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used
only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2
are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger.
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
x y z Uiso*/Ueq
O1A 0.4019 (5) 0.49649 (15) −0.2515 (2) 0.0736 (11)
H1A 0.4861 0.5162 −0.2429 0.110*
O2A 0.4217 (6) 0.45492 (17) 0.5085 (3) 0.1053 (15) O3A 0.4272 (6) 0.40694 (19) 0.6381 (3) 0.1025 (14)
H3A 0.4460 0.4378 0.6623 0.154*
C1A 0.4333 (5) 0.4231 (2) −0.0165 (3) 0.0514 (13)
H1B 0.4978 0.3905 0.0097 0.062*
H1C 0.4729 0.4538 0.0263 0.062*
C2A 0.4665 (6) 0.4361 (2) −0.1140 (3) 0.0598 (14)
H2B 0.5855 0.4441 −0.1062 0.072*
H2C 0.4390 0.4038 −0.1549 0.072*
C3A 0.3638 (6) 0.4847 (2) −0.1605 (3) 0.0550 (13)
H3B 0.4002 0.5169 −0.1194 0.066*
C4A 0.1742 (6) 0.47895 (19) −0.1730 (3) 0.0474 (12) C5A 0.1445 (5) 0.46214 (17) −0.0730 (3) 0.0405 (11)
H5A 0.1862 0.4941 −0.0322 0.049*
C6A −0.0405 (5) 0.4583 (2) −0.0718 (3) 0.0478 (12)
H6A −0.0883 0.4244 −0.1030 0.057*
H6B −0.1016 0.4893 −0.1068 0.057*
C7A −0.0605 (6) 0.45899 (19) 0.0304 (3) 0.0463 (11)
H7A −0.0235 0.4948 0.0584 0.056*
H7B −0.1793 0.4552 0.0289 0.056*
C8A 0.0369 (5) 0.41387 (18) 0.0942 (3) 0.0375 (10) C9A 0.2225 (5) 0.41276 (18) 0.0839 (3) 0.0365 (10)
H9A 0.2696 0.4484 0.1103 0.044*
C10A 0.2469 (5) 0.41271 (18) −0.0206 (3) 0.0373 (10) C11A 0.3273 (6) 0.3697 (2) 0.1496 (3) 0.0490 (12)
H11A 0.4439 0.3730 0.1459 0.059*
H11B 0.2880 0.3331 0.1271 0.059*
C12A 0.3200 (5) 0.3752 (2) 0.2539 (3) 0.0463 (11)
H12A 0.3776 0.3438 0.2895 0.056*
H12B 0.3794 0.4086 0.2804 0.056*
C13A 0.1401 (5) 0.37762 (18) 0.2656 (3) 0.0396 (10)
H13A 0.0862 0.3426 0.2406 0.048*
C15A −0.1435 (6) 0.4277 (2) 0.2165 (3) 0.0531 (13)
H15A −0.1959 0.4616 0.1873 0.064*
H15B −0.2074 0.3968 0.1827 0.064*
C16A −0.1581 (6) 0.4262 (2) 0.3206 (3) 0.0601 (14)
H16A −0.1139 0.4604 0.3521 0.072*
H16B −0.2760 0.4233 0.3218 0.072*
C17A −0.0614 (6) 0.3773 (2) 0.3745 (3) 0.0549 (13) C18A 0.1245 (5) 0.3822 (2) 0.3694 (3) 0.0447 (11)
H18A 0.1615 0.4197 0.3910 0.054*
C19A 0.1199 (6) 0.48195 (18) 0.2365 (3) 0.0499 (12)
H19A 0.1220 0.4872 0.3026 0.075*
H19B 0.0550 0.5110 0.1995 0.075*
H19C 0.2331 0.4829 0.2285 0.075*
C20A −0.0540 (6) 0.3579 (2) 0.0622 (3) 0.0532 (13) H20A −0.0753 0.3546 −0.0058 0.080*
H20B −0.1591 0.3569 0.0808 0.080*
H20C 0.0162 0.3278 0.0918 0.080*
C21A 0.2002 (6) 0.35597 (19) −0.0707 (3) 0.0543 (13)
H21A 0.2604 0.3514 −0.1196 0.081*
H21B 0.0810 0.3547 −0.0987 0.081*
H21C 0.2304 0.3267 −0.0248 0.081*
C22A 0.2223 (6) 0.3418 (2) 0.4466 (3) 0.0542 (13)
H22A 0.2230 0.3053 0.4169 0.065*
C23A 0.1075 (7) 0.3379 (2) 0.5189 (3) 0.0699 (16)
H23A 0.0661 0.3003 0.5213 0.084*
H23B 0.1705 0.3484 0.5824 0.084*
C24A −0.0391 (7) 0.3779 (3) 0.4828 (3) 0.0712 (16)
H24A −0.0117 0.4148 0.5085 0.085*
H24B −0.1406 0.3651 0.4997 0.085*
C25A 0.4062 (6) 0.35740 (19) 0.4938 (3) 0.0550 (13)
H25A 0.4637 0.3636 0.4426 0.066*
C26A 0.4983 (8) 0.3097 (2) 0.5556 (4) 0.092 (2)
H26A 0.6115 0.3210 0.5856 0.138*
H26B 0.5014 0.2779 0.5162 0.138*
H26C 0.4396 0.3005 0.6035 0.138*
C27A 0.4182 (6) 0.4103 (2) 0.5498 (3) 0.0550 (13) C28A −0.1458 (7) 0.3220 (2) 0.3346 (4) 0.0782 (17)
H28A −0.2565 0.3200 0.3463 0.117*
H28B −0.0787 0.2916 0.3656 0.117*
H28C −0.1547 0.3203 0.2673 0.117*
C29A 0.0973 (6) 0.4379 (2) −0.2531 (3) 0.0623 (15)
H29A 0.1003 0.4538 −0.3135 0.093*
H29B −0.0175 0.4303 −0.2520 0.093*
H29C 0.1611 0.4041 −0.2439 0.093*
C30A 0.0924 (7) 0.5359 (2) −0.2004 (3) 0.0719 (16)
H30A 0.1336 0.5513 −0.2516 0.108*
H30B 0.1204 0.5602 −0.1463 0.108*
O1B −0.3919 (4) 0.06052 (18) −0.5555 (2) 0.0700 (11) H1D −0.4123 0.0275 −0.5507 0.13 (3)* O2B 0.3652 (5) 0.07827 (15) 0.2089 (2) 0.0718 (10) O3B 0.3763 (5) 0.12571 (16) 0.3420 (2) 0.0810 (12)
H3C 0.4566 0.1053 0.3643 0.122*
C1B −0.2454 (5) 0.1336 (2) −0.3199 (3) 0.0446 (12)
H1E −0.2252 0.1004 −0.2810 0.054*
H1F −0.3023 0.1601 −0.2880 0.054*
C2B −0.3622 (5) 0.1191 (2) −0.4166 (3) 0.0539 (13)
H2E −0.4652 0.1024 −0.4073 0.065*
H2F −0.3923 0.1526 −0.4540 0.065*
C3B −0.2750 (6) 0.0791 (2) −0.4693 (3) 0.0489 (12)
H3D −0.2443 0.0464 −0.4284 0.059*
C4B −0.1127 (6) 0.1016 (2) −0.4883 (3) 0.0490 (12) C5B 0.0020 (5) 0.1188 (2) −0.3901 (3) 0.0417 (11)
H5B 0.0215 0.0840 −0.3541 0.050*
C6B 0.1782 (6) 0.1380 (2) −0.3928 (3) 0.0635 (15)
H6C 0.1739 0.1763 −0.4141 0.076*
H6D 0.2204 0.1155 −0.4376 0.076*
C7B 0.2961 (5) 0.1331 (3) −0.2939 (3) 0.0643 (16)
H7C 0.4076 0.1456 −0.2968 0.077*
H7D 0.3050 0.0944 −0.2755 0.077*
C8B 0.2397 (5) 0.16637 (19) −0.2169 (3) 0.0426 (11) C9B 0.0475 (5) 0.15561 (17) −0.2245 (2) 0.0345 (10)
H9B 0.0428 0.1168 −0.2051 0.041*
C10B −0.0743 (5) 0.15809 (18) −0.3263 (3) 0.0376 (10) C11B −0.0128 (5) 0.1885 (2) −0.1481 (3) 0.0468 (12) H11C −0.0070 0.2279 −0.1610 0.056* H11D −0.1297 0.1793 −0.1515 0.056* C12B 0.0939 (5) 0.1762 (2) −0.0475 (3) 0.0439 (11)
H12C 0.0583 0.2005 −0.0026 0.053*
H12D 0.0735 0.1383 −0.0309 0.053*
C13B 0.2814 (5) 0.18404 (18) −0.0382 (3) 0.0355 (10)
H13B 0.2968 0.2228 −0.0545 0.043*
C14B 0.3431 (5) 0.14795 (18) −0.1129 (3) 0.0380 (10) C15B 0.5351 (5) 0.1556 (2) −0.1021 (3) 0.0537 (13)
H15C 0.5736 0.1273 −0.1395 0.064*
H15D 0.5535 0.1913 −0.1288 0.064*
C16B 0.6443 (5) 0.1527 (2) 0.0006 (3) 0.0547 (13)
H16C 0.6442 0.1150 0.0242 0.066*
H16D 0.7598 0.1626 0.0013 0.066*
C17B 0.5776 (5) 0.1921 (2) 0.0662 (3) 0.0465 (12) C18B 0.3953 (5) 0.17462 (18) 0.0612 (3) 0.0349 (10)
H18B 0.3973 0.1345 0.0727 0.042*
C19B 0.3177 (5) 0.08591 (18) −0.0951 (3) 0.0481 (12)
H19D 0.3947 0.0747 −0.0368 0.072*
H19E 0.3385 0.0645 −0.1470 0.072*
C20B 0.2710 (6) 0.2288 (2) −0.2339 (3) 0.0671 (16)
H20D 0.2366 0.2368 −0.3008 0.101*
H20E 0.3890 0.2370 −0.2108 0.101*
H20F 0.2069 0.2512 −0.2005 0.101*
C21B −0.1063 (6) 0.21777 (19) −0.3646 (3) 0.0590 (13) H21D −0.2039 0.2183 −0.4173 0.089* H21E −0.0099 0.2306 −0.3854 0.089* H21F −0.1249 0.2416 −0.3150 0.089* C22B 0.3501 (5) 0.20229 (18) 0.1478 (3) 0.0390 (10)
H22B 0.2932 0.2375 0.1270 0.047*
C23B 0.5246 (5) 0.2153 (2) 0.2166 (3) 0.0527 (12)
H23C 0.5441 0.2550 0.2213 0.063*
H23D 0.5300 0.2007 0.2797 0.063*
C24B 0.6561 (5) 0.1867 (2) 0.1722 (3) 0.0551 (13)
H24C 0.7643 0.2054 0.1900 0.066*
H24D 0.6709 0.1481 0.1912 0.066*
C25B 0.2342 (5) 0.16724 (18) 0.1960 (3) 0.0411 (11)
H25B 0.1442 0.1514 0.1454 0.049*
C26B 0.1519 (6) 0.2037 (2) 0.2579 (3) 0.0652 (15)
H26D 0.0817 0.1813 0.2875 0.098*
H26E 0.0843 0.2316 0.2191 0.098*
H26F 0.2379 0.2212 0.3063 0.098*
C27B 0.3324 (5) 0.1198 (2) 0.2512 (3) 0.0425 (11) C28B 0.5975 (6) 0.2518 (2) 0.0357 (4) 0.0666 (15)
H28D 0.7151 0.2598 0.0428 0.100*
H28E 0.5518 0.2766 0.0749 0.100*
H28F 0.5383 0.2565 −0.0297 0.100*
C29B −0.1482 (7) 0.1483 (2) −0.5636 (3) 0.0664 (16) H29D −0.2040 0.1331 −0.6244 0.100* H29E −0.0435 0.1650 −0.5678 0.100* H29F −0.2191 0.1759 −0.5453 0.100* C30B −0.0267 (7) 0.0540 (2) −0.5289 (3) 0.0792 (18) H30D −0.1064 0.0371 −0.5811 0.119*
H30E 0.0133 0.0269 −0.4801 0.119*
H30F 0.0667 0.0684 −0.5510 0.119*
Atomic displacement parameters (Å2)
U11 U22 U33 U12 U13 U23
C22B 0.040 (3) 0.039 (3) 0.034 (2) −0.004 (2) 0.0004 (18) −0.005 (2) C23B 0.047 (3) 0.060 (3) 0.046 (3) −0.011 (2) 0.002 (2) −0.013 (2) C24B 0.034 (3) 0.078 (4) 0.047 (3) −0.009 (3) −0.003 (2) −0.004 (3) C25B 0.033 (2) 0.053 (3) 0.035 (2) 0.004 (2) 0.0053 (19) −0.005 (2) C26B 0.061 (3) 0.079 (4) 0.059 (3) 0.020 (3) 0.020 (2) 0.001 (3) C27B 0.038 (3) 0.053 (3) 0.035 (3) −0.004 (2) 0.005 (2) −0.001 (2) C28B 0.052 (3) 0.076 (4) 0.067 (3) −0.029 (3) 0.005 (2) 0.005 (3) C29B 0.073 (4) 0.097 (4) 0.027 (2) −0.007 (3) 0.007 (2) 0.009 (3) C30B 0.081 (4) 0.103 (5) 0.045 (3) 0.026 (4) −0.001 (3) −0.026 (3)
Geometric parameters (Å, º)
O1A—C3A 1.452 (5) O1B—C3B 1.445 (5)
O1A—H1A 0.82 O1B—H1D 0.82
O2A—C27A 1.234 (6) O2B—C27B 1.236 (5)
O3A—C27A 1.264 (5) O3B—C27B 1.284 (5)
O3A—H3A 0.82 O3B—H3C 0.82
C1A—C10A 1.529 (6) C1B—C2B 1.530 (5)
C1A—C2A 1.532 (6) C1B—C10B 1.540 (5)
C1A—H1B 0.97 C1B—H1E 0.97
C1A—H1C 0.97 C1B—H1F 0.97
C2A—C3A 1.501 (7) C2B—C3B 1.511 (6)
C2A—H2B 0.97 C2B—H2E 0.97
C2A—H2C 0.97 C2B—H2F 0.97
C3A—C4A 1.521 (6) C3B—C4B 1.517 (6)
C3A—H3B 0.98 C3B—H3D 0.98
C4A—C30A 1.536 (7) C4B—C30B 1.534 (6)
C4A—C29A 1.537 (6) C4B—C29B 1.545 (6)
C4A—C5A 1.577 (5) C4B—C5B 1.555 (5)
C5A—C6A 1.518 (6) C5B—C6B 1.522 (6)
C5A—C10A 1.545 (6) C5B—C10B 1.556 (6)
C5A—H5A 0.98 C5B—H5B 0.98
C6A—C7A 1.527 (5) C6B—C7B 1.524 (6)
C6A—H6A 0.97 C6B—H6C 0.97
C6A—H6B 0.97 C6B—H6D 0.97
C7A—C8A 1.520 (6) C7B—C8B 1.531 (6)
C7A—H7A 0.97 C7B—H7C 0.97
C7A—H7B 0.97 C7B—H7D 0.97
C8A—C20A 1.555 (6) C8B—C20B 1.557 (6)
C8A—C9A 1.559 (5) C8B—C9B 1.568 (5)
C8A—C14A 1.588 (5) C8B—C14B 1.599 (5)
C9A—C11A 1.522 (5) C9B—C11B 1.535 (5)
C9A—C10A 1.572 (5) C9B—C10B 1.566 (5)
C9A—H9A 0.98 C9B—H9B 0.98
C10A—C21A 1.552 (6) C10B—C21B 1.541 (6) C11A—C12A 1.529 (5) C11B—C12B 1.533 (6)
C11A—H11A 0.97 C11B—H11C 0.97
C12A—C13A 1.521 (6) C12B—C13B 1.516 (5)
C12A—H12A 0.97 C12B—H12C 0.97
C12A—H12B 0.97 C12B—H12D 0.97
C13A—C18A 1.541 (5) C13B—C18B 1.528 (5) C13A—C14A 1.565 (6) C13B—C14B 1.563 (5)
C13A—H13A 0.98 C13B—H13B 0.98
C14A—C15A 1.548 (6) C14B—C19B 1.541 (6) C14A—C19A 1.557 (6) C14B—C15B 1.549 (6) C15A—C16A 1.540 (6) C15B—C16B 1.538 (5)
C15A—H15A 0.97 C15B—H15C 0.97
C15A—H15B 0.97 C15B—H15D 0.97
C16A—C17A 1.522 (7) C16B—C17B 1.533 (6)
C16A—H16A 0.97 C16B—H16C 0.97
C16A—H16B 0.97 C16B—H16D 0.97
C17A—C24A 1.532 (6) C17B—C24B 1.518 (6) C17A—C18A 1.542 (6) C17B—C28B 1.525 (7) C17A—C28A 1.547 (7) C17B—C18B 1.532 (5) C18A—C22A 1.549 (6) C18B—C22B 1.541 (5)
C18A—H18A 0.98 C18B—H18B 0.98
C19A—H19A 0.96 C19B—H19D 0.96
C19A—H19B 0.96 C19B—H19E 0.96
C19A—H19C 0.96 C19B—H19F 0.96
C20A—H20A 0.96 C20B—H20D 0.96
C20A—H20B 0.96 C20B—H20E 0.96
C20A—H20C 0.96 C20B—H20F 0.96
C21A—H21A 0.96 C21B—H21D 0.96
C21A—H21B 0.96 C21B—H21E 0.96
C21A—H21C 0.96 C21B—H21F 0.96
C22A—C25A 1.538 (6) C22B—C25B 1.554 (6) C22A—C23A 1.569 (6) C22B—C23B 1.559 (6)
C22A—H22A 0.98 C22B—H22B 0.98
C23A—C24A 1.528 (7) C23B—C24B 1.541 (6)
C23A—H23A 0.97 C23B—H23C 0.97
C23A—H23B 0.97 C23B—H23D 0.97
C24A—H24A 0.97 C24B—H24C 0.97
C24A—H24B 0.97 C24B—H24D 0.97
C25A—C27A 1.502 (7) C25B—C27B 1.509 (6) C25A—C26A 1.536 (6) C25B—C26B 1.524 (6)
C25A—H25A 0.98 C25B—H25B 0.98
C26A—H26A 0.96 C26B—H26D 0.96
C26A—H26B 0.96 C26B—H26E 0.96
C26A—H26C 0.96 C26B—H26F 0.96
C28A—H28A 0.96 C28B—H28D 0.96
C28A—H28B 0.96 C28B—H28E 0.96
C28A—H28C 0.96 C28B—H28F 0.96
C29A—H29A 0.96 C29B—H29D 0.96
C29A—H29B 0.96 C29B—H29E 0.96
C30A—H30A 0.96 C30B—H30D 0.96
C30A—H30B 0.96 C30B—H30E 0.96
C30A—H30C 0.96 C30B—H30F 0.96
C3A—O1A—H1A 109.5 C3B—O1B—H1D 109.5
C27A—O3A—H3A 109.5 C27B—O3B—H3C 109.5 C10A—C1A—C2A 113.2 (3) C2B—C1B—C10B 113.9 (3)
C10A—C1A—H1B 108.9 C2B—C1B—H1E 108.8
C2A—C1A—H1B 108.9 C10B—C1B—H1E 108.8
C10A—C1A—H1C 108.9 C2B—C1B—H1F 108.8
C2A—C1A—H1C 108.9 C10B—C1B—H1F 108.8
H1B—C1A—H1C 107.7 H1E—C1B—H1F 107.7
C3A—C2A—C1A 111.6 (4) C3B—C2B—C1B 110.0 (4)
C3A—C2A—H2B 109.3 C3B—C2B—H2E 109.7
C1A—C2A—H2B 109.3 C1B—C2B—H2E 109.7
C3A—C2A—H2C 109.3 C3B—C2B—H2F 109.7
C1A—C2A—H2C 109.3 C1B—C2B—H2F 109.7
H2B—C2A—H2C 108.0 H2E—C2B—H2F 108.2
O1A—C3A—C2A 109.9 (4) O1B—C3B—C2B 109.8 (4) O1A—C3A—C4A 110.2 (4) O1B—C3B—C4B 112.2 (4) C2A—C3A—C4A 115.1 (4) C2B—C3B—C4B 113.7 (4)
O1A—C3A—H3B 107.1 O1B—C3B—H3D 106.9
C2A—C3A—H3B 107.1 C2B—C3B—H3D 106.9
C4A—C3A—H3B 107.1 C4B—C3B—H3D 106.9
C3A—C4A—C30A 108.4 (4) C3B—C4B—C30B 107.7 (4) C3A—C4A—C29A 111.5 (4) C3B—C4B—C29B 111.5 (4) C30A—C4A—C29A 107.9 (4) C30B—C4B—C29B 107.4 (4) C3A—C4A—C5A 107.1 (3) C3B—C4B—C5B 106.8 (3) C30A—C4A—C5A 108.2 (4) C30B—C4B—C5B 108.6 (4) C29A—C4A—C5A 113.6 (4) C29B—C4B—C5B 114.7 (4) C6A—C5A—C10A 111.7 (3) C6B—C5B—C4B 114.9 (3) C6A—C5A—C4A 113.7 (3) C6B—C5B—C10B 110.3 (4) C10A—C5A—C4A 117.6 (4) C4B—C5B—C10B 117.7 (3)
C6A—C5A—H5A 104.0 C6B—C5B—H5B 104.0
C10A—C5A—H5A 104.0 C4B—C5B—H5B 104.0
C4A—C5A—H5A 104.0 C10B—C5B—H5B 104.0
C5A—C6A—C7A 110.6 (3) C5B—C6B—C7B 109.7 (4)
C5A—C6A—H6A 109.5 C5B—C6B—H6C 109.7
C7A—C6A—H6A 109.5 C7B—C6B—H6C 109.7
C5A—C6A—H6B 109.5 C5B—C6B—H6D 109.7
C7A—C6A—H6B 109.5 C7B—C6B—H6D 109.7
H6A—C6A—H6B 108.1 H6C—C6B—H6D 108.2
C8A—C7A—C6A 114.7 (4) C6B—C7B—C8B 114.2 (4)
C8A—C7A—H7A 108.6 C6B—C7B—H7C 108.7
C6A—C7A—H7A 108.6 C8B—C7B—H7C 108.7
C8A—C7A—H7B 108.6 C6B—C7B—H7D 108.7
C6A—C7A—H7B 108.6 C8B—C7B—H7D 108.7
C7A—C8A—C20A 107.4 (3) C7B—C8B—C20B 107.5 (4) C7A—C8A—C9A 109.4 (3) C7B—C8B—C9B 109.6 (3) C20A—C8A—C9A 111.2 (4) C20B—C8B—C9B 110.4 (4) C7A—C8A—C14A 111.0 (3) C7B—C8B—C14B 110.8 (4) C20A—C8A—C14A 109.1 (3) C20B—C8B—C14B 110.3 (3) C9A—C8A—C14A 108.9 (3) C9B—C8B—C14B 108.1 (3) C11A—C9A—C8A 111.3 (3) C11B—C9B—C10B 114.6 (3) C11A—C9A—C10A 114.7 (3) C11B—C9B—C8B 110.8 (3) C8A—C9A—C10A 116.4 (3) C10B—C9B—C8B 117.1 (3) C11A—C9A—H9A 104.2 C11B—C9B—H9B 104.2
C8A—C9A—H9A 104.2 C10B—C9B—H9B 104.2
C10A—C9A—H9A 104.2 C8B—C9B—H9B 104.2
C23A—C24A—C17A 103.5 (4) C17B—C24B—C23B 102.5 (4) C23A—C24A—H24A 111.1 C17B—C24B—H24C 111.3 C17A—C24A—H24A 111.1 C23B—C24B—H24C 111.3 C23A—C24A—H24B 111.1 C17B—C24B—H24D 111.3 C17A—C24A—H24B 111.1 C23B—C24B—H24D 111.3 H24A—C24A—H24B 109.0 H24C—C24B—H24D 109.2 C27A—C25A—C26A 111.3 (4) C27B—C25B—C26B 112.3 (4) C27A—C25A—C22A 112.1 (4) C27B—C25B—C22B 110.3 (3) C26A—C25A—C22A 111.3 (4) C26B—C25B—C22B 110.7 (4) C27A—C25A—H25A 107.3 C27B—C25B—H25B 107.8 C26A—C25A—H25A 107.3 C26B—C25B—H25B 107.8 C22A—C25A—H25A 107.3 C22B—C25B—H25B 107.8 C25A—C26A—H26A 109.5 C25B—C26B—H26D 109.5 C25A—C26A—H26B 109.5 C25B—C26B—H26E 109.5 H26A—C26A—H26B 109.5 H26D—C26B—H26E 109.5 C25A—C26A—H26C 109.5 C25B—C26B—H26F 109.5 H26A—C26A—H26C 109.5 H26D—C26B—H26F 109.5 H26B—C26A—H26C 109.5 H26E—C26B—H26F 109.5 O2A—C27A—O3A 122.9 (5) O2B—C27B—O3B 123.8 (5) O2A—C27A—C25A 119.1 (4) O2B—C27B—C25B 120.2 (4) O3A—C27A—C25A 118.1 (5) O3B—C27B—C25B 116.0 (4) C17A—C28A—H28A 109.5 C17B—C28B—H28D 109.5 C17A—C28A—H28B 109.5 C17B—C28B—H28E 109.5 H28A—C28A—H28B 109.5 H28D—C28B—H28E 109.5 C17A—C28A—H28C 109.5 C17B—C28B—H28F 109.5 H28A—C28A—H28C 109.5 H28D—C28B—H28F 109.5 H28B—C28A—H28C 109.5 H28E—C28B—H28F 109.5 C4A—C29A—H29A 109.5 C4B—C29B—H29D 109.5 C4A—C29A—H29B 109.5 C4B—C29B—H29E 109.5 H29A—C29A—H29B 109.5 H29D—C29B—H29E 109.5 C4A—C29A—H29C 109.5 C4B—C29B—H29F 109.5 H29A—C29A—H29C 109.5 H29D—C29B—H29F 109.5 H29B—C29A—H29C 109.5 H29E—C29B—H29F 109.5 C4A—C30A—H30A 109.5 C4B—C30B—H30D 109.5 C4A—C30A—H30B 109.5 C4B—C30B—H30E 109.5 H30A—C30A—H30B 109.5 H30D—C30B—H30E 109.5 C4A—C30A—H30C 109.5 C4B—C30B—H30F 109.5 H30A—C30A—H30C 109.5 H30D—C30B—H30F 109.5 H30B—C30A—H30C 109.5 H30E—C30B—H30F 109.5
C7A—C8A—C14A—C19A −59.0 (4) C7B—C8B—C14B—C15B 57.8 (5) C20A—C8A—C14A—C19A −177.1 (4) C20B—C8B—C14B—C15B −61.1 (5) C9A—C8A—C14A—C19A 61.4 (4) C9B—C8B—C14B—C15B 178.0 (4) C7A—C8A—C14A—C13A −179.8 (3) C7B—C8B—C14B—C13B 179.1 (4) C20A—C8A—C14A—C13A 62.1 (4) C20B—C8B—C14B—C13B 60.1 (4) C9A—C8A—C14A—C13A −59.4 (4) C9B—C8B—C14B—C13B −60.7 (4) C19A—C14A—C15A—C16A −71.5 (5) C19B—C14B—C15B—C16B −73.1 (5) C13A—C14A—C15A—C16A 46.7 (5) C13B—C14B—C15B—C16B 46.3 (5) C8A—C14A—C15A—C16A 168.0 (4) C8B—C14B—C15B—C16B 166.0 (4) C14A—C15A—C16A—C17A −51.8 (6) C14B—C15B—C16B—C17B −51.9 (6) C15A—C16A—C17A—C24A 168.9 (4) C15B—C16B—C17B—C24B 170.5 (4) C15A—C16A—C17A—C18A 57.6 (5) C15B—C16B—C17B—C28B −65.7 (5) C15A—C16A—C17A—C28A −67.2 (5) C15B—C16B—C17B—C18B 58.7 (5) C12A—C13A—C18A—C17A −173.3 (4) C12B—C13B—C18B—C17B −171.3 (4) C14A—C13A—C18A—C17A 60.1 (5) C14B—C13B—C18B—C17B 61.8 (4) C12A—C13A—C18A—C22A −50.1 (6) C12B—C13B—C18B—C22B −48.2 (5) C14A—C13A—C18A—C22A −176.7 (4) C14B—C13B—C18B—C22B −175.1 (3) C16A—C17A—C18A—C13A −63.2 (5) C24B—C17B—C18B—C13B 172.3 (4) C24A—C17A—C18A—C13A 175.2 (4) C28B—C17B—C18B—C13B 55.4 (5) C28A—C17A—C18A—C13A 59.6 (5) C16B—C17B—C18B—C13B −65.6 (5) C16A—C17A—C18A—C22A 165.1 (4) C24B—C17B—C18B—C22B 41.5 (4) C24A—C17A—C18A—C22A 43.6 (5) C28B—C17B—C18B—C22B −75.4 (4) C28A—C17A—C18A—C22A −72.1 (5) C16B—C17B—C18B—C22B 163.6 (3) C13A—C18A—C22A—C25A 84.3 (5) C13B—C18B—C22B—C25B 89.9 (4) C17A—C18A—C22A—C25A −149.8 (4) C17B—C18B—C22B—C25B −144.0 (3) C13A—C18A—C22A—C23A −150.9 (4) C13B—C18B—C22B—C23B −146.4 (4) C17A—C18A—C22A—C23A −25.0 (5) C17B—C18B—C22B—C23B −20.3 (4) C25A—C22A—C23A—C24A 123.5 (4) C18B—C22B—C23B—C24B −8.2 (5) C18A—C22A—C23A—C24A −3.3 (5) C25B—C22B—C23B—C24B 116.9 (4) C22A—C23A—C24A—C17A 30.4 (5) C28B—C17B—C24B—C23B 75.0 (5) C16A—C17A—C24A—C23A −161.0 (4) C18B—C17B—C24B—C23B −46.0 (4) C18A—C17A—C24A—C23A −45.1 (5) C16B—C17B—C24B—C23B −161.1 (4) C28A—C17A—C24A—C23A 74.3 (5) C22B—C23B—C24B—C17B 33.9 (5) C18A—C22A—C25A—C27A 64.5 (5) C18B—C22B—C25B—C27B 73.3 (4) C23A—C22A—C25A—C27A −55.1 (5) C23B—C22B—C25B—C27B −45.7 (5) C18A—C22A—C25A—C26A −170.1 (4) C18B—C22B—C25B—C26B −161.8 (3) C23A—C22A—C25A—C26A 70.3 (5) C23B—C22B—C25B—C26B 79.2 (4) C26A—C25A—C27A—O2A 148.1 (5) C26B—C25B—C27B—O2B 159.2 (4) C22A—C25A—C27A—O2A −86.5 (6) C22B—C25B—C27B—O2B −76.9 (5) C26A—C25A—C27A—O3A −30.6 (7) C26B—C25B—C27B—O3B −20.4 (6) C22A—C25A—C27A—O3A 94.7 (5) C22B—C25B—C27B—O3B 103.6 (4)
Hydrogen-bond geometry (Å, º)
D—H···A D—H H···A D···A D—H···A
O1A—H1A···O2Bi 0.82 1.91 2.705 (6) 161
O3B—H3C···O1Biv 0.82 1.82 2.625 (5) 163
C18A—H18A···O2A 0.98 2.52 3.262 (6) 132 C18B—H18B···O2B 0.98 2.46 3.206 (5) 133 C23A—H23B···O3A 0.97 2.49 3.222 (7) 132 C23B—H23D···O3B 0.97 2.49 3.237 (6) 134 C30A—H30A···O1A 0.96 2.56 2.957 (7) 105 C30B—H30D···O1B 0.96 2.51 2.917 (7) 105