The evolution of the structure of tubulin and its potential consequences for the role and function of microtubules in cells and embryos
Full text
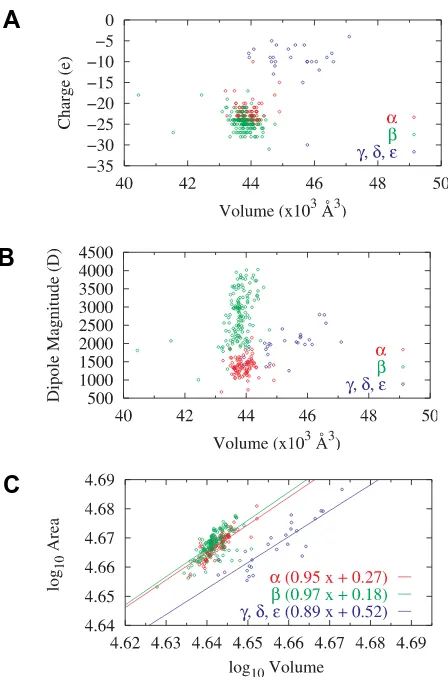
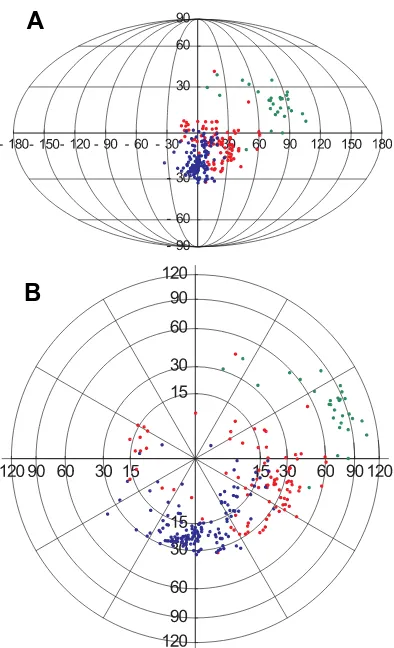
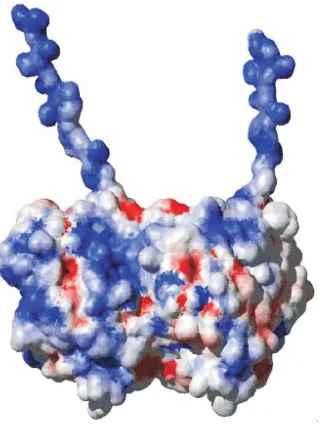
Figure

Related documents
From the above discussions, it is found that Schiff base, 2-[(2-hydroxyphenylimino) methyl]-4-nitrophenol [HPIMNP] is a good sensitive reagent for development of rapid and
The aim of the study was to measure behavioral data (such as eye and head movements) and to assess how the two age groups differ in terms of number of safe street crossings,
It was decided that with the presence of such significant red flag signs that she should undergo advanced imaging, in this case an MRI, that revealed an underlying malignancy, which
ings were observed in the renal tissue of the Cis group with significantly increased levels of oxidants and IL-1β, such as wide hemorrhage areas, dilated and congested blood
19% serve a county. Fourteen per cent of the centers provide service for adjoining states in addition to the states in which they are located; usually these adjoining states have
The high level of coastal litter in the study region adds to the findings of the Report on the State of the Mediterranean Marine and Coastal Environment, which
The objective of this study was to develop Fourier transform infrared (FTIR) spectroscopy in combination with multivariate calibration of partial least square (PLS) and