organic papers
o2972
Zhonget al. C16H7Cl2F3N4O doi:10.1107/S1600536805025699 Acta Cryst.(2005). E61, o2972–o2973 Acta Crystallographica Section E
Structure Reports Online
ISSN 1600-5368
1-[2,6-Dichloro-4-(trifluoromethyl)phenyl]-5-[(2-furyl)methyleneamino]-1
H
-pyrazole-3-carbonitrile
Ping Zhong,* Zhiping Yang,‡ Qian Shi, Maolin Hu, Shuyan Li and Riyuan Tang
Department of Chemistry, Wenzhou Normal College, 325027 Wenzhou, People’s Republic of China
‡ Present address: Zhangzhou Vocational and Technical College, 363000 Zhangzhou, People’s Republic of China
Correspondence e-mail: zhongp0512@163.com
Key indicators
Single-crystal X-ray study T= 298 K
Mean(C–C) = 0.005 A˚ Rfactor = 0.069 wRfactor = 0.204
Data-to-parameter ratio = 12.8
For details of how these key indicators were automatically derived from the article, see http://journals.iucr.org/e.
#2005 International Union of Crystallography Printed in Great Britain – all rights reserved
The title compound, C16H7Cl2F3N4O, is a tricyclic imide with
an overall U-shape. There are –interactions between the pyrazole and furyl rings.
Comment
The title compound, (I), is an important starting material for the synthesis of cyano-1-[2,6-dichloro-4-(trifluoro-methyl)phenyl]- 4-(trifluoromethyl)thiopyrazole, 5-amino-3-
cyano-1-[2,6-dichloro-4-(trifluoromethyl)phenyl]-4-(trifluoro-methylsulfenyl)pyrazole and
5-amino-3-cyano-1-[2,6- dichloro-4-(trifluoromethyl)phenyl]-4-(trifluoromethylsulfon-yl)pyrazole, which are all good insecticides (Hatton et al., 1993).
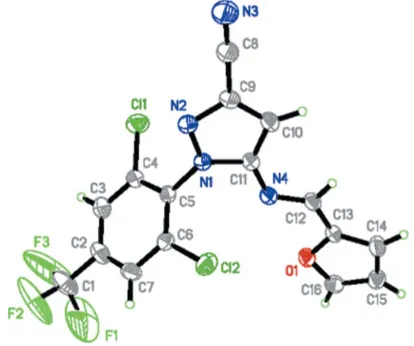
The structure of (I) is shown in Fig. 1, with the atom-numbering scheme. The molecule contains three planar moieties, forming an overall U-shape. The dihedral angles between the pyrazole and the furyl and benzene rings are
[image:1.610.281.382.336.448.2] [image:1.610.227.434.538.711.2]Received 28 February 2005 Accepted 11 August 2005 Online 17 August 2005
Figure 1
19.8 (2) and 67.9 (1), respectively. The plane-to-plane
separation of 3.8411 (1) A˚ between the furyl and pyrazole rings indicates the presence of a weak–interaction. In the crystal structure, the molecules are stacked along thebaxis, as shown in Fig. 2.
Experimental
Following the method of Hattonet al.(1993), reaction of 2,6-dichloro-4-(trifluoromethyl)amine with a suspension of nitrosyl sulfuric acid, followed by reaction with a solution of ethyl 2,3-dicyanopropionate in acetic acid, gave 5-amino-3-cyano-1-[2,6-dichloro-4-(trifluorometh-yl)phenyl]pyrazole, which was then reacted with 2-furanal to give (I). Single crystals suitable for X-ray analysis were obtained by slow evaporation of a methanol solution (m.p. 449–451 K). IR (KBr,
cm1): 3129, 2240, 1611, 1558, 1395, 1310, 1133, 873, 818;1H NMR (CDCl3):8.81 (s, 1H), 8.12 (s, 2H), 7.83 (s, 1H), 7.24 (m, 2H), 6.69
(m, 1H);13C NMR (CDCl3): 154.2 (1C), 153.3 (1C), 152.1 (1C), 149.2
(1C), 136.6 (1C), 134.4 (q, J= 34.3 Hz, 1C), 128.2 (1C), 127.05 (1C), 127.01 (1C), 126.95 (1C), 126.91 (1C), 123.3 (q, J= 271.6 Hz, 1C), 122.0 (1C), 114.2 (1C), 114.0 (1C), 98.4 (1C).
Crystal data
C16H7Cl2F3N4O
Mr= 399.16 Monoclinic,P21=n
a= 11.8828 (9) A˚
b= 6.7072 (5) A˚
c= 21.1191 (16) A˚ = 92.084 (1) V= 1682.1 (2) A˚3
Z= 4
Dx= 1.576 Mg m
3 MoKradiation Cell parameters from 3298
reflections = 3.2–25.0
= 0.43 mm1
T= 298 (2) K Block, colorless 0.380.310.29 mm
Data collection
Bruker APEX area-detector diffractometer
’and!scans
Absorption correction: multi-scan (SADABS; Bruker, 2002)
Tmin= 0.854,Tmax= 0.885 8571 measured reflections
3013 independent reflections 2571 reflections withI> 2(I)
Rint= 0.019
max= 25.2
h=13!14
k=8!6
l=24!25
Refinement
Refinement onF2
R[F2> 2(F2)] = 0.069
wR(F2) = 0.204
S= 1.05 3013 reflections 235 parameters
H-atom parameters constrained
w= 1/[2(F
o2) + (0.1164P)2 + 2.0094P]
whereP= (Fo2+ 2Fc2)/3 (/)max< 0.001
max= 1.12 e A˚
3
min=0.64 e A˚
3
Table 1
Selected geometric parameters (A˚ ,).
Cl1—C4 1.724 (4) F1—C1 1.271 (9) O1—C16 1.352 (4) O1—C13 1.362 (4) N1—N2 1.348 (4) N1—C11 1.374 (4) N2—C9 1.337 (5) N3—C8 1.147 (5)
N4—C12 1.273 (4) N4—C11 1.382 (4) C9—C10 1.389 (5) C10—C11 1.374 (5) C13—C14 1.350 (5) C14—C15 1.403 (5) C15—C16 1.340 (6)
C16—O1—C13 106.2 (3) N2—N1—C11 113.0 (3) C9—N2—N1 103.2 (3) F3—C1—F2 111.0 (7) N3—C8—C9 179.1 (5) N2—C9—C10 113.1 (3)
C11—C10—C9 104.9 (3) C10—C11—N1 105.7 (3) C14—C13—O1 109.3 (3) C13—C14—C15 107.6 (3) C16—C15—C14 105.6 (3) C15—C16—O1 111.4 (3)
All H atom were initially observed in a difference Fourier map and were then placed in geometrically idealized positions and constrained to ride on their parent atoms, with C—H = 0.93 A˚ andUiso(H) =
1.22eq(C). The low Ueqvalue of atom C1 compared with its
neigh-bours may be attributed to the three possibly disordered F atoms. The highest peak is located 1.27 A˚ from atoms C1 and F.
Data collection:SMART(Bruker, 2002); cell refinement:SAINT (Bruker, 2002); data reduction:SAINT; program(s) used to solve structure:SHELXS97(Sheldrick, 1997); program(s) used to refine structure: SHELXL97 (Sheldrick, 1997); molecular graphics: XP (Bruker, 2002); software used to prepare material for publication: SHELXL97.
This work was supported by the National Nature Science Foundation of China (No. 20272043) and the Nature Science Foundation of Zhejiang Province (No. M203001).
References
Bruker (2002).SMART,SAINT,SADABS,SHELXLandXP. Bruker AXS Inc., Madison, Wisconsin, USA.
Hatton, L. R., Buntain, I. G., Hawkins, D. W., Parnell, E. W., Pearson C. J. & Roberts, D. A. (1993).US Patent No.5232940.
[image:2.610.44.291.69.307.2]Sheldrick, G. M. (1997). SHELXL97 and SHELXS97. University of Go¨ttingen, Germany.
Figure 2
supporting information
sup-1
Acta Cryst. (2005). E61, o2972–o2973
supporting information
Acta Cryst. (2005). E61, o2972–o2973 [https://doi.org/10.1107/S1600536805025699]
1-[2,6-Dichloro-4-(trifluoromethyl)phenyl]-5-[(2-furyl)methyleneamino]-1
H
-pyrazole-3-carbonitrile
Ping Zhong, Zhiping Yang, Qian Shi, Maolin Hu, Shuyan Li and Riyuan Tang
1-[2,6-Dichloro-4-(trifluoromethyl)phenyl]-5-[(2-furyl)methyleneamino]- 1H-pyrazole-3-carbonitrile
Crystal data
C16H7Cl2F3N4O
Mr = 399.16 Monoclinic, P21/n
Hall symbol: -P 2yn
a = 11.8828 (9) Å
b = 6.7072 (5) Å
c = 21.1191 (16) Å
β = 92.084 (1)°
V = 1682.1 (2) Å3
Z = 4
F(000) = 800
Dx = 1.576 Mg m−3
Melting point: 450(1) K Mo Kα radiation, λ = 0.71073 Å Cell parameters from 3298 reflections
θ = 3.2–25.0°
µ = 0.43 mm−1
T = 298 K Block, colorless 0.38 × 0.31 × 0.29 mm
Data collection
Bruker APEX area-detector diffractometer
Radiation source: fine-focus sealed tube Graphite monochromator
φ and ω scans
Absorption correction: multi-scan (SADABS; Bruker, 2002)
Tmin = 0.854, Tmax = 0.885
8571 measured reflections 3013 independent reflections 2571 reflections with I > 2σ(I)
Rint = 0.019
θmax = 25.2°, θmin = 1.9°
h = −13→14
k = −8→6
l = −24→25
Refinement
Refinement on F2
Least-squares matrix: full
R[F2 > 2σ(F2)] = 0.069
wR(F2) = 0.204
S = 1.05 3013 reflections 235 parameters 0 restraints
Primary atom site location: structure-invariant direct methods
Secondary atom site location: difference Fourier map
Hydrogen site location: inferred from neighbouring sites
H-atom parameters constrained
w = 1/[σ2(F
o2) + (0.1164P)2 + 2.0094P]
where P = (Fo2 + 2Fc2)/3
(Δ/σ)max < 0.001
Δρmax = 1.12 e Å−3
Special details
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes.
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2,
conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used
only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2
are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger.
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
x y z Uiso*/Ueq
Cl1 0.58604 (10) 0.09721 (15) 0.17137 (5) 0.0678 (4)
Cl2 0.60827 (9) 0.75002 (15) 0.02417 (5) 0.0637 (4)
F1 0.2558 (5) 0.7800 (15) 0.1718 (6) 0.293 (6)
F2 0.1880 (3) 0.6094 (13) 0.1130 (2) 0.206 (3)
F3 0.2255 (5) 0.5229 (17) 0.2002 (4) 0.308 (7)
O1 0.8328 (2) 0.9534 (4) 0.21212 (12) 0.0528 (6)
N1 0.6953 (2) 0.3706 (4) 0.08051 (14) 0.0438 (7)
N2 0.7008 (2) 0.2090 (4) 0.04282 (14) 0.0485 (7)
N3 0.8761 (4) −0.1086 (6) −0.0323 (2) 0.0843 (13)
N4 0.8054 (2) 0.6190 (4) 0.13362 (14) 0.0450 (7)
C1 0.2601 (4) 0.5992 (10) 0.1540 (3) 0.0871 (16)
C2 0.3750 (3) 0.5423 (7) 0.13571 (18) 0.0580 (10)
C3 0.4202 (3) 0.3669 (6) 0.15846 (19) 0.0561 (9)
H3 0.3797 0.2856 0.1851 0.067*
C4 0.5269 (3) 0.3135 (5) 0.14114 (17) 0.0466 (8)
C5 0.5876 (3) 0.4327 (5) 0.10036 (16) 0.0412 (7)
C6 0.5386 (3) 0.6063 (5) 0.07742 (17) 0.0455 (8)
C7 0.4320 (3) 0.6633 (6) 0.09581 (19) 0.0560 (9)
H7 0.3999 0.7819 0.0811 0.067*
C8 0.8478 (3) 0.0251 (6) −0.0030 (2) 0.0597 (10)
C9 0.8111 (3) 0.1898 (6) 0.03400 (17) 0.0499 (8)
C10 0.8752 (3) 0.3377 (6) 0.06397 (18) 0.0513 (9)
H10 0.9529 0.3542 0.0636 0.062*
C11 0.7988 (3) 0.4543 (5) 0.09428 (16) 0.0430 (7)
C12 0.8959 (3) 0.7205 (5) 0.13453 (16) 0.0456 (8)
H12 0.9534 0.6794 0.1088 0.055*
C13 0.9146 (3) 0.8937 (5) 0.17273 (16) 0.0437 (8)
C14 1.0007 (3) 1.0250 (6) 0.17631 (19) 0.0568 (10)
H14 1.0670 1.0174 0.1544 0.068*
C15 0.9719 (3) 1.1751 (6) 0.21907 (18) 0.0545 (9)
H15 1.0145 1.2862 0.2309 0.065*
C16 0.8701 (3) 1.1248 (6) 0.23926 (19) 0.0551 (9)
supporting information
sup-3
Acta Cryst. (2005). E61, o2972–o2973
Atomic displacement parameters (Å2)
U11 U22 U33 U12 U13 U23
Cl1 0.0748 (7) 0.0525 (6) 0.0762 (7) 0.0074 (5) 0.0035 (5) 0.0156 (5)
Cl2 0.0673 (7) 0.0498 (6) 0.0738 (7) −0.0100 (4) −0.0003 (5) 0.0125 (4)
F1 0.102 (4) 0.302 (9) 0.480 (15) 0.047 (5) 0.068 (6) −0.226 (11)
F2 0.0508 (19) 0.412 (10) 0.155 (4) 0.073 (4) −0.001 (2) −0.024 (5)
F3 0.159 (5) 0.473 (14) 0.304 (8) 0.202 (7) 0.174 (6) 0.256 (10)
O1 0.0416 (13) 0.0535 (15) 0.0642 (15) 0.0001 (11) 0.0142 (11) −0.0077 (12)
N1 0.0393 (15) 0.0376 (15) 0.0547 (16) 0.0007 (11) 0.0041 (12) −0.0068 (12)
N2 0.0458 (16) 0.0419 (16) 0.0579 (17) 0.0016 (12) 0.0053 (13) −0.0091 (13)
N3 0.080 (3) 0.074 (3) 0.102 (3) 0.000 (2) 0.034 (2) −0.034 (2)
N4 0.0385 (14) 0.0428 (16) 0.0538 (16) 0.0012 (12) 0.0023 (12) −0.0051 (13)
C1 0.056 (3) 0.124 (5) 0.082 (3) 0.034 (3) 0.017 (3) −0.004 (3)
C2 0.0409 (19) 0.079 (3) 0.054 (2) 0.0093 (18) 0.0019 (16) −0.009 (2)
C3 0.046 (2) 0.066 (2) 0.056 (2) −0.0039 (17) 0.0063 (16) −0.0006 (18)
C4 0.0426 (18) 0.0459 (19) 0.0511 (19) 0.0003 (15) −0.0008 (14) −0.0011 (15)
C5 0.0346 (16) 0.0391 (17) 0.0497 (18) 0.0000 (13) 0.0009 (13) −0.0075 (14)
C6 0.0439 (18) 0.0389 (18) 0.0536 (19) −0.0037 (14) −0.0027 (15) −0.0060 (15)
C7 0.051 (2) 0.050 (2) 0.066 (2) 0.0145 (17) −0.0075 (17) −0.0089 (18)
C8 0.055 (2) 0.056 (2) 0.069 (2) 0.0019 (18) 0.0138 (18) −0.010 (2)
C9 0.0489 (19) 0.047 (2) 0.054 (2) 0.0075 (16) 0.0067 (15) −0.0052 (16)
C10 0.0387 (17) 0.055 (2) 0.061 (2) 0.0033 (16) 0.0083 (15) −0.0065 (17)
C11 0.0367 (16) 0.0439 (18) 0.0487 (18) 0.0017 (13) 0.0036 (13) −0.0011 (14)
C12 0.0354 (16) 0.051 (2) 0.0507 (19) 0.0041 (14) 0.0065 (14) −0.0046 (15)
C13 0.0349 (16) 0.0479 (19) 0.0486 (18) 0.0029 (13) 0.0051 (13) −0.0006 (15)
C14 0.0438 (19) 0.062 (2) 0.066 (2) −0.0085 (17) 0.0127 (16) −0.0113 (19)
C15 0.054 (2) 0.051 (2) 0.058 (2) −0.0050 (17) 0.0017 (16) −0.0067 (17)
C16 0.056 (2) 0.051 (2) 0.059 (2) 0.0043 (17) 0.0096 (17) −0.0107 (17)
Geometric parameters (Å, º)
Cl1—C4 1.724 (4) C3—H3 0.9300
Cl2—C6 1.716 (4) C4—C5 1.395 (5)
F1—C1 1.271 (9) C5—C6 1.382 (5)
F2—C1 1.198 (7) C6—C7 1.393 (5)
F3—C1 1.188 (7) C7—H7 0.9300
O1—C16 1.352 (4) C8—C9 1.430 (5)
O1—C13 1.362 (4) C9—C10 1.389 (5)
N1—N2 1.348 (4) C10—C11 1.374 (5)
N1—C11 1.374 (4) C10—H10 0.9300
N1—C5 1.423 (4) C12—C13 1.427 (5)
N2—C9 1.337 (5) C12—H12 0.9300
N3—C8 1.147 (5) C13—C14 1.350 (5)
N4—C12 1.273 (4) C14—C15 1.403 (5)
N4—C11 1.382 (4) C14—H14 0.9300
C1—C2 1.483 (6) C15—C16 1.340 (6)
C2—C3 1.373 (6) C16—H16 0.9300
C3—C4 1.380 (5)
C16—O1—C13 106.2 (3) C2—C7—H7 120.5
N2—N1—C11 113.0 (3) C6—C7—H7 120.5
N2—N1—C5 118.2 (3) N3—C8—C9 179.1 (5)
C11—N1—C5 128.7 (3) N2—C9—C10 113.1 (3)
C9—N2—N1 103.2 (3) N2—C9—C8 118.1 (3)
C12—N4—C11 117.7 (3) C10—C9—C8 128.8 (3)
F3—C1—F2 111.0 (7) C11—C10—C9 104.9 (3)
F3—C1—F1 98.6 (8) C11—C10—H10 127.5
F2—C1—F1 97.1 (7) C9—C10—H10 127.5
F3—C1—C2 116.9 (5) C10—C11—N1 105.7 (3)
F2—C1—C2 117.8 (5) C10—C11—N4 135.3 (3)
F1—C1—C2 111.8 (6) N1—C11—N4 118.9 (3)
C7—C2—C3 121.9 (3) N4—C12—C13 123.9 (3)
C7—C2—C1 119.3 (4) N4—C12—H12 118.0
C3—C2—C1 118.8 (4) C13—C12—H12 118.0
C2—C3—C4 118.8 (4) C14—C13—O1 109.3 (3)
C2—C3—H3 120.6 C14—C13—C12 131.6 (3)
C4—C3—H3 120.6 O1—C13—C12 119.0 (3)
C3—C4—C5 121.0 (3) C13—C14—C15 107.6 (3)
C3—C4—Cl1 119.1 (3) C13—C14—H14 126.2
C5—C4—Cl1 119.9 (3) C15—C14—H14 126.2
C6—C5—C4 118.6 (3) C16—C15—C14 105.6 (3)
C6—C5—N1 121.0 (3) C16—C15—H15 127.2
C4—C5—N1 120.3 (3) C14—C15—H15 127.2
C5—C6—C7 120.6 (3) C15—C16—O1 111.4 (3)
C5—C6—Cl2 119.7 (3) C15—C16—H16 124.3
C7—C6—Cl2 119.6 (3) O1—C16—H16 124.3