organic papers
o2022
Wanget al. C30H48O2 doi:10.1107/S1600536805017216 Acta Cryst.(2005). E61, o2022–o2023
Acta Crystallographica Section E
Structure Reports
Online
ISSN 1600-5368
6a-Hydroxy-12-oleanen-3-one: a triterpenoid
from the stalks of
Celastrus hypoleucus
Kui-Wu Wang, Cui-Rong Sun, Xiao-Ji Cao and Yuan-Jiang Pan*
Department of Chemistry, Zhejiang University, Hangzhou 310027, People’s Republic of China
Correspondence e-mail: cheyjpan@css.zju.edu.cn
Key indicators
Single-crystal X-ray study
T= 293 K
Mean(C–C) = 0.003 A˚
Rfactor = 0.037
wRfactor = 0.107 Data-to-parameter ratio = 9.7
For details of how these key indicators were automatically derived from the article, see http://journals.iucr.org/e.
#2005 International Union of Crystallography Printed in Great Britain – all rights reserved
The title compound, C30H48O2, is a triterpenoid which was
isolated from the stalks ofCelastrus hypoleucus(Oliv.)Warb. The molecule contains five six-membered rings adopting distorted boat, half-chair and chair conformations. The hydroxy groups serve as hydrogen-bond donors and the carbonyl groups as acceptors, forming molecular chains parallel to thebaxis.
Comment
Celastraceae plants have been the subject of continued and growing interest, due to the range of biological activities shown by many members of this family (Bruning & Wagner, 1978; Tu, 1990, 1991; Jianget al., 1996). Pharmaceutical studies and clinical practice have demonstrated that sesquiterpenes and triterpenes possess notable antibacterial, antitumour, insect antifeedant and cytoxic activities (Chen & Liang, 1999). Our investigation of the bioactive constituents of Celastrus hypoleucus(Oliv.) Warb., a perennial plant belonging to the Celastraceae family, led to the isolation of 6 -hydroxyl-12-oleanene-3-one, (I). The structure of (I) was elucidated by spectroscopic analysis, including two-dimensional NMR spectroscopy, and was confirmed by single-crystal X-ray diffraction analysis, the results of which are presented here.
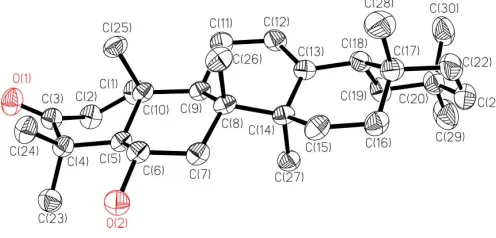
The molecular structure of (I) and the atom-numbering scheme are shown in Fig. 1. The molecule contains five six-membered rings (A atoms C1–C5/C10, BC5–C10, CC8/C9/ C11–C14, D C13–C18 and E C17–C22). Rings B, D and E
adopt chair conformations, while ring A adopts a slightly distorted boat conformation and ring C a slightly distorted half-chair conformation, as a result of the C3 carbonyl group and the C12 C13 double bond, respectively. All rings are
trans fused, except for the D/E junction, which is cis. The orientation of the hydroxy group is axial.
In the crystal structure of (I), the hydroxy groups serve as hydrogen-bond donors and the carbonyl groups as acceptors, forming molecular chains running parallel to thebaxis.
Experimental
Stalks ofCelastrus hypoleucus(Oliv.) Warb. were collected in Jiujiang city, Jiangxi Province, China, in September 2002. The shade-dried powder of the stalks (10 kg) was extracted at room temerature three times with methanol (3 20 l). The extracts were evaporatedin vacuo affording a gummy residue (514 g). This residue was parti-tioned in H2O and extracted at room temerature with petroleum ether (4 3000 ml). The petroleum ether extract (103 g) was adsorbed on to silica gel (100 g) and then subjected to column chromatography (silica gel, 1 kg, 200–300 mesh), eluted with petro-leum ether–AcOEt (gradients 10:0–0:10). The eluted fractions were evaluated by thin-layer chromatography and combined to give 16 main fractions. Fraction 8 (4 g) was rechromatographed on a silica-gel (80 g) column with petroleum ether–acetone (5:1) to afford the pure title compound, (I) (m. p. 492–494 K).13C NMR (125 MHz, CDCl
3,, p.p.m.): 219.6 (C3), 144.8 (C13), 121.9 (C12), 68.0 (C6), 59.0 (C5), 47.5 (C9), 47.4 (C4), 47.0 (C19), 45.9 (C18), 43.4 (C7), 42.3 (C8), 41.0 (C14), 39.2 (C1), 38.3 (C10), 37.3 (C22), 34.9 (C21), 33.5 (C29), 33.3 (C2), 32.8 (C17), 32.0 (C23), 31.3 (C20), 28.7 (C28), 27.2 (C16), 26.3 (C15), 26.0 (C27), 23.9 (C11), 23.9 (C30), 20.4 (C24), 17.5 (C25), 17.5 (C26). Crystals of (I) suitable for X-ray structure analysis were obtained by slow evaporation of a methanol solution at room temperature.
Crystal data
C30H48O2 Mr= 440.68
Orthorhombic,P212121 a= 11.5254 (14) A˚
b= 14.5357 (18) A˚
c= 15.769 (2) A˚
V= 2641.7 (6) A˚3
Z= 4
Dx= 1.108 Mg m
3
MoKradiation Cell parameters from 3643
reflections
= 2.2–21.5 = 0.07 mm1
T= 293 (2) K Prism, colourless 0.460.400.33 mm
Refinement
Refinement onF2 R[F2> 2(F2)] = 0.037 wR(F2) = 0.107
S= 1.07 2915 reflections 299 parameters
H-atom parameters constrained
w= 1/[2
(Fo2) + (0.0518P)2 + 0.1992P]
whereP= (Fo2+ 2Fc2)/3 (/)max< 0.001
max= 0.13 e A˚
3
min=0.10 e A˚
3
Extinction correction:SHELXL97
(Sheldrick, 1997)
[image:2.610.45.293.73.192.2]Extinction coefficient: 0.0025 (6)
Table 1
Selected geometric parameters (A˚ ,).
O1—C3 1.215 (3) O2—C6 1.432 (3) C2—C3 1.491 (3)
C3—C4 1.524 (3) C12—C13 1.330 (3) C3—C2—C1 112.2 (2)
O1—C3—C2 122.3 (2) O1—C3—C4 122.3 (2) C2—C3—C4 115.3 (2) C3—C4—C24 109.9 (2)
C3—C4—C23 103.8 (2) O2—C6—C7 109.01 (17) O2—C6—C5 108.53 (18) C7—C6—C5 110.97 (17) C1—C2—C3—O1 121.6 (3)
C1—C2—C3—C4 60.7 (3) O1—C3—C4—C24 24.4 (3) C2—C3—C4—C24 157.9 (2) O1—C3—C4—C23 90.4 (3) C2—C3—C4—C23 87.3 (2) O1—C3—C4—C5 152.0 (2)
C2—C3—C4—C5 30.4 (3) C3—C4—C5—C6 159.00 (19) C10—C5—C6—O2 178.24 (17) C4—C5—C6—O2 52.2 (2) C10—C5—C6—C7 58.5 (2) C4—C5—C6—C7 171.98 (18) O2—C6—C7—C8 178.33 (17)
Table 2
Hydrogen-bond geometry (A˚ ,).
D—H A D—H H A D A D—H A
O2—H2O O1i
0.82 2.08 2.901 (2) 175
Symmetry code: (i)xþ1;y1 2;zþ
3 2.
H atoms were placed in calculated positions, with C—H = 0.93– 0.97 A˚ and O—H = 0.82 A˚, and refined as riding atoms, withUiso(H) set equal to 1.2Ueq(C) or 1.5Ueq(O). The absolute configuration could not be determined from the X-ray analysis, as no strong anomalous scatterers are present; 2242 Friedel pairs were therefore merged before refinement.
Data collection:SMART(Bruker, 1999); cell refinement:SAINT
(Bruker, 1999); data reduction:SAINT; program(s) used to solve structure:SHELXS97(Sheldrick, 1997); program(s) used to refine structure: SHELXL97 (Sheldrick, 1997); molecular graphics:
SHELXTL (Bruker, 1998); software used to prepare material for publication:SHELXTL.
This work was supported by the Natural Science Founda-tion of China (grant No. 20375036).
References
Bruker (1998). SHELXTL. Version 5.10. Bruker AXS Inc., Madison, Wisconsin, USA.
[image:2.610.314.567.206.345.2]Bruker (1999).SMARTandSAINT. Bruker AXS Inc., Madison, Wisconsin,
Figure 1
supporting information
sup-1 Acta Cryst. (2005). E61, o2022–o2023
supporting information
Acta Cryst. (2005). E61, o2022–o2023 [https://doi.org/10.1107/S1600536805017216]
6
α
-Hydroxy-12-oleanen-3-one: a triterpenoid from the stalks of
Celastrus
hypoleucus
Kui-Wu Wang, Cui-Rong Sun, Xiao-Ji Cao and Yuan-Jiang Pan
6α-Hydroxy-12-oleanen-3-one
Crystal data
C30H48O2
Mr = 440.68
Orthorhombic, P212121
Hall symbol: P 2ac 2ab
a = 11.5254 (14) Å
b = 14.5357 (18) Å
c = 15.769 (2) Å
V = 2641.7 (6) Å3
Z = 4
F(000) = 976
Dx = 1.108 Mg m−3
Mo Kα radiation, λ = 0.71073 Å Cell parameters from 3643 reflections
θ = 2.2–21.5°
µ = 0.07 mm−1
T = 293 K Prism, colourless 0.46 × 0.40 × 0.33 mm
Data collection
Bruker SMART CCD area-detector diffractometer
Radiation source: fine-focus sealed tube Graphite monochromator
φ and ω scans
14520 measured reflections 2915 independent reflections
2407 reflections with I > 2σ(I)
Rint = 0.031
θmax = 26.0°, θmin = 1.9°
h = −14→14
k = −8→17
l = −19→19
Refinement
Refinement on F2
Least-squares matrix: full
R[F2 > 2σ(F2)] = 0.037
wR(F2) = 0.107
S = 1.07 2915 reflections 299 parameters 0 restraints
Primary atom site location: structure-invariant direct methods
Secondary atom site location: difference Fourier map
Hydrogen site location: inferred from neighbouring sites
H-atom parameters constrained
w = 1/[σ2(F
o2) + (0.0518P)2 + 0.1992P]
where P = (Fo2 + 2Fc2)/3
(Δ/σ)max < 0.001
Δρmax = 0.13 e Å−3
Δρmin = −0.10 e Å−3
Extinction correction: SHELXL97 (Sheldrick, 1997), Fc*=kFc[1+0.001xFc2λ3/sin(2θ)]-1/4
Special details
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes.
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2,
conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used
only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2
are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger.
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
x y z Uiso*/Ueq
O1 0.50968 (19) 0.79481 (12) 0.82828 (11) 0.0785 (5) O2 0.39668 (17) 0.47970 (10) 0.68236 (11) 0.0678 (5) H2O 0.4191 0.4264 0.6779 0.102* C1 0.4647 (3) 0.80461 (15) 0.61170 (16) 0.0682 (7) H1A 0.5264 0.8487 0.6019 0.082* H1B 0.4059 0.8145 0.5686 0.082* C2 0.4114 (2) 0.82328 (16) 0.69866 (15) 0.0669 (7) H2A 0.4239 0.8873 0.7136 0.080* H2B 0.3283 0.8129 0.6958 0.080* C3 0.4621 (2) 0.76347 (16) 0.76592 (15) 0.0584 (6) C4 0.4458 (2) 0.66042 (15) 0.75275 (14) 0.0544 (5) C5 0.44156 (18) 0.63853 (13) 0.65508 (13) 0.0482 (5)
H5 0.3608 0.6484 0.6378 0.058*
C6 0.4693 (2) 0.53857 (13) 0.63255 (14) 0.0518 (5)
H6 0.5509 0.5258 0.6455 0.062*
C7 0.4468 (2) 0.52001 (14) 0.53911 (13) 0.0534 (5) H7A 0.3648 0.5290 0.5281 0.064* H7B 0.4645 0.4560 0.5276 0.064* C8 0.51612 (18) 0.58009 (13) 0.47699 (14) 0.0473 (5) C9 0.50315 (19) 0.68179 (12) 0.50502 (13) 0.0483 (5)
H9 0.4233 0.6985 0.4902 0.058*
supporting information
sup-3 Acta Cryst. (2005). E61, o2022–o2023
Atomic displacement parameters (Å2)
U11 U22 U33 U12 U13 U23
O1 0.1128 (15) 0.0545 (9) 0.0683 (10) −0.0094 (10) −0.0166 (11) −0.0097 (8) O2 0.0940 (12) 0.0396 (8) 0.0697 (10) −0.0122 (9) −0.0022 (9) 0.0085 (8) C1 0.0982 (19) 0.0355 (11) 0.0710 (15) 0.0029 (12) −0.0052 (14) 0.0007 (11) C2 0.0828 (17) 0.0419 (12) 0.0760 (16) 0.0101 (12) −0.0044 (14) −0.0038 (11) C3 0.0650 (14) 0.0471 (12) 0.0631 (13) −0.0011 (11) −0.0011 (12) −0.0030 (11) C4 0.0614 (13) 0.0418 (11) 0.0599 (12) −0.0030 (10) −0.0062 (11) 0.0001 (10) C5 0.0493 (11) 0.0359 (10) 0.0593 (12) 0.0001 (9) −0.0086 (9) 0.0015 (9) C6 0.0599 (13) 0.0337 (10) 0.0620 (13) −0.0010 (9) −0.0071 (11) 0.0067 (9) C7 0.0656 (13) 0.0296 (9) 0.0651 (13) −0.0025 (10) −0.0070 (11) 0.0012 (9) C8 0.0451 (10) 0.0339 (10) 0.0631 (12) −0.0001 (9) −0.0047 (10) 0.0010 (9) C9 0.0507 (11) 0.0334 (9) 0.0607 (12) −0.0026 (9) −0.0040 (10) 0.0040 (9) C10 0.0564 (12) 0.0316 (9) 0.0613 (12) −0.0049 (9) −0.0076 (10) 0.0011 (9) C11 0.0857 (17) 0.0421 (12) 0.0710 (14) −0.0181 (12) 0.0024 (13) 0.0009 (11) C12 0.0749 (15) 0.0423 (12) 0.0649 (13) −0.0109 (11) 0.0088 (12) 0.0038 (10) C13 0.0496 (11) 0.0400 (10) 0.0634 (12) 0.0020 (9) 0.0036 (10) 0.0000 (10) C14 0.0463 (11) 0.0369 (10) 0.0627 (13) −0.0004 (9) −0.0006 (10) 0.0000 (9) C15 0.0727 (15) 0.0386 (11) 0.0799 (16) −0.0022 (11) −0.0083 (13) −0.0063 (11) C16 0.0727 (16) 0.0462 (12) 0.0799 (16) −0.0022 (12) −0.0013 (14) −0.0134 (12) C17 0.0615 (14) 0.0572 (14) 0.0796 (16) 0.0000 (12) 0.0089 (12) −0.0195 (13) C18 0.0642 (14) 0.0518 (12) 0.0639 (14) −0.0076 (12) 0.0147 (11) −0.0085 (11) C19 0.0849 (17) 0.0651 (15) 0.0578 (13) 0.0067 (14) 0.0069 (13) 0.0018 (12) C20 0.093 (2) 0.0814 (19) 0.0578 (14) −0.0102 (16) 0.0082 (14) 0.0015 (14) C21 0.103 (2) 0.095 (2) 0.0653 (16) −0.0250 (19) 0.0050 (16) −0.0129 (15) C22 0.104 (2) 0.0766 (18) 0.0804 (18) −0.0126 (19) 0.0229 (18) −0.0274 (16) C23 0.0774 (16) 0.0710 (16) 0.0704 (15) −0.0080 (14) 0.0040 (13) 0.0004 (14) C24 0.0870 (18) 0.0547 (14) 0.0681 (15) 0.0038 (13) −0.0186 (14) 0.0018 (12) C25 0.0633 (14) 0.0671 (15) 0.0754 (15) −0.0175 (12) −0.0100 (12) −0.0024 (13) C26 0.0563 (13) 0.0582 (14) 0.0840 (17) 0.0108 (11) −0.0095 (12) −0.0035 (13) C27 0.0491 (12) 0.0725 (15) 0.0643 (14) 0.0023 (11) −0.0030 (11) 0.0035 (12) C28 0.0663 (16) 0.099 (2) 0.128 (3) 0.0157 (17) 0.0121 (17) −0.036 (2) C29 0.123 (3) 0.159 (4) 0.0707 (19) 0.017 (3) −0.0072 (18) 0.019 (2) C30 0.162 (3) 0.092 (2) 0.0694 (17) −0.041 (2) 0.028 (2) −0.0069 (16)
Geometric parameters (Å, º)
O1—C3 1.215 (3) C16—H16A 0.9700
O2—C6 1.432 (3) C16—H16B 0.9700
O2—H2O 0.8200 C17—C28 1.533 (4)
C1—C2 1.527 (3) C17—C22 1.539 (4)
C1—C10 1.558 (3) C17—C18 1.540 (3)
supporting information
sup-5 Acta Cryst. (2005). E61, o2022–o2023
C3—C4 1.524 (3) C20—C29 1.513 (5)
C4—C24 1.528 (3) C20—C21 1.528 (4)
C4—C23 1.550 (4) C20—C30 1.549 (4)
C4—C5 1.573 (3) C21—C22 1.515 (5)
C5—C6 1.530 (3) C21—H21A 0.9700
C5—C10 1.547 (3) C21—H21B 0.9700
C5—H5 0.9800 C22—H22A 0.9700
C6—C7 1.520 (3) C22—H22B 0.9700
C6—H6 0.9800 C23—H23A 0.9600
C7—C8 1.536 (3) C23—H23B 0.9600
C7—H7A 0.9700 C23—H23C 0.9600
C7—H7B 0.9700 C24—H24A 0.9600
C8—C26 1.547 (3) C24—H24B 0.9600
C8—C9 1.550 (3) C24—H24C 0.9600
C8—C14 1.596 (3) C25—H25A 0.9600
C9—C11 1.540 (3) C25—H25B 0.9600
C9—C10 1.560 (3) C25—H25C 0.9600
C9—H9 0.9800 C26—H26A 0.9600
C10—C25 1.537 (3) C26—H26B 0.9600
C11—C12 1.485 (3) C26—H26C 0.9600
C11—H11A 0.9700 C27—H27A 0.9600
C11—H11B 0.9700 C27—H27B 0.9600
C12—C13 1.330 (3) C27—H27C 0.9600
C12—H12 0.9300 C28—H28A 0.9600
C13—C18 1.526 (3) C28—H28B 0.9600
C13—C14 1.535 (3) C28—H28C 0.9600
C14—C15 1.543 (3) C29—H29A 0.9600
C14—C27 1.552 (3) C29—H29B 0.9600
C15—C16 1.522 (3) C29—H29C 0.9600
C15—H15A 0.9700 C30—H30A 0.9600
C15—H15B 0.9700 C30—H30B 0.9600
C16—C17 1.538 (4) C30—H30C 0.9600
C6—O2—H2O 109.5 C17—C16—H16B 108.9 C2—C1—C10 114.19 (19) H16A—C16—H16B 107.7 C2—C1—H1A 108.7 C28—C17—C16 109.3 (3) C10—C1—H1A 108.7 C28—C17—C22 107.8 (2) C2—C1—H1B 108.7 C16—C17—C22 111.3 (2) C10—C1—H1B 108.7 C28—C17—C18 109.5 (2) H1A—C1—H1B 107.6 C16—C17—C18 108.4 (2) C3—C2—C1 112.2 (2) C22—C17—C18 110.6 (2) C3—C2—H2A 109.2 C13—C18—C19 112.03 (19) C1—C2—H2A 109.2 C13—C18—C17 112.7 (2) C3—C2—H2B 109.2 C19—C18—C17 113.4 (2)
C1—C2—H2B 109.2 C13—C18—H18 106.0
C2—C3—C4 115.3 (2) C20—C19—H19A 108.5 C3—C4—C24 109.9 (2) C18—C19—H19A 108.5 C3—C4—C23 103.8 (2) C20—C19—H19B 108.5 C24—C4—C23 107.5 (2) C18—C19—H19B 108.5 C3—C4—C5 109.64 (18) H19A—C19—H19B 107.5 C24—C4—C5 115.2 (2) C29—C20—C21 109.8 (3) C23—C4—C5 110.11 (19) C29—C20—C19 108.9 (3) C6—C5—C10 110.95 (18) C21—C20—C19 106.7 (2) C6—C5—C4 114.39 (17) C29—C20—C30 109.3 (3) C10—C5—C4 113.19 (17) C21—C20—C30 111.5 (3) C6—C5—H5 105.8 C19—C20—C30 110.7 (3) C10—C5—H5 105.8 C22—C21—C20 113.4 (3)
C4—C5—H5 105.8 C22—C21—H21A 108.9
O2—C6—C7 109.01 (17) C20—C21—H21A 108.9 O2—C6—C5 108.53 (18) C22—C21—H21B 108.9 C7—C6—C5 110.97 (17) C20—C21—H21B 108.9 O2—C6—H6 109.4 H21A—C21—H21B 107.7 C7—C6—H6 109.4 C21—C22—C17 115.4 (2)
C5—C6—H6 109.4 C21—C22—H22A 108.4
C6—C7—C8 115.36 (18) C17—C22—H22A 108.4 C6—C7—H7A 108.4 C21—C22—H22B 108.4 C8—C7—H7A 108.4 C17—C22—H22B 108.4 C6—C7—H7B 108.4 H22A—C22—H22B 107.5
C8—C7—H7B 108.4 C4—C23—H23A 109.5
H7A—C7—H7B 107.5 C4—C23—H23B 109.5 C7—C8—C26 108.51 (18) H23A—C23—H23B 109.5 C7—C8—C9 108.07 (17) C4—C23—H23C 109.5 C26—C8—C9 111.58 (18) H23A—C23—H23C 109.5 C7—C8—C14 110.87 (16) H23B—C23—H23C 109.5 C26—C8—C14 110.20 (19) C4—C24—H24A 109.5 C9—C8—C14 107.60 (16) C4—C24—H24B 109.5 C11—C9—C8 109.37 (18) H24A—C24—H24B 109.5 C11—C9—C10 112.48 (17) C4—C24—H24C 109.5 C8—C9—C10 118.76 (16) H24A—C24—H24C 109.5 C11—C9—H9 105.0 H24B—C24—H24C 109.5
C8—C9—H9 105.0 C10—C25—H25A 109.5
supporting information
sup-7 Acta Cryst. (2005). E61, o2022–o2023
H11A—C11—H11B 107.8 C14—C27—H27B 109.5 C13—C12—C11 126.5 (2) H27A—C27—H27B 109.5 C13—C12—H12 116.8 C14—C27—H27C 109.5 C11—C12—H12 116.8 H27A—C27—H27C 109.5 C12—C13—C18 118.4 (2) H27B—C27—H27C 109.5 C12—C13—C14 121.0 (2) C17—C28—H28A 109.5 C18—C13—C14 120.60 (18) C17—C28—H28B 109.5 C13—C14—C15 110.55 (18) H28A—C28—H28B 109.5 C13—C14—C27 108.07 (18) C17—C28—H28C 109.5 C15—C14—C27 105.69 (19) H28A—C28—H28C 109.5 C13—C14—C8 109.42 (16) H28B—C28—H28C 109.5 C15—C14—C8 111.50 (17) C20—C29—H29A 109.5 C27—C14—C8 111.52 (18) C20—C29—H29B 109.5 C16—C15—C14 114.07 (19) H29A—C29—H29B 109.5 C16—C15—H15A 108.7 C20—C29—H29C 109.5 C14—C15—H15A 108.7 H29A—C29—H29C 109.5 C16—C15—H15B 108.7 H29B—C29—H29C 109.5 C14—C15—H15B 108.7 C20—C30—H30A 109.5 H15A—C15—H15B 107.6 C20—C30—H30B 109.5 C15—C16—C17 113.3 (2) H30A—C30—H30B 109.5 C15—C16—H16A 108.9 C20—C30—H30C 109.5 C17—C16—H16A 108.9 H30A—C30—H30C 109.5 C15—C16—H16B 108.9 H30B—C30—H30C 109.5
C7—C8—C9—C11 175.25 (18) C15—C16—C17—C18 −59.5 (3) C26—C8—C9—C11 56.0 (2) C12—C13—C18—C19 −90.1 (3) C14—C8—C9—C11 −65.0 (2) C14—C13—C18—C19 86.5 (3) C7—C8—C9—C10 44.3 (3) C12—C13—C18—C17 140.7 (2) C26—C8—C9—C10 −74.9 (3) C14—C13—C18—C17 −42.7 (3) C14—C8—C9—C10 164.09 (17) C28—C17—C18—C13 −68.5 (3) C6—C5—C10—C25 −74.7 (2) C16—C17—C18—C13 50.7 (3) C4—C5—C10—C25 55.4 (2) C22—C17—C18—C13 172.9 (2) C6—C5—C10—C1 167.65 (19) C28—C17—C18—C19 163.0 (2) C4—C5—C10—C1 −62.2 (2) C16—C17—C18—C19 −77.9 (2) C6—C5—C10—C9 51.7 (2) C22—C17—C18—C19 44.3 (3) C4—C5—C10—C9 −178.11 (17) C13—C18—C19—C20 178.4 (2) C2—C1—C10—C25 −87.7 (3) C17—C18—C19—C20 −52.7 (3) C2—C1—C10—C5 31.9 (3) C18—C19—C20—C29 174.5 (3) C2—C1—C10—C9 150.5 (2) C18—C19—C20—C21 56.1 (3) C11—C9—C10—C25 −52.0 (2) C18—C19—C20—C30 −65.4 (3) C8—C9—C10—C25 77.5 (3) C29—C20—C21—C22 −173.7 (2) C11—C9—C10—C5 −176.72 (18) C19—C20—C21—C22 −55.9 (3) C8—C9—C10—C5 −47.2 (3) C30—C20—C21—C22 65.0 (3) C11—C9—C10—C1 65.7 (2) C20—C21—C22—C17 54.4 (3) C8—C9—C10—C1 −164.7 (2) C28—C17—C22—C21 −165.9 (3) C8—C9—C11—C12 42.9 (3) C16—C17—C22—C21 74.3 (3) C10—C9—C11—C12 177.1 (2) C18—C17—C22—C21 −46.2 (3)
Hydrogen-bond geometry (Å, º)
D—H···A D—H H···A D···A D—H···A
O2—H2O···O1i 0.82 2.08 2.901 (2) 175