J
OURNAL OFC
LINICALM
ICROBIOLOGY, Mar. 2011, p. 808–813
Vol. 49, No. 3
0095-1137/11/$12.00
doi:10.1128/JCM.01149-10
Copyright © 2011, American Society for Microbiology. All Rights Reserved.
Identification of
Fusarium
Species in Formalin-Fixed and
Paraffin-Embedded Sections by
In Situ
Hybridization
Using Peptide Nucleic Acid Probes
䌤
†
Minoru Shinozaki,
1Yoichiro Okubo,
1Daisuke Sasai,
1Haruo Nakayama,
1Somay Yamagata Murayama,
2Tadashi Ide,
1Megumi Wakayama,
1Nobuyuki Hiruta,
1and Kazutoshi Shibuya
1*
Department of Surgical Pathology, Toho University School of Medicine, Tokyo, Japan,
1and Laboratory of Molecular Epidemiology for
Infectious Agents, Graduate School of Infection Control Sciences, Kitasato University, Tokyo, Japan
2Received 7 June 2010/Returned for modification 21 July 2010/Accepted 17 November 2010
Fusarium
has recently emerged as an opportunistic pathogen of humans, but the histological differentiation
of
Fusarium
from
Aspergillus
and
Scedosporium
is particularly difficult because these fungi may induce similar
clinical features and exhibit filamentous development in host tissues. Thus, there is a need to establish rapid
and reliable methods that are applicable to pathological diagnoses. The aim of this study was to evaluate and
establish
in situ
hybridization (ISH) using peptide nucleic acid (PNA) probes targeting the 28S rRNA to
identify
Fusarium
species in tissue sections. This technique was validated using both formalin-fixed and
paraffin-embedded pulmonary tissues from mice infected with seven different species of fungi and cell blocks
from fungal cultures of 30 strains. As a result, strong positive signals were observed within fungal organisms
present in tissues of the lung from mice infected with
Fusarium solani
. Furthermore, this probe reacted strongly
with both
F. solani
and
Fusarium oxysporum
in sections from cell blocks. Although some cross-reactivity
occurred with the
Pseudallescheria boydii
in sections from cell blocks, the signal intensity was low and most
hyphae were not reactive. In conclusion, it was confirmed that ISH with PNA probes is accurate and is a
valuable tool for identifying
Fusarium
spp. among organisms that have identical morphological features in
formalin-fixed and paraffin-embedded sections.
Early diagnosis of invasive fungal infection is essential
be-cause the disease mostly occurs in patients with severely
im-paired defense mechanism.
Fusarium
species are widely
dis-tributed in soil, subterranean, aerial plant parts, plant debris,
and other organic substrates, and mycotoxins produced by
these organisms have often been associated with animal and
human diseases (16). In humans,
Fusarium
spp. cause a broad
spectrum of infections, including superficial (keratitis and
onychomycosis), locally invasive, and disseminated infections
in immunocompromised patients (18). In addition to that
in-vasive and disseminated infections caused by
Fusarium
spp. are
being diagnosed with increasing frequency in patients with
hematological malignancies (3), it has been accepted that
Fusarium
spp. are resistant to most antifungal agents (5).
Therefore, an early diagnosis of the infection is now required
to improve the outcome of treatment for seriously debilitating
conditions. Because of morphological similarities among
molds in histopathological specimens, it has been difficult to
differentiate histologically
Fusarium
spp. from other molds.
Recently, sensitive and rapid molecular detection assays that
use PCR-based methods have been introduced to detect
Fusar-ium
DNA in serum, total blood, and tissue samples (10).
How-ever, there have been a few attempts to use
in situ
hybridization
(ISH) to identify
Fusarium
spp. in tissue sections for
histolog-ical diagnosis (8, 14). We are describing the first report of ISH
using peptide nucleic acid (PNA) as the probe targeting the
28S rRNA of
Fusarium
spp. to identify the fungus in
formalin-fixed and paraffin-embedded tissue sections that are widely
used as routine preparations for surgical and anatomical
pa-thology in hospitals.
MATERIALS AND METHODS
Preparation of infected animals and tissue specimens.To verify the specificity
of probes, sections of formalin-fixed and paraffin-embedded tissues of lung were prepared from mice experimentally infected with seven different fungi. Lung has been understood as one of the commonest organ involved by invasive fungal infection. A part of this may be explained by the fact that the lung serves as a porta of infection. Therefore, in the present study, lungs from mice with intra-tracheal infection were used as a tissue specimen to evaluate the ISH procedure. Six-week-old, male Institute of Cancer Research (ICR) mice (Sankyo Labo Service Corp., Inc., Tokyo, Japan) were used in the present study. Immune suppression was achieved by intraperitoneal injections of cyclophosphamide (Shionogi and Co., Ltd., Osaka, Japan) at a dose of 150 mg/kg (body weight) 3 days prior to infection. To prevent bacterial infection, the animals were also intraperitoneally administered with imipenem/cilastatin sodium (Banyu Pharma-ceutical, Tokyo, Japan). Prior to inoculation, the animals were anesthetized intraperitoneally with 80 mg of ketamine (Daiichi Sankyo Co., Ltd., Tokyo, Japan) and 10 mg of xylazine (Bayer Health Care, Tokyo, Japan)/kg. The conidiae or yeast cells were injected intratracheally as previously described (19). A 25-l aliquot of the conidiae or yeast cell suspension was injected into the trachea via a clinically used intravascular catheter (24G, Insyte-W; Becton Dick-inson, Hollister, CA). The mice were infected with 3⫻105
conidiae or yeast cells ofAspergillus fumigatus(TIMM1776),Aspergillus terreus(TIMM2929), Aspergil-lus flavus (TIMM2935), Candida albicans, (TIMM1768), Rhizopus oryzae (TIMM1326), Fusarium solani (TIMM1303), and Pseudallescheria boydii (TIMM0952). The animals were sacrificed on the third day after infection, and
* Corresponding author. Mailing address: Department of Surgical
Pathology, Toho University School of Medicine, 6-11-1 Omori-Nishi,
Ota-Ku, Tokyo 143-8541, Japan. Phone: 3762-4151. Fax:
81-3-3767-1567. E-mail: kaz@med.toho-u.ac.jp.
† Supplemental material for this article may be found at http://jcm
.asm.org/.
䌤
Published ahead of print on 24 November 2010.
808
on May 16, 2020 by guest
http://jcm.asm.org/
the lungs were removed and fixed in 10% formalin, followed by dehydration with ethanol and embedding in paraffin. Tissue sections (3m) were mounted on aminoalkylsilane-coated slide glasses (Dako Japan, Tokyo, Japan). Pulmonary lesions induced by this procedure were confirmed by histological examination using these sections stained with hematoxylin and eosin (H&E) and Grocott’s stains.
Strains tested.For specificity testing of the probes, cells from the following molds
and yeasts were tested:A. flavusvar.flavus(NBRC 33021),Aspergillus niger(NBRC 33023),A. terreus(NBRC 33026),A. fumigatus(NBRC 6344),A. fumigatusvar. fumigatus(NBRC 33022),F. solani(NBRC 5232),Fusarium oxysporum(NBRC 7152),P. boydii(NBRC 8078),R. oryzae(NBRC 5780),Cunninghamella elegansvar. elegans(NBRC 4446),Rhizomucor pusillus(NBRC 9744),Mucor circinelloidesf. sp. circinelloides(NBRC 4554),Penicillium commune(NBRC 5763),Pseudocochliobolus spicifer (NBRC 100222), C. albicans (ATCC 10231), Trichosporon asahii (CBS2479T), C. albicans var. stellatoidea (TIMM0310), Candida glabrata (TIMM1064),Candida guilliermondii(TIMM0260), Candida kefyr(TIMM0302), Candida krusei(TIMM0269),Candida lusitaniae(TIMM1668),Candida parapsilosis (TIMM0292), Candida tropicalis (TIMM0313), Cryptococcus neoformans (TIMM0354), Debaryomyces polymorphus (TIMM2937), Hansenula anomala (JCM3585), Pichia subpelliculosa (IFO0808), Saccharomyces cerevisiae (TIMM0925), andSchizosaccharomyces pombe(TIMM3376).
Preparation of cell blocks from cultured fungal cells.Mold strains were grown
for 48 to 72 h at 25°C in potato dextrose broth (Sigma Aldrich, St. Louis, MO). Yeast strains exceptCandidaspp. were grown overnight at 25°C in YMPD broth (0.3% yeast extract, 0.3% malt extract, 0.5% peptone, 1.0% glucose) (Becton Dickinson), andCandidaspp. were grown overnight at 37°C in Medium 199 (Nissui Pharmaceutical Co., Ltd., Tokyo, Japan) with 10% calf fetal serum (Sigma-Aldrich Co., St. Louis, MO). From cultures of the above-mentioned molds or yeasts, fungal cell suspensions were prepared in 10% formalin for fixation. Fixed molds were collected with centrifugation for 10 min at 2,000 rpm.
The cluster of molds was carefully transferred onto filter paper (Advantec Toyo, Ltd., Tokyo, Japan) with pointed forceps. This was also wrapped with same filter paper and placed in an embedding cassette (Murazumi Industrial Co., Ltd., Hyogo, Japan) and then penetrated with paraffin by using an automated tissue processor (Tissue-Tek VIP Premier; Sakura Finetek Japan Co., Ltd., Tokyo, Japan) (12). The cluster of molds penetrated with paraffin was transferred to bottom of an embedding stainless dish, followed by filling solidifying of paraffin, and cut into 3-m sections that were then mounted on aminoalkylsilane-coated slide glasses (Dako Japan, Tokyo, Japan).
To prevent the diffusion of yeast cells in suspension and to obtain a high density in paraffin blocks, we used agarose gel as an intermediate embedding medium. Yeast cells were collected with centrifugation for 10 min at 2,000 rpm and added with 50l of 2% liquid agarose at 65°C. This agarose gel was heaped onto the surface of cover glass (Matsunami Glass Ind., Ltd., Osaka, Japan), and solidified at room temperature. The solidified gel was penetrated with paraffin by using an automated tissue processor (Tissue-Tek VIP Premier), as well, and sections were prepared in same way (12).
PNA probes.The design of the PNA probes forFusariumspp. (N
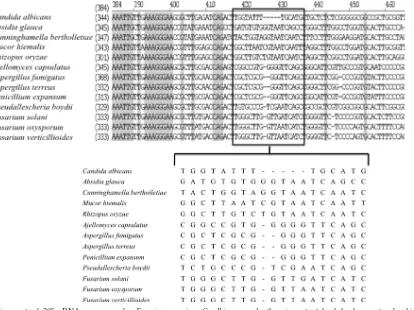
[image:2.585.88.502.69.379.2]terminus-GAT terminus-GAT CAA CCA AGC CCA) and panfungal species (N terminus-TAC TTG TGC GCT ATC GGT) was derived from a comparison of 28S rRNA genes in the GenBank database. The alignment of the DNA sequence was performed by using Vector NTI Advance TM10 (Invitrogen, Carlsbad, CA). After align-ment and visual assessalign-ment of theFusariumand non-Fusariumsequences, the probe forFusariumspp. was designed. As shown in Fig. 1, aFusariumsp. antisense PNA probe targeting the 28S rRNA could be designed and evaluated on a genus-specific level. The sequence revealed at least 9 to 14 mismatches within the target region in sequences of nontarget organisms. Furthermore, to assess the retention of RNA in samples, we also designed a panfungal antisense PNA probe in the same way. Each selected sequence was checked for specificity against the GenBank database by using the Basic Local Alignment Search Tool
FIG. 1. Alignment of 28S rRNA sequences for
Fusarium
species,
C. albicans
, and other important hyalohyphomycetes for histological
differentiation. The binding regions of the antisense probe used in the present study to detect
Fusarium
rRNA were boxed and magnified.
Homologous regions were highlighted in gray. Species and GenBank accession numbers were as follows:
C. albicans
(AB436387),
Absidia glauca
(AF113447),
Cunninghamella bertholletiae
(AF113459),
Mucor hiemalis
(AF113468),
Rhizopus oryzae
(DQ466617),
Ajellomyces capsulatus
(AB176493),
A. fumigatus
(AB354577),
A. terreus
(AF454185),
Penicillium expansum
(AJ519347),
P. boydii
(EF151324),
F. solani
(AF178377),
F.
oxysporum
(AF060383), and
Fusarium verticillioides
(U34526).
V
OL. 49, 2011
IDENTIFICATION OF
FUSARIUM
BY
IN SITU
HYBRIDIZATION
809
on May 16, 2020 by guest
http://jcm.asm.org/
(BLAST; http//www.ncbi.nlm.nih.gov/BLAST/). The selected sequence was then synthesized, and the N terminus of the oligomer was conjugated to fluorescein isothiocyanate (FITC) via a double aminoethoxyethoxyacetate (AEEA) linker (Fasmac Co., Ltd., Kanagawa, Japan).
ISH.The ISH procedure was performed as described previously (24). Briefly, sections were deparaffinized and rehydrated according to standard procedures. To expose target nucleic acids in the formalin-fixed tissue, the sections were treated with a 1 mM concentration of EDTA buffer (pH 8.0) in a water bath (Thermo Fisher Scientific K.K., Yokohama, Kanagawa) for 20 min at 98°C and digested with a 10-g/ml concentration of proteinase K (Nippon Gene Co., Ltd., Tokyo, Japan) for 10 min at 37°C. Hybridization was performed at 56°C for 90 min with 1g of PNA probe/ml dissolved in hybridization medium (Dako Japan, Tokyo, Japan). After repeated washings with 2⫻standard saline citrate (SSC) at 56°C, the signals were detected by enzyme immunohistochemistry using an anti-FITC antibody (Roche Diagnostics K.K., Tokyo, Japan) and horseradish perox-idase-labeled polymer solution (Nichirei Biosciences, Inc., Tokyo, Japan). Fi-nally, the sites of peroxidase were visualized by 3,3⬘-diaminobenzidine tetrahydrochloride (DAB; Dojindo Laboratories, Kumamoto, Japan) in the pres-ence of H2O2and nickel and cobalt ions (1). As negative controls, ISH
proce-dures were performed with aC. albicansPNA probe (20).
RESULTS
Paraffin sections of both lungs of mice infected with different
fungi and cell blocks mounted on the slide glasses were H&E
and/or Grocott’s stained, processed with ISH, and observed
under light microscopy for evaluation of our ISH procedure.
Specificity of ISH for
Fusarium
spp. in infected animal
mod-els.
Histological examination revealed an extensive fungal
growth both in alveoli and terminal bronchus with necrosis and
minor polymorphonuclear leukocyte infiltrate (Fig. 2). The
establishment of experimental pulmonary lesions was
con-firmed in mice infected with all seven of the different fungi that
we examined.
Formalin-fixed and paraffin-embedded pulmonary tissues
from mice infected with seven different fungi were tested to
assess whether the probe hybridized specifically with
Fusarium
spp. Strong positive signals against 28S rRNA of
Fusarium
spp.
were observed within fungal organisms present in lung tissue
from mice infected with
F. solani
(Fig. 3d). Positive organisms
typically exhibited a signal visualized by a DAB reaction that
was limited in a large part of cytoplasm and can be recognized
as black fine dots. The signal intensity varied within and
be-tween fungal organisms in tissue sections. No substantial
back-ground signal was observed in any tissue. In addition, no
hy-bridization was found in other fungi tested.
Specificity of ISH for
Fusarium
spp. in cell blocks of
cul-tured fungi.
In total, 30 cell block sections from formalin-fixed
and paraffin-embedded fungi of 30 strains were studied (see
Fig. S1 in the supplemental material). Within the panel of 30
fungi, the
Fusarium
sp. PNA probe reacted strongly with both
F. solani
(Fig. 4d) and
F. oxysporum
in sections of cell blocks.
The signal intensity and distribution in fungal organisms were
similar to those observed in animal models. With the exception
of the
Fusarium
spp., the
P. boydii
in sections of cell blocks (not
of tissue sections) showed positive reactivity for the probe, but
their signal intensity was low and most of the hyphae were
negative (Fig. 4c). No hybridization was observed in other
fungi tested.
[image:3.585.114.472.68.330.2]Control experiments.
To confirm the specificity of the 28S
rRNA signals, adjacent sections were hybridized with a
C.
albicans
PNA probe. A strong positive signal was detected in
C.
albicans
in a tissue section, whereas no hybridization signal was
found in the other fungi tested. The panfungal PNA probe
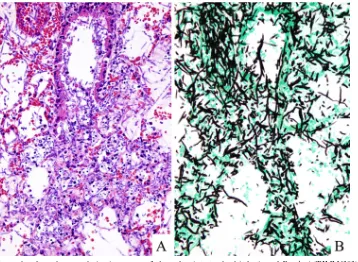
FIG. 2. Microphotographs of a pulmonary lesion in a mouse 3 days after intratracheal infection of
F. solani
(TIMM1303). (A) Histological
examination revealed an extensive fungal growth both in alveoli and terminal bronchus with necrosis and minor polymorphonuclear leukocyte
infiltrate with scattering nuclear debris (H&E stain; original magnification,
⫻
100). (B) There is extensive hyphal growth of invading mold showing
dichotomous branching (Grocott’s stain; original magnification,
⫻
100).
810
SHINOZAKI ET AL.
J. C
LIN. M
ICROBIOL.
on May 16, 2020 by guest
http://jcm.asm.org/
reacted with all fungi tested (Fig. 3e to h, Fig. 4e to h). The
intensity of ISH signals for the panfungal probe was similar to
that with species-specific probes.
DISCUSSION
Recently, several genome databases have provided new
in-formation that can be used for field studies for the molecular
identification and epidemiology of pathogenic fungi.
Accord-ingly, sensitive and rapid molecular detection assays have been
established by using PCR-based methods to detect fungal
DNA (7, 11, 27). The application of these molecular
tech-niques to formalin-fixed and paraffin-embedded tissue has also
been reported (2, 13, 21). Although there have been a few
attempts to use ISH to detect fungal agents in
histopatholog-ical specimens (6, 8, 9, 14, 15), the use of ISH for the diagnosis
of fungal infection in formalin-fixed and paraffin-embedded
sections has not been systematically assessed. We have
previ-ously reported that a combination of high-temperature heating
in solutions of high pH, followed by a 10-min proteinase K
digestion step, gave better ISH results (24). The heating
pre-treatment used in the present study was adapted from antigen
retrieval techniques used in conventional
immunohistochem-istry (23).
Our purpose was to evaluate and establish an ISH procedure
for the detection of
Fusarium
spp. in formalin-fixed and
par-affin-embedded sections. Diagnosis of fusariosis from cultures
remains a difficult and time-consuming task, relying on
mor-phological and physiological examinations and requiring some
degree of expertise.
Fusarium
spp. are phylogenetically
heter-ogeneous with variable antifungal susceptibilities (25). An
ap-proach based on PCR methods has been used to detect
Fusar-ium
DNA (10). Although there have been a few attempts to
use ISH to identify
Fusarium
spp. in histopathological
speci-mens (8, 14), to our knowledge there has been no report of
ISH using a PNA probe. Our results obtained with mice
ex-perimentally infected with seven different fungi showed that
F.
solani
can be specifically detected in infected tissues by ISH
with a PNA probe targeting 28S rRNA of
Fusarium
spp. On
the other hand, using cell block sections from formalin-fixed
and paraffin-embedded fungi, this probe reacted strongly with
both
F. solani
and
F. oxysporum
, but some cross-reactivity was
observed in
P. boydii
hyphae. Part of this result may be
ex-plained by the fact that
P. boydii
has a sequence similar to the
target of our probe.
[image:4.585.82.504.71.351.2]PNA molecules are DNA mimics in which the negatively
charged sugar-phosphate backbone is replaced by a neutral
polyamide backbone, formed by repetitive units of
N
-glycine.
This structure enables PNA probes to hybridize to
complemen-tary nucleic acid targets with high specificity and rapid binding
kinetics (4, 17). Due to the novel properties of its
hybridiza-tion, PNA is beginning to be applied in ISH to detect fungal
nucleic acids (20, 22, 26, 28); however, there has been no
report of the application of such a probe to formalin-fixed and
paraffin-embedded tissues. Better outcomes are obtained with
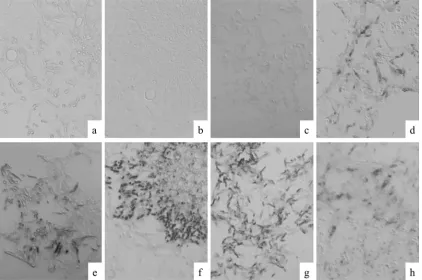
FIG. 3. Specificity verification of the
Fusarium
sp. PNA probe and assessments of rRNA retention and its hybridizability in experimentally
infected mice. The tissue sections were hybridized with
Fusarium
sp. PNA probe (a to d) or with panfungal PNA probe (e to h). Strong positive
signals against 28S rRNA of
Fusarium
spp. were observed in lung tissues from mice infected with
F. solani
(d). The panfungal PNA probe reacted
with all fungi tested (e to h). (a and e)
A. fumigatus
(TIMM1776); (b and f)
A. terreus
(TIMM2929); (c and g)
P. boydii
(TIMM0952); (d and h)
F. solani
(TIMM1303). Original magnification,
⫻
400.
V
OL. 49, 2011
IDENTIFICATION OF
FUSARIUM
BY
IN SITU
HYBRIDIZATION
811
on May 16, 2020 by guest
http://jcm.asm.org/
PNA probes compared to conventional DNA probes (26). In
our first approach, we confirmed that PNA probes required
shorter hybridization times than double-stranded DNA
probes. From the standpoint of decreased assay turnaround
time, the application of PNA probes is especially attractive.
Recently, Montone reported that the use of dual
fluoro-genic-labeled locked nucleic acids (LNA) probes of ISH were
able to differentiate
Fusarium
from
Aspergillus
organisms (14)
and that the LNA probe produced a stronger signal compared
to a DNA probe with the same sequence (15). Our probe could
differentiate
Fusarium
from 23 fungal species other than
As-pergillus
. These novel findings demonstrate the feasibility of
the approach and strongly suggest that LNA and PNA can be
widely used as probes of ISH in the near future.
The 28S rRNA sequence was selected as a detection target
because its large size may reveal adequate differences in
dis-tinguishing closely related organisms. In addition, it has been
accepted that multiple copies of ribosomal genes are present in
fungi, which can be transcribed into rRNA. It is essential that
assessment of retention of rRNA and its hybridizability should
be performed, because loss of rRNA or failure of the
accessi-bility of probes in processed tissue sections can lead to
mis-leading results. In the present study, we designed a panfungal
PNA probe and confirmed that the intensity of ISH signals of
this probe was similar to those of species-specific probes.
These findings suggest that ISH with the panfungal probe may
be useful for the estimation of hybridizable rRNA for the
specific detection of human pathogenic fungi.
In conclusion, we have shown the superiority and the
use-fulness of ISH with PNA probes for identifying
Fusarium
spp.
in formalin-fixed and paraffin-embedded sections. Further
studies are needed to establish ISH with PNA probes as an
accurate and rapid diagnostic procedure for tissue sections
from patients with suspected fusariosis.
ACKNOWLEDGMENTS
This study was supported by Health Science Research grants for
Research on Emerging and Re-Emerging Infectious Diseases
(H16-Shinko-6, H19-Shinko-8, and H22-Shinko-8) and Measures for
Intrac-table Diseases (H20 Nannchi Ippann 35) from the Ministry of Health,
Labor, and Welfare of Japan and by the Grant of the Strategic Basis on
Research Grounds for Non-Governmental Schools at Heisei 20th from
the Ministry of Education, Culture, Sports, Science, and Technology of
Japan to K.S.
We are grateful to K Makimura, K Uchida, and H Yamaguchi for
kindly providing important advice.
REFERENCES
1.Adams, J. C.1981. Heavy metal intensification of DAB-based HRP reaction
product. J. Histochem Cytochem.29:775.
2.Bialek, R., et al.2005. PCR based identification and discrimination of agents
of mucormycosis and aspergillosis in paraffin wax embedded tissue. J. Clin. Pathol.58:1180–1184.
3.Boutati, E. I., and E. J. Anaissie.1997.Fusarium, a significant emerging
[image:5.585.82.504.70.354.2]pathogen in patients with hematologic malignancy: ten years’ experience at a cancer center and implications for management. Blood90:999–1008.
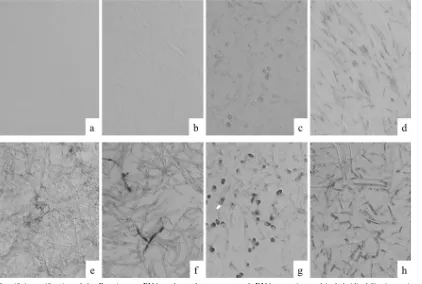
FIG. 4. Specificity verification of the
Fusarium
sp. PNA probe and assessments of rRNA retention and its hybridizability in strains of molds.
Cell block sections from formalin-fixed and paraffin-embedded fungi were hybridized with
Fusarium
sp. PNA probe (a to d) or with panfungal PNA
probe (e to h).
Fusarium
sp. PNA probe hybridized strongly with
F. solani
(d), whereas some cross-hybridization occurred with the
P. boydii
(c).
The panfungal PNA probe reacted with all fungi tested (e to h). (a and e)
A. fumigatus
(NBRC 6344); (b and f)
A. terreus
(NBRC 33026); (c and
g)
P. boydii
(NBRC 8078); (d and h)
F. solani
(NBRC 5232). Original magnification,
⫻
400.
812
SHINOZAKI ET AL.
J. C
LIN. M
ICROBIOL.
on May 16, 2020 by guest
http://jcm.asm.org/
4.Demidov, V. V., M. V. Yavnilovich, B. P. Belotserkovskii, M. D.
Frank-Kamenetskii, and P. E. Nielsen.1995. Kinetics and mechanism of polyamide
(“peptide”) nucleic acid binding to duplex DNA. Proc. Natl. Acad. Sci. U. S. A.92:2637–2641.
5.Guinea, J., T. Pela´ez, S. Recio, M. Torres-Narbona, and E. Bouza.2008. In
vitro antifungal activity of isavuconazole (BAL4815), voriconazole, and flu-conazole against 1,007 isolates of zygomycetes,Candida,Aspergillus, Fusar-ium, andScedosporiumspecies. Antimicrob. Agents Chemother.52:1396– 1400.
6.Hanazawa, R., S. Y. Murayama, and H. Yamaguchi.2000. In-situ detection
ofAspergillus fumigatus. J. Med. Microbiol.49:285–290.
7.Hata, D. J., S. P. Buckwalter, B. S. Pritt, G. D. Roberts, and N. L.
Wen-genack.2008. Real-time PCR method for detection of zygomycetes. J. Clin.
Microbiol.46:2353–2358.
8.Hayden, R. T., et al.2003. In situ hybridization for the differentiation of
Aspergillus,Fusarium, andPseudallescheriaspecies in tissue section. Diagn. Mol. Pathol.12:21–26.
9.Hayden, R. T., X. Qian, G. W. Procop, G. D. Roberts, and R. V. Lloyd.2002.
In situ hybridization for the identification of filamentous fungi in tissue section. Diagn. Mol. Pathol.11:119–126.
10.Hue, F. X., M. Huerre, M. A. Rouffault, and C. de Bievre.1999. Specific
detection ofFusariumspecies in blood and tissues by a PCR technique. J. Clin. Microbiol.37:2434–2438.
11.Hummel, M., et al.2006. Detection ofAspergillusDNA in cerebrospinal fluid
from patients with cerebral aspergillosis by a nested PCR assay. J. Clin. Microbiol.44:3989–3993.
12.Kerstens, H. M., et al.2000. AgarCyto: a novel cell-processing method for
multiple molecular diagnostic analyses of the uterine cervix. J. Histochem. Cytochem.48:709–718.
13.Lau, A., et al.2007. Development and clinical application of a panfungal
PCR assay to detect and identify fungal DNA in tissue specimens. J. Clin. Microbiol.45:380–385.
14.Montone, K. T.2009. Differentiation ofFusariumfromAspergillusspecies by
colorimetric in situ hybridization in formalin-fixed, paraffin-embedded tissue sections using dual fluorogenic-labeled LNA probes. Am. J. Clin. Pathol.
132:866–870.
15.Montone, K. T., and M. D. Feldman.2009. In situ detection ofAspergillus18S
rRNA Sequences using a terminally biotinylated locked nucleic acid (LNA) probe. Diagn. Mol. Pathol.18:239–242.
16.Nelson, P. E., M. C. Dignani, and E. J. Anaissie.1994. Taxonomy, biology,
and clinical aspects ofFusariumspecies. Clin. Microbiol. Rev.7:479–504.
17.Nielsen, P. E., M. Egholm, R. H. Berg, and O. Buchardt.1991.
Sequence-selective recognition of DNA by strand displacement with a thymine-substi-tuted polyamide. Science254:1497–1500.
18.Nucci, M., and E. Anaissie.2002. Cutaneous infection byFusariumspecies in
healthy and immunocompromised hosts: implications for diagnosis and man-agement. Clin. Infect. Dis.35:909–920.
19.Ochiai, E., et al.2008. Inhalation ofStachybotrys chartarumcauses
pulmo-nary arterial hypertension in mice. Int. J. Exp. Pathol.89:201–208.
20.Oliveira, K., G. Haase, C. Kurtzman, J. J. Hyldig-Nielsen, and H. Stender.
2001. Differentiation ofCandida albicansandCandida dubliniensisby fluo-rescent in situ hybridization with peptide nucleic acid probes. J. Clin. Mi-crobiol.39:4138–4141.
21.Paterson, P. J., S. Seaton, J. McLaughlin, and C. C. Kibbler.2003.
Devel-opment of molecular methods for the identification of aspergillus and emerging moulds in paraffin wax embedded tissue sections. Mol. Pathol.
56:368–370.
22.Rigby, S., et al.2002. Fluorescence in situ hybridization with peptide nucleic
acid probes for rapid identification ofCandida albicansdirectly from blood culture bottles. J. Clin. Microbiol.40:2182–2186.
23.Shi, S. R., M. E. Key, and K. L. Kalra.1991. Antigen retrieval in
formalin-fixed, paraffin-embedded tissues: an enhancement method for immunohis-tochemical staining based on microwave oven heating of tissue sections. J. Histochem. Cytochem.39:741–748.
24.Shinozaki, M., et al. 2009. Application of in situ hybridization to tissue
sections for identification of molds causing invasive fungal infection. Nippon Ishinkin Gakkai Zasshi50:75–83.
25.Stanzani, M., F. Tumietto, N. Vianelli, and M. Baccarani.2007. Update on
the treatment of disseminated fusariosis: focus on voriconazole. Ther. Clin. Risk Manag.3:1165–1173.
26.Teertstra, W. R., L. G. Lugones, and H. A. Wo¨sten.2004. In situ hybridisation
in filamentous fungi using peptide nucleic acid probes. Fungal Genet. Biol.
41:1099–1103.
27.Vollmer, T., M. Sto¨rmer, K. Kleesiek, and J. Dreier.2008. Evaluation of
novel broad-range real-time PCR assay for rapid detection of human patho-genic fungi in various clinical specimens. J. Clin. Microbiol.46:1919–1926.
28.Wilson, D. A., et al.2005. Multicenter evaluation of aCandida albicans
peptide nucleic acid fluorescent in situ hybridization probe for characteriza-tion of yeast isolates from blood cultures. J. Clin. Microbiol.43:2909–2912.