metal-organic papers
Acta Cryst.(2006). E62, m523–m525 doi:10.1107/S1600536806004843 Cuiet al. [Cu(NCS)
2(C7H8N4)2]
m523
Acta Crystallographica Section EStructure Reports Online
ISSN 1600-5368
catena
-Poly[[bis(isothiocyanato-
j
N
)copper(II)]-bis(
l
-1,1
000-methylenedi-1
H
-imidazole-j
N
3:
j
N
3000)]
Guang-Hua Cui,a* Gui-Ying
Dongaand Joan Ribasb
a
College of Chemical Engineering and Biotechnology, Hebei Polytechnic University, Tangshan 063009, People’s Republic of China, andbDepartament de Quı´mica Inorga`nica,
Universitat de Barcelona, Diagonal 6487, 08028-Barcelona, Spain
Correspondence e-mail: tscghua@126.com
Key indicators
Single-crystal X-ray study
T= 293 K
Mean(C–C) = 0.003 A˚
Rfactor = 0.030
wRfactor = 0.071
Data-to-parameter ratio = 14.6
For details of how these key indicators were automatically derived from the article, see http://journals.iucr.org/e.
Received 1 February 2006 Accepted 9 February 2006
#2006 International Union of Crystallography
All rights reserved
The title crystal structure, [Cu(NCS)2(C7H8N4)2]n, comprises
one-dimensional chains propagating in the a-axis direction. The CuII atoms, which have crystallographic 2/m site symmetry, are coordinated by six N atoms from four 1,10 -methylenedi-1H-imidazole ligands and two NCS anions, giving a slightly disorted octahedral coordination geometry. The 1,10-methylenedi-1H-imidazole ligand adopts a bis-monodentate bridging mode, linking the CuIIatoms.
Comment
Copper complexes with imidazole ligands are of considerable interest as models for the active site of copper proteins such as haemocyanin, azurin and plastocyanin (Chen et al., 1994; Kirchner & Krebs, 1987). Some metal complexes with alkyl-linked bisimidazole ligands have been used successfully to construct three-dimensional networks (Wuet al., 1997; Maet al., 2004; Cui et al., 2005). In order to further examine the structural features of copper complexes with this class of ligand, the crystal structure determination of the title compound, (I), was carried out.
The structure of (I) comprises a one-dimensional neutral double [Cu(NCS)2(L)2] chain [L is 1,10-methylenedi-1H
-imidazole]. Within a chain, the coordination geometry of each CuIIatom is slightly distorted octahedral (Fig. 1 and Table 1). The CuIIatom, lying on a position of site symmetry 2/m, is six-coordinated by four N atoms from four symmetry-related L
ligands, with a unique Cu—N distance of 2.0440 (17) A˚ . The coordination is completed by two N atoms from two NCS anions. The NCS anions are trans-coordinated to the CuII atoms, occupying the axial positions with a unique Cu—N distance of 2.425 (3) A˚ ; this is elongated, most likely due to Jahn–Teller effects. The cis-N—Cu—N bond angles deviate only slightly from ideal octahedral values (90) ranging from 87.55 (9) to 92.45 (9). EachLligand coordinates to two CuII
closest CuII atoms within a chain is 8.877 (2) A˚ and the dihedral angle between the two imidazole planes in one L
ligand is 71.5 (2).
Experimental
The ligand L was prepared according to the reported procedure (Schu¨tze & Schubert, 1959). A mixture of Cu(NO3)2 (0.126 g,
1 mmol) and KSCN (0.196 g, 2 mmol) in water (30 ml) was stirred at room temperature for 30 min.L(0.296 g, 2 mmol) was added and the resulting solution was refluxed for 3 h. The blue precipitate that was obtained was filtered off, washed with water and dissolved in a minimum amount of aqueous ammonia (14M). Blue single crystals were obtained by slow evaporation of the ammoniacal solution of the solid at room temperature over three days (68% yield based on Cu).
Analysis calculated for C16H16CuN10S2: C 40.37, H 3.39, N 29.42%;
found: C 40.31, H 3.34, N 29.21%.
Crystal data
[Cu(NCS)2(C7H8N4)2] Mr= 476.05
Orthorhombic,Cmca a= 8.877 (3) A˚
b= 15.602 (6) A˚
c= 14.170 (4) A˚
V= 1962.5 (12) A˚3 Z= 4
Dx= 1.611 Mg m 3
MoKradiation Cell parameters from 932
reflections
= 2.6–26.2 = 1.35 mm1 T= 293 (2) K Block, blue
0.200.180.16 mm
Data collection
Bruker SMART CCD area-detector diffractometer
’and!scans
Absorption correction: multi-scan (SADABS; Sheldrick, 1996)
Tmin= 0.774,Tmax= 0.813
5534 measured reflections
1082 independent reflections 872 reflections withI> 2(I)
Rint= 0.044
max= 26.5
h=11!10
k=19!19
l=17!14
Refinement
Refinement onF2 R[F2> 2(F2)] = 0.030 wR(F2) = 0.071 S= 1.05 1082 reflections 74 parameters
H-atom parameters constrained
w= 1/[2(F
o2) + (0.0384P)2
+ 0.5699P]
whereP= (Fo2+ 2Fc2)/3
(/)max< 0.001
max= 0.24 e A˚
3
min=0.30 e A˚
3
Table 1
Selected geometric parameters (A˚ ,).
Cu1—N1i
2.0440 (17) Cu1—N3 2.425 (3) S1—C5 1.638 (4)
N1—C3 1.313 (2) N2—C1 1.375 (3) N2—C4 1.455 (2)
N1i—Cu1—N1ii 92.45 (9) N1i —Cu1—N1iii 87.55 (9) N1ii —Cu1—N1iii 180.00 (8) N1i—Cu1—N3 89.23 (7)
N1iii—Cu1—N3 90.77 (7) C3—N1—C2 105.88 (16) C3—N1—Cu1 126.34 (13) C5—N3—Cu1 154.8 (2)
Symmetry codes: (i)xþ1;yþ1;z; (ii)x;yþ1;z; (iii)xþ1;y;z.
H atoms were placed in calculated positions, with C—H = 0.93 or 0.97 A˚ , and included in the final cycles of refinement using a riding model, withUiso(H) = 1.2Ueq(C).
Data collection:SMART(Bruker, 1998); cell refinement:SAINT
(Bruker, 1999); data reduction:SAINT; program(s) used to solve structure:SHELXS97(Sheldrick, 1997); program(s) used to refine structure: SHELXL97 (Sheldrick, 1997); molecular graphics:
SHELXTL (Bruker, 1999); software used to prepare material for publication:SHELXTL.
The authors thank Hebei Polytechnic University for supporting this work. JR acknowledges financial support from the Spanish Government (grant No. BQU2003/00539).
References
Bruker (1998).SMART. Bruker AXS Inc., Madison, Wisconsin, USA. Bruker (1999). SAINT and SHELXTL. Bruker AXS Inc., Madison,
Wisconsin, USA.
Chen, S., Richardson, J. F. & Buchanan, R. M. (1994).Inorg. Chem.33, 2376– 2382.
metal-organic papers
m524
Cuiet al. [Cu(NCS) [image:2.610.45.298.72.167.2]2(C7H8N4)2] Acta Cryst.(2006). E62, m523–m525
Figure 1
Part of the structure of (I), showing the coordination environment of atom Cu1. Displacement ellipsoids are drawn at the 30% probability level. [symmetry codes: (A)x+ 1,y+ 1,z; (B)x,y+ 1,z; (C)
[image:2.610.94.246.235.512.2]x+ 1,y,z; (D)x,y,z].
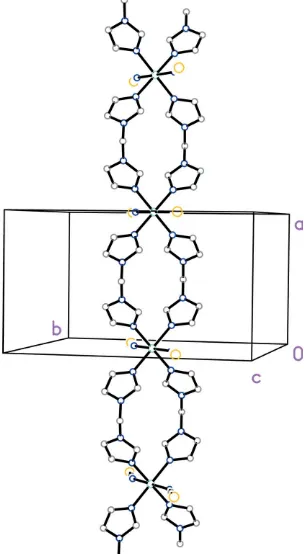
Figure 2
Cui, G.-H., Li, J.-R., Tian, J.-L., Bu, X.-H. & Batten, S. R. (2005).Cryst. Growth Des.5, 1775–1780.
Kirchner, C. & Krebs, B. (1987).Inorg. Chem.26, 3569–3576.
Ma, J.-F., Yang, J., Zheng, G.-L., Zhang, Y.-M., Li, F.-F. & Liu, J.-F. (2004).
Polyhedron,23, 553–559.
Schu¨tze, W. & Schubert, H. (1959).J. Prakt. Chem.8, 306–313.
Sheldrick, G. M. (1996).SADABS. University of Go¨ttingen, Germany. Sheldrick, G. M. (1997). SHELXS97 and SHELXL97. University of
Go¨ttingen, Germany.
Wu, L.-P., Yamagiwa, Y., Kuroda-Sowa, K., Kamikawa, T. & Munakata, M. (1997).Inorg. Chim. Acta,256, 155–159.
metal-organic papers
Acta Cryst.(2006). E62, m523–m525 Cuiet al. [Cu(NCS)
supporting information
sup-1 Acta Cryst. (2006). E62, m523–m525
supporting information
Acta Cryst. (2006). E62, m523–m525 [https://doi.org/10.1107/S1600536806004843]
catena
-Poly[[bis(isothiocyanato-
κ
N
)copper(II)]bis(
µ
-1,1
′
-methylenedi-1
H
-imidazole-
κ
N
3:
κ
N
3′)]
Guang-Hua Cui, Gui-Ying Dong and Joan Ribas
catena-Poly[[bis(isothiocyanato-κN)copper(II)]bis(µ-1,1′-methylenedi-1H- imidazole-κN3:κN3′)]
Crystal data
[Cu(NCS)2(C7H8N4)2]
Mr = 476.05
Orthorhombic, Cmca
Hall symbol: -C 2bc 2
a = 8.877 (3) Å
b = 15.602 (6) Å
c = 14.170 (4) Å
V = 1962.5 (12) Å3
Z = 4
F(000) = 972
Dx = 1.611 Mg m−3
Mo Kα radiation, λ = 0.71073 Å
Cell parameters from 932 reflections
θ = 2.6–26.2°
µ = 1.35 mm−1
T = 293 K
Block, blue
0.20 × 0.18 × 0.16 mm
Data collection
Bruker SMART CCD area-detector diffractometer
Radiation source: fine-focus sealed tube Graphite monochromator
φ and ω scans
Absorption correction: multi-scan (SADABS; Sheldrick, 1996)
Tmin = 0.774, Tmax = 0.813
5534 measured reflections 1082 independent reflections
872 reflections with I > 2σ(I)
Rint = 0.044
θmax = 26.5°, θmin = 2.6°
h = −11→10
k = −19→19
l = −17→14
Refinement
Refinement on F2
Least-squares matrix: full
R[F2 > 2σ(F2)] = 0.030
wR(F2) = 0.071
S = 1.05
1082 reflections 74 parameters 0 restraints
Primary atom site location: structure-invariant direct methods
Secondary atom site location: difference Fourier map
Hydrogen site location: inferred from neighbouring sites
H-atom parameters constrained
w = 1/[σ2(F
o2) + (0.0384P)2 + 0.5699P]
where P = (Fo2 + 2Fc2)/3
(Δ/σ)max < 0.001
Δρmax = 0.24 e Å−3
Δρmin = −0.30 e Å−3
Special details
supporting information
sup-2 Acta Cryst. (2006). E62, m523–m525
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2,
conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used
only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2
are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger.
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
x y z Uiso*/Ueq
Cu1 0.5000 0.5000 0.0000 0.02288 (16)
S1 0.5000 0.66345 (6) 0.31088 (7) 0.0567 (3)
N1 0.33373 (18) 0.43299 (10) 0.06719 (11) 0.0242 (4)
N2 0.13513 (18) 0.40312 (10) 0.15403 (11) 0.0240 (4)
N3 0.5000 0.60686 (18) 0.1242 (2) 0.0419 (7)
C1 0.1915 (2) 0.32644 (14) 0.12179 (17) 0.0386 (6)
H1A 0.1537 0.2720 0.1344 0.046*
C2 0.3122 (2) 0.34561 (13) 0.06839 (17) 0.0369 (5)
H2A 0.3720 0.3058 0.0370 0.044*
C3 0.2247 (2) 0.46553 (13) 0.11852 (14) 0.0247 (4)
H3A 0.2108 0.5238 0.1291 0.030*
C4 0.0000 0.4136 (2) 0.2110 (2) 0.0273 (7)
H4A 0.0000 0.3715 0.2614 0.033*
H4B 0.0000 0.4701 0.2395 0.033*
C5 0.5000 0.62989 (18) 0.2014 (2) 0.0310 (7)
Atomic displacement parameters (Å2)
U11 U22 U33 U12 U13 U23
Cu1 0.0153 (2) 0.0240 (3) 0.0293 (3) 0.000 0.000 0.0071 (2)
S1 0.0852 (8) 0.0423 (5) 0.0424 (5) 0.000 0.000 −0.0108 (4)
N1 0.0201 (8) 0.0227 (9) 0.0300 (9) −0.0017 (7) −0.0002 (7) 0.0026 (7)
N2 0.0173 (8) 0.0275 (9) 0.0273 (9) 0.0000 (6) 0.0008 (7) 0.0028 (7)
N3 0.0444 (18) 0.0398 (17) 0.0414 (17) 0.000 0.000 −0.0049 (14)
C1 0.0295 (12) 0.0217 (11) 0.0647 (16) −0.0015 (9) 0.0104 (11) 0.0047 (11)
C2 0.0276 (12) 0.0232 (11) 0.0598 (15) 0.0009 (9) 0.0114 (11) −0.0016 (10)
C3 0.0207 (10) 0.0239 (10) 0.0295 (11) 0.0006 (8) −0.0010 (8) 0.0020 (8)
C4 0.0180 (14) 0.0394 (17) 0.0244 (15) 0.000 0.000 0.0007 (12)
C5 0.0278 (16) 0.0186 (14) 0.047 (2) 0.000 0.000 0.0037 (14)
Geometric parameters (Å, º)
Cu1—N1i 2.0440 (17) N2—C1 1.375 (3)
Cu1—N1ii 2.0440 (17) N2—C4 1.455 (2)
Cu1—N1iii 2.0440 (17) N3—C5 1.150 (4)
Cu1—N1 2.0440 (17) C1—C2 1.345 (3)
Cu1—N3 2.425 (3) C1—H1A 0.9300
Cu1—N3i 2.425 (3) C2—H2A 0.9300
S1—C5 1.638 (4) C3—H3A 0.9300
supporting information
sup-3 Acta Cryst. (2006). E62, m523–m525
N1—C2 1.377 (3) C4—H4A 0.9700
N2—C3 1.354 (2) C4—H4B 0.9700
N1i—Cu1—N1ii 92.45 (9) C3—N2—C4 127.6 (2)
N1i—Cu1—N1iii 87.55 (9) C1—N2—C4 125.59 (19)
N1ii—Cu1—N1iii 180.00 (8) C5—N3—Cu1 154.8 (2)
N1i—Cu1—N1 180.0 C2—C1—N2 106.46 (19)
N1ii—Cu1—N1 87.55 (9) C2—C1—H1A 126.8
N1iii—Cu1—N1 92.45 (9) N2—C1—H1A 126.8
N1i—Cu1—N3 89.23 (7) C1—C2—N1 109.75 (19)
N1ii—Cu1—N3 89.23 (7) C1—C2—H2A 125.1
N1iii—Cu1—N3 90.77 (7) N1—C2—H2A 125.1
N1—Cu1—N3 90.77 (7) N1—C3—N2 111.14 (17)
N1i—Cu1—N3i 90.77 (7) N1—C3—H3A 124.4
N1ii—Cu1—N3i 90.77 (7) N2—C3—H3A 124.4
N1iii—Cu1—N3i 89.23 (7) N2iv—C4—N2 111.1 (2)
N1—Cu1—N3i 89.23 (7) N2iv—C4—H4A 109.4
N3—Cu1—N3i 180.00 (13) N2—C4—H4A 109.4
C3—N1—C2 105.88 (16) N2iv—C4—H4B 109.4
C3—N1—Cu1 126.34 (13) N2—C4—H4B 109.4
C2—N1—Cu1 127.78 (14) H4A—C4—H4B 108.0
C3—N2—C1 106.77 (17) N3—C5—S1 179.6 (3)
N1ii—Cu1—N1—C3 −56.28 (15) C3—N2—C1—C2 0.0 (2)
N1iii—Cu1—N1—C3 123.72 (15) C4—N2—C1—C2 176.93 (19)
N3—Cu1—N1—C3 32.91 (17) N2—C1—C2—N1 0.6 (3)
N3i—Cu1—N1—C3 −147.09 (17) C3—N1—C2—C1 −1.0 (2)
N1ii—Cu1—N1—C2 123.7 (2) Cu1—N1—C2—C1 179.05 (15)
N1iii—Cu1—N1—C2 −56.3 (2) C2—N1—C3—N2 1.0 (2)
N3—Cu1—N1—C2 −147.11 (18) Cu1—N1—C3—N2 −179.05 (12)
N3i—Cu1—N1—C2 32.89 (18) C1—N2—C3—N1 −0.6 (2)
N1i—Cu1—N3—C5 −133.77 (5) C4—N2—C3—N1 −177.48 (18)
N1ii—Cu1—N3—C5 133.77 (5) C3—N2—C4—N2iv 100.9 (3)
N1iii—Cu1—N3—C5 −46.23 (5) C1—N2—C4—N2iv −75.4 (3)
N1—Cu1—N3—C5 46.23 (5)