organic papers
Acta Cryst.(2006). E62, o47–o48 doi:10.1107/S1600536805039462 Wang and Zhu C
20H15FN4O
o47
Acta Crystallographica Section E Structure Reports Online
ISSN 1600-5368
4-(4-Fluorophenyl)-3-methyl-6-oxo-1-phenyl-4,5,6,7-tetrahydro-1
H
-pyrazolo[3,4-
b
]pyridine-5-carbonitrile
Jing Wang and Song-Lei Zhu*
Department of Chemistry, Xuzhou Medical College, Xuzhou 221002, People’s Republic of China
Correspondence e-mail: songleizhu@sina.com.cn
Key indicators
Single-crystal X-ray study
T= 298 K
Mean(C–C) = 0.005 A˚
Rfactor = 0.057
wRfactor = 0.165
Data-to-parameter ratio = 13.0
For details of how these key indicators were automatically derived from the article, see http://journals.iucr.org/e.
#2006 International Union of Crystallography Printed in Great Britain – all rights reserved
The title compound, C20H15FN4O, was synthesized by the
reaction of 5-amino-3-methyl-1-phenylpyrazole with ethyl 2-cyano-3-(4-fluorophenyl)-1-acylate in glycol under microwave irradiation. The tetrahydropyridine ring adopts a distorted envelope conformation. The pyrazole ring forms a dihedral angle of 39.2 (3)with the attached phenyl ring.
Comment
The pyrazolo[3,4-b]pyridine system has many interesting
biological and pharmacological properties and is used in the treatment of a wide variety of stress-related illnesses (Seki-kawaet al., 1973; Kuczynskiet al., 1979; El-Deanet al., 1991). As part of our program aimed at employing microwave irra-diation for the preparation of heterocyclic compounds (Tuet al., 2004), we have recently synthesized the title
pyrazolo[3,4-b]pyridine derivative, (I), under microwave irradiation and we report here its crystal structure.
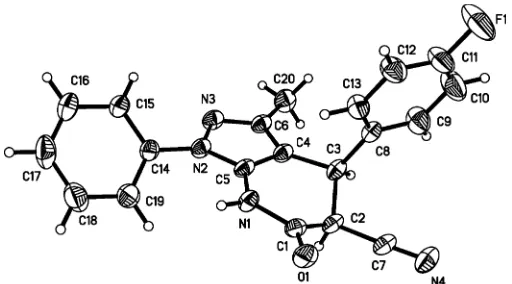
The molecular structure of (I) is shown in Fig. 1. The tetrahydropyridine ring adopts a distorted envelope confor-mation, with atom C1 and C2 deviating from the C3/C4/C5/N1 plane by 0.231 (1) and 0.731 (1) A˚ , respectively, so that C2 is the main flap atom. The pyrazole ring forms a dihedral angle of 39.2 (3) with the attached phenyl ring.
Experimental
A dry flask (25 ml) was charged with 5-amino-3-methyl-1-phenyl-pyrazole (2 mmol), ethyl 2-cyano-3-(4-fluorophenyl)-1-acylate (2 mmol) and glycol (1 ml). The unsealed reaction vessel was put into a modified household microwave oven and connected to refluxing equipment. After microwave irradiation for 5 min (250 W), the reaction mixture was cooled and washed with a small amount of ethanol. The crude product was filtered off and single crystals of (I) were obtained by recrystallization from a 95% ethanol solution (yield
80%). Spectroscopic analysis: IR (KBr,, cm1): 3360, 3074 (NH2),
2228 (CN), 1703 (CO);1H NMR (300 MHz, DMSO-d
6): 2.04 (3H,s,
CH3), 4.96–5.18 (1H,m, CH), 4.68–4.77 (1H,m, CH), 7.16–7.81 (9H, m, ArH), 11.30(1H,s, NH).
Crystal data
C20H15FN4O Mr= 346.36 Orthorhombic,Pbca a= 10.856 (5) A˚
b= 8.245 (4) A˚
c= 38.898 (17) A˚
V= 3482 (3) A˚3 Z= 8
Dx= 1.322 Mg m
3
MoKradiation Cell parameters from 2536
reflections = 2.1–21.1
= 0.09 mm1 T= 298 (2) K Block, colorless 0.450.410.35 mm
Data collection
Bruker SMART CCD area-detector diffractometer
’and!scans
Absorption correction: none 17020 measured reflections 3069 independent reflections
1712 reflections withI> 2(I)
Rint= 0.061
max= 25.0 h=12!12
k=8!9
l=34!46
Refinement
Refinement onF2 R[F2> 2(F2)] = 0.057 wR(F2) = 0.165 S= 1.03 3069 reflections 236 parameters
H-atom parameters constrained
w= 1/[2(F
o2) + (0.054P)2
+ 3.1712P]
whereP= (Fo2+ 2Fc2)/3
(/)max< 0.001
max= 0.17 e A˚ 3
min=0.20 e A˚ 3
Table 1
Selected geometric parameters (A˚ ,).
N1—C1 1.368 (4)
N1—C5 1.389 (4)
C1—C2 1.532 (5)
C2—C3 1.561 (4)
C3—C4 1.503 (4)
C4—C5 1.351 (4)
C1—N1—C5 119.8 (3)
N1—C1—C2 114.5 (3)
C7—C2—C1 109.1 (3)
C7—C2—C3 111.5 (3)
C1—C2—C3 115.3 (2)
C4—C3—C8 115.3 (3)
C4—C3—C2 104.9 (3)
C8—C3—C2 113.8 (3)
C5—C4—C3 121.5 (3)
C4—C5—N1 124.9 (3)
Methyl H atoms were placed in calculated positions, with C—H = 0.96 A˚ and torsion angles refined to fit the electron density, with
Uiso(H) = 1.5Ueq(C). Other H atoms were placed in idealized
posi-tions, with C—H = 0.930.98 A˚ and N—H = 0.86 A˚, and allowed to ride on their parent atoms, withUiso(H) = 1.2Ueq(carrier).
Data collection:SMART(Bruker, 1999); cell refinement:SAINT
(Bruker, 1999); data reduction:SAINT; program(s) used to solve structure:SHELXS97(Sheldrick, 1997); program(s) used to refine structure: SHELXL97 (Sheldrick, 1997); molecular graphics:
SHELXTL (Bruker, 1999); software used to prepare material for publication:SHELXTL.
The authors are deeply indebted to Professor S.-J. Tu and Professor D.-Q. Shi for their invaluable help. We also thank the Key Laboratory of Biotechnology for Medicinal Plants of Jiangsu Province (grant No. 01AXL14) for financial support.
References
Bruker (1999).SMART,SAINTandSHELXTL. Bruker AXS Inc., Madison, Wisconsin, USA.
El-Dean, A. M. K., Atalla, A. A., Mohamed, T. A. & Geies, A. A. (1991).Z. Naturforsch. Teil B,46, 541–546.
Kuczynski, L., Mrozikiewic, A., Banaszkiewicz, W. & Poreba, K. (1979).J. Pharmacol. Pharm.31, 217–225.
Sekikawa, I., Nishie, J., Tono-oka, S., Tanaka, Y. & Kakimoto, S. (1973).J. Heterocycl. Chem.10, 931–932.
Sheldrick, G. M. (1997). SHELXS97 and SHELXL97. University of Go¨ttingen, Germany.
Tu, S.-J., Fang, F., Zhu, S.-L., Li, T.-J., Zhang, X.-J. & Zhuang, Q.-Y. (2004).
[image:2.610.312.568.70.212.2]Synlett, pp. 537–539. Figure 1
supporting information
sup-1
Acta Cryst. (2006). E62, o47–o48
supporting information
Acta Cryst. (2006). E62, o47–o48 [doi:10.1107/S1600536805039462]
4-(4-Fluorophenyl)-3-methyl-6-oxo-1-phenyl-4,5,6,7-tetrahydro-1
H
-pyrazolo-[3,4-
b
]pyridine-5-carbonitrile
Jing Wang and Song-Lei Zhu
S1. Comment
The pyrazolo[3,4-b]pyridine system has many interesting biological and pharmacological properties and is used in the
treatment of a wide variety of stress-related illnesses (Sekikawa et al., 1973; Kuczynski et al., 1979; El-Dean et al.,
1991). As part of our program aimed at employing microwave irradiation for the preparation of heterocyclic compounds
(Tu et al., 2004), we have recently synthesized the title pyrazolo[3,4-b]pyridine derivative, (I), under microwave
irradiation and we report here its crystal structure.
The molecular structure of (I) is shown in Fig. 1. The tetrahydropyridine ring adopts a distorted envelope conformation,
with atom C1 and C2 deviating from the C3/C4/C5/N1 plane by 0.231 (1) and 0.731 (1) Å, respectively. The dihedral
angle of 0.62 (3)° between the C3/C4/C5/N1 plane and the fused pyrazole plane indicates they are coplanar. The pyrazole
ring forms a dihedral angle of 39.2 (3)° with the attached phenyl ring.
S2. Experimental
A dry flask (25 ml) was charged with 5-amino-3-methyl-1-phenylpyrazole (2 mmol), ethyl
2-cyano-3-(4-fluorophenyl)-1-acylate (2 mmol) and 1 ml glycol as energy transfer. The unsealed reaction vessel was put into a modified household
microwave oven and connected with refluxing equipment. After microwave irradiation for 5 min (250 W), the reaction
mixture was cooled and washed with small amount ethanol. The crude product was filtered and single crystals of (I) were
obtained by recrystallization from a 95% ethanol solution (yield 80%). Spectroscopic analysis: IR (KBr, ν, cm−1): 3360,
3074 (NH2), 2228 (CN), 1703 (CO); 1H NMR (300 MHz, DMSO-d6): 2.04 (3H, s, CH3), 4.96–5.18 (1H, m, CH), 4.68–
4.77 (1H, m, CH), 7.16–7.81 (9H, m, ArH), 11.30(1H, s, NH).
S3. Refinement
Methyl H atoms were placed in calculated positions, with C—H = 0.96 Å and torsion angles refined to fit the electron
density, with Uiso(H) = 1.5Ueq(C). Other H atoms were placed in idealized positions, with C—H = 0.93 − 0.98 Å and N—
Figure 1
The molecular structure of (I), showing 40% probability displacement ellipsoids (arbitrary spheres for H atoms).
4-(4-Fluorophenyl)-3-methyl-6-oxo-1-phenyl-4,5,6,7-tetrahydro-1H- pyrazolo[3,4-b]pyridine-5-carbonitrile
Crystal data C20H15FN4O
Mr = 346.36
Orthorhombic, Pbca Hall symbol: -P 2ac 2ab a = 10.856 (5) Å b = 8.245 (4) Å c = 38.898 (17) Å V = 3482 (3) Å3
Z = 8
F(000) = 1440
Dx = 1.322 Mg m−3
Melting point = 497–498 K Mo Kα radiation, λ = 0.71073 Å Cell parameters from 2536 reflections θ = 2.1–21.1°
µ = 0.09 mm−1
T = 298 K Block, colorless 0.45 × 0.41 × 0.35 mm
Data collection
Bruker SMART CCD area-detector diffractometer
Radiation source: fine-focus sealed tube Graphite monochromator
φ and ω scans
17020 measured reflections 3069 independent reflections
1712 reflections with I > 2σ(I) Rint = 0.061
θmax = 25.0°, θmin = 2.1°
h = −12→12 k = −8→9 l = −34→46
Refinement Refinement on F2
Least-squares matrix: full R[F2 > 2σ(F2)] = 0.057
wR(F2) = 0.165
S = 1.03 3069 reflections 236 parameters 0 restraints
Secondary atom site location: difference Fourier map
Hydrogen site location: inferred from neighbouring sites
H-atom parameters constrained w = 1/[σ2(F
o2) + (0.054P)2 + 3.1712P]
where P = (Fo2 + 2Fc2)/3
supporting information
sup-3
Acta Cryst. (2006). E62, o47–o48
Special details
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes.
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2,
conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used
only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2
are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger.
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
x y z Uiso*/Ueq
F1 0.5985 (3) 0.3427 (4) 0.49399 (7) 0.1356 (12)
N1 0.4743 (2) 0.1085 (3) 0.66799 (7) 0.0479 (7)
H1 0.4430 0.1726 0.6831 0.058*
N2 0.2834 (2) −0.0461 (3) 0.66026 (7) 0.0468 (7)
N3 0.2384 (2) −0.1592 (3) 0.63736 (7) 0.0511 (7)
N4 0.8660 (3) 0.0407 (4) 0.61601 (9) 0.0696 (9)
O1 0.6632 (2) 0.2236 (3) 0.67158 (6) 0.0589 (7)
C1 0.5971 (3) 0.1169 (4) 0.66039 (8) 0.0462 (8)
C2 0.6450 (2) −0.0233 (4) 0.63825 (9) 0.0463 (8)
H2 0.6520 −0.1179 0.6534 0.056*
C3 0.5599 (3) −0.0737 (4) 0.60773 (8) 0.0450 (8)
H3 0.5860 −0.1817 0.6001 0.054*
C4 0.4338 (3) −0.0907 (4) 0.62334 (8) 0.0447 (8)
C5 0.4001 (3) −0.0040 (4) 0.65128 (8) 0.0443 (8)
C6 0.3311 (3) −0.1871 (4) 0.61542 (9) 0.0477 (8)
C7 0.7699 (3) 0.0152 (4) 0.62641 (9) 0.0525 (9)
C8 0.5677 (3) 0.0389 (4) 0.57698 (9) 0.0472 (8)
C9 0.6277 (4) −0.0099 (5) 0.54775 (10) 0.0795 (13)
H9 0.6622 −0.1131 0.5468 0.095*
C10 0.6378 (5) 0.0929 (7) 0.51946 (11) 0.1000 (16)
H10 0.6781 0.0592 0.4996 0.120*
C11 0.5878 (4) 0.2421 (6) 0.52155 (10) 0.0813 (13)
C12 0.5286 (4) 0.2951 (5) 0.54961 (10) 0.0719 (11)
H12 0.4950 0.3989 0.5502 0.086*
C13 0.5183 (3) 0.1936 (4) 0.57759 (9) 0.0581 (9)
H13 0.4774 0.2296 0.5972 0.070*
C14 0.2059 (3) 0.0187 (4) 0.68625 (9) 0.0489 (8)
C15 0.0829 (3) 0.0493 (5) 0.67888 (10) 0.0618 (10)
H15 0.0513 0.0240 0.6573 0.074*
C16 0.0082 (4) 0.1170 (6) 0.70340 (12) 0.0799 (13)
H16 −0.0749 0.1343 0.6988 0.096*
C17 0.0563 (4) 0.1593 (6) 0.73481 (13) 0.0911 (15)
H17 0.0064 0.2096 0.7511 0.109*
C18 0.1774 (4) 0.1280 (6) 0.74243 (11) 0.0914 (15)
C19 0.2525 (4) 0.0561 (5) 0.71824 (9) 0.0683 (11)
H19 0.3342 0.0331 0.7235 0.082*
C20 0.3188 (3) −0.3058 (5) 0.58706 (9) 0.0667 (11)
H20A 0.2540 −0.3810 0.5923 0.100*
H20B 0.3949 −0.3637 0.5843 0.100*
H20C 0.2996 −0.2497 0.5661 0.100*
Atomic displacement parameters (Å2)
U11 U22 U33 U12 U13 U23
F1 0.201 (3) 0.136 (3) 0.0688 (17) 0.016 (2) 0.0285 (19) 0.0418 (18)
N1 0.0346 (14) 0.0499 (17) 0.0592 (18) −0.0046 (13) 0.0005 (12) −0.0043 (14)
N2 0.0359 (14) 0.0485 (17) 0.0560 (17) −0.0087 (13) −0.0018 (12) 0.0018 (13)
N3 0.0400 (15) 0.0506 (18) 0.0628 (18) −0.0125 (13) −0.0065 (14) 0.0014 (14)
N4 0.0414 (17) 0.066 (2) 0.101 (3) −0.0054 (16) 0.0015 (17) 0.0100 (18)
O1 0.0483 (13) 0.0606 (16) 0.0679 (16) −0.0215 (12) −0.0073 (11) −0.0004 (13)
C1 0.0380 (18) 0.049 (2) 0.052 (2) −0.0054 (16) −0.0073 (15) 0.0135 (16)
C2 0.0297 (16) 0.0400 (19) 0.069 (2) −0.0011 (14) −0.0055 (15) 0.0145 (16)
C3 0.0347 (16) 0.0344 (18) 0.066 (2) −0.0003 (14) 0.0011 (15) −0.0011 (16)
C4 0.0349 (16) 0.0435 (19) 0.056 (2) −0.0051 (15) −0.0046 (15) 0.0039 (16)
C5 0.0306 (16) 0.045 (2) 0.057 (2) −0.0051 (14) −0.0063 (15) 0.0077 (17)
C6 0.0386 (17) 0.045 (2) 0.059 (2) −0.0042 (15) −0.0061 (16) 0.0054 (17)
C7 0.0363 (19) 0.048 (2) 0.074 (2) −0.0001 (16) −0.0085 (17) 0.0060 (18)
C8 0.0363 (17) 0.047 (2) 0.058 (2) −0.0011 (16) 0.0007 (15) −0.0040 (17)
C9 0.092 (3) 0.076 (3) 0.071 (3) 0.030 (3) 0.020 (2) −0.003 (2)
C10 0.129 (4) 0.114 (4) 0.056 (3) 0.027 (4) 0.030 (3) 0.004 (3)
C11 0.105 (3) 0.086 (3) 0.053 (3) 0.003 (3) 0.005 (2) 0.021 (2)
C12 0.090 (3) 0.064 (3) 0.062 (3) 0.009 (2) 0.004 (2) 0.013 (2)
C13 0.065 (2) 0.053 (2) 0.056 (2) 0.0035 (19) 0.0079 (18) 0.0020 (18)
C14 0.0394 (17) 0.055 (2) 0.053 (2) −0.0078 (16) 0.0052 (15) 0.0059 (17)
C15 0.0405 (19) 0.084 (3) 0.061 (2) −0.0038 (19) 0.0048 (17) 0.008 (2)
C16 0.047 (2) 0.106 (4) 0.087 (3) −0.001 (2) 0.020 (2) 0.002 (3)
C17 0.074 (3) 0.114 (4) 0.085 (3) −0.019 (3) 0.038 (3) −0.013 (3)
C18 0.080 (3) 0.133 (4) 0.061 (3) −0.028 (3) 0.012 (2) −0.011 (3)
C19 0.052 (2) 0.090 (3) 0.063 (2) −0.017 (2) 0.002 (2) 0.002 (2)
C20 0.059 (2) 0.070 (3) 0.071 (3) −0.020 (2) 0.0012 (19) −0.006 (2)
Geometric parameters (Å, º)
F1—C11 1.360 (5) C9—C10 1.393 (6)
N1—C1 1.368 (4) C9—H9 0.9300
N1—C5 1.389 (4) C10—C11 1.347 (6)
N1—H1 0.8600 C10—H10 0.9300
N2—C5 1.359 (4) C11—C12 1.340 (6)
N2—N3 1.379 (4) C12—C13 1.377 (5)
N2—C14 1.420 (4) C12—H12 0.9300
supporting information
sup-5
Acta Cryst. (2006). E62, o47–o48
O1—C1 1.216 (4) C14—C15 1.388 (5)
C1—C2 1.532 (5) C15—C16 1.371 (5)
C2—C7 1.466 (4) C15—H15 0.9300
C2—C3 1.561 (4) C16—C17 1.374 (6)
C2—H2 0.9800 C16—H16 0.9300
C3—C4 1.503 (4) C17—C18 1.372 (6)
C3—C8 1.516 (5) C17—H17 0.9300
C3—H3 0.9800 C18—C19 1.379 (5)
C4—C5 1.351 (4) C18—H18 0.9300
C4—C6 1.403 (4) C19—H19 0.9300
C6—C20 1.481 (5) C20—H20A 0.9600
C8—C9 1.370 (5) C20—H20B 0.9600
C8—C13 1.385 (5) C20—H20C 0.9600
C1—N1—C5 119.8 (3) C10—C9—H9 119.5
C1—N1—H1 120.1 C11—C10—C9 118.4 (4)
C5—N1—H1 120.1 C11—C10—H10 120.8
C5—N2—N3 109.7 (3) C9—C10—H10 120.8
C5—N2—C14 129.7 (3) C12—C11—C10 122.7 (4)
N3—N2—C14 120.3 (2) C12—C11—F1 118.9 (5)
C6—N3—N2 105.2 (2) C10—C11—F1 118.4 (4)
O1—C1—N1 122.3 (3) C11—C12—C13 119.0 (4)
O1—C1—C2 123.2 (3) C11—C12—H12 120.5
N1—C1—C2 114.5 (3) C13—C12—H12 120.5
C7—C2—C1 109.1 (3) C12—C13—C8 121.0 (3)
C7—C2—C3 111.5 (3) C12—C13—H13 119.5
C1—C2—C3 115.3 (2) C8—C13—H13 119.5
C7—C2—H2 106.8 C19—C14—C15 119.9 (3)
C1—C2—H2 106.8 C19—C14—N2 120.6 (3)
C3—C2—H2 106.8 C15—C14—N2 119.4 (3)
C4—C3—C8 115.3 (3) C16—C15—C14 120.0 (4)
C4—C3—C2 104.9 (3) C16—C15—H15 120.0
C8—C3—C2 113.8 (3) C14—C15—H15 120.0
C4—C3—H3 107.5 C15—C16—C17 119.8 (4)
C8—C3—H3 107.5 C15—C16—H16 120.1
C2—C3—H3 107.5 C17—C16—H16 120.1
C5—C4—C6 105.1 (3) C18—C17—C16 120.6 (4)
C5—C4—C3 121.5 (3) C18—C17—H17 119.7
C6—C4—C3 133.4 (3) C16—C17—H17 119.7
C4—C5—N2 108.9 (3) C17—C18—C19 120.0 (4)
C4—C5—N1 124.9 (3) C17—C18—H18 120.0
N2—C5—N1 126.2 (3) C19—C18—H18 120.0
N3—C6—C4 111.1 (3) C14—C19—C18 119.7 (4)
N3—C6—C20 121.3 (3) C14—C19—H19 120.2
C4—C6—C20 127.5 (3) C18—C19—H19 120.2
N4—C7—C2 177.0 (4) C6—C20—H20A 109.5
C9—C8—C13 117.9 (3) C6—C20—H20B 109.5
C13—C8—C3 121.9 (3) C6—C20—H20C 109.5
C8—C9—C10 121.0 (4) H20A—C20—H20C 109.5
C8—C9—H9 119.5 H20B—C20—H20C 109.5
C5—N2—N3—C6 1.9 (3) C3—C4—C6—N3 179.6 (3)
C14—N2—N3—C6 176.4 (3) C5—C4—C6—C20 −179.8 (3)
C5—N1—C1—O1 172.2 (3) C3—C4—C6—C20 −0.7 (6)
C5—N1—C1—C2 −10.7 (4) C4—C3—C8—C9 −132.2 (4)
O1—C1—C2—C7 −13.8 (4) C2—C3—C8—C9 106.5 (4)
N1—C1—C2—C7 169.2 (3) C4—C3—C8—C13 49.4 (4)
O1—C1—C2—C3 −140.1 (3) C2—C3—C8—C13 −71.8 (4)
N1—C1—C2—C3 42.8 (4) C13—C8—C9—C10 −0.4 (6)
C7—C2—C3—C4 −173.9 (3) C3—C8—C9—C10 −178.9 (4)
C1—C2—C3—C4 −48.8 (3) C8—C9—C10—C11 0.4 (8)
C7—C2—C3—C8 −47.1 (4) C9—C10—C11—C12 −0.3 (8)
C1—C2—C3—C8 78.0 (3) C9—C10—C11—F1 179.6 (4)
C8—C3—C4—C5 −97.6 (4) C10—C11—C12—C13 0.1 (7)
C2—C3—C4—C5 28.3 (4) F1—C11—C12—C13 −179.7 (4)
C8—C3—C4—C6 83.4 (4) C11—C12—C13—C8 −0.1 (6)
C2—C3—C4—C6 −150.6 (3) C9—C8—C13—C12 0.3 (5)
C6—C4—C5—N2 0.6 (4) C3—C8—C13—C12 178.7 (3)
C3—C4—C5—N2 −178.5 (3) C5—N2—C14—C19 −41.8 (5)
C6—C4—C5—N1 −179.9 (3) N3—N2—C14—C19 144.9 (3)
C3—C4—C5—N1 0.9 (5) C5—N2—C14—C15 136.0 (4)
N3—N2—C5—C4 −1.6 (4) N3—N2—C14—C15 −37.2 (5)
C14—N2—C5—C4 −175.4 (3) C19—C14—C15—C16 0.1 (6)
N3—N2—C5—N1 179.0 (3) N2—C14—C15—C16 −177.8 (4)
C14—N2—C5—N1 5.1 (5) C14—C15—C16—C17 2.2 (7)
C1—N1—C5—C4 −11.9 (5) C15—C16—C17—C18 −2.8 (7)
C1—N1—C5—N2 167.5 (3) C16—C17—C18—C19 1.2 (8)
N2—N3—C6—C4 −1.5 (4) C15—C14—C19—C18 −1.8 (6)
N2—N3—C6—C20 178.8 (3) N2—C14—C19—C18 176.1 (4)