9
Genomic Approaches for Abiotic
Stress Tolerance in Sorghum
Santosh Deshpande, Sujay Rakshit, K.G. Manasa,
Sarita Pandey and Rajeev Gupta
Abstract
Although sorghum is a crop grown under harsh environments, its productivity is adversely affected by various abiotic stresses including drought, temperature extremes, low fertility, and mineral toxicity among others. In recent years a large number of genetic and genomic resources have become available in sorghum, which provide researchers opportunities to relate sequence variations with phenotypic traits of interest and their utilization in sorghum improvement programs. The application of the molecular marker and genomic technologies has shown promise for efficient breeding. However, very few successful examples are available in the public domain of research in this direction. Some of these successes specifically related to application of molecular marker technologies for improving abiotic stresses are explained in this chapter. With recent advances in next-generation sequencing technologies and high-throughput phenotyping platforms/technologies, utilizing the new/advanced mapping populations such as nested-association mapping (NAM), backcross-derived NAM has shown great potential. These recent advancements will be the drivers for integration of genomics technologies in routine breeding programs in the immediate future.
1
Introduction
Sorghum [Sorghum bicolorL. (Moench)] is vital to the food security of many of the world’s poorest people living in fragile agroecological zones. Sorghum is produced by about 100 countries in the world and mainly used as staple food in parts of Asia and Africa, whereas in the United States, Mexico, and Australia it is used as a major feed crop (Rakshit et al. 2014). The genome of sorghum is unique as there is only*3 % differential sorghum–rice gene loss
S. Deshpande (&)K.G. ManasaS. Pandey R. Gupta
International Crops Research Institute for the Semi-Arid Tropics (ICRISAT), Patancheru 502 324, Telangana, India
e-mail: s.deshpande@cgiar.org S. Rakshit
ICAR-Indian Institute of Millets Research, Rajendranagar, Hyderabad, Telangana, India ©Springer International Publishing AG 2016
S. Rakshit and Y.-H. Wang (eds.),The Sorghum Genome,
Compendium of Plant Genomes, DOI 10.1007/978-3-319-47789-3_9
and comparatively less structural rearrangement for sorghum–rice in 50 million years than sor-ghum–maize rearrangement (with genome duplication events) in*15 million years in evolutionary lineage (Bowers et al. 2003). The sorghum karyotype shows 70 million years of
“abstinence” from genome duplication. Researchers have exploited this great opportunity to study functions of sorghum genes which may still resemble those of the common cereal ancestor. Accordingly, the sequenced*740 Mb sorghum genome (Paterson et al. 2009) is a logical complement to that of Oryza (rice) for grass functional genomics.
In addition to its importance as a model crop, sorghum is the most drought-resistant among the world’s top five cereal crops, and an important dual-purpose crop. Sorghum cultivation ranges from the equator (approximately 50° latitude) to elevations of 2500 m (Rao et al.2015). Sorghum is exceptionally tolerant to low input levels (especially in West Africa), which is an impor-tant characteristic for the areas receiving little rainfall. Thus, sorghum plays a vital role in feeding the world’s most vulnerable population under the context of increasing demand for lim-ited fresh water supply, increasing use of mar-ginal farmland, and climatic trends. These interesting facts about sorghum along with recent advances in genomics with the advent of the next-generation sequencing (NGS) platforms for genotyping, expression studies, and high-throughput phenotyping facilities make it a model plant species not only to study evolu-tionary relationships across other grass species but also to help understand the mechanisms and functions of the genome that will lead to better adaptation to climate change, especially for abi-otic stresses such as drought, salinity, cold tol-erance, and nutrient use efficiency. This is more relevant in the current scenario where global food production has to be increased by 70 % in the background of ever-shrinking cultivable land and water resources (MacIntyre et al. 2009), and increased incidences of environmental extremes such as floods, drought, and extreme tempera-tures (Mickelbart et al. 2015). Wang et al. (2014a,b) made a detailed review of the status of
abiotic stress genomics in sorghum vis-à-vis other cereals, and Anami et al. (2015) reviewed biotic and abiotic stress resistance with specific reference to sweet sorghum. In this chapter, efforts have been made to update the status in this area, keeping the developments in perspective.
2
Stay-Green as a Post-flowering
Drought Tolerance Trait
2.1
Importance of Stay-Green
Among the various abiotic stress factors affecting crop growth and productivity, water stress is the single greatest factor and this will be the most important factor under a changed climatic regime (Araus et al.2002). The impact of drought may largely be addressed through genetic improve-ment for drought response (Mutava et al.2011a, b). In sorghum, two distinct drought-stress responses have been identified based on the time of occurrence: a pre-flowering drought response occurring prior to anthesis and a post-flowering drought response during the grain-filling stage (Harris et al. 2007). Even though a number of commonly grown cultivars show tolerance at the pre-anthesis stage, they are not tolerant at post-anthesis stages (Sanchez et al. 2002), which is of more economic consequence. Post-flowering drought susceptibility symptoms are characterized by premature leaf and plant senescence, stalk lodging, charcoal rot, and reduction in seed sizes (Rosenow and Clark 1995). Retention of green leaf area during the grain-filling stage has been found to be associ-ated with post-flowering drought tolerance in sorghum (Rakshit et al. 2016). This capacity of certain genotypes in sorghum is referred as the
“stay-green”phenotype, and is one of the most well-characterized and utilized traits in sorghum improvement (Rosenow and Clark1982; Borrell et al. 2000a, b; Borrell and Hammer 2000; Rakshit et al. 2016). The expression of stay-green has been reported to improve the quality of crop residues (van Oosterom et al. 1996), support the continuation of carbon fi xa-tion and supply of starch to the sink (McBee
et al.1983), prevent premature death and lodging of crop (Rosenow and Clark1982), sustain grain filling under water stress (Rosenow et al.1983), and improve grain yield under moisture stress (Borrell and Douglas1996). Even though some progress has been reported in identification of factors underpinning stay-green expression (Vadez et al. 2011; Kholová et al. 2014), understanding the genetic regulation of the mechanisms that lead to the expression of the stay-green phenotype in sorghum is incomplete. Early works considered the benefit of stay-green in terms of extending the period during which a leaf could actively fix carbon (McBee et al. 1983). Subsequently, it was related to the carbon economy of the plant, addressing the nitrogen status of the plant and, in particular, the balance between nitrogen demand and nitrogen capture (Borrell et al.2001). Owing to its importance as a most widely characterized drought tolerance component trait in sorghum breeding, several quantitative trait locus (QTL) mapping studies (Tuinstra et al.1997; Crasta et al.1999; Subudhi et al. 2000; Tao et al. 2000; Xu et al. 2000; Kebede et al. 2001; Haussmann et al. 2002; Sanchez et al.2002; Harris et al.2007; Srinivas et al. 2009; Habyarimana et al. 2010; Sabadin et al. 2012) for stay-green are reported in sor-ghum and are well documented (Mace and Jor-dan 2010; Kiranmayee et al. 2015). There are several well-documented reviews about the pro-gress of research on stay-green in sorghum (Vadez et al.2013; Wang et al. 2014a,b).
2.2
QTL Analysis for Stay-Green
and Associated Root Traits
Overall, seven sources of the stay-green trait have thus far been used for identification of QTLs for this phenotype, and these are B35 (Tuinstra et al.1997; Crasta et al.1999; Subudhi et al.2000; Xu et al.2000; Sanchez et al.2002; Harris et al. 2007), E36-1 (Haussmann et al. 2002), QL41 (Tao et al. 2000), SC56 (Kebede et al.2001), 296B (Srinivas et al. 2009), SC283 (Sabadin et al. 2012), and SDS 1948-3
(Habyarimana et al. 2010). Among these, the most commonly used source is B35 [BTx642, a BC1 derivative of IS12555, a durra sorghum from Ethiopia (Rosenow et al.1983)]. All studies showed that stay-green is quantitatively inher-ited, and the QTLs varied across environments and years. However, six major stay-green QTLs, stgC(SBI-01),stg3Aandstg3B(on SBI-02),stg1 andstg2(on SBI-03), andstg4(on SBI-05) have been detected across several studies (Tuinstra et al. 1997; Crasta et al. 1999; Subudhi et al. 2000; Tao et al. 2000; Xu et al. 2000; Hauss-mann et al. 2002; Sanchez et al. 2002; Harris et al. 2007). Although the mapped QTLs are from different mapping populations, their physi-cal positions are consistent across the maps (Wang et al.2014a,b). A simple sequence repeat (SSR) framework map aligned to sorghum gen-ome assembly and the consensus QTL intervals for stay-green traits from these studies (Mace and Jordan 2011a, b) are integrated (Fig.1; http:// cmap.icrisat.ac.in/cmap/) for selecting additional markers in the intervals, especially to support introgression work. Many studies have attempted to use the identified stay-green QTLs for devel-oping drought-tolerant cultivars through marker-assisted backcrossing (Kassahun et al. 2010; Jordan et al.2012; Vadez et al.2013).
Root traits are assumed to play a major role towards improved drought tolerance. (For details refer to Chap. 11.) The nodal root angle in sor-ghum is hypothesized to influence both hori-zontal and vertical exploration of roots in the soil (Kato et al. 2006; Hammer et al. 2009; Singh et al. 2010; Mace et al. 2012). A putative asso-ciation between narrow root angle and moderate to high levels of stay-green was made while studying genetic diversity for nodal root angle in a set of 44 diverse sorghum genotypes (Singh et al. 2011). As sorghum produces only one seminal root and the major root system forms nodal root axes, the root angle measured on nodal roots, the first flush of which appears approximately at thefive-leaf stage, is suggested to be the most important parameter (Singh et al. 2010). Thus, small soil-filled root chambers can be used effectively to grow plants for a few
weeks to study the nodal root angles (Singh et al. 2010,2012). A seedling root angle tends to have medium to high heritability (Mace et al. 2012; Singh et al. 2011). Using the rapid root angle screening strategy, Mace et al. (2012) identified four QTLs for nodal root angle in sorghum, explaining 29.78, 11.65, 10.01, and 6.72 % of total phenotypic variation, respectively. These were further confirmed in a backcross-derived
nested association mapping population. Three of the four identified QTLs showed homology to previously identified root angle QTL in rice and maize, whereas all four QTLs colocalized with previously identified QTL for stay-green in sor-ghum. This has opened up the scope to use these reported QTLs towards developing sorghum genotypes with better rooting systems, making them water- and nutrient-efficient genotypes. The
Fig. 1 Consensus QTL intervals for stay-green QTLs from Mace and Jordan (2011a,b) aligned to physically aligned SSRs (vertical bars indicate QTL interval tracks; red tracks for maturity genes/QTLs; aqua for panicle
traits, and grey tracks for stay-green).aQTL tracks for sorghum chromosome SBI-01 to SBI-05.b QTL tracks for sorghum chromosome SBI-06 to SBI-10
projection of the six additional significant mar-ker–trait associations for nodal root angle iden-tified in the association mapping study identified thatfive out of six (83 %) of these new potential QTLs were also found to colocate with regions of the genome containing previously mapped stay-green QTLs (Mace et al.2012).
In a recent study at the ICAR-Indian Institute of Millets Research, Hyderabad, India, a refer-ence set for drought study comprising 96 genotypes has been developed through a com-bination of detailed multilocation drought response phenotyping of 258 diverse sorghum genotypes and their molecular diversity analysis using 39 polymorphic SSR markers (Rakshit et al.2014). Through double digest restriction-site associated DNA ddRAD sequencing of this reference set, 235,009 genomewide single nucleotide polymorphisms (SNPs) have been identified, 1439 of which fall within the coor-dinates of reported QTLs associated with drought tolerance in sorghum (Table1; Sujay Rakshit unpublished data).
2.3
Stay-Green QTL Introgression:
Successes and Lessons
Learned
The initial introgression work carried out at ICRISAT, Patancheru, India, has further dis-sected the genetic function of the stay-green QTLs. These studies suggest that stg3A and st-g3B QTLs are responsible for transpiration effi -ciency and vapor pressure deficit response, and are also found to be most stable across several genetic backgrounds and across years (personal communication: Santosh Deshpande). The marker-assisted backcross (MABC) research at ICRISAT till the late 2000s were based on SSRs alone (Vadez et al.2013), and a very large QTL interval that almost covered the complete chro-mosome arm was used. These datasets are pub-lically available (www.icrisat/cmap/ac.in). With progress in the understanding of the genetic regulation of the stay-green phenotype in sor-ghum, the current introgression of genomic regions involved in expression of the stay-green
Table 1 SNPs associated with drought-related QTLs (Sujay Rakshit personal communication unpublished data) Chromosome
no.
Trait QTL Physical position No. of
SNP Starting
point
Ending point
SB01 Shoot dry weight qSDW1_1# 59236915 64917463 75
SB02 Stay-green Stg3@ 26010527 28923791 42
Root dry weight qRDW1_2# 71213786 77627131 124
SB03 Stay-green Stg2@ 19344798 22773106 35
Stay-green Stg1@ 25307987 25969626 5
SB05 Stay-green Stg4@ 5166739 12984849 111
Nodal root angle qRA1_5# 13464752 45972507 278
Root dry weight, Shoot dry weight
qRDW1_5# 50053880 51943001 15
Nodal root angle qRA2_5# 54167643 55807231 28
SB08 Nodal root angle qRA1_8# 8026484 41653341 358
Root dry weight, Total leaf area
qRDW1_8, qTLA1_8, qTLA2_8#
47089939 50926404 66
Total leaf area qTLA3_8# 54033969 54997392 20
SB10 Bloom BLMC$ 56693 11980745 257
Nodal root angle qRA1_10# 57009865 58763524 25
phenotype will further lead a well-targeted manipulation of individual components. For instance, it is reported that B35 donor parent alleles at stay-green QTL stg1 contribute to increased water extraction by the moderately senescent caudatum line, S35 (Vadez et al. 2011); however, stg1 failed to show the same phenomenon in the genetic background of the highly senescent durra line, R16 (Vadez et al. 2011). Therefore, it is necessary to identify better donors for each of the component traits of the stay-green phenotype, which may vary with the genetic backgrounds, specific soil, water, and temperature regimes in which improved drought tolerance is desired. Among the investigated six QTLs, stg3A and stg3B QTLs are more stable
across genetic backgrounds and environments. The stg3A and stg3B QTLs are located next to each other on SBI-02, and the mesocarp gene,Z, which governs the grain seed coat color, is located between the two QTL intervals (Mace and Jordan 2010). This poses a problem in selection for grain color coupled with stay-green traits. Similarly, location of the maturity gene Ma5, just after thestg3Binterval, also adversely
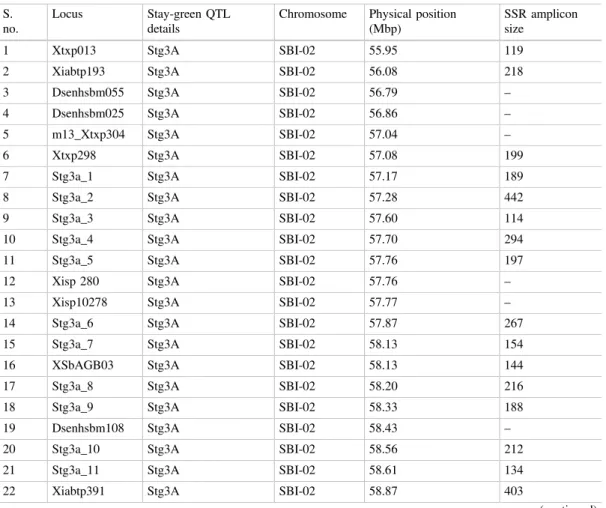
affectsfixation of theflowering time (Kim2003). QTL analysis (Srinivas et al.2008,2009) along with in silico studies (Ramu et al. 2010) helped to add several SSRs in the region represented by the mentioned QTLs (Table2). By exploiting existing backcross generations in ongoing MABC projects, the QTL interval forstg3Aand
Table 2 A list of SSRs available for stg3A and stg3B QTL region including the original SSRs listed in Vadez et al. (2013)
S. no.
Locus Stay-green QTL
details
Chromosome Physical position (Mbp)
SSR amplicon size
1 Xtxp013 Stg3A SBI-02 55.95 119
2 Xiabtp193 Stg3A SBI-02 56.08 218
3 Dsenhsbm055 Stg3A SBI-02 56.79 –
4 Dsenhsbm025 Stg3A SBI-02 56.86 –
5 m13_Xtxp304 Stg3A SBI-02 57.04 –
6 Xtxp298 Stg3A SBI-02 57.08 199
7 Stg3a_1 Stg3A SBI-02 57.17 189
8 Stg3a_2 Stg3A SBI-02 57.28 442
9 Stg3a_3 Stg3A SBI-02 57.60 114
10 Stg3a_4 Stg3A SBI-02 57.70 294
11 Stg3a_5 Stg3A SBI-02 57.76 197
12 Xisp 280 Stg3A SBI-02 57.76 –
13 Xisp10278 Stg3A SBI-02 57.77 –
14 Stg3a_6 Stg3A SBI-02 57.87 267
15 Stg3a_7 Stg3A SBI-02 58.13 154
16 XSbAGB03 Stg3A SBI-02 58.13 144
17 Stg3a_8 Stg3A SBI-02 58.20 216
18 Stg3a_9 Stg3A SBI-02 58.33 188
19 Dsenhsbm108 Stg3A SBI-02 58.43 –
20 Stg3a_10 Stg3A SBI-02 58.56 212
21 Stg3a_11 Stg3A SBI-02 58.61 134
22 Xiabtp391 Stg3A SBI-02 58.87 403
Table 2 (continued) S.
no.
Locus Stay-green QTL
details
Chromosome Physical position (Mbp)
SSR amplicon size
23 Xiabtp265 Stg3A SBI-02 58.91 246
24 Stg3a_12 Stg3A SBI-02 58.95 166
25 Stg3a_13 Stg3A SBI-02 59.07 166
26 Xcup63 Stg3A SBI-02 59.10 145
27 Xtxp464 Stg3A SBI-02 59.20 140
28 Xiabtp80 Stg3A SBI-02 59.20 176
29 Stg3a_14 Stg3A SBI-02 59.75 153
30 Stg3a_15 Stg3A SBI-02 59.82 201
31 Stg3a_16 Stg3A SBI-02 59.88 156
32 Stg3a_17 Stg3A SBI-02 59.95 253
33 Stg3a_18 Stg3A SBI-02 59.97 204
34 msbcir339 Stg3A SBI-02 60.19 176
35 Stg3a_19 Stg3A SBI-02 60.19 131
36 Stg3a_20 Stg3A SBI-02 60.21 202
37 Stg3a_21 Stg3A SBI-02 60.23 227
38 Stg3a_22 Stg3A SBI-02 60.28 208
39 Stg3a_23 Stg3A SBI-02 60.28 176
40 Stgnhsbm34 Stg3A SBI-02 60.44 –
41 Xtxp214 Stg3A SBI-02 60.44 220
42 Xtxp445 Stg3A SBI-02 60.45 238
43 Xcup29 Stg3A SBI-02 60.45 191
44 Stg3a_24 Stg3A SBI-02 60.46 125
45 Stg3a_25 Stg3A SBI-02 60.48 224
46 Stg3a_26 Stg3A SBI-02 60.51 162
47 Xtxp466 Stg3A SBI-02 60.65 159
48 Xtxp465 Stg3A SBI-02 60.67 177
49 Xiabtp509 Stg3A SBI-02 60.82 240
50 Stg3a_27 Stg3A SBI-02 60.84 103
51 Xtxp430 Stg3A SBI-02 61.09 158
52 Xtxp430 Stg3A SBI-02 61.09 158
53 Xsbarslbk2.61 Stg3A SBI-02 61.09 –
54 Stg3a_28 Stg3A SBI-02 61.13 221
55 Stg3a_29 Stg3A SBI-02 61.13 113
56 Xisep0934 Stg3A SBI-02 61.22 196
57 Stg3a_30 Stg3A SBI-02 61.26 148
58 Stg3a_31 Stg3A SBI-02 61.33 169
59 Xtxp001 Stg3A SBI-02 61.37 211
60 Stg3a_32 Stg3A SBI-02 61.37 184
Table 2 (continued) S.
no.
Locus Stay-green QTL
details
Chromosome Physical position (Mbp)
SSR amplicon size
61 Stg3a_33 Stg3A SBI-02 61.47 99
62 Dsenhsbm032 Stg3A SBI-02 61.47 –
63 Xisep0926 Stg3A SBI-02 61.54 191
64 Xtxp056 Stg3A SBI-02 61.57 347
65 Stgnhsbm35 Stg3A SBI-02 61.66 242
66 Stg3a_34 Stg3A SBI-02 61.83 268
67 Stg3a_35 Stg3A SBI-02 61.84 146
68 Stgnhsbm36 Stg3A SBI-02 61.95 180
69 Stg3a_36 overlap region SBI-02 62.13 212
70 Stg3a_37 overlap region SBI-02 62.29 186
71 Xisep0913 overlap region SBI-02 62.40 208
72 Xisp10336 overlap region SBI-02 62.69 –
73 Xisp 336 overlap region SBI-02 62.69 –
74 Xisep0941 overlap region SBI-02 63.12 188
75 Dsenhsbm045 overlap region SBI-02 63.13 –
76 Stgnhsbm31 overlap region SBI-02 63.13 –
77 Xiabtp231 overlap region SBI-02 63.13 159
78 Stgnhsbm39 overlap region SBI-02 63.20 –
79 Xtxp286 overlap region SBI-02 63.39 –
80 Xgap84 overlap region SBI-02 63.39 –
81 Stgnhsbm40 overlap region SBI-02 63.44 –
82 Xisep0938 overlap region SBI-02 63.44 205
83 Xisep0938 overlap region SBI-02 63.44 –
84 Xiabtp226 overlap region SBI-02 63.49 283
85 Xcup41 overlap region SBI-02 63.62 231
86 Stgnhsbm42 overlap region SBI-02 64.87 –
87 Xisep0849 overlap region SBI-02 64.91 –
88 Xisp10334 overlap region SBI-02 65.28 –
89 Xisep0944 overlap region SBI-02 65.30 –
90 Xisep0939 overlap region SBI-02 65.48 –
91 Xisep1022 overlap region SBI-02 65.48 –
92 Xisep0942 overlap region SBI-02 65.83 –
93 Xisep0910 overlap region SBI-02 65.95 194
94 Xgpsb128 overlap region SBI-02 66.17 285
95 Ungnhsbm49 overlap region SBI-02 67.12 –
96 XmSbCIR187 overlap region SBI-02 67.19 –
97 Xtxp348 overlap region SBI-02 67.32 –
98 Xisep0935 overlap region SBI-02 67.41 –
Table 2 (continued) S.
no.
Locus Stay-green QTL
details
Chromosome Physical position (Mbp) SSR amplicon size 99 Xisp10200 Stg3B SBI-02 67.80 – 100 Xiabtp323 Stg3B SBI-02 68.22 191 101 Dsenhsbm015 Stg3B SBI-02 68.22 – 102 Xtxp428 Stg3B SBI-02 68.39 213 103 Xisp10259 Stg3B SBI-02 68.39 – 104 Xisp 259 Stg3B SBI-02 68.39 – 105 Xiabtp397 Stg3B SBI-02 68.69 188 106 Xtxp429 Stg3B SBI-02 68.85 214 107 Stg3b_1 Stg3B SBI-02 69.31 152 108 Stg3b_2 Stg3B SBI-02 69.61 209 109 Xtxp100 Stg3B SBI-02 69.64 116 110 Stg3b_3 Stg3B SBI-02 70.09 188 111 Stg3b_4 Stg3B SBI-02 70.20 164 112 Xtxp207 Stg3B SBI-02 70.26 184 113 Xtxp007 Stg3B SBI-02 70.26 230 114 Xcup26 Stg3B SBI-02 70.26 220 115 Stg3b_5 Stg3B SBI-02 70.42 188 116 Stg3b_6 Stg3B SBI-02 70.60 99 117 Stg3b_7 Stg3B SBI-02 70.66 175 118 Stg3b_8 Stg3B SBI-02 70.67 151 119 Xisep0733 Stg3B SBI-02 70.75 330 120 Stg3b_9 Stg3B SBI-02 70.85 225 121 Xiabtp190 Stg3B SBI-02 70.89 215 122 Xisep0841 Stg3B SBI-02 70.89 215 123 Xiabtp388 Stg3B SBI-02 71.03 129 124 Xtxp296 Stg3B SBI-02 71.11 168 125 Stg3b_10 Stg3B SBI-02 71.20 277 126 Stg3b_11 Stg3B SBI-02 71.43 219 127 Xiabtp484 Stg3B SBI-02 71.60 117 128 Xiabtp205 Stg3B SBI-02 71.79 165 129 Xiabtp076 Stg3B SBI-02 72.17 292 130 Xiabtp317 Stg3B SBI-02 72.33 236 131 Xcup40 Stg3B SBI-02 75.36 193 132 Stg3b_1 Stg3B SBI-02 77.29 140 133 Xtxp019 Stg3B SBI-02 – – 134 Xtxp008 Stg3B SBI-02 – –
stg3B QTL has been reduced. At ICRISAT additional efforts are currently being made to identify SNPs representing this region of the genome and currently a set of*70 SNPs from genic regions have already been identified (San-tosh Deshpande, personal communication).
These additional SSRs facilitate not only the selection for the stay-green QTLs independently but also for stringent mesocarp color in segre-gating populations. Similarly, as elaborated ear-lier, a major nodal root angle QTL distal tostg3B (Mace et al. 2012) has provided further oppor-tunities for improvement of drought tolerance by selecting these QTLs simultaneously or in com-bination. The QTL intervals reported by Vadez et al. (2013) also overlap this root angle QTL within the stg3B region. This in turn is an opportunity to evaluate the available introgres-sion lines (Vadez et al.2011) for the combined effort of stay-green QTL(s) and root angle QTL in different combinations. Recent advances in the area of accurate and precise phenotyping plat-forms are adding values to dissect complex traits such as stay-green. Vadez et al. (2011) utilized the lysimeter-based estimations for the mecha-nism responsible for stg3A and stg3B as water-use efficiency under water-limited condi-tions and vapor pressure deficit for better tran-spiration efficiency. These traits help direct trait value measurement rather than stay-green expression. But they are cumbersome to mea-sure for a large breeding or segregating popula-tion. At ICRISAT, a high-throughput phenotyping platform called the “LeasyScan” facility has been established to measure leaf area in a quicker way to access the dynamics of leaf development and leaf conductance (Vadez et al. 2015). This further helps to develop an early stage assay for screening the large breeding populations. The technique is based on a novel 3D scanning technique to capture leaf area development continuously, increasing imaging throughput and analytical scales by combining gravimetric transpiration measurements. Utiliza-tion of these new phenotyping advances has a great scope for high-throughput trait screening and breeding selections in large breeding populations.
3
Aluminum (Al) Tolerance
3.1
Importance and Mechanism
of Al-Tolerance
Soil acidity imposes one of the most severe constraints on crop productivity in the tropics and subtropics (Wang et al. 2014a, b). Many tropical soils are acidic because percolating rainwater leaches the cations such as calcium, magnesium, and potassium, which are replaced by aluminum (Al), manganese, and hydrogen, leading to toxic levels of these elements (Rao et al. 1993). In acidic soils, particularly at pH below 5, Al is solubilized into ionic forms (Al3+) and this leads to phosphorus (P) deficiency as P isfixed as aluminum phosphate (a highly insol-uble and unavailable form to plants; Rao et al. 1993). Plants growing in Al toxicity display stunted growth and become susceptible to drought (Marschner 1991; Kochian et al. 2004; Wang et al.2014a,b). This happens mainly due to inhibition of cell division, cell elongation, or both under low availability of P leading to poor root growth (Delhaize et al. 2004, 2009; Magalhaes et al.2004). Al toxicity is the single most important factor affecting crop production on 2/3 of the acid soil affected area (Eswaran et al.1997), and it is a major constraint for sor-ghum production in tropical and subtropical regions (Doumbia et al. 1993, 1998). Plant adaptation to acid soils is mainly through toler-ance and avoidtoler-ance, of which avoidtoler-ance occurs more commonly. Avoidance is achieved either through changes in the rhizosphere ecology such as increase in pH through root exudates or release of chelators for Al (such as citrate or malate), or increase in root surface area via mycorrhizae (Marschner1991).
3.2
Genes for Al-Tolerance
in Sorghum
Al tolerance in sorghum is controlled by a single gene (AltSB/SbMATE) in a cross between toler-ant, SC283, and sensitive, BR007, which has been mapped to sorghum chromosome 3
(Magalhaes et al. 2004). Positional cloning identified a gene coding for an aluminum-activated citrate transporter, which is a member of the multidrug and toxic compound extrusion (MATE) family, the causal component leading Al tolerance (Magalhaes et al. 2007). Markers from this region have been deployed by breeders to introgress favorable SbMATE alleles in sus-ceptible sorghum genotypes (Anami et al.2015). SbMATE expression is reported to be induced with time of exposure to Al and the expression is higher in the root apex compared to the rest of the root (Magalhaes et al.2007). The region 1– 3 mm behind the root tip where transition from cell division to cell elongation occurs was reported to be the most sensitive area (Sivaguru et al. 2013). After exposure to Al, sensitive genotypes accumulate several-fold more Al in their root apex compared to Al-tolerant geno-types (Delhaize et al.1993). The coding region ofSbMATEis identical between Al-tolerant and Al-sensitive genotypes. However, the second intron ofSbMATEshows polymorphism between the two types. In the promoter region of SbMATE, a tourist-like miniature inverted repeat transposable element (MITE) transposon has been reported, whose repeat numbers are observed to be positively correlated with Al tol-erance (Magalhaes et al. 2007). With this it is assumed that the causative mutations underlying aluminum tolerance may have a regulatory nature (Anami et al.2015).
4
QTL/Marker Analysis for Other
Abiotic Stresses
4.1
Early-Season Cold Tolerance
Sorghum, in general, cannot grow well under soil temperature below 15 °C. However, if early-season cold tolerance can be induced in sorghum cultivars, the area under it can be extended to more northern latitudes, and may allow early planting, particularly in the United States (Yu and Tuinstra2001). Under this situ-ation, improved emergence and early-season vigor are needed, which will ensureestablishment of better crop stand, and prevent loss of seedlings due to cold. The “Kaoliangs” sorghum germplasm from temperate regions of China shows better cold tolerance than tropical germplasm (Cisse and Ejeta 2003). Thus, this particular germplasm is a promising source for improvement of cold tolerance (Yu and Tuistra 2001; Franks et al. 2006). However, these germplasms are in general not agronomically promising, and thus require extensive back-crossing to transfer cold tolerance in the breeding program. Gunaratna (2002) found that genetic control of seedling vigor traits under cold stress and under optimal temperatures is similar. In order to identify QTLs for seedling vigor traits under cold stress, Knoll et al. (2008) developed a recombinant inbred line (RIL) population of 153 RILs from a cross between cold-tolerant Chinese Kaoliang “Shan Qui Red” (SQR) and a cold stress-susceptible African caudatum SRN39. They could identify two QTLs for germination. One among them on linkage group SBI-03a was significant under cold and optimal temperature. The other, on SBI-07b, was contributed by SQR, and showed higher significance under cold tem-peratures. They also identified a region on SBI-01a from SQR, favoring seedling emergence and seedling vigor under early and late field plantings. Knoll and Ejeta (2008) further demonstrated favorable effects of SQR alleles towards seedling vigor and/or emergence in two new populations. Subsequently, Burow et al. (2011) reported an additional 16 QTLs for cold germinability, field emergence, and early seed-ling vigor in a mapping population consisting of 171 RILs derived from the cross between RTx430 (cold-sensitive) and PI610727 (cold-tolerant) using 141 SSR markers. The most promising region for improvingfield emergence identified in this study is located on SBI-01. Other QTL-rich regions were located on SBI-03, SBI-04, SBI-06, SBI-08, and SBI-09. Utilizing a genetic map based on an F8 RIL population
reported by Shiringani et al. (2010) and by phe-notyping a subset of the same population for cold tolerance, Bekele et al. (2013) identified highly interactive epistatic QTL hotspots having a sig-nificant effect on prolonged chilling survival. The
major QTL regions on chromosome SBI06 har-bor candidate genes that govern tolerance to abiotic stress. They identified several genes conferring maintenance of cell division and growth under early chilling stress within QTL hotspot regions, which can be a potential candi-date for breeding cold tolerance.
In sorghum, a higher respiration rate has been correlated with higher germination under cold stress (Balota et al.2010). Washburn et al. (2013) correlated a rhizome formation trait to overwin-tering ability of sorghum. The phenomenon of overwintering and rhizomatousness are reported to be controlled by seven QTLs (Paterson et al. 1995; Washburn et al.2013) that were identified in a mapping population of BTx623/S. propein-quum. The ability of sorghum to overwinter and form rhizome has been suggested to be useful for biofuel sorghum production (Anami et al.2015). Through a differentially expressed gene (DEG) analysis between contrasting genotypes, HongkeiZi (cold tolerant) and BTx623 (cold sensitive), Chopra et al. (2015) identified 41,603 SNPs. They could validate 89 % of the 114 selected SNPs using endpoint genotyping tech-nology. By combining expression profile data and gene-based SNP information they generated a searchable database, which turned out to be an important resource for sorghum cold stress genomics research (http://www.csrl.ars.usda.gov/ psgd/index-sorghum.aspx.html). All these mark-ers are important resources to introgress cold tolerance in sorghum. Recently Upadhyaya et al. (2016) conducted association mapping in a sor-ghum mini-core collection (Upadhyaya et al. 2009) using 162,177 SNPs and identified only one marker locus (Locus 7-2) to be significantly associated with low-temperature germination and none with vigor. The Locus 7-2 was found to representfield early-emergence QTL flanked by Xtxp278 and Xtxp295 (Burow et al.2011). The locus was found next to three overlapping emergence QTLs close to sPb-5796 (Fiedler et al. 2012). Furthermore, the syntenic region of this locus colocalizes with two cold-tolerance rice QTLs, and it is found that when its wheat
homologue is overexpressed in tobacco, cold-tolerance and germination rate are increased.
4.2
Salinity Tolerance
Salinity is also an emerging problem in sorghum cultivation in different parts of the world. It has been reported that upon Na+stress a high-affinity potassium transporter gene family in sorghum, SbHKT1;4, is strongly upregulated in salt-tolerant sorghum genotypes, leading to better Na+/K+ ratio and optimum plant growth (Wang et al.2014b). A total of 38 QTLs influencing salt tolerance has been reported from an RIL popu-lation comprising 181 lines derived from a cross Shihong137/L-Tian (Wang et al.2014a). Out of these, six are reported to be major QTLs explaining above 10 % phenotypic variation. Studies suggested that the mechanism of salt tolerance at the germination and seedling stages is different. Further research in this direction is essential before deploying the identified QTLs in MAS.
4.3
Nitrogen Use Efficiency
The demand for nitrogen (N) fertilizer in agri-culture across the globe currently stands at*110 million metric tons per year and is projected to reach up to*250 million metric tons by the year 2050 (www.fao.org). Because of the high mobility of nitrate in the soil, up to 50 % of applied N is lost by the processes of leaching, runoff, and denitrification. This not only increa-ses the cost of crop production but also adds to the pollution of the groundwater and adversely affects the soil structure. The process has detri-mental effects on the environment by increasing greenhouse gases such as nitric oxide in the atmosphere. Hence, developing crop varieties with improved N absorption and utilization can mitigate these problems of modern agriculture. Sorghum is predominantly cultivated under low
fertility conditions, particularly in Africa and Asia. However, in developed countries it is cul-tivated under high fertility conditions. Significant genotypic differences in terms of nitrogen-use efficiency (NUE) have been reported in sorghum (Maranville et al.1980; Youngquist et al.1992). Expression analysis between four low-N tolerant sorghum genotypes (San Chi San, China17, KS78, and high-NUE bulk) and three sensitive genotypes (CK60, BTx623, and low-NUE bulk) under of low-N (LN, 0 kg ha−1) and normal N (NN, 100 kg ha−1) regimes revealed that in sensitive genotypes, N-stress increased the abundance of DEG-transcripts associated with stress responses including oxidative stress, whereas the tolerant genotypes produced greater root mass for efficient uptake of nutrients (Gelli et al. 2014). Higher abundance of transcripts related to high-affinity nitrate transporters (NRT2.2, NRT2.3, NRT2.5, and NRT2.6) and lysine histidine transporter 1 (LHT1) were detected in tolerant genotypes, which indicates possible improved uptake of inorganic and organic forms of nitrogen by these genotypes. Higher abundance of SEC14 cytosolic factor family protein transcript in tolerant genotypes could lead to increased membrane stability and tolerance to N-stress.
Gelli et al. (2016) sequenced a population of 131 RILs derived from a cross between CK60 (inefficient N user) and China17 (efficient N user) using GbS. Following the composite interval mapping (CIM) technique using 642 polymor-phic SNPs they could identify 38 QTLs for 11 agronomic traits under normal (100 kg ha−1) and no N application condition on chromosomes 1, 5, 6, 7, and 9. Phenotypic variation explained by each QTL ranged from 6.2–50.8 %. Using Illu-mina RNA sequencing on seedling root tissues they could identify 726 differentially expressed transcripts between the parents. Among these 108 were mapped close to the QTL regions. Differentially expressed transcripts were related to nitrogen metabolism (Ferredoxin-nitrate reductase), glycolysis (Phosphofructo-2-kinase), seed storage proteins, plant hormone metabolism, and membrane transport. The study indicated that the differentially expressed transcripts underlying
the pleiotropic QTL regions could be potential targets for improving sorghum performance under limited N fertilizer through marker-assisted selection.
5
Molecular Insight into Abiotic
Stress Response
5.1
Noncoding RNAs
MicroRNAs (miRNAs) have been associated with regulations of different classes of genes across plant species. Pasini et al. (2014) reported upregulation of miRNA associated with the regulation of tran-scription, signal transduction, carbon metabolism, detoxification, osmoprotection mechanisms, and stability of protein membranes upon imposition of low moisture stress in four-leaf-old sorghum genotype, IS1945. The study suggested the possi-ble utility of these drought-related genes in identi-fying drought-tolerant sorghum lines. In fact, miRNA 169 g, as reported to be upregulated under drought stress in rice, has five sorghum homo-logues (Zhao et al. 2007). Qi et al. (2013) also reported long noncoding RNAs (lncRNAs) from foxtail millet in response to drought stress, having sequence conservation and collinearity with sor-ghum. Through an in silico study, Ram and Sharma (2013) further indicated a probable role of miRNAs in water stress response in sorghum.
5.2
Role of Genes Governing Auxin
and Transcription Factors
Over 50 differentially expressed drought-responsive gene orthologues with enriched ABREs and CGTCA-motifs or motifs responsive to ABA specific to sorghum have been identified (Dugas et al. 2011). A large number of auxin-related gene families including SbGH3, SbLBD, SbIAA1, SbGH3-13, SbLBD32, and others have been reported that are highly induced by salt or drought stress (Wang et al. 2010). Moisture stress has been observed to trigger upregulation of transcription factor genes of MADS-box, Auxin Responsive Factors, Heme
Activator Protein 2, and so on, specifically in root tissues (Aglawe et al.2012). Sorghum auxin transporters have been observed to be up- or downregulated in response to abiotic stresses, depending on the class of transporters (Shen et al. 2010).
Over 100 ethylene response factor (ERF) ge-nes have been reported in sorghum (SbERF), which have been grouped into 12 classes (A-1 to A-6 and B-1 to B-6) on the basis of sequence homology (Yan et al. 2013). The ERF super-family plays an important role in both abiotic and biotic stress response in plant systems. In addi-tion to ERF, chloroplast glutathione reductase (cpGRs), G-protein complexes, drought response element-binding (DREB) proteins, andSbESTare reported to play very important roles in abiotic stress responses not only in sorghum but in other plant species as well. In a recent study, Chopra et al. (2015) identified a total of 1910 DEGs under cold and control temperature from cold-tolerant genotype HongkeZi and cold-sensitive genotype BTx623. They could identify TFs including DREB,C-repeat binding factors, and ERF TFs to be upregulated under cold stress in tolerant genotype HongkeZi. They also identified specific genes such as plant cytochromes, glutathione s-transferases, and heat shock proteins to be differentially regulated under cold stress between cold-tolerant and sus-ceptible genotypes.
5.3
Role of Compatible Solutes
Betaise aldehyde dehydrogenase (BADH1 and BADH15) is reported to be induced in response to moisture deficit leading to accumulation of glycine betaine (Wood et al. 1996). SbGRRNP (glycine-rich RNA-binding protein) expression is regulated by salinity and ABA, as well as blue and red light (Aneeta et al.2002). This suggests that in sorghum a possible crosstalk between abiotic stress and light signal may exist. By engineering higher mannitol synthesis, Mah-eswari et al. (2010) could induce salinity toler-ance in transgenic sorghum cv. SPV462. Studies further suggest thatSbP5CS1 and SbP5CS2 are
upregulated in response to drought, salt, and jasmonic acid treatment (Su et al. 2011). These two are important regulatory genes controlling proline synthesis. Thus, the study suggested a role of proline biosynthesis in imparting abiotic stress tolerance. All the above reports suggest the possibility to develop abiotic stress-tolerant genotypes by modulating the role of these genes, transcription factors, or miRNAs either through the transgenic route or through genome editing.
6
Next-Generation Sequencing
(NGS) Tools and Next-Generation
Populations
When combined with high-throughput and pre-cise phenotyping platforms developed at ICRI-SAT (Vadez et al. 2015), NGS technologies provide a powerful and rapid tool for identifying the genetic basis of agriculturally important traits and for predicting the breeding value of indi-viduals in a plant breeding population (Varshney et al.2014). Next-generation populations such as nested-association mapping (NAM) populations (Yu et al. 2008; Buckler et al. 2009) and backcross-derived NAM (BCNAM) populations (Jordan et al. 2012) are two most potent next-generation populations to dissect genomics of complex traits. All these new tools play a major role in advancing breeding efficiencies by simultaneously aiding genetic studies and also providing access to new variability in compara-tively elite backgrounds. These new-generation populations are required for linking phenotypic variations with sequence variations at high res-olution. The BCNAM scheme was already well established by Jordan et al. (2011) to benefit both trait mapping and infusing diversity that was not accessible previously in the traditional plant breeding populations. In combination with gen-omewide sequencing approaches, these popula-tions provide access to insights into the genetic architecture of important traits (Yang et al. 2014a,b). The major limitation in implementing these new advances routinely in the breeding programs is lack of data management skills and
database resources. In terms of applications of these tools in molecular breeding, the NGS-based genotyping platforms such as genotyping-by-sequencing (GbS) have the potential for background selection in traditional introgression breeding and genomic selection (GS) which is an emerging methodology in modern breeding pro-grams. The main advantage and driving force for its implementation are availability of low-cost high-throughput genotyping platforms and access to high-throughput phenotyping facilities. These in turn feed the improved capacities of performance prediction of individual genotypes for quantitative traits. The genomic selection, to realize its full potential, will need to address specific issues related to the theory of GS such as design of training populations, predicting effi -cacy under an altered marker and population variable scenario, and an approach for integration of GS in ongoing breeding schemes (both in scale and dimension). Similarly, considering the different level of advancement of constituent technologies involved in GS, an appropriate resource investment strategy for every single breeding program needs to be developed for maximizing the returns in terms of genetic gain.
7
Future Prospects
Sorghum is an important failsafe crop providing food–feed–fodder–fuel to most of the resource-poor farming communities in drylands where it is a staple crop. The wide repertoire of germplasm, genetic, genomic, and breeding information/resources positions sorghum as one of the C4-model crop species. Advancement in
NGS, high-throughput phenotyping, and recent developments in advanced breeding populations will drive the new cycle of genetic gain in sor-ghum across agroecological zones. The sorsor-ghum research community has access to large genetic (germplasm with unique traits) and genomic (SSRs, SNPs, high-density genetic maps, gen-ome sequence) resources, and many QTLs/candidate genes associated with agro-nomic traits are known in sorghum. These mar-ker–trait associations need to be validated
independently in breeding populations and suit-able marker assays such as the KASPar SNP assay need to be generated. The integration of these new tools along with new-generation breeding populations will further help accumu-late the required information to develop the framework for implementing genomic selection for sorghum improvement and thus historical division between breeding and genomics will become increasingly blurred.
References
Aglawe SB, Fakrudin B, Patole CB, Bhairappanavar SB, Koti RV, Krishnaraj PU (2012) Quantitative RT-PCR analysis of 20 transcription factor genes of MADS, ARF, HAP2, MBF and HB families in moisture stressed shoot and root tissues of sorghum. Physiol Mol Biol Plants 18(4):287–300
Anami SE, Zhang LM, Xia Y, Zhang YM, Liu ZQ, Jing HC (2015) Sweet sorghum ideotypes: genetic improvement of stress tolerance. Food Energy Secur 4 (1):3–24
Aneeta, Sanan-Mishra N, Tuteja N, Kumar Sopory S (2002) Salinity- and ABA-induced up-regulation and light-mediated modulation of mRNA encoding glycine-rich RNA-binding protein from Sorghum bicolor. Biochem Biophys Res Commun 296:1063– 1068
Araus JL, Slafer GA, Reynolds MP, Royo C (2002) Plant breeding and drought in C3 cereals: what should we breed for? Ann Bot 89(7):925–940
Balota M, Payne W, Veeragoni S, Stewart B, Rosenow D (2010) Respiration and its relationship to germination, emergence, and early growth under cool temperatures in sorghum. Crop Sci 50:1414–1422
Bekele WA, Wieckhorst S, Friedt W, Snowdon RJ (2013) High-throughput genomics in sorghum: from whole-genome resequencing to a SNP screening array. Plant Biotechnol J 11(9):1112–1125
Borrell A, Hammer G, Oosterom E (2001) Stay-green: a consequence of the balance between supply and demand for nitrogen during grainfilling? Ann Appl Biol 138(1):91–95
Borrell AK, Hammer GL (2000) Nitrogen dynamics and the physiological basis of stay-green in sorghum. Crop Sci 40(5):1295–1307
Borrell AK, Hammer GL, Douglas AC (2000a) Does maintaining green leaf area in sorghum improve yield under drought? I. Leaf growth and senescence. Crop Sci 40(4):1026–1037
Borrell AK, Hammer GL, Henzell RG (2000b) Does maintaining green leaf area in sorghum improve yield under drought? II. Dry matter production and yield. Crop Sci 40(4):1037–1048
Borrell AK, Douglas ACL (1996) Maintaining green leaf area in grain sorghum increases yield in a water-limited environment. In: Proceedings of the third australian sorghum conference. Melbourne, Australian Institute of Agricultural Science, Occa-sional Publication (No. 93), pp 323–326
Bowers JE, Abbey C, Anderson S, Chang C, Draye X, Hoppe AH, Jessup R, Lemke C, Lennington J, Li Z, Lin YR (2003) A high-density genetic recombination map of sequence-tagged sites for sorghum, as a framework for comparative structural and evolution-ary genomics of tropical grains and grasses. Genetics 165(1):367–386
Buckler ES, Holland JB, Bradbury PJ, Acharya CB, Brown PJ, Browne C, Ersoz E, Flint-Garcia S, Garcia A, Glaubitz JC, Goodman MM (2009) The genetic architecture of maize f owering time. Science 325:714–718
Burow G, Burke JJ, Xin Z, Franks CD (2011) Genetic dissection of early- season cold tolerance in sorghum (Sorghum bicolor(L.) Moench). Mol Breed 28:391–402 Burow GB, Franks CD, Acosta-Martinez V, Xin Z (2009) Molecular mapping and characterization of BLMC, a locus for profue wax (bloom) and enhanced cuticular features of Sorghum (Sorghum bicolor( L.) Moench.). Theor Appl Genet 118:423–431
Chopra R, Burow G, Hayes C, Emendack Y, Xin Z, Burke J (2015) Transcriptome profiling and validation of gene based single nucleotide polymorphisms (SNPs) in sorghum genotypes with contrasting responses to cold stress. BMC Genom 16(1):1 Cisse ND, Ejeta G (2003) Genetic variation and
relation-ships among seedling vigor traits in sorghum. Crop Sci 43(3):824–828
Crasta OR, Xu WW, Rosenow DT, Mullet J, Nguyen HT (1999) Mapping of post-flowering drought resistance traits in grain sorghum: association between QTLs influencing premature senescence and maturity. Mol Gen Genet 262(3):579–588
Delhaize E, Craig S, Beaton CD, Bennet RJ, Jagadish VC, Randall PJ (1993) Aluminum tolerance in wheat (Triticum aestivum L.)(I. Uptake and distribution of aluminum in root apices). Plant Physiol 103(3):685– 693
Delhaize E, Ryan PR, Hebb DM, Yamamoto Y, Sasaki T, Matsumoto H (2004) Engineering high-level alu-minum tolerance in barley with the ALMT1 gene. Proc Natl Acad Sci USA 101(42):15249–15254 Delhaize E, Taylor P, Hocking PJ, Simpson RJ, Ryan PR,
Richardson AE (2009) Transgenic barley (Hordeum vulgare L.) expressing the wheat aluminium resistance gene (TaALMT1) shows enhanced phosphorus nutri-tion and grain producnutri-tion when grown on an acid soil. Plant Biotechnol J 7(5):391–400
Doumbia MD, Hossner LR, Onken AB (1993) Variable sorghum growth in acid soils of subhumid West Africa. Arid Land Res Manag 7(4):335–346 Doumbia MD, Hossner LR, Onken AB (1998) Sorghum
growth in acid soils of West Africa: variations in soil
chemical properties. Arid Land Res Manage 12 (2):179–190
Dugas DV, Monaco MK, Olson A, Klein RR, Kumari S, Ware D, Klein PE (2011) Functional annotation of the transcriptome of Sorghum bicolor in response to osmotic stress and abscisic acid. BMC Genom 12(1):1 Eswaran H, Reich P, Beinroth F (1997) Global distribu-tion of soils with acidity. In: Moniz AC, Furlani AMC, Schaffert RE, Fageria NK, Rosolem CA, Cantarella H (eds) Plant-soil interactions at low pH. Brazilian Soil Science Society, pp. 159–164
Fiedler K, Bekele WA, Friedt W, Snowdon R, Stützel H, Zacharias A, Uptmoor R (2012) Genetic dissection of the temperature dependent emergence processes in sorghum using a cumulative emergence model and stability parameters. Theor Appl Genet 125(8):1647– 1661
Franks TK, Powell KS, Choimes S, Marsh E, Iocco P, Sinclair BJ, Ford CM, Van Heeswijck R (2006) Consequences of transferring three sorghum genes for secondary metabolite (cyanogenic glucoside) biosyn-thesis to grapevine hairy roots. Transgen Res 15 (2):181–195
Gelli M, Duo Y, Konda AR, Zhang C, Holding D, Dweikat I (2014) Identification of differentially expressed genes between sorghum genotypes with contrasting nitrogen stress tolerance by genome-wide transcriptional profiling. BMC Genom 15(1):1 Gelli M, Mitchell SE, Liu K, Clemente TE, Weeks DP,
Zhang C, Holding DR, Dweikat IM (2016) Mapping QTLs and association of differentially expressed gene transcripts for multiple agronomic traits under differ-ent nitrogen levels in sorghum. BMC Plant Biol 16 (1):1
Gunaratna N (2002) Early season cold tolerance in sorghum. MS Thesis, Purdue University, West Lafay-ette, IN, USA
Habyarimana E, Lorenzoni C, Busconi M (2010) Search for new stay-green sources inSorghum bicolor(L.) Moench. Maydica 55(3):187
Hammer GL, Dong Z, McLean G, Doherty A, Messina C, Schussler J, Zinselmeier C, Paszkiewicz S, Cooper M (2009) Can changes in canopy and/or root system architecture explain historical maize yield trends in the US corn belt? Crop Sci 49(1):299–312
Harris K, Subudhi PK, Borrell A, Jordan D, Rosenow D, Nguyen H, Klein P, Klein R, Mullet J (2007) Sorghum stay-green QTL individually reduce post-flowering drought-induced leaf senescence. J Exp Bot 58 (2):327–338
Haussmann B, Mahalakshmi V, Reddy B, Seetharama N, Hash C, Geiger H (2002) QTL mapping of stay-green in two sorghum recombinant inbred populations. Theor Appl Genet 106(1):133–142
Jordan DR, Hunt CH, Cruickshank AW, Borrell AK, Henzell RG (2012) The relationship between the stay-green trait and grain yield in elite sorghum hybrids grown in a range of environments. Crop Sci 52(3):1153–1161
Jordan DR, Mace ES, Cruickshank AW, Hunt CH, Henzell RG (2011) Exploring and exploiting genetic variation from unadapted sorghum germplasm in a breeding program. Crop Sci 51(4):1444–1457 Kassahun B, Bidinger FR, Hash CT, Kuruvinashetti MS
(2010) Stay-green expression in early generation sorghum [Sorghum bicolor(L.) Moench] QTL intro-gression lines. Euphytica 172(3):351–362
Kato Y, Kamoshita A, Yamagishi J, Abe J (2006) Growth of three rice (Oryza sativaL.) cultivars under upland conditions with different levels of water supply. Plant Prod Sci 9(4):422–434
Kebede H, Subudhi PK, Rosenow DT, Nguyen HT (2001) Quantitative trait loci influencing drought tolerance in grain sorghum (Sorghum bicolor L. Moench). Theor Appl Genet 103(2–3):266–276 KholováJ, Murugesan T, Kaliamoorthy S, Malayee S,
Baddam R, Hammer GL, McLean G, Deshpande S, Hash CT, Craufurd PQ, Vadez V (2014) Modelling the effect of plant water use traits on yield and stay-green expression in sorghum. Funct Plant Biol 41 (11):1019–1034
Kim JS (2003) Genomic analysis of sorghum byfl uores-cence in situ hybridisation. Texas A&M University, College Station, TX, USA
Kiranmayee KNS, Hash CT, Deshpande SP, Vara-prasad KVGK, Kishor PB (2015) Biotechnological approaches to evolve sorghum (Sorghum bicolor(L.) Moench) for drought stress tolerance and shoot fly resistance. Curr Trends Biotechnol Pharm 9(3):281– 292
Knoll J, Ejeta G (2008) Marker-assisted selection for early-season cold tolerance in sorghum: QTL valida-tion across populavalida-tions and environments. Theor Appl Genet 116(4):541–553
Knoll J, Gunaratna N, Ejeta G (2008) QTL analysis of early-season cold tolerance in sorghum. Theor Appl Genet 116(4):577–587
Kochian LV, Hoekenga OA, Pineros MA (2004) How do crop plants tolerate acid soils? Mechanisms of aluminum tolerance and phosphorous efficiency. Annu Rev Plant Biol 55:459–493
Mace ES, Jordan DR (2010) Location of major effect genes in sorghum (Sorghum bicolor (L.) Moench). Theor Appl Genet 121(7):1339–1356
Mace ES, Jordan DR (2011a) Integrating sorghum whole genome sequence information with a compendium of sorghum QTL studies reveals uneven distribution of QTL and of gene-rich regions with significant implications for crop improvement. Theor Appl Genet 123(1):169–191 Mace ES, Jordan DR (2011b) Integrating sorghum whole
genome sequence information with a compendium of sorghum QTL studies reveals uneven distribution of QTL and of gene-rich regions with significant impli-cations for crop improvement. Theor Appl Genet 123 (1):169–191
Mace ES, Singh V, Van Oosterom EJ, Hammer GL, Hunt CH, Jordan DR (2012) QTL for nodal root angle in sorghum (Sorghum bicolor L. Moench) co-locate
with QTL for traits associated with drought adapta-tion. Theor Appl Genet 124(1):97–109
MacIntyre BD, Herren HR, Wakhungu J, Watson RT (2009) Agriculture at a crossroads: international assessment of agricultural science and technology for development (IAASTD) global report. IAASTD, Washington DC, USA
Magalhaes JV, Garvin DF, Wang Y, Sorrells ME, Klein PE, Schaffert RE, Li L, Kochian LV (2004) Comparative mapping of a major aluminum tolerance gene in sorghum and other species in the Poaceae. Genetics 167(4):1905–1914
Magalhaes JV, Liu J, Guimaraes CT, Lana UG, Alves VM, Wang YH, Schaffert RE, Hoekenga OA, Pineros MA, Shaff JE, Klein PE (2007) A gene in the multidrug and toxic compound extrusion (MATE) family confers aluminum tolerance in sorghum. Nat Genet 39(9):1156–1161
Maheswari M, Varalaxmi Y, Vijayalakshmi A, Yadav S, Sharmila P, Venkateswarlu B et al (2010) Metabolic engineering using mtlD gene enhances tolerance to water deficit and salinity in sorghum. Biol Plant 54:647–652
Maranville JW, Clark RB, Ross WM (1980) Nitrogen efficiency in grain sorghum. J Plant Nutr 2(5):577–589 Marschner H (1991) Mechanisms of adaptation of plants to acid soils. In: Wright RJ, Baligar VC, Murrmann RP (eds) Plant-soil interactions at low pH. Springer, Netherlands, pp 683–702
McBee GG, Waskom RM, Creelman RA (1983) Effect of senescene on carbohydrates in sorghum during late kernel maturity states. Crop Sci 23(2):372–376 Mickelbart MV, Hasegawa PM, Bailey-Serres J (2015)
Genetic mechanisms of abiotic stress tolerance that translate to crop yield stability. Nat Rev Genet 16 (4):237–251
Mutava RN, Prasad PVV, Tuinstra MR, Kofoid KD, Yu J (2011a) Characterization of sorghum genotypes for traits related to drought tolerance. Field Crops Res 123 (1):10–18. doi:10.1016/j.fcr.2011.04.006
Mutava RN, Prasad PVV, Tuinstra MR, Kofoid KD, Yu J (2011b) Characterization of sorghum genotypes for traits related to drought tolerance. Field Crops Research 123(1):10–18
Pasini L, Bergonti M, Fracasso A, Marocco A, Ama-ducci S (2014) Microarray analysis of differentially expressed mRNAs and miRNAs in young leaves of sorghum under dry-down conditions. J Plant Physiol 171(7):537–548
Paterson AH, Bowers JE, Bruggmann R, Dubchak I, Grimwood J, Gundlach H, Haberer G, Hellsten U, Mitros T, Poliakov A, Schmutz J (2009) TheSorghum bicolor genome and the diversification of grasses. Nature 457(7229):551–556
Paterson AH, Schertz KF, Lin YR, Liu SC, Chang YL (1995) The weediness of wild plants: molecular analysis of genes influencing dispersal and persistence of johnsongrass,Sorghum halepense (L.) Pers. Proc Natl Acad Sci USA 92(13):6127–6131
Qi X, Xie S, Liu Y, Yi F, Yu J (2013) Genome- wide annotation of genes and noncoding RNAs of foxtail millet in response to simulated drought stress by deep sequencing. Plant Mol Biol 83:459–473
Rakshit S, Hariprasanna K, Gomashe S, Ganapathy KN, Das IK, Ramana OV, Dhandapani A, Patil JV (2014) Changes in area, yield gains, and yield stability of sorghum in major sorghum-producing countries, 1970 to 2009. Crop Sci 54(4):1571–1584
Rakshit S, Swapna M, Dalal M, Sushma G, Ganapa-thy KN, Dhandapani A, Karthikeyan M, Talwar HS (2016) Post-flowering drought stress response of post-rainy sorghum genotypes. Indian J Plant Physiol 21(1):8–14
Ram G, Sharma AD (2013) In silico analysis of putative miRNAs and their target genes in sorghum (Sorghum bicolor). Int J Bioinformat Res Applicat 9(4):349–364 Ramu P, Deshpande SP, Senthilvel S, Jayashree B, Billot C, Deu M, Reddy LA, Hash CT (2010) In silico mapping of important genes and markers available in the public domain for efficient sorghum breeding. Mol Breed 26(3):409–418
Rao IM, Zeigler RS, Vera R, Sarkarung S (1993) Selection and breeding for acid-soil tolerance in crops. Bioscience 43(7):454–465
Rao PS, Prakasham RS, Rao PP, Chopra S, Jose S (2015) Sorghum as a sustainable feedstock for biofuels. In: Shibu J, Thallada B (eds.) Biomass and biofuels: advanced biorefineries for sustainable production and distribution. CRC Press 2015, Pages 27–48, Print ISBN: 978–1-4665-9531-6, eBook ISBN: 978-1-4665-9532-3
Rosenow DT, Clark LE (1982) Drought tolerance in sorghum [through breeding]. In: Proceedings of the 37th annual corn and sorghum industry research conference-American Seed Trade Association, Corn and Sorghum Division, Corn and Sorghum Research Conference, Chicago, IL, USA, pp 18–30
Rosenow DT, Quisenberry JE, Wendt CW, Clark LE (1983) Drought tolerant sorghum and cotton germ-plasm. Agric Water Manag 7(1):207–222
Rosenow DT, Clark LE (1995) December. Drought and lodging resistance for a quality sorghum crop. In: Proceedings of the 5th annual corn and sorghum industry research conference (Chicago, IL, 6–7 December 1995), American Seed Trade Association, Chicago, IL, USA, pp 82–97
Sabadin PK, Malosetti M, Boer MP, Tardin FD, San-tos FG, Guimaraes CT, Gomide RL, Andrade CLT, Albuquerque PEP, Caniato FF, Mollinari M (2012) Studying the genetic basis of drought tolerance in sorghum by managed stress trials and adjustments for phenological and plant height differences. Theor Appl Genet 124(8):1389–1402
Sanchez AC, Subudhi PK, Rosenow DT, Nguyen HT (2002) Mapping QTLs associated with drought resis-tance in sorghum (Sorghum bicolorL. Moench). Plant Mol Biol 48(5–6):713–726
Shen C, Bai Y, Wang S, Zhang S, Wu Y, Chen M, Jiang D, Qi Y (2010) Expression profile of PIN,
AUX/LAX and PGP auxin transporter gene families in Sorghum bicolor under phytohormone and abiotic stress. FEBS J 277(14):2954–2969
Shiringani AL, Frisch M, Friedt W (2010) Genetic mapping of QTLs for sugar-related traits in a RIL population of Sorghum bicolor L. Moench. Theor Appl Genet 121(2):323–336
Singh V, van Oosterom EJ, Jordan DR, Messina CD, Cooper M, Hammer GL (2010) Morphological and architectural development of root systems in sorghum and maize. Plant Soil 333(1–2):287–299
Singh V, van Oosterom EJ, Jordan DR, Hunt CH, Hammer GL (2011) Genetic variability and control of nodal root angle in sorghum. Crop Sci 51(5) Singh V, van Oosterom EJ, Jordan R, Hammer GL (2012)
Genetic control of root angle in sorghum and its implication in water extraction. Eur J Agron 42: 3–10 Sivaguru M, Liu J, Kochian LV (2013) Targeted expres-sion of SbMATE in the root distal transition zone is responsible for sorghum aluminum resistance. Plant J 76(2):297–307
Srinivas G, Satish K, Madhusudhana R, Seetharama N (2009) Exploration and mapping of microsatellite mark-ers from subtracted drought stress ESTs in Sorghum bicolor(L.) Moench. Theor Appl Genet 118(4):703–717 Srinivas G, Satish K, Mohan SM, Reddy RN, Madhusud-hana R, Balakrishna D, Bhat BV, Howarth CJ, Seetharama N (2008) Development of genic-microsatellite markers for sorghum staygreen QTL using a comparative genomic approach with rice. Theor Appl Genet 117(2):283–296
Su M, Li X-F, Ma X-Y, Peng X-J, Zhao A-G, Cheng L-Q et al (2011) Cloning twoP5CS genes from bioenergy sorghum and their expression profiles under abiotic stresses and MeJA treatment. Plant Sci 181:652–659 Subudhi PK, Rosenow DT, Nguyen HT (2000)
Quanti-tative trait loci for the stay green trait in sorghum (Sorghum bicolor L. Moench): consistency across genetic backgrounds and environments. Theor Appl Genet 101(5–6):733–741
Tao YZ, Henzell RG, Jordan DR, Butler DG, Kelly AM, McIntyre CL (2000) Identification of genomic regions associated with stay green in sorghum by testing RILs in multiple environments. Theor Appl Genet 100 (8):1225–1232
Tuinstra MR, Grote EM, Goldsbrough PB, Ejeta G (1997) Genetic analysis of post-flowering drought tolerance and components of grain development in Sorghum bicolor(L.) Moench. Mol Breed 3(6):439–448 Upadhyaya HD, Pundir RPS, Dwivedi SL, Gowda CLL,
Reddy VG, Singh S (2009) Developing a mini core collection of sorghum for diversified utilization of germplasm. Crop Sci 49(5):1769–1780
Upadhyaya HD, Wang YH, Sastry DV, Dwivedi SL, Prasad PV, Burrell AM, Klein RR, Morris GP, Klein PE (2016) Association mapping of germinabil-ity and seedling vigor in sorghum under controlled low-temperature conditions. Genome 59(2):137–145 Vadez V, Deshpande S, Kholova J, Ramu P, Hash CT
and challenges in sorghum. In: Translational genomics for crop breeding: abiotic stress, yield and quality. Wiley Inc., pp. 125–141. ISBN 9781118728482 Vadez V, Deshpande SP, Kholova J, Hammer GL,
Borrell AK, Talwar HS, Hash CT (2011) Stay-green quantitative trait loci’s effects on water extraction, transpiration efficiency and seed yield depend on recipient parent background. Funct Plant Biol 38 (7):553–566
Vadez V, Kholová J, Hummel G, Zhokhavets U, Gupta SK, Hash CT (2015) LeasyScan: a novel concept combining 3D imaging and lysimetry for high-throughput phenotyping of traits controlling plant water budget. J Exp Bot 251
Van Oosterom EJ, Jayachandran R, Bidinger FR (1996) Diallel analysis of the stay-green trait and its compo-nents in sorghum. Crop Sci 36(3):549–555
Varshney RK, Terauchi R, McCouch SR (2014) Harvest-ing the promisHarvest-ing fruits of genomics: applyHarvest-ing genome sequencing technologies to crop breeding. PLoS Biol 12(6):e1001883
Wang H, Chen G, Zhang H, Liu B, Yang Y, Qin L, Chen E, Guan Y (2014a) Identification of QTLs for salt tolerance at germination and seedling stage of Sorghum bicolorL. Moench. Euphytica 196(1):117– 127
Wang TT, Ren ZJ, Liu ZQ, Feng X, Guo RQ, Li BG et al (2014b) SbHKT1; 4, a member of the high- affinity potassium transporter gene family from Sorghum bicolor, functions to maintain optimal Na+/K+ bal-ance under Na + stress. J Integr Plant Biol 56:315– 332
Wang S, Bai Y, Shen C, Wu Y, Zhang S, Jiang D, Guilfoyle TJ, Chen M, Qi Y (2010) Auxin-related gene families in abiotic stress response in Sorghum bicolor. Funct Integr Genom 10(4):533–546 Washburn JD, Murray SC, Burson BL, Klein RR,
Jessup RW (2013) Targeted mapping of quantitative trait locus regions for rhizomatousness in chromosome
SBI-01 and analysis of overwintering in a Sorghum bicolorS. propinquum population. Mol Breed 31 (1):153–162
Wood AJ, Saneoka H, Rhodes D, Joly RJ, Golds-brough PB (1996) Betaine aldehyde dehydrogenase in sorghum (molecular cloning and expression of two related genes). Plant Physiol 110:1301–1308 Xu W, Subudhi PK, Crasta OR, Rosenow DT, Mullet JE,
Nguyen HT (2000) Molecular mapping of QTLs conferring stay-green in grain sorghum (Sorghum bicolorL. Moench). Genome 43(3):461–469 Yan H, Hong L, Zhou Y, Jiang H, Zhu S, Fan J et al
(2013) A genome- wide analysis of the ERF gene family in sorghum. Genet Mol Res 12:2038–2055 Yang R, Yang T, Zhang H, Qi Y, Xing Y, Zhang N, Li R,
Weeda S, Ren S, Ouyang B, Guo YD (2014a) Hormone profiling and transcription analysis reveal a major role of ABA in tomato salt tolerance. Plant Physiol Biochem 77:23–34
Yang W, Guo Z, Huang C, Duan L, Chen G, Jiang N, Fang W, Feng H, Xie W, Lian X, Wang G (2014b) Combining high-throughput phenotyping and genome-wide association studies to reveal natural genetic variation in rice. Nat Commun 5
Youngquist JB, Bramel-Cox P, Maranville JW (1992) Evaluation of alternative screening criteria for select-ing nitrogen-use efficient genotypes in sorghum. Crop Sci 32(6):1310–1313
Yu J, Tuinstra MR (2001) Genetic analysis of seedling growth under cold temperature stress in grain sorghum. Crop Sci 41(5):1438–1443
Yu JM, Holland JB, McMullen MD, Buckler ES (2008) Genetic design and statistical power of nested asso-ciation mapping in maize. Genetics 178:539–551 Zhao B, Liang R, Ge L, Li W, Xiao H, Lin H, Ruan K,
Jin Y (2007) Identification of drought-induced micro-RNAs in rice. Biochem Biophys Res Commun 354 (2):585–590