© Mary Ann Liebert, Inc. Pp. 181–188
DNA Fingerprinting in the Criminal Justice System:
An Overview
VARSHA
ABSTRACT
DNA fingerprinting is a powerful technology that has revolutionized forensic science. No two individuals can
have an identical DNA pattern except identical twins. Such DNA-based technologies have enormous social
im-plications and can help in the fight against crime. This technology has experienced many changes over time
with many advancements occurring. DNA testing is a matter of serious concern as it involves ethical issues.
This article describes various trends in DNA fingerprinting and the current technology used in DNA
profil-ing, possible uses and misuses of DNA databanks and ethical issues involved in DNA testing. Limitations and
problems prevailing in this field are highlighted.
181
INTRODUCTION
K
INGSOLOMON’S STORYis well known for his famous de-cision in a maternity dispute, but today, parentage deci-sions are relatively easy with the assistance of DNA technol-ogy. DNA technology was first developed in England in 1985 by Sir Alec Jeffreys. This technology, so named because DNA is used for identification rather than latent physical fingerprints, gives a unique and specific profile similar to a thumb impres-sion (Jeffreys et al., 1985). DNA fingerprinting technology to-day has made it possible to identify the source of biological samples found at a crime scene and also to resolve disputes of paternity and other criminal cases.Ninety-five percent of the human genome is noncoding DNA, with only 5% coding for protein region. These noncod-ing regions are also called “junk” DNA, and are abundant where the number of repeats show variations among individuals. These DNA polymorphisms change the length of the DNA fragments produced by the digestion of restriction enzymes.
A DNA fingerprint is constructed by first extracting a DNA sample from biological sample. The sample is then cut at spe-cific sites with restriction enzymes, and the segments are arranged by their molecular size using electrophoresis. The seg-ments are labeled with probes and exposed to X-ray film, where they form a characteristic pattern, that is, the DNA fingerprint. If the DNA fingerprints produced from two different sam-ples match, the two samsam-ples probably came from the same per-son. The exact number and size of the fragments produced by
a specific restriction enzyme digestion varies from individual to individual.
In 1985, police in the UK for the first time used forensic DNA profiling (http://www.crimtrac.gov.au/dnahistory.htm), and in the United States, the first criminal conviction based on DNA evidence occurred in 1988. In 1992, the National Re-search Council of the United States found that DNA testing was a reliable method for the identification of criminal suspects, and the technology rapidly entered the mainstream court system. The DNA evidence obtained is powerful, since it can implicate or exonerate a suspect. Currently, DNA testing is routinely used in civil and criminal cases.
DNA ANALYSIS
Early markers
Karl Landsteiner, in 1901, discovered ABO blood grouping in humans (Landsteiner, 1901). A set of blood group markers (red cell antigens and serum protein) were also used to identify victims and suspects in crime cases. As the polymorphism was very limited in ABO blood grouping, it was neither informa-tive nor very suitable for excluding a person as a suspect in a criminal case. The advent of DNA-based markers has now rev-olutionized the field of forensic science, as it can measure an unlimited extent of polymorphisms sufficient to differentiate one individual from another.
Hybridization based multilocus and single locus DNA
profiling
The use of DNA fingerprinting in forensic science had been suggested in the early 1980s by Prof. Alec Jeffreys using re-striction fragment-length polymorphism (RFLP). There was great enthusiasm among the forensic science community for the discovery of the minisatellite approach for specific identifica-tion (Wong et al., 1987).
Eukaryotic genomes contain a large number of repeated DNA sequences. These regions are often referred to as satellite DNA. The core repeat unit for a medium-length repeat, some-times referred to as a minisatellite or variable number of tan-dem repeats (VNTR) is in the range of approximately 10–100 bases in length.
VNTR patterns are inherited and are unique (Weising, et al., 1995). The more VNTR probes used to analyze a person’s VNTR pattern, the more distinctive and individualized that pat-tern, or DNA fingerprint, will be. The children will inherit the different combinations of paternal and maternal VNTRs, and therefore their profile will be different from each other except identical twins as shown in Figure 1.
DNA regions with repeat units that are 2–6 bp in length are called mcrosatellites, simple sequence repeats (SSRs) or short tandem repeats (STRs). STRs consisted of microsatellites of small repeating units, two to five bases in length, with a maxi-mum length of around 300 bases, which show a high degree of polymorphism. Specific microsatellites can be isolated using hy-bridized probes followed by their sequencing. SSRs can be de-tected by specific dyes or by radiolabeling using gel elec-trophoresis. The advantage of using SSRs as molecular markers is the extent of polymorphism shown, which enables the detec-tion of differences at multiple loci between strains (Butler, 2001). Hybridization-based DNA fingerprinting involves the use of two types of probes called multilocus probes (MLP), which iden-tify repeating sequences in genomic DNA digested by restric-tion enzymes followed by Southern blot analysis at low strin-gency conditions. The radio labeled probes hybridize with a series of tandemly repeated sequences with a size range greater than 20 kb. The resulting autoradiographs consist of a ladder-like series of bands that will be identical only for identical twins as shown in Figure 2. The fact that DNA is a very stable mol-ecule makes this technology more effective, and even old cases
also can be reviewed. This is equally good for archived speci-mens. The MLP system, however, was not very successful. The method was demanding and the results were not suited for in-clusion in computer databases. After the advent of the single lo-cus probe (SLP) the technology substantially progressed. In the SLP system probes were isolated, which identified repeat se-quences in individual minisatellites using high stringency con-ditions to produce a maximum of two bands per probe. Although the targeted minisatellites showed considerable variation in their repeat numbers, a single locus was not sufficient for individu-alization. A number of such loci were analyzed (Lalji, 1991).
PCR-based DNA profiling
In the early 1990s, the DNA fingerprinting technology took a dramatic change with the advent of the polymerase chain re-action (PCR), a technique that enabled amplification from tiny amounts of template DNA. The method was fast, reliable, and was efficient to work even with degraded samples. In the PCR-based technique the time required to analyze a sample was sig-nificantly reduced (Weber and May, 1989). It was found to be more suitable than the previously used MLP and SLP systems. There was a period when both SLP and PCR-based methods were used to identify a person in parallel. PCR technology was used to type many VNTRs region for forensic purposes. Many polymorphic loci like D1S80 and HLADQ were used for analysis. There were many reports claiming that it was not much useful for forensic purposes (Varsha et al., 1999).
In 1992, the use of a new compatible form of genetic pro-filing called STRs was proposed. Due to their small size they were extremely stable, and could be detected even with old and degraded samples.
In 1998, STR analysis became an established feature of forensic sciences, which basically involved study of a panel of polymorphic loci comprising THO1, VWA, FGA, D21S11, D8S1179, and D18S51, as well as D7S820, D13S317, D5S818, CSF1P0, D16S539, D2S1338, D19S433, TPOX, with recently introduced Penta E and Penta D (Edwards et al., 1991).
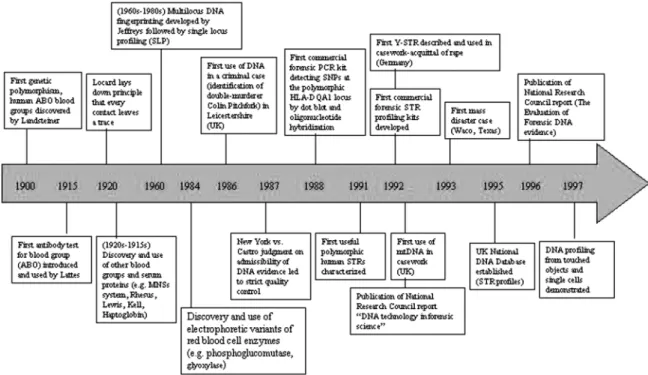
A flow diagram (Fig. 3) and Table 1 show the progressive advancements in forensic genetics.
HOW IS DNA TYPING PERFORMED?
In criminal cases, samples obtained from crime scene evi-dence and a suspect are used for DNA extraction and analysis for the presence of a set of specific DNA markers.DNA markers are small DNA probes that bind to a comple-mentary sequence in the DNA sample. A distinctive pattern for an individual is created by a series of probes bound to a DNA VARSHA 182
FIG. 2. Multilocus DNA profiling of (a) Identical and (b) nonidentical twins.
sample. Comparing such DNA profiles determines whether the suspect’s sample matches the evidence sample. If the patterns match, the suspect may have contributed the evidence sample and if profiles do not match, the person is assumed innocent. Scien-tists believe that DNA evidence is far superior to eyewitness ac-counts, where the chance for correct identification is about 50:50. A combination of probes used in DNA analysis, gives greater reliability for profiling. Four to six probes are recommended for a good DNA profile. The recent new innovation in DNA chip technology (in which thousands of short DNA sequences are embedded onto a tiny silicon chip) enables a more rapid, and less expensive analysis using many more probes (Linacre and Graham, 2002).
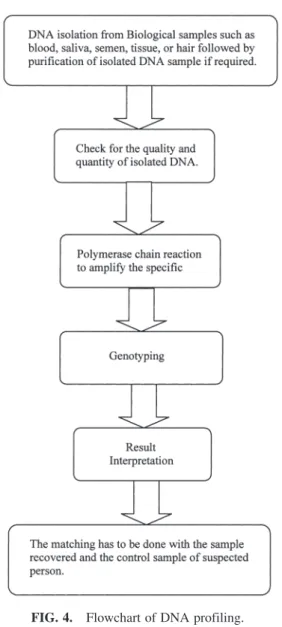
A flowchart is presented in Figure 4, which shows the steps involved in DNA profiling.
DNA TECHNOLOGIES CURRENTLY USED IN
FORENSIC INVESTIGATION
RFLP is a technique for analyzing the variable lengths of DNA fragments that result from digesting a DNA sample with a special kind of restriction endonucleases. Amplification frag-ment Length polymorphism (AFLP) is a PCR-based derivative method of RFLP in which sequences are selectively amplified using primers. It is a reliable and efficient method of detecting molecular markers. Random amplified polymorphic DNA (RAPD) is one of the most commonly used primary assays for screening the differences in DNA sequences of two species of plants. It involves random amplification (Cooper, 2000). HLA-DQ analysis was also used for identification for quite long time (Saiki et al., 1986). With the advancement, these nologies became obsolete and were replaced by various
tech-niques like microsatellite analysis, single nucleotide polymor-phisms, etc.
Capillary electrophoresis
Capillary electrophoresis, or CE, is a group of techniques used to separate a variety of samples. These analyses are performed in narrow tubes, and can result in the rapid separation of dif-ferent compounds. DNA typing with STR markers is now widely used for a variety of applications including human identifica-tion. CE instruments, such as genetic analyzers, are the method of choice for many laboratories which perform STR analysis. The Genetic Analyzer (Applied Biosystems, Foster City, CA) is a fluorescence based DNA analysis system utilizing cap-illary electrophoresis. It has multiple color fluorescence detec-tion capability. This capillary machine is capable of running se-quence and fragment analysis (Butler et al., 2004) and is a very popular method for STR typing. ABI 310 is a single-capillary
TABLE 1. YEAR-WISEADVANCEMENT INFORENSIC DNA TYPING
Year Developments
1985 RFLP approach was developed by Alec Jeffreys 1986 Automated DNA sequencing was first described 1988 FBI started using RFLP probes for case studies 1989 TWGDAM (Technical Working Group on DNA
Analysis Methods) was established
1995 O.J. Simpson case, which spread the awareness of DNA; UK DNA database established
1997 Y chromosome STR marker was first described Source: http://www.denverda.org/legalResource/Overview.pdf. FIG. 3. The progress in Forensic genetics. (Diagram reprinted by permission of Mark A. Jobling and Tanita Casci from paper “Encoded evidence: DNA in Forensic analysis, Mark A. Jobling, Nature reviews/Genetics, Vol 5, Oct 2004.)
instrument. Similarly ABI 3100 is a 16-capillary instrument and is used in large-scale applications.
Short tandem repeat analysis
STRs have discrete alleles. Variability in STR regions can be used to distinguish one DNA profile from another (Edwards
et al., 1991). The chance that two individuals will have the same 13-loci DNA profile is about one in one billion. Microsatellite typing can be multiplexed and semiautomated.
Sophisticated software has been developed for STR analysis to take sample electrophoretic data through the genotyping process (Ziegle et al., 1992). This is done in two steps by two different software programs. Genescan software (Applied Biosystems) is used to spectrally resolve the dye colors for each peak and to size the DNA fragments. The resulting pherograms are then imported into the second software program, Genotyper. This program de-termines each samples genotype by comparing the sizes of alle-les observed in a standard allelic ladder sample to those obtained at each locus in the DNA sample. Genotyping can be performed on the Genetic Analyzer with different available models (Ziegle
et al., 1992). These are capillary electrophoresis systems.
A Genescan program was used to compare the individual’s profile (Fig. 5). In genotyping the alleles are denoted by peaks rather than bands unlike Genescan and peak height is also taken into consideration while analyzing the results (Fig. 6, Probabil-ity of paternProbabil-ity is 99.999%). The match probabilities obtained with STR multiplexes are very low, and is more than the world population. The preferred system for STR genetic typing was via fluorescent dye-labeled primers and automated DNA sequencers. Multiplexes are analyzed and typed using automated sequencing equipment. First multiplex used was quadruplex, which had four STR regions. The recent multiplex amplifies for 16 loci in a sin-gle reaction one out of that is Amelogenin, a gender marker. STR base provides complete information about different STR mark-ers available (Butler, 2001). Simultaneous amplification and sep-aration of a number of STR loci in a single step is possible here. Other methods include the use of single nucleotide poly-morphs (SNPs), DNA amplification fingerprinting (DAF) and their offshoots. DNA amplification fingerprinting (DAF) in-volves amplification of genomic DNA, which is directed by only one oligonucleotide primer of arbitrary sequence (Caetano-Anolles et al., 1991). While the major efforts were made for de-tecting markers in genomic DNA, there has also been a consid-erable effort expanded on methods for typing sequences within the mitochondrial DNA (mtDNA). Although these results are not as discriminating as those of the STRs, they can be very use-ful in cases where skeletal remains, hairs, etc., are the only sam-ples available. There have also been advances in the use of mark-ers which are made specific (Y chromosome), and these have been used successfully particularly in cases of sexual assault.
Mitochondrial DNA analysis
Degraded samples such as hair, bones, and teeth quite often lack nucleated cellular material and cannot be analyzed with STR and RFLP. Mitochondrial DNA analysis (mtDNA) can be used to examine such DNA samples. mtDNA is extremely valu-VARSHA 184
FIG. 4. Flowchart of DNA profiling.
FIG. 5. Genescan analysis of STR markers in a paternity case. Lane 1. DNA profile of the mother; Lane 2. DNA profile of the child; Lane 3. DNA profile of the father.
able in such cases, as it has high copy number and has no re-combinational events.
Mitochondrial DNA is inherited maternally and does segre-gate independently. Therefore, it gives an excellent record of matrilineage. The father’s sperm contributes only nuclear DNA. Occurrence of heteroplasmy is the special feature of mtDNA. Comparing the mtDNA profile of unidentified remains with the profile of a potential maternal relative can be an important tech-nique in investigations on missing persons. Simultaneously, it can also be used as an important tool for matrilineage and evo-lutionary studies. Mitochondrial DNA can also help in studying the variety of human diseases, associated with mitochondrial dis-orders (Wallace, 1999). SNP outside the hypervariable segments along with mtDNA can provide us a better tool for typing.
Y-chromosome analysis
Analysis of genetic markers on the Y chromosome is espe-cially useful for tracing relationships among males or for ana-lyzing biological evidence involving multiple male contributors as in rape cases. There are a total of 219 useful STR markers available on Y chromosome (Kayser et al., 2004).
Single nucleotide polymorphism typing
SNP are DNA sequence variations that occur when a single nucleotide (A, T, C, or G) in the genome sequence is changed. SNPs create mutations in human and can be used as markers. SNPs have lower heterozygosity compared to STR. It will be not of much use when the samples are mixed, as SNP is a bi-nary marker. Forensic SNP typing provides considerable assis-tance to forensic scientists in major disasters (Budowle et al., 2005). Many polymorphic sites need to be analyzed for the iden-tification of SNPs. Mutation analysis is one of the major thrusts of functional genomics. Validation of a 21-locus autosomal SNP multiplex has been performed for forensic identification purposes (Dixon et al., 2005). Microarrays are used very
effi-ciently for mutation analysis, which helps in monitoring the whole genome on a single chip and for studying the interac-tions of thousands of genes simultaneously. In the scientific lit-erature various terminologies have been used to describe this technology as such as DNA chip, DNA microarray, etc. (Linacre and Graham, 2002).
Forensic studies are mainly done by commercially developed autosomal STR multiplexes, autosomal SNPs, and markers on the Y chromosome and mitochondrial DNA (Jobling and Gill, 2004).
ESTABLISHMENT OF DATABANKS
DNA intelligence databases are modern tools for the execu-tive force investigating crime (Werret, 1997; Hoyle, 1998; Par-soh et al., 1998; Peeren boom, 1998; Schneider, 1998).DNA forensics databases
The first of the national DNA databases involving STR data was formed in the UK in 1995. The usefulness of such types of databases was established during the first 5 years of use of this technology (Werrett and Sparkes, 1998). By 1998, three other European countries, Austria, Germany, and The Nether-lands had introduced national DNA databases (Martin et al., 2001). The STR loci used to form these databases were THOI, VWA, FGA, D8S1179, D18S51, and D21S11. Finland and Nor-way have introduced a national database in 1999. Sweden has already a database of unknown crime samples. Belgium, Den-mark, Switzerland, France, Italy, Spain, and some east Euro-pean countries also took steps in this direction. In 1989, the FBI proposed a national DNA database for North America.
The Austrian DNA intelligence database was inaugurated in 1997. In the United States, the Federal Bureau of Investigation (FBI) has created a national database of genetic information called the National DNA Index System. The database contains FIG. 6. Electropherogram of genotype of STR markers in samples of a paternity case. 1. Genotype of the mother; 2. Genotype of the child; 3. Genotype of the father.
DNA obtained from convicted criminals and from evidence found at crime scenes.
National DNA databank: CODIS
The COmbined DNA Index System, CODIS, utilizes com-puter and DNA technologies together. All DNA profiles stored in CODIS are generated using STR analysis. The current ver-sion of CODIS uses two indexes where biological evidence is recovered from the crime scene. The Convicted Offender in-dex contains DNA profiles of individuals convicted of sex of-fenses and other violent crimes. The Forensic index contains DNA profiles developed from crime scene evidence. A com-puter software will search these two indexes for matching DNA profiles. Law enforcement agencies compare DNA from bio-logical evidence collected from crime scenes that have no sus-pect with the DNA profiles stored in the CODIS systems. If a match is made between a sample and a stored profile, CODIS can identify the culprit.
This technology is authorized by the DNA Identification Act of 1994. As of January 2003, the database contained more than a million DNA profiles in its Convicted Offender Index and about 48,000 DNA profiles collected from crime scenes but which have not been connected to a particular offender. This database can also be used to identify criminals who are re-sponsible for crime in other countries. The FBI uses a standard set of 13 specific STR regions for CODIS (Budowle et al., 1998). CODIS is a software program that operates local, state, and national databases of DNA profiles from convicted crimi-nals and missing persons.
Advantages of DNA databanking
One advantage of a comprehensive and modern national DNA databank is the capability to perform DNA identification tests of victims and missing persons in the case of disaster.
It is just a matter of comparison of the profile of the sample with the databank, as the data is already available and the re-sults are more reliable. Time of investigation as well as cost is reduced. It is also beneficial to society as it helps the investi-gator to catch the criminal and exonerate the innocent person.
APPLICATIONS OF DNA FINGERPRINTING
TO PLANTS AND ANIMALS
Plant DNA fingerprinting is defined here as the application of molecular marker techniques to identify cultivars, but this also can be helpful in providing evidence in crime cases. RAPD can be a better choice for identification of plant strains. DNA profiling of plants can be used in solving civil disputes over the identity of commercially important cultivars (Kumar et al., 2001).
The genotypic information is used for identification of ani-mals (Barallon, 1998), forensic analysis (Zehner et al., 1998) and for tracing specific animals to finished meat products (Var-sha et al., 2003), pedigree analysis, selection, and breeding pro-grams. Forensic analysis can be of immense help for poaching cases and to check the illegal trade of endangered species. The cytochrome bregion had been used for identification of the en-dangered animal species (Branicki et al., 2003). DNA typing in plants and animals is again a vast topic, and needs a separate
discussion. DNA fingerprinting has tremendous application es-pecially in forensic sciences, which is mentioned in Table 2.
ETHICAL, LEGAL, AND SOCIAL CONCERNS
INVOLVING DNA FINGERPRINTING
The primary ethical, legal, and social issues concerning DNA fingerprinting is privacy. DNA profiles are different from latent fingerprints, which are useful only for identification. DNA can provide insights into many aspects of a person and their families including susceptibility to particular genetic disorders, legitimacy of birth, and fertility. This increases the potential for genetic dis-crimination by government, society and others. Therefore, DNA typing should be carried out in a very sophisticated way, and should meet all the international standards and follow and abide all the ethical, legal, and social concerns involved with DNA typ-ing. Databanking also involves some ethical issues and requires a balance between human rights and justice. Retention of sam-ples for long periods of time is also risky, since this provides considerable accessibility to private genetic information.RELIABILITY OF DNA TECHNOLOGY
Generally, the legal system and courts have accepted the re-liability of DNA testing everywhere, and admitted DNA test results as evidence. However, DNA fingerprinting is still con-troversial because of some important aspects. One is the accu-racy of the results, as result will always be in the form of some probability value. Another aspect is the high cost associated with analysis. Some of the other important aspects such as the possibility of misuse of the technique, chances of manual er-ror, and contamination cannot be ignored.The accuracy of DNA fingerprinting has been challenged be-cause DNA segments rather than complete DNA strands are an-alyzed, which may not be necessarily unique. More research work is needed to confirm the uniqueness of DNA fingerprint-ing has not been conducted. In addition, DNA ffingerprint-ingerprintfingerprint-ing is often performed in private laboratories that may not follow uni-form testing standards and quality controls. Therefore, human error could lead to false results. Awareness is still lacking about this technology among law enforcing agencies and judiciaries, requiring a lot of interaction between these and forensic scien-tists. DNA identification can be quite effective if used intelli-VARSHA 186
TABLE2. SOMEEXAMPLES OFDNA USE FORFORENSIC IDENTIFICATION
Criminal identification.
Exonerate persons wrongly accused of crimes.
Identify crime and catastrophe victims in mass disasters. Establish paternity and other family relationships.
Identify endangered species and can help in wild-life poach-ing cases.
Match organ donors with recipients in organ transplantation as kidney transplantation.
Determine pedigree for seed or livestock breeds.
To identify different strains and cultivars and can help in pro-tecting the rights of plant breeders.
gently. Portions of the DNA sequence that vary the most among humans must be used; also, the number of loci studied should be sufficient enough to make the results more reliable.
PROBLEMS WITH DNA FINGERPRINTING
DNA fingerprinting is not fully assured. The term DNA fin-gerprint is, in a true sense, a misnomer. We cannot compare it with latent fingerprinting, as it is not as specific as fingerprints. Actually, a VNTR pattern gives a probability value that the per-son in question is indeed the perper-son to whom the VNTR pat-tern (of the child, the criminal evidence, or whatever else) be-longs. The probability decides whether the person can be reasonably matched with the DNA fingerprint.1. Generation and determination of high probability is a req-uisite for forensic cases. The probability of a DNA finger-print belonging to a specific person needs to be reasonably high, particularly in criminal cases. High probability can be achieved by increasing the number of polymorphic loci an-alyzed. Population structure can cause variation in allele fre-quencies between subpopulations (Lander and Budowle, 1994). VNTRs are not distributed evenly across all of hu-man population. A given VNTR does not have a stable prob-ability of occurrence, and not enough is known about the VNTR frequency distributions among ethnic groups to de-termine accurate probabilities for individuals within those groups. Some technical difficulties, like errors in the hy-bridization and probing process, will also affect the proba-bility. Population studies should be carried out and repro-ducibility of the results should be confirmed.
2. DNA testing faces important problem of impurities of the sam-ple. The sample collected from a crime scene most of the time is mixed with soil and other potential contaminants and con-siderable time and effort in purification processes purification on samples not directly amenable to analysis. Another impor-tant challenge to amplifying DNA samples from crime scenes is the fact that the PCR amplification process can be affected by inhibitors present in the samples themselves. The sensitiv-ity of PCR with its abilsensitiv-ity to amplify low quantities of DNA can be a problem if proper care is not taken. Mixed samples can be challenging to detect and interpret. If mutation occurs at the primer binding site it can result in failure to amplify, which is known as null allele or allele dropout. Sometimes, stutter bands are noted in the electropherogram of STR analy-sis, which is nothing but small peaks several bases smaller than STR allele peaks, which occurs because of strand slippage. These bands effect the interpretation of DNA profiles specially in mixed samples, which requires good understanding of the behavior of stutter products (John M. Butler, forensic DNA typing). Proper training, better understanding, extensive expe-rience, and careful analysis are required.
3. Very often investigators are not properly trained for sample collection and encounter problems in identification and col-lection of proper DNA evidence from the scene of crime. Contamination of the samples and using the correct sample, proper labeling, etc., are a few important aspects, which should be taken into consideration. The DNA laboratory should follow the international standard and should be ac-credited.
4. Proper awareness about the subject is required among the investigators, but unfortunately, interaction and proper com-munication is lacking between law enforcement agencies and DNA laboratories.
5. Limited resources and poor infrastructure facility of differ-ent laboratories effects the efficiency of the DNA typing lab-oratories.
6. Risk of false or misleading results from DNA testing and chances of tampering with the evidence cannot be ignored. Strict confidentiality is required as far as identity of the sam-ple is concerned, and each samsam-ple should be tested by more than one examiner to confirm the results. Irrespective of hav-ing all these limitations, DNA fhav-ingerprinthav-ing is a very reliable technique for identification if used efficiently and intelligently. There are many international organizations, which are in-volved in quality control and uniformity of DNA fingerprint-ing such as the Technical Workfingerprint-ing Group on DNA Analysis Methods, The DNA Advisory Board, National Institute of Stan-dards and Technology in United States, and International So-ciety for Forensic Genetics in Europe.
SOME HIGH-PROFILE CASES SOLVED BY
DNA FINGERPRINTING
1. Identification of the remains of the last Russian Czar. 2. O.J. Simpson case in Los Angeles.
3. Clinton-Lewinsky affair.
4. Thomas Jefferson-Sally Hemings affair.
5. Identification of the bodies in World Trade Center attack: A mass disaster.
6. Assassination case of Sri Rajeev Gandhi, Prime Minister of India.
CONCLUSION AND FUTURE DIRECTIONS
The power of DNA technology will be of immense help in the future. The limitations of the technique have to be given se-rious thought. Forensic DNA testing will improve day by day, and will be benefited with the outcome of the Human Genome Project. Microarray-based analysis will be in much use for its more reliability and fast analysis. This technology has the po-tential to review the cases, which were previously reported with older methods. There is a real need for public awareness of the new DNA technologies and their implications. For instance, the collection of the correct sample and its proper preservation is very crucial. Productive interactions are required between the forensic scientists and the law enforcement and other investiga-tors. It is necessary to make this technology more popular and cost effective so as to enable even developing and undeveloped third world countries to take advantage of the technology as well.ACKNOWLEDGMENTS
My special thanks to all those authors whose names are not quoted here because of space constraints, and those who have made the information freely available on the internet. I am
thankful to Dr. S. E. Hasnain and Dr. J. Nagaraju, for their con-stant support and encouragement. A draft of this manuscript was significantly improved through consultation with Dr. T. Ramasarma, Dr. Gayatri Ramakrishna, and the comments of the anonymous reviewer, and I thank them for their valuable sug-gestions. I am also thankful to Dr. Mark A. Jobling, author, and Dr. Tanita Casci, senior editor, Nature Reviews Genetics for permitting me to take Figure 3 from the published paper.
REFERENCES
BARALLON, R. (1998). Species determination by analysis of the Cy-tochrome bgene. In Forensic DNA Profiling Protocols. P.J. Lincoln and J. Thompson, ed. (Humana Press, Totowa, NJ).
BRANICKI, W., KUPIEC, T., and PAWLOWSKI, R. (2003).Valida-tion of cytochrome bsequence analysis as a method of species iden-tification. J. Forensic Sci. 48,83–87.
BUDOWLE, B., BIEBER, F.R., and EISENBERG, A.J. (2005). Foren-sic aspects of mass disasters: Strategic considerations for DNA-based human identification. Legal Med. 7,230–243.
BUDOWLE, B., MORETTI, T.R., NIEZGODA, S.J., and BROWN, B.L. (1998). Proceedings of the Second European Symposium on Hu-man Identification. (Promega Corporation, Madison, WI), pp. 73–88. BUTLER, J.M. (2001). Forensic DNA typing: Biology and technology
behind STR markers (Academic Press, New York).
BUTLER, J.M., BUEL, E., CRIVELLENTE, F., and MCCORD, B.R. (2004). Forensic DNA typing by capillary electrophoresis using the ABI Prism 310 and 3100 genetic analyzers for STR analysis. Elec-trophoresis 25,1397–1412.
CAETANO-ANOLLES, G., BASSAM, B.J., and GRESSHOFF, P.M. (1991). DNA amplification fingerprinting using very short arbitrary oligonucleotide primers. Biotechnology (NY) 9,553–557. COOPER, M.L. (2000). RAPD analysis of southern brown bandicoots
(Isodon obesulus) populations in West Australia reveals genetic dif-ferentiation elated to environmental variables. Mol. Ecol. 9,469–479. DIXON, L.A., MURRAY, C.M., ARCHER, E.J., DOBBINS, A.F., KOUMBI, P., and GILL, P. (2005). Validation of a 21-locus auto-somal SNP multiplex for forensic identification purposes. Forensic Sci. Int. 154,62–77.
EDWARDS, A., CIVITELLO, A., HAMMOND, H.A., and CASKEY, C.T. (1991). DNA typing & genetic mapping with trimeric & tetremeric tandem repeats. Am. J. Hum. Genet. 49,746–756. HOYLE, R. (1998). Forensics. The FBI’s national DNA database. Nat.
Biotechnol. 16,987.
JEFFREYS, A.J., WILSON, V., and THEIN, S.L. (1985). Individual specific “fingerprints” of human DNA. Nature 316,76–79. JOBLING, M.A., and GILL, P. (2004). Encoded evidence: DNA in
forensic analysis. Nat. Rev. 5,739–751.
KAYSER, M., et al. (2004). A comprehensive survey of human Y chro-mosomal microsatellite. Am. J. Hum. Genet. 74,1183–1197. LALJI S. (1991). DNA profiling and its applications. Curr. Sci. 60,
580–585.
KUMAR, L.D., KATHIRVEL, M., RAO, G.V., and NAGARAJU, J. 2001. DNA profiling of disputed chilli samples (Capsicus annum) using ISSR-PCR and FISSR-PCR marker assays. Forensic Sci. Int. 116,63–68.
LANDER, E.S., and BUDOWLE, B. (1994). DNA Fingerprinting dis-pute laid to rest. Nature 371,735–738.
LANDSTEINER, K. (1901). Uber agglutinationserscheinungen nor-malen menschlichen blutes. Wien Klin Wschr. 14,1132–1134.
LINACRE, A., and GRAHAM, D. (2002). Role of molecular diagnos-tics in forensic science. Expert Rev. Mol. Diagn. 2,346–353. MARTIN, P.D., HERMANN, S., and SCHNEIDER, P.M. (2001). A
brief history of the formation of DNA database in forensic science within Europe. Forensic Sci. Int. 119,225–231.
PARSOH, W., STEINLECHNER, M., SCHEITHAUER, R., and SCHNEIDER, D.M. (1998). National DNA intelligence database in Europe—Report on the current situation. In Proc 9th Symp Human identification(Promega Corporation, Madison, WI). pp. 52–54. PEEREN BOOM, E. (1998). Central DNA database created in
Ger-many. Nat. Biotechnol. 16,510–511.
SAIKI, R.K., BUGAWAN, T.L., HORN, G.T., MULLIS, K.B., and ERLICH, H.A. (1986). Analysis of enzymetically amplified -glo-bin and HLA-DQDNA with allele–specific oligonucleotide probes. Nature 324,163–166.
SCHNEIDER, P.M. (1998). DNA databases for offender identification in Europe—The need for technical, legal & political harmonization. In Proc Europ 2nd Symp Human identification(Promega Corpora-tion, Madison, WI) pp. 40–44.
VARSHA, et al. (1999). D1S80 variation in random Indian popula-tions. Proceedings of X Intl. Symposium of Human Identification, FL, USA.
VARSHA, et al. (2003). Identification of cooked meat sample by 12Sr-RNA sequence analysis. Proceedings of 55th annual meeting of
American Academy of Forensic Sciences,Chicago, IL.
WALLACE, D.C. (1999). Mitochondrial diseases in man and mouse. Science 283,1482–1488.
WEBER, J.L., and MAY, P.E. (1989). Abundant class of human DNA polymorphisms which can be typed using polymerase chain reaction. Am. J. Hum. Genet. 44,388–396.
WEISING, K., NYMBOM, H., WOLFF, K., and MAYER, W. (1995).
DNA Fingerprinting in Plants and Fungi(CRL Press, Boco Raton, FL).
WERRET, D.J. (1997). The National DNA database. Forensic Sci. Int. 88,33–42.
WERRET, D.J., and SPARKES, R. (1998). Proceedings of Ninth In-ternational Symposium on Human Identification(Promega Corpora-tion, Madison, WI) pp. 55–62.
WONG, Z., WILSON, V., PATEL, I., POVEY, S., JEFFREYS, A.J. (1987). Characterization of a panel of highly variable mini-satellites, cloned from human DNA. Ann. Hum. Genet. 51,269–288. ZEHNER, R., ZIMMERMAN, S., and MEBS, D. (1998). RFLP and
se-quence analysis of the cytochrome bgene of selected animals and man: Methodology and forensic application. Int. J. Legal Med. 111, 323–327.
ZIEGLE, J.S., SU, Y., CORCORAN, K.P., NIE, L., MAYRAND, P.E., HOFF, L.B., MCBRIDE, L.J., KRONICK, M.N., and DIEHL, S.R. (1992). Ligase-based detection of monoclonal repeat sequences. Ge-nomics 14,1026–1031.
Address reprint requests to:
Varsha Laboratory of DNA Fingerprinting Services Centre for DNA Fingerprinting and Diagnostics ECIL Road Nacharam, Hyderabad-76, AP, India E-mail:varsha@cdfd.org.in Received for publication November 4, 2005; received in revised form December 14, 2005; accepted December 15, 2005.
VARSHA 188