0095-1137/10/$12.00
doi:10.1128/JCM.00156-10
Copyright © 2010, American Society for Microbiology. All Rights Reserved.
Rapid Genus- and Species-Specific Identification of
Cronobacter
spp.
by Matrix-Assisted Laser Desorption Ionization–Time of
Flight Mass Spectrometry
䌤
Roger Stephan,
1* Dominik Ziegler,
2Valentin Pflu
¨ger,
2Guido Vogel,
2and Angelika Lehner
1Institute for Food Safety and Hygiene, Vetsuisse Faculty University of Zurich, Zurich, Switzerland,
1and Mabritec AG, Riehen, Switzerland
2Received 25 January 2010/Returned for modification 15 March 2010/Accepted 9 June 2010
Cronobacter
spp. are Gram-negative opportunistic food-borne pathogens and are known as rare but
impor-tant causes of life-threatening neonatal infections. Rapid and reliable identification of
Cronobacter
species and
their differentiation from phenotypically similar, nonpathogenic
Enterobacter turicensis
,
Enterobacter helveticus
,
and
Enterobacter pulveris
have become increasingly important. We evaluated here the application of
matrix-assisted laser desorption ionization–time of flight mass spectrometry (MALDI-TOF MS) for rapid genus and
species identification of the six
Cronobacter
species recognized so far. To this end, we developed a reference MS
database library that includes 54
Cronobacter
target strains as well as 17 nontarget strains. The strains
provided reproducible and unique mass spectra profiles covering a wide molecular mass range (2,000 to 30,000
Da). Genus- and species-specific biomarker protein mass patterns were determined. The defined biomarker
mass patterns (Spectral Archive and Microbial Identification System [SARAMIS] SuperSpectrum) were
validated using 36 strains from various
Cronobacter
species as well as eight nontarget strains. For all strains
the mass spectrometry-based identification scheme yielded identical results as with a PCR-based identification
system. All strains were correctly identified, and no nontarget strain was misidentified as
Cronobacter
. Our
study demonstrates that MALDI-TOF MS is a reliable and powerful tool for the rapid identification of
Cronobacter
strains to the genus and species level.
The genus
Cronobacter
comprises six species,
Cronobacter
sakaza-kii
,
Cronobacter malonaticus
,
Cronobacter muytjensii
,
Cronobacter
dublinensis
,
Cronobacter turicensis
, and
Cronobacter
genomo-species 1, which have been recognized on the basis of a
polyphasic approach using extensive geno- and phenotypic
evaluations (8, 9). The definition of six species within the genus
Cronobacter
was further supported by a recent study in which
a multilocus sequence analysis (MLSA) approach was used
(11).
Cronobacter
spp. are Gram-negative opportunistic
food-borne pathogens and are known as rare but important causes
of life-threatening neonatal infections which can lead to severe
disease manifestations such as brain abscesses, meningitis,
ne-crotizing enterocolitis, and systemic sepsis (12). Neonates and
infants under 2 months of age which are born prematurely are
at greater risk of
Cronobacter
infections from consuming
Cronobacter
-contaminated powdered infant formulas (8).
Rapid and reliable identification of strains of the genus
Cronobacter
and differentiation from phenotypically similar,
apathogenic
Enterobacter turicensis
,
Enterobacter helveticus
,
and
Enterobacter pulveris
are important for surveillance,
pre-vention, and control of food-borne diseases. Moreover, for
Cronobacter
a species differentiation is relevant for
epidemio-logical studies, and the different species show differences in
sensitivities to chemical agents and antibiotics. However,
iden-tification of
Cronobacter
species by biochemical tests is very
laborious. Although very recently Stoop et al. (14) described
for the first time
rpoB
based PCR systems which enable the
identification to the species level strains previously confirmed
to belong to the genus
Cronobacter
, there is still a need for
alternative procedures that allow rapid and reliable
identifica-tion.
In recent years, several reports have shown the feasibility of
using matrix-assisted laser desorption ionization–time of flight
mass spectrometry (MALDI-TOF MS) for identifying
micro-organisms (for a review, see reference 13). The detection of
protein mass patterns has become a convenient tool for the
rapid analysis of bacteria (5). The method analyzes the profiles
of proteins that are extracted from whole bacteria. A MALDI
mass spectrometer can efficiently detect numerous molecules
simultaneously. Protein mass patterns can be used for
identi-fication of bacteria at the genus, species, and in some cases the
subspecies level. The mass pattern detection method has been
validated recently in several studies for food-borne pathogens,
including the classification and identification of
Arcobacter
,
Campylobacter
,
Clostridia
,
Listeria
,
Salmonella
,
Staphylococcus
species and
Vibrio parahaemolyticus
(1, 2, 3, 4, 6, 7).
The aim of the present study was to develop a reference MS
database library of biomarker protein mass patterns (Spectral
Archive and Microbial Identification System [SARAMIS]
SuperSpectrum), including
Cronobacter
target strains as well as
nontarget strains, and to perform thereafter a blind test using
strains from various
Cronobacter
species and nontarget strains
to validate the MALDI-TOF-based identification system.
MATERIALS AND METHODS
Bacterial strains.A first set of strains was used to define the superspectrum:
30C. sakazakiistrains (E266, E269, E272, E274, E280, E283, E292, E302, E309,
* Corresponding author. Mailing address: Institute for Food Safety
and Hygiene, Vetsuisse Faculty University of Zurich, Winterthurerstr.
272, CH-8057 Zurich, Switzerland. Phone: 6358651. Fax:
41-44-6358908. E-mail: stephanr@fsafety.uzh.ch.
䌤
Published ahead of print on 16 June 2010.
2846
on May 16, 2020 by guest
http://jcm.asm.org/
E314, E328, E393, E423, E468, E601, E602, E604, E607, E620, E622, E624, E627, E632, E736, E739, E750, E761, E768, E796, and E828), fourC. malonati-cusstrains (E265, E621, E825, and E829), sixC. turicensisstrains (3032, E609, E626, E676, E681, and E688), twoC. genomospeciesstrains (E680 and E797), six
C. muytjensiistrains (E456, E488, E603, E616, E769, and E888), and six C. dublinensisstrains (E464, E465, E515, E791, E798, and E799), with isolates originating from human, food, and environmental origins included in this study. The selected strains were part of a taxonomy study (9, 10). This factor guaran-teed the correct identification of all strains used for the definition of the super-spectrum.
Moreover, several nontarget strains were included in the study (Escherichia coliATCC 25922,Escherichia hermaniiDSM 4560,Escherichia vulnerisDSM 4564,Enterobacter cloacaeDSM 30054,Enterobacter kobeiDSM 13645, Entero-bacter ludwigiiDSM 166889,Enterobacter cancerogenusDSM 17580,Enterobacter asburiaeDSM 17506,Enterobacter radicincitansDSM 16656,Leclerica adecaboxy-lataDSM 5077,Enterobacter pyrinusDSM 12410,Enterobacter aerogenesDSM 30053,Enterobacter turicensis 508/05T
[DSM 18397T
], Enterobacter helveticus
513/05T[DSM 18396T]),Enterobacter pulveris601/05T[DSM 19144T]),
Entero-bacter amnigenusDSM 4486, andE. cowaniiDSM 18146).
Second set of strains for validation purposes.After defining the superspectra
for the different species, 44 additional strains isolated within a field study and identified by the recently described PCR system (14) were used for validation of the system in a blind test: 24C. sakazakii, 7C. malonaticus, 1C. turicensis, 2C. dublinensis, 2C. muytensii, and 8 nontarget strains. The selected strains origi-nated from food and environmental samples.
Culture conditions.Standard conditions were used for bacterial culturing. All
strains were streaked onto sheep blood agar (Difco Laboratories, Detroit, MI; 5% sheep blood from Oxoid, Hampshire, United Kingdom) and on tryptic soy agar (TSA) agar (BD, Sparks, MD) and incubated at 37°C for 24 and 48 h, and single colonies were selected.
Preparation of samples for MALDI-TOF MS of whole bacterial cells.Cells
from representative single bacterial colonies were directly smeared onto a target spot of a steel target plate by using a disposable loop, overlaid with 1l of matrix consisting of a saturated solution of sinapic acid (49508; Sigma-Aldrich, Buchs, Switzerland) in 60% acetonitrile (154601; Sigma-Aldrich, Buchs, Switzerland)– 0.3% trifluoroacetic acid (T6508; Sigma-Aldrich, Buchs, Switzerland), and air dried within minutes at room temperature. A first subset of strains grown on TSA and blood agar was tested after 24 and 48 h to compare the reproducibility of masses within eight replicates and to choose the optimized growth conditions for further studies. Additionally, replicates of each isolate were transferred onto two independent steel targets for mass accuracy and proof of external instrument
calibration. All strains included in this study were cultivated for 48 h on blood agar and TSA and were measured in eight replicates. For each strain, four distinct single bacterial colonies were spotted in duplicate onto two independent steel target plates.
MALDI-TOF MS parameters.Protein mass fingerprints were obtained using a
MALDI-TOF mass spectrometry Axima Confidence machine (Shimadzu-Bio-tech Corp., Kyoto, Japan), with detection in the linear, positive mode at a laser frequency of 50 Hz and within a mass range of 2,000 to 30,000 Da. Acceleration voltage was 20 kV, and the extraction delay time was 200 ns. A minimum of 10 laser shots per sample was used to generate each ion spectrum. For each bac-terial sample, a total of 100 protein mass fingerprints were averaged and pro-cessed using the Launchpad v. 2.8 software (Shimadzu-Biotech Corp., Kyoto, Japan). This software was also used for peak processing of all raw spectra with the following settings: the advanced scenario was chosen from the Parent Peak Cleanup menu, peak width was set at 80 channels, the smoothing filter width was set at 50 channels, the baseline filter width was set at 500 channels and for peak detection method the threshold apex was chosen. For the threshold apex peak detection, the threshold type was set as dynamic and the threshold offset was set to 0.025 mV with a threshold response factor of 1.25. The processed spectra were exported as peak lists withm/zvalues and signal intensities for each peak in the ASCII format. Calibration was conducted for each target plate using spectra of the reference strainEscherichia coliDH5␣.
MALDI-TOF MS spectral analysis.Reference spectra for the first set of 54
Cronobacterstrains and 17 nontarget strains were analyzed in duplicates for each of the four representative single bacterial colonies. Generated protein mass fingerprints were analyzed with SARAMIS (AnagnosTec, Potsdam-Golm, Ger-many). In the first step biomarker mass patterns, called superspectra, were calculated for the genusCronobacteras well as for each of the sixCronobacter
species, using the SARAMIS SuperSpectrum tool. Therefore, peak lists for all 54
[image:2.585.44.283.69.285.2]Cronobacterisolates (reference set) were imported into the SARAMIS software in octuplicates. Peak lists were trimmed to a mass range of 2 to 30 kDa, and peaks with a relative intensity below 1% were removed. Peak lists were binned, and average masses were calculated using the SARAMIS SuperSpectrum tool with an error of 800 ppm. For the identification ofCronobactergenus, respec-tively, species-specific protein mass patterns, allEnterobacteriaceaefamily bio-marker masses were excluded by using the SARAMISEnterobacteriaceaefamily exclusion list. From the remaining peaks only masses present in at least 50% of the spectra were selected for the genus superspectrum; alternatively, masses
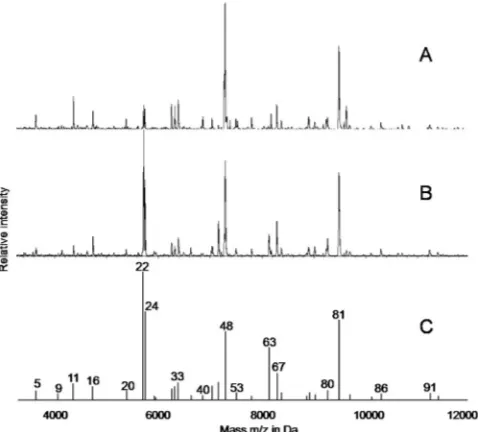
FIG. 1. Representative spectra of
C. sakazakii
E393 grown on
blood agar (A) or TSA (B) and the corresponding superspectrum
computed with the SARAMIS software (C). Selected biomarker
masses (4,000 to 12,000 Da) and the corresponding peak numbers are
as shown in Table 1.
genic
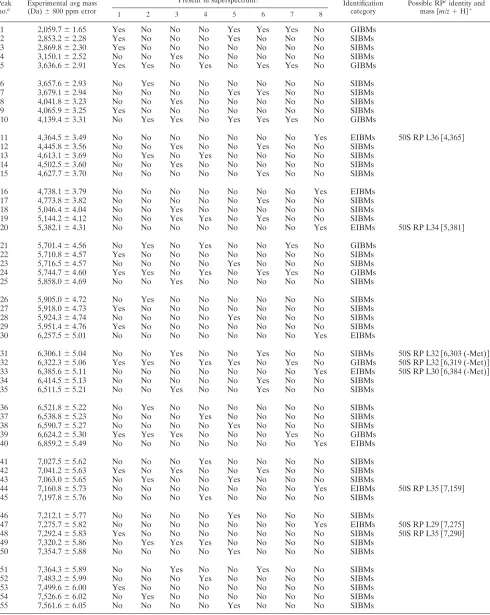
FIG. 2. Spectra of different
Enterobacter
species, grown on TSA. The spectra display high
Cronobacter
species and three
apatho-overall levels of similarity but slight mass shifts of peaks in the protein
profiles (4,000 to 12,000 Da).
on May 16, 2020 by guest
http://jcm.asm.org/
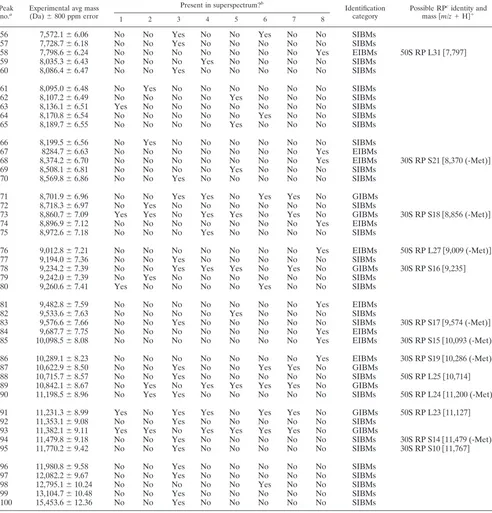
[image:2.585.301.544.70.318.2]TABLE 1. Selected genus- and species-identifying biomarker masses obtained by whole-cell MALDI-TOF-MS of
Cronobacter
spp.
Peak
no.a Experimental avg mass
(Da)⫾800 ppm error
Present in superspectrum?b
Identification category
Possible RPc
identity and mass关m/z⫹H兴⫹
1 2 3 4 5 6 7 8
1
2,059.7
⫾
1.65
Yes
No
No
No
Yes
Yes
Yes
No
GIBMs
2
2,853.2
⫾
2.28
Yes
No
No
No
Yes
No
No
No
SIBMs
3
2,869.8
⫾
2.30
Yes
No
No
No
No
No
No
No
SIBMs
4
3,150.1
⫾
2.52
No
No
Yes
No
No
No
No
No
SIBMs
5
3,636.6
⫾
2.91
Yes
Yes
No
Yes
No
Yes
Yes
No
GIBMs
6
3,657.6
⫾
2.93
No
Yes
No
No
No
No
No
No
SIBMs
7
3,679.1
⫾
2.94
No
No
No
No
Yes
Yes
No
No
SIBMs
8
4,041.8
⫾
3.23
No
No
Yes
No
No
No
No
No
SIBMs
9
4,065.9
⫾
3.25
Yes
No
No
No
No
No
No
No
SIBMs
10
4,139.4
⫾
3.31
No
Yes
Yes
No
Yes
Yes
Yes
No
GIBMs
11
4,364.5
⫾
3.49
No
No
No
No
No
No
No
Yes
EIBMs
50S RP L36
关
4,365
兴
12
4,445.8
⫾
3.56
No
No
Yes
No
No
Yes
No
No
SIBMs
13
4,613.1
⫾
3.69
No
Yes
No
Yes
No
No
No
No
SIBMs
14
4,502.5
⫾
3.60
No
No
Yes
No
No
No
No
No
SIBMs
15
4,627.7
⫾
3.70
No
No
No
No
No
Yes
No
No
SIBMs
16
4,738.1
⫾
3.79
No
No
No
No
No
No
No
Yes
EIBMs
17
4,773.8
⫾
3.82
No
No
No
No
No
Yes
No
No
SIBMs
18
5,046.4
⫾
4.04
No
No
Yes
No
No
No
No
No
SIBMs
19
5,144.2
⫾
4.12
No
No
Yes
Yes
No
Yes
No
No
SIBMs
20
5,382.1
⫾
4.31
No
No
No
No
No
No
No
Yes
EIBMs
50S RP L34
关
5,381
兴
21
5,701.4
⫾
4.56
No
Yes
No
Yes
No
No
Yes
No
GIBMs
22
5,710.8
⫾
4.57
Yes
No
No
No
No
No
No
No
SIBMs
23
5,716.5
⫾
4.57
No
No
No
No
Yes
No
No
No
SIBMs
24
5,744.7
⫾
4.60
Yes
Yes
No
Yes
No
Yes
Yes
No
GIBMs
25
5,858.0
⫾
4.69
No
No
Yes
No
No
No
No
No
SIBMs
26
5,905.0
⫾
4.72
No
Yes
No
No
No
No
No
No
SIBMs
27
5,918.0
⫾
4.73
Yes
No
No
No
No
No
No
No
SIBMs
28
5,924.3
⫾
4.74
No
No
No
No
Yes
No
No
No
SIBMs
29
5,951.4
⫾
4.76
Yes
No
No
No
No
No
No
No
SIBMs
30
6,257.5
⫾
5.01
No
No
No
No
No
No
No
Yes
EIBMs
31
6,306.1
⫾
5.04
No
No
Yes
No
No
Yes
No
No
SIBMs
50S RP L32
关
6,303 (-Met)
兴
32
6,322.3
⫾
5.06
Yes
Yes
No
Yes
Yes
No
Yes
No
GIBMs
50S RP L32
关
6,319 (-Met)
兴
33
6,385.6
⫾
5.11
No
No
No
No
No
No
No
Yes
EIBMs
50S RP L30
关
6,384 (-Met)
兴
34
6,414.5
⫾
5.13
No
No
No
No
No
Yes
No
No
SIBMs
35
6,511.5
⫾
5.21
No
No
Yes
No
No
Yes
No
No
SIBMs
36
6,521.8
⫾
5.22
No
Yes
No
No
No
No
No
No
SIBMs
37
6,538.8
⫾
5.23
No
No
No
Yes
No
No
No
No
SIBMs
38
6,590.7
⫾
5.27
No
No
No
No
Yes
No
No
No
SIBMs
39
6,624.2
⫾
5.30
Yes
Yes
Yes
No
No
No
Yes
No
GIBMs
40
6,859.2
⫾
5.49
No
No
No
No
No
No
No
Yes
EIBMs
41
7,027.5
⫾
5.62
No
No
No
Yes
No
No
No
No
SIBMs
42
7,041.2
⫾
5.63
Yes
No
Yes
No
No
Yes
No
No
SIBMs
43
7,063.0
⫾
5.65
No
Yes
No
No
Yes
No
No
No
SIBMs
44
7,160.8
⫾
5.73
No
No
No
No
No
No
No
Yes
EIBMs
50S RP L35
关
7,159
兴
45
7,197.8
⫾
5.76
No
No
No
Yes
No
No
No
No
SIBMs
46
7,212.1
⫾
5.77
No
No
No
No
Yes
No
No
No
SIBMs
47
7,275.7
⫾
5.82
No
No
No
No
No
No
No
Yes
EIBMs
50S RP L29
关
7,275
兴
48
7,292.4
⫾
5.83
Yes
No
No
No
No
No
No
No
SIBMs
50S RP L35
关
7,290
兴
49
7,320.2
⫾
5.86
No
Yes
Yes
Yes
No
No
No
No
SIBMs
50
7,354.7
⫾
5.88
No
No
No
No
Yes
No
No
No
SIBMs
51
7,364.3
⫾
5.89
No
No
Yes
No
No
Yes
No
No
SIBMs
52
7,483.2
⫾
5.99
No
No
No
Yes
No
No
No
No
SIBMs
53
7,499.6
⫾
6.00
Yes
No
No
No
No
No
No
No
SIBMs
54
7,526.6
⫾
6.02
No
Yes
No
No
No
No
No
No
SIBMs
55
7,561.6
⫾
6.05
No
No
No
No
Yes
No
No
No
SIBMs
Continued on following page
on May 16, 2020 by guest
http://jcm.asm.org/
present in at least 75% of the spectra were selected for species superspectra. The specificity of these potential biomarker masses was determined by comparison against the whole SARAMIS spectral archive. Fourteen masses for the genus and between 19 and 33 masses for the different species were weighted and used as superspectrum for automatedCronobactergenus and species identification.
For dendrogram generation, the SARAMIS Premium software package was used. The dendrogram was based on whole spectra, including all signals passing the peak detection criteria of the Launchpad software. A binary mass list was
calculated with an error of 800 ppm, and a single-link clustering algorithm was applied.
RESULTS AND DISCUSSION
[image:4.585.45.537.83.596.2]In order to establish a standardized analytical protocol,
sam-ple preparation and mass spectrometric parameters that could
TABLE 1—
Continued
Peak
no.a Experimental avg mass
(Da)⫾800 ppm error
Present in superspectrum?b
Identification category
Possible RPc
identity and mass关m/z⫹H兴⫹
1 2 3 4 5 6 7 8
56
7,572.1
⫾
6.06
No
No
Yes
No
No
Yes
No
No
SIBMs
57
7,728.7
⫾
6.18
No
No
Yes
No
No
No
No
No
SIBMs
58
7,798.6
⫾
6.24
No
No
No
No
No
No
No
Yes
EIBMs
50S RP L31
关
7,797
兴
59
8,035.3
⫾
6.43
No
No
No
Yes
No
No
No
No
SIBMs
60
8,086.4
⫾
6.47
No
No
Yes
No
No
No
No
No
SIBMs
61
8,095.0
⫾
6.48
No
Yes
No
No
No
No
No
No
SIBMs
62
8,107.2
⫾
6.49
No
No
No
No
Yes
No
No
No
SIBMs
63
8,136.1
⫾
6.51
Yes
No
No
No
No
No
No
No
SIBMs
64
8,170.8
⫾
6.54
No
No
No
No
No
Yes
No
No
SIBMs
65
8,189.7
⫾
6.55
No
No
No
No
Yes
No
No
No
SIBMs
66
8,199.5
⫾
6.56
No
Yes
No
No
No
No
No
No
SIBMs
67
8284.7
⫾
6.63
No
No
No
No
No
No
No
Yes
EIBMs
68
8,374.2
⫾
6.70
No
No
No
No
No
No
No
Yes
EIBMs
30S RP S21
关
8,370 (-Met)
兴
69
8,508.1
⫾
6.81
No
No
No
No
Yes
No
No
No
SIBMs
70
8,569.8
⫾
6.86
No
No
Yes
No
No
No
No
No
SIBMs
71
8,701.9
⫾
6.96
No
No
Yes
Yes
No
Yes
Yes
No
GIBMs
72
8,718.3
⫾
6.97
No
Yes
No
No
No
No
No
No
SIBMs
73
8,860.7
⫾
7.09
Yes
Yes
No
Yes
Yes
No
Yes
No
GIBMs
30S RP S18
关
8,856 (-Met)
兴
74
8,896.9
⫾
7.12
No
No
No
No
No
No
No
Yes
EIBMs
75
8,972.6
⫾
7.18
No
No
No
Yes
No
No
No
No
SIBMs
76
9,012.8
⫾
7.21
No
No
No
No
No
No
No
Yes
EIBMs
50S RP L27
关
9,009 (-Met)
兴
77
9,194.0
⫾
7.36
No
No
Yes
No
No
No
No
No
SIBMs
78
9,234.2
⫾
7.39
No
No
Yes
Yes
Yes
No
Yes
No
GIBMs
30S RP S16
关
9,235
兴
79
9,242.0
⫾
7.39
No
Yes
No
No
No
No
No
No
SIBMs
80
9,260.6
⫾
7.41
Yes
No
No
No
No
Yes
No
No
SIBMs
81
9,482.8
⫾
7.59
No
No
No
No
No
No
No
Yes
EIBMs
82
9,533.6
⫾
7.63
No
No
No
No
Yes
No
No
No
SIBMs
83
9,576.6
⫾
7.66
No
No
Yes
No
No
No
No
No
SIBMs
30S RP S17
关
9,574 (-Met)
兴
84
9,687.7
⫾
7.75
No
No
No
No
No
No
No
Yes
EIBMs
85
10,098.5
⫾
8.08
No
No
No
No
No
No
No
Yes
EIBMs
30S RP S15
关
10,093 (-Met)
兴
86
10,289.1
⫾
8.23
No
No
No
No
No
No
No
Yes
EIBMs
30S RP S19
关
10,286 (-Met)
兴
87
10,622.9
⫾
8.50
No
No
Yes
No
No
Yes
Yes
No
GIBMs
88
10,715.7
⫾
8.57
No
No
Yes
No
No
No
No
No
SIBMs
50S RP L25
关
10,714
兴
89
10,842.1
⫾
8.67
No
Yes
No
Yes
Yes
Yes
Yes
No
GIBMs
90
11,198.5
⫾
8.96
No
Yes
Yes
No
No
No
No
No
SIBMs
50S RP L24
关
11,200 (-Met)
兴
91
11,231.3
⫾
8.99
Yes
No
Yes
Yes
No
Yes
Yes
No
GIBMs
50S RP L23
关
11,127
兴
92
11,353.1
⫾
9.08
No
No
Yes
No
No
No
No
No
SIBMs
93
11,382.1
⫾
9.11
Yes
Yes
No
Yes
Yes
Yes
Yes
No
GIBMs
94
11,479.8
⫾
9.18
No
No
Yes
No
No
No
No
No
SIBMs
30S RP S14
关
11,479 (-Met)
兴
95
11,770.2
⫾
9.42
No
No
Yes
No
No
No
No
No
SIBMs
30S RP S10
关
11,767
兴
96
11,980.8
⫾
9.58
No
No
Yes
No
No
No
No
No
SIBMs
97
12,082.2
⫾
9.67
No
No
Yes
No
No
No
No
No
SIBMs
98
12,795.1
⫾
10.24
No
No
No
No
No
Yes
No
No
SIBMs
99
13,104.7
⫾
10.48
No
No
Yes
No
No
No
No
No
SIBMs
100
15,453.6
⫾
12.36
No
No
Yes
No
No
No
No
No
SIBMs
aData show whether or not the indicated species was found in the superspectrum, as follows: 1,C. sakazakii; 2,C. malonaticus; 3,C. turicensis; 4,C. dublinensis; 5,
C. muytjensii; 6,C. genomospecies1; 7,Cronobactergenus, 8,Enterobacteriaceaefamily exclusion list.
bRP, ribosomal protein.
on May 16, 2020 by guest
http://jcm.asm.org/
affect the reproducibility and accuracy of data were first
eval-uated. The influence of growth conditions on the
MALDI-TOF MS patterns of different
Cronobacter
species was
ana-lyzed by subculturing strains on two different media.
Depending on the culture medium, some variations in the
pattern compositions and the measured relative intensities
were observed (Fig. 1). Results were most reproducible when
using bacterial strains grown on TSA agar for 48 h by applying
the direct smear method. These conditions also showed a
slightly better discrimination of species in a cluster analysis
than samples grown on blood agar (data not shown). However,
genus- and species-identifying biomarker masses were found to
be present in all strains analyzed, independent of growth time
and culture conditions.
In a first step, 54
Cronobacter
and 17 nontarget strains were
analyzed eight times each. The spectra displayed high overall
levels of similarity, but slightly different mass patterns for
dif-ferent
Cronobacter
species were detected (Fig. 2). The
SARA-MIS software was used to identify biomarker masses and to
assign them to the following three categories: (i) biomarker
masses that were present in most of the
Cronobacter
spectra
but also in other
Enterobacteriaceae
and thus had no
discrim-inatory power to identify the genus
Cronobacter
; these masses
with discriminatory power only on the family level were
des-ignated
Enterobacteriaceae
-identifying
biomarker
masses
(EIBMs); (ii) biomarker masses that were calculated as part of
a protein mass pattern for the identification of the genus
Cronobacter
; these masses were reproducibly detected in the
genus
Cronobacter
and were designated genus-identifying
bio-marker masses (GIBMs); (iii) biobio-marker masses that were
calculated as part of protein mass patterns for the
identifica-tion of
Cronobacter
species; these species identifying masses
were assigned accordingly as SIBMs. In total, 17 masses were
designated EIBMs. Fourteen biomarker masses were selected
as biomarkers to unambiguously discriminate
Cronobacter
spp.
from other genera (GIBMs), and 69 SIBMs were reproducibly
detected, resulting in specific protein mass patterns and a clear
discrimination of the six species. The designations of these
EIBMs, GIBMs, and SIBMs are shown in Table 1. Based on
the GIBMs and the SIBMs superspectra for
Cronobacter
sp.,
Cronobacter sakazakii
,
Cronobacter malonaticus
,
Cronobacter
muytjensii
,
Cronobacter dublinensis
,
Cronobacter turicensis
, and
Cronobacter
genomospecies 1 were defined. Additionally, from
sequence database information (ExPASy), several masses
could be expected for selected ribosomal proteins: the large
ribosomal protein (RP) L36 (4,365 Da) was detectable in all
Cronobacter
species and is known as a prominent marker of
Enterobacteriaceae
. The small RP S18 (8,856 Da) was
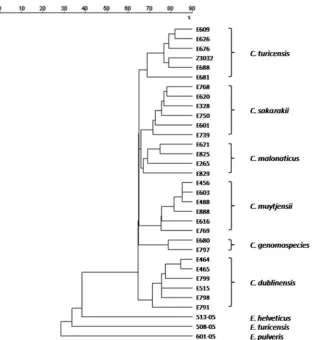
detect-FIG. 3. Cluster analysis of different target strains and three relevant nontarget strains (
E. turicensis
,
E. helveticus
, and
E. pulveris
), based on
whole spectra (2,000 to 15,000 Da). Isolates of
C. malonaticus
, which was first described as a subspecies of
C. sakazakii
, show a distinct subcluster
and confirm recent results reported by Iversen et al. (10).
on May 16, 2020 by guest
http://jcm.asm.org/
[image:5.585.138.453.65.405.2]able in four
Cronobacter
species and thus is a marker for
Cronobacter
. Finally, the large RP L32 observed in
C. turicensis
(6,303 Da) carried a mutation in 1 amino acid, resulting in a
measurable mass shift of 16 Da in comparison to
C. sakazakii
(6,319 Da) (Table 1).
In a next step, the defined superspectra were used to identify
36
Cronobacter
and 8 nontarget strains grown on TSA as well
as on blood agar in a blind test. For all strains the mass
spectrometry-based identification scheme yielded identical
re-sults as a PCR-based identification system (14). All these 44
strains were correctly identified as
Cronobacter
on the genus as
well as on the species level or as nontarget strains, respectively.
Moreover, a dendrogram was calculated using MALDI-TOF
MS whole spectra of
Cronobacter
strains, and three nontarget
type strains were identified by using the SARAMIS software
(Fig. 3). The topology obtained using MALDI-TOF MS
pro-filing strongly resembled the topologies of dendrograms
con-structed using a 16S rRNA gene phylogenetic tree, fluorescent
amplified fragment length polymorphism analysis, and
ri-botype dendrograms of
Cronobacter
(9).
Supporting recent results, and visible in the dendrogram, is
C. malonaticus
, which was first described as a subspecies of
C.
sakazakii
(9) and defined as a separate species by DNA:DNA
analysis (10) only later. It is in close relation to
C. sakazakii
but
in a distinct subcluster.
In summary, our study demonstrates that MALDI-TOF MS
is a reliable and powerful tool for the rapid, reliable, and
relatively inexpensive identification of
Cronobacter
strains to
the genus and species level. The major advantages of
MALDI-TOF MS-based bacterial identification compared to other
identification methods are the ease and speed of the procedure
and the possibility of automation and high-throughput analysis.
The costs of consumables are minimal, and the whole process
takes less than 5 min per sample.
REFERENCES
1.Alispahic, M., K. Hummel, D. Jandreski-Cvetkovic, K. No¨bauer, E.
Razzazi-Fazeli, M. Hess, and C. Hess.2010. Species specific identification and
dif-ferentiation of Arcobacter, Helicobacter and Campylobacter by full-spectral matrix-associated laser desorption/ionization time of flight spectrometry analysis. J. Med. Microbiol.59:295–301.
2.Barbuddhe, S. B., T. Maier, G. Schwarz, M. Kostrzewa, H. Hof, E. Domann,
T. Chakraborty, and T. Hain.2008. Rapid identification and typing of
Lis-teria species by matrix-assisted laser desorption ionization–time of flight mass spectrometry. Appl. Environ. Microbiol.74:5402–5407.
3.Carbonnelle, E., J. L. Beretti, S. Cottyn, G. Quesne, P. Berche, X. Nassif, and
A. Ferroni.2007. Rapid identification of Staphylococci isolated in clinical
microbiology laboratories by matrix-assisted laser desorption ionization– time of flight mass spectrometry. J. Clin. Microbiol.45:2156–2161.
4.Dieckmann, R., R. Helmuth, M. Erhard, and B. Malorny.2008. Rapid
classification and identification of salmonellae at the species and subspecies levels by whole-cell matrix-assisted laser desorption ionization–time of flight mass spectrometry. Appl. Environ. Microbiol.74:7767–7778.
5.Freiwald, A., and S. Sauer.2009. Phylogenetic classification and
identifica-tion of bacteria by mass spectrometry. Nat. Protoc.4:732–742.
6.Grosse-Herrenthey, A., T. Maier, F. Gessler, R. Schaumann, H. Bo¨hnel, M.
Kostrzewa, and M. Kru¨ger.2008. Challenging the problem of clostridial
identification with matrix-assisted laser desorption and ionization–time-of-flight mass spectrometry (MALDI-TOF MS). Anaerobe14:242–249.
7.Hazen, T. H., R. J. Martinez, Y. Chen, P. C. Lafon, N. M. Garrett, M. B.
Parsons, C. A. Bopp, M. C. Sullards, and P. A. Sobecky. 2009. Rapid
identification ofVibrio parahaemolyticusby whole-cell matrix-assisted laser desorption ionization–time of flight mass spectrometry. Appl. Environ. Mi-crobiol.75:6745–6756.
8.Hunter, C. J., M. Petrosyan, H. R. Ford, and N. V. Prasadarao.2008.
Enterobacter sakazakii: an emerging pathogen in infants and neonates. Surg. Infect.9:533–539.
9.Iversen, C., A. Lehner, N. Mullane, E. Bidlas, I. Cleenwerck, J. Marugg,
S. Fanning, R. Stephan, and H. Joosten.2007. The taxonomy of
Entero-bacter sakazakii: proposal of a new genus Cronobactergen. nov. and descriptions ofCronobacter sakazakiicomb. nov.Cronobacter sakazakii
subsp.sakazakii comb. nov.,Cronobacter sakazakii subsp. malonaticus
subsp. nov., Cronobacter turicensis sp. nov.,Cronobacter muytjensiisp. nov.,Cronobacter dublinensissp. nov. andCronobactergenomospecies 1. BMC Evol. Biol.7:64.
10.Iversen, C., N. Mullane, B. Mc Cardell, B. D. Tall, A. Lehner, S. Fanning, R.
Stephan, and H. Joosten.2008.Cronobacter gen. nov., a new genus to
accommodate the biogroups of Enterobacter sakazakii, and proposal of
Cronobacter sakazakii gen. nov. comb. nov.,C. malonaticussp. nov., C. turicensissp. nov.,C. muytjensiisp. nov.,C. dublinensissp. nov.,Cronobacter
genomospecies 1, and of three subspecies, C. dublinensissp. nov. subsp.
dublinensissubsp. nov.,C. dublinensis sp. nov. subsp.lausannensissubsp. nov., andC. dublinensissp. nov. subsp.lactaridisubsp. nov. Int. J. Syst. Evol. Microbiol.58:1442–1447.
11.Kuhnert, P., B. M. Korczak, R. Stephan, H. Joosten, and C. Iversen.2009.
Phylogeny and whole genome DNA-DNA similarity ofCronobacter( Entero-bacter sakazakii) and related species by multilocus sequence analysis (MLSA). Int. J. Food Microbiol.136:152–158.
12.Lehner, A., and R. Stephan.2004. Microbiological, epidemiological and food
safety aspects ofEnterobacter sakazakii. J. Food Prot.67:2850–2857.
13.Sauer, S., and M. Kliem.2010. Mass spectrometry tools for the classification
and identification of bacteria. Nat. Rev. Microbiol.8:74–82.
14.Stoop, B., A. Lehner, C. Iversen, S. Fanning, and R. Stephan.2009.
Devel-opment and evaluation ofrpoBbased PCR systems to differentiate the six proposed species within the genus Cronobacter. Int. J. Food Microbiol.
136:165–168.