Copyright01974 AmericanSocietyforMicriobiology Printed in U.SA.
Inhibition
of Protein
Synthesis
in Cell-Free
Systems
from
Interferon-Treated,
Infected Cells: Further Characterization
and Effect of
Formylmethionyl-tRNAF
IAN M. KERR, R. M. FRIEDMAN, R. E. BROWN, L. A. BALL, AND J. C. BROWN
National InstituteforMedicalResearch,MillHill, London NW71AA, andDepartment of Microbiology, University of Virginia, Charlottesville, Virginia, 22901
Received forpublication3August 1973
The translation of
encephalomyocarditis
virus (EMC) RNA is markedlyinhibited in cell-free systems from
interferon-treated,
vaccinia virus-infectedL-cells (10, 11). The polypeptide products synthesized inresponse toEMC RNA
incell-free systemsfromthese and
untreated
infected cells have beenanalyzed
by electrophoresison
polyacrylamide
gels. Qualitatively,
thesameEMC-specific
polypeptides were
synthesized throughout.
Inexperiments using
preincubatedmicrosomes from normal Krebs cells to assay cell sap from L-cells which had
been
exposed
to interferonprior
toinfection,
only
the amount of theEMC-specific
polypeptide
products
was reduced. This resultsuggeststhat there isaninhibitionvery
early
intranslation ininterferon-treated,
infectedcells. Initiationseems a
priori
the more attractive site for thisinhibition,
butan effectshortly
after initiation cannot be excluded. With unfractionated cell-freesystemsfrom
interferon-treated infected
L-cells, however,
thereappeared
tobe anadditionalminor inhibitory effect on
polypeptide
chain elongation, in that theEMC-specific polypeptides
synthesized
showednotonly
areductionin amountbut alsoabias towardslower molecular
weight.
Theformylated
methionyl initiator tRNA(Fmet-tRNAF) was usedasafurther
probe
intotheapparent effect onintiation.With thisreagent wehaveconfirmed that there isonemajor initiation site for the
translation of EMC RNA in these cell-freesystems.In
addition,
theresults haveshown that
EMC-specific
polypeptide
chains initiated with Fmet escape themajor
interferon-mediated inhibition
at orshortly
after initiation.In the
interferon-treated,
vacciniavirus-infectedL-cell, vaccinia virus mRNA is
synthe-sized but not translated (16,
21).
Recently,
wehave shown that the translation of
enceph-alomyocarditis virus
(EMC)
RNA is inhibitedincell-freesystemsfromsuch cells (10, 11). It is
nowwell established fromour ownwork(6, 11,
18) and that of others (1, 2, 8, 23, 27) that the
EMC RNAgenomeistranslated withfidelityin
cell-free systems from mouse L- and Krebs 2
ascitestumourcells. The
products
are aseriesofhigh-molecular-weight
EMC-specific
polypep-tides. The majority of these ariseas aresult of
prematuretermination atpreferred sites during
the translation of this large mRNA (molecular
weight 2.7 x 10') (2, 18). Here we have made
useof ourknowledge ofthesepolypeptide prod-ucts to analyze the nature of the inhibitory
event(s) in the translation of EMC RNA in
cell-free systems from interferon-treated,
vac-cinia virus-infected L-cells. In addition, we have used the formylated methionyl initiator 9
tRNA (Fmet-tRNAF) (14, 28) to confirm (23,
27)
that initiation oftranslation ofEMC RNAinthe cell-free system is at aunique site as in
the intact cell. With this established for the
cell-freesystemswith whichweareworking,we
have used the Fmet-tRNAF as a probe in our
investigation ofthe translationof EMC RNA in
thesesystems.Theresults ofthe electrophoretic
analysis
oftheproducts suggestthat the majorinhibition in the translation of EMC RNA in
cell-free systems from interferon-treated,
in-fected cells isat orshortlyafter initiation, with
a minor secondary effect on chain elongation.
With EMC RNAas messenger, however,
poly-peptidechains initiated with Fmet from
Fmet-tRNAF appear to escape the major inhibition
occurring early intranslation.
MATERIALS AND METHODS
Materials. Chemicals for use in the cell-free system, for the isolation of cell fractions and viral RNA and forthepreparation of tryptic digests were as
on November 10, 2019 by guest
http://jvi.asm.org/
KERR ET AL.
described previously (6, 19). Allradiochemicals were supplied by the Radiochemical Centre, Amersham, Buckinghamshire, England. The calcium leucovorin was from Lederle Laboratories Division, American Cyanamid Company, Pearle River, N.Y. Some of the EMC virus was obtained from Products for Research, G.D. Searle and Co. Ltd., High Wycombe, Bucking-hamshire,England.
Methods. The Krebs 2 mouse ascites tumour cells and the preparation of cytoplasmic extracts from them (6, 19), the preparation andpurification of EMC virus(20) and the extraction of EMC RNA (19), the labeling of polypeptide products in thecell-free sys-tem, theperformate oxidation and tryptic digestion of theseproducts, and the analysis of the tryptic digests byelectrophoresis and chromatography on thin-layer silica gel plates (6) have already been described. Qualitative and quantitative analysis of the polypep-tide products by electrophoresis on 5 or7.5% polya-crylamide gels after treatment with sodium dodecyl sulphate (SDS) was as described previously (18). Furtherdigestion of the tryptic digests with Pronase wasovernight at 37 Cin0.2 Mammonium bicarbon-ate, 10mM CaCl2.
Preparation of L-cellextracts. Nonpreincubated post-mitochondrial supernatant-S10-fractions were prepared as before (11) from interferon-treated (50 to 100 reference units/ml) and control, vaccinia virus-infected L-cells. Centrifugationofthis material at100,000xg for90 minyielded the supernatant cell sap fractions used in the mixed system assays with Krebs cell microsomes.
Amino acid incorporation assays. For the Krebs andL-cell systems the assaysof aminoacid incorpora-tion inresponsetosaturatingamountsofEMC RNA (20to 40Mg/ml)werecarriedout aspreviously (6, 11). Assays in the mixed system in which L-cell sapwas assayed with preincubated microsomes from normal Krebs cellswere incubated at 30C under the same conditionsasfor theL-cell system(11) butusing200
jtgof Krebsmicrosomes and 10 ulitersof L-cellsap (approximately 150jtgofprotein) per 50-Mliter assay. Assays involving the use of35S-Fmet- or met-tRNAF and/orAUG(U)n,wereroutinelycarriedout inafinal volume of25Mliters.Tosaturatethese systems2pmol (35,000 to 250,000 counts/min per pmol for methio-nineat aspecific activityof 20to150Ci/mmolat80% efficiency of counting) of 36S-met- or Fmet-tRNAF were added per
25-Aliter
sample. Unlabeled methio-nine(50AM)
wasincludedinthe amino acid mixture used in all such assays. AUG(U)ri was added at a saturating concentrationof 80Mg/ml(1.6OD260units/ ml) and was assayed at the same magnesium ion concentration (4.35to4.85mM) asforEMCRNAintheL-cell system. It was prepared according to the methodofSundararajan and Thach(30).
Interferon preparation andassay. Mouse L-cell interferon (> 106 reference units per milligram of protein) was purified bythe method ofPaucker (25) andassayedaspreviously described (11).
Charging and formylation of Krebs
tRNANet.
The enzyme fraction used for the charging and formylation ofthe tRNA was prepared from Esche-richia coli (MRE 600) accordingtoMuenchandBerg (22).Thedialyzed andconcentrated DEAE-cellulose
fraction in 50% glycerol was employed throughout. Theactivity of the enzymes was monitored by using pure E. coli
tRNAFM,t
(thegenerous gift of Kellmers and Novelli, Oak Ridge National Laboratory). 100% charging and >80% formylation of this tRNA was routinely observed. E. coli Fmet-tRNAF has been reported (14) to be active at a low level in the rabbit reticulocyte system, but with the Krebs and L-cell systems employed here, although there was some incorporation of label from the E. coli36S-Fmet-tRNA,
inresponse to EMC RNA, analysis of theFriiet trypticpeptides showed it to be nonspecific. Accord-ingly, the E. coli enzymes were used to charge and formylate Krebs celltRNAFMel.The Krebs cell tRNA was obtained by phenol extraction of whole cells followed by salt fractionation (31) and chromatogra-phy onDEAE-cellulose (32). Initially, it was fraction-ated on BD-cellulose to yield the tRNAFMet and tRNAFM"t species (27). Having confirmed (12) that the E.colienzymepreparation was inactive with the mammalian noninitiatingtRNAlNIet
species, how-ever,unfractionated tRNAwasused in all of the work described here. Charging and formylation were car-ried outsimultaneouslyin areaction mixture contain-ingthe followingcomponents: 50 mM HEPES buffer, pH 7.5; 50 mM KCI; 5 mM Mg acetate; 2 mM dithiothreitol, 4 mM ATP, 1 mM CTP; 10 MAM Ca leucovorin;200Mg
ofKrebs cell tRNA per ml (approx-imately 7,000 pmol of total tRNAequivalent to 140 pmoloftRNAFMet at 2%)and 1 MM1-35S-methionine (40 to 150 Ci/mmol). Incubation was for 15 min at 30 C. For the preparation of nonformylated met-tRNAF the formyl donor Ca leucovorinwas omitted. TheCa leucovorinwasmade up and storedin0.25N HCI. On additiontothe cell-freesystem, the change in pH converts it to theunstable active form (26). The amount of E. coli enzyme required for optimum charging and formylation varied considerably from preparation topreparation and with storage andwas determinedon a smallscalepriortopreparativeuse. The charged and formylated tRNA was extracted withphenol before purification at pH4.5 on asmall DEAE-cellulose column (32). Afterethanol precipita-tion the RNAwasdissolvedin waterat2x 105counts perminperMlliter.
Toassaytheextentofformylation a sample of the 3`S-Fmet-tRNAF was precipitated with 5% trichloroacetic acidat 0C, washed with cold acid on a cellulose nitrate filter (13-mm diameter, Sartorius Membranfilter GMBH, Gottingen, Ger-many),rinsed withethanol,and theaminoacyl-tRNA wasdechargedby incubation of the filter for 30 minat37Cin 0.4 NNH40H.The eluted Fmet andmetwere fractionated by electrophoresis for 1 h at 400 V on
Whatman 3 MM paperinpyridineacetatebuffer, pH 6.5. The amounts of Fmet and metweredetermined by usingastripscannerorbycuttingthedriedpaper into0.5-cmstripsandcountingthese inascintillation counter. Charging was to a level of2 to 2.4pmol of Met-tRNA, per 100 pmol of total tRNA. Only70 to
80%formylationoftheKrebsmet-tRNAFwasusually
obtained. However, the residual (20to 30%) nonfor-mylated methionine in these preparations was not
incorporatedintoacid-insolubleproductsinresponse toEMCRNA.AnynonformylatedN-terminal methi-onine is rapidly cleaved (14, 15) from the nascent
10
J. VIROL.on November 10, 2019 by guest
http://jvi.asm.org/
polypeptide chain. Inaddition,althoughwhen work-ing with completely nonformylated initiator met-tRNAF there is a significantincorporationof methio-nine intointernal sites in thepolypeptidechain(7,13, and our own unpublished results), no significant incorporation of this type occurred from the nonfor-mylated fraction (20 to 30%) of the Fmet-tRNAF preparation used here. Forexample, complete diges-tion ofthe EMC RNA-stimulatedpolypeptide prod-ucts with trypsin and Pronase consistently yielded >95% Fmetand <5% methionine. It couldbe, there-fore, that the 20to30%ofapparently nonformylatable met-tRNAF wasartefactual orrepresented damaged nonfunctional RNA.
RESULTS
Throughout
theexperiments
tobedescribed,
the L-cell extracts
employed
in the cell-freesystems were neither
preincubated
norsub-jected to
Sephadex
chromatography.
With suchnonpreincubated
material,
amino acidincorpo-ration inresponse toEMC RNA inthe cell-free
system is not reduced
by
prior infection ofthecells and interferon treatment alone has
only
asmall inhibitory effect (11). A clear-cut 50 to
90% inhibition of the translation of the added
EMC RNA isseen,
however,
insystemsemploy-ingmaterial ofthistypefromL-cellsexposedto
both interferon treatmentand infection
(10,
11,I. M. Kerr et
al.,
Advances in theBiosciences,
vol. 11, in press). It is with the further
charac-terization ofthis enhanced inhibitionof
transla-tion triggered
by
infection that we will becon-cerned here.
Characterization
of thepolypeptide
prod-uctssynthesized in responsetoEMC RNAin
cell-freesystemsfrom
interferon-treated,
in-fected cells. An analysis by
electrophoresis
onSDS
polyacrylamide
gels of the EMC-specificpolypeptide
products
synthesizedinresponse toEMC RNA in cell-free systems from
vaccinia-virus-infected and
interferon-treated,
vaccinia-infected cells is shown in Fig. 1. A qualitative
autoradiographic analysis
oftheproducts
syn-thesized after incubation for 30, 60, 120, and
240
min in the presence ofEMC RNA yielded the
results shown inFig. 1A,
whereas
aquantitativeanalysis
ofthedistribution
ofthe products at120 min is shown in Fig. 1B. It
should
beemphasized that in this experiment
pretreat-ment ofthe cells with interferon prior to
infec-tion resulted in a 70% inhibition of
incorpora-tion in response to EMC RNA in the cell-free
system (Fig. 1C). To compensate for this the
autoradiographs of gels 6 to 9 (Fig. 1A)
(inter-ferontreated) were exposedfor longerthanthe
corresponding controls (gels 1 to 4, Fig. 1A).
Similarly,
for ease of comparison, the data inFig. 1B areexpressedaspercentageofrecovered
radioactivity rather than in terms ofabsolute
counts. From thisdata it canbc seen that the sameseries
(A
toG, Fig. 1A)
ofhigh-molecular-weight EMC-specific polypeptides
wassynthe-sized in both systems. These are
apparently
identical in molecular weight to those
synthe-sized incell-free systems fromuninfected Krebs
and L-cells. The nomenclature A to G is that
which was used to
identify
theEMC-specific
polypeptides
inourprevious work(Fig.3,refer-ence
18).
Inaddition,
theresults for the earlier 30-min(gels
1 and6)
and60-min(gels
2 and7)
time
points
indicate that there is nostriking
differencein the timeat which the
polypeptide
chains first reachasize of130,000in molecular
weight,
(polypeptide G), for example. This inturn suggests that there is no gross overall
reduction in the rate of polypeptide chain
elongation
in the inhibited systems.Neverthe-less,
there does appearto be aslightinhibitory
effectonchaingrowthinthattheamountof the
major
polypeptideGaccumulatingatlater timepoints (gels
3 and4)
is significantly reduced(gels
8 and9).
This is shownquantitatively forthe120-mintimepointinFig. 1B, inwhich the
bias in the distribution of the
polypeptide
products towards lower molecular weight is
obvious in theinhibited system..Thenature of
this
relatively
smalleffect isuncertain, butitisclearly insufficient to accountfor the 70%
inhi-bition of total incorporation (Fig. 1C).
Accord-ingly,
the results show that themajorinhibitionmustoccurpriortotheproducts reachingasize
detectableonthesegels, i.e., mostprobably at,
orpossibly shortly after, initiation.
Characterization of the
polypeptide
prod-uctssynthesizedinresponsetoEMC RNA in
mixed cell-free systems. In general, on
frac-tionation ofthe L-cell extracts, the amount of
the interferon-mediated inhibitory effect
re-coveredwith the cellsap (S100) andmicrosome
fractions has proved variable. It may be that
this variability iscorrelated, forexample, with
the degree ofpolysome
disaggregation
orribo-some dissociation in the system. The results
obtained in preliminary assay of the crude
microsome fractionwith normal Krebs cellsap
were qualitatively similar to those for the
un-fractionated system (Fig. 1). It should be
em-phasized that thismaymerely reflectthe
crude-ness of the fractionation procedure. Different
resultswereobtained, however, onassay of cell
sap from
interferon-treated,
vacciniavirus-infected cells with preincubated microsomes
from normal Krebs cells. Withthissystem only
the majorinhibition at or shortly after initiation
was observed, with no secondary effect on
elongation. This is shown in Fig. 2. In this
experiment interferontreatment prior to
on November 10, 2019 by guest
http://jvi.asm.org/
KERR ET AL.
[image:4.495.119.400.70.449.2]Cf .P ... .11
A
-
II2
74C3
7 wS
- e.
000 34f
1
2
3
4
B
6
7
8
9
RFC)
r
.6k
ap..
-i
&- --.:> ..
i t.
;.
s , *0F
.; s ¢&.Z.i
v
},} t^'.S.F
*5,*As'
.,.._.
Atu; :: :: 4 Lj
jP _ <
FIG. 1. Characterization ofthe polypeptideproductssynthesized in response toEMC RNA with time in cell-free systems from interferon-treated, vaccinia virus-infected cells. Unfractionated postmitochondrial supernatant (S10) fractions were used throughout. Incubation in the cell-free system in the presence of 35S-methionine(100MCi/ml,100Ci/mmol)andEMC RNA (40
,g/ml)
was asalreadydescribed(Materialsand Methods). A,Analysis ofthepolypeptide products by electrophoresisinSDS-polyacrylamide gels. Gels6to9 werewithinterferon-treated,vaccinia-infected cell extracts; Gels1 to 4 werewithcorrespondingextractsfrom infectedcells withoutinterferonpretreatment.Afterincubationofthecell-freesystemsfor30min(gels1and6), 60 min(gels2and7),120min(gels3and8)and240min(gels4and9),50ulitersamplesweretreatedwithSDS andsubjectedtoelectrophoresison SDS-polyacrylamide gels. Thefigureshowsautoradiographs ofthedried gelsand,as anindexof molecular weight,aphotographofastainedgelonwhich thepolypeptides of purified reoviruswereelectrophoresedinparallel.Thefigurestotherightrepresent the molecularweightinthousandsof themajor reoviruspolypeptides Xl, A2, p2,and q3(29). Theautoradiogr:aphsofgels 1 to4 wereexposed for4 days; those for gels 6 to 9for20days. B, Quantitative analysis ofthe molecularweight distribution ofthe products at 120 min. Gels corresponding to3 and 8 (Fig. 1A) werefixed, sliced, prepared for radioactive counting, and countedaspreviously (18). Theabscissa shows the distance migrated fromthe top ofthegel(1-mm slices)with theorigintotheleft; themajor peak(fractions8to10)correspondstoG in thegels3and8. Theordinategivestheradioactivityincorporatedintopolypeptiderecoveredineachfractionas apercentageof
the totalrecoveredfrom thecompletegel.Symbols:0, nointerferonpretreatment ofthecells; 0, interferon
pretreated. The total recoveries from the gels (100% values) were 53,000 counts/min without and 15,000 counts/min with interferon pretreatment. C, Direct quantitative assay of83S-methionine incorporation in responsetoEMC RNA with timeinthe cell-freesystemsused in IA andIB above.Afterincubationforthe indicated time
5-pgliter
sampleswereassayed forincorporation of radioactivity intoproteinin the usual way. Symbols:*,vaccinia-infected cellextracts; 0, interferon-treated, vaccinia-infected cellextracts.12 J. VIROL.
2f-..
z
4". fl.-..;
on November 10, 2019 by guest
http://jvi.asm.org/
INTERFERON
A
..
.155
-140
-
-72
am
-34
1
2
3
4
5
6
REO
C
I;
-~~ ~ ~ ~ ~ ~ R.
2*Z i 8 i
3C 35 4
FRArT .ON No
16, 0
12/
PM/P
I, f>.
/
./"
30 60
TIME(m:ns)
i20
FIG. 2. Characterizationofthepolypeptide products synthesizedinresponseto EMC RNA with timeinthe
cell-free systems in the presence of cell sap fractions from vaccinia virus-infected and interferon-treated vaccinia virus-infectedcells. The cell sapfractionswereassayedwithpreincubated microsomesfrom Krebs
ascites tumourcells. Incubation in the cell-free systemin the presence of3"S-methionine (100pCi/ml, 100
Ci/mmol) and EMC RNA (40ug/ml) was asalreadydescribed.A,Analysis ofthepolypeptide products by electrophoresis inSDS-polyacrylamidegels. Gels 1 to 3employedcellsapfrom vacciniavirus-infectedcells.
Gels 4to 6were with cellsapfrom interferon-treated, vacciniavirus-infected cells.Afterincubationofthe
cell-freesystemsfor30min (gels1 and4),60min (gels2 and5),and 120min (gels3 and6),40-plitersamples weretreatedwith SDS andsubjectedtoelectrophoresisonSDS-polyacrylamidegels inparallelwith reovirus polypeptidesasdescribedinFig.1.Thefigurestotheright representthe molecularweightsinthousandsofthe
major reoviruspolypeptides (Fig. 1).Theautoradiographswereexposed for4days. B, Quantitative analysis of the molecularweightdistributionoftheproductsat 120min.Thiswascarried out withgels correspondingto 3
and6 in A aboveasdescribedinFig. lB. The dataareexpressedasinFig. lB.Symbols: 0,Nointerferon;0,
interferon pretreated. The total recoveriesfrom the gels (100%/o values) were 75,000counts/minwithout and
21,000 counts/minwithinterferonpretreatment.C,Adirectquantitativeassayof"'S-methionineincorporation inresponsetoEMCRNA withtime in thecell-freesystems usedinA and BabovewascarriedoutasinFig.1C. Symbols: 0, vaccinia virus-infected cellextracts; 0, interferon treated, vaccinia virus-infected cellextracts.
tion reduced the level of translation of EMC RNA to 50% of the control value (Fig. 2C) without apparently affecting the nature ofthe EMC-specific polypeptides formed (Fig. 2A and B). In the inhibited system identical products
were synthesized (Fig. 2B) at apparently the
same rate ofpolypeptide chain growth (gels 1 and 4, Fig. 2A) but in reduced amounts (Fig. 2C).
Taken together these results are consistent
G%*.
F
%B--A-O
B
VOL. 13, 1974
tomftw
1 .:
i
IW
A&gk;
MPF,
on November 10, 2019 by guest
http://jvi.asm.org/
[image:5.495.108.393.61.444.2]KERR ET AL.
with a major block at orshortlyafter
initiation,
with a minor inhibition of polypeptide chain
elongation, the latter being seen only in the
unfractionated system.
IncorporationofFmet fromFmet-tRNAFin
response to EMC RNA. The initiation of
polypeptide chain synthesis in eukaryotes is
thought to occur exclusively with methionine
fromoneof themethioninetRNAs: the initiator
tRNAF\et.
Normally, this N-terminal methio-nine is cleaved from the nascent polypeptidechain when it isstillquite short (14, 15). Thus,
35S-methionine from labeled initiator
35S-met-tRNAF is removed from the N-terminus of the
nascentpolypeptide chains almost immediately
after it is incorporated and is not normally
detected. If, however, the methionine on the
initiator tRNA is formylated, the Fmet once
incorporatedN-terminally onthe nascent chain
is not cleaved and is readily detected (14, 27).
Here, therefore, wehave used Krebs cell
initia-tor
tRNA,NIet
loaded and formylated with en-zymesfromE. coli as a probe in our studiesoninitiation.The E. colienzymesdonotrecognize
the noninitiator mammalian methionine tRNA species (tRNAlet )responsiblefor thedonation
ofmethionineintointernal sites(12,and
Mate-rials and Methods). Moreover, the presence of
the formyl group both prevents the proteolytic
cleavageofthe N-terminal (formyl) methionine
and removes any possibility ofincorporation of
the methionine into internal sites in the
poly-peptide chain. Incubation of
35S-methionine-labeled Fmet-tRNAF with EMC RNA in the
Krebs cell-freesystemprovidesaproductwhich
on digestion with trypsin yields a single major
EMC-specific Fmet-peptide (23, 27). Further
analysis of this peptide
by
Smith has shown ittobe identical (with the additionof the
formyl
group) totheunique methionineinitiation
pep-tide formedinthe translationoftheentireEMC
RNA genome in the intact EMC-infected cell
(27). Thus, initiation ofprotein synthesiswith
Fmet-tRNAF in response to EMC RNA in the
Krebs cell-freesystem occursatthesamesiteas
does initiationofEMCpolypeptide synthesisin the intact cell.
N-terminal incorporation of 35S-methionine
labeled Fmet from
Fmet-tRNAF,
as a measureofinitiation, and a mixture of free "C-amino
acidsor
3H-leucine,
as anindex of totalsynthe-sis, werecompared forthesystems under
study
here. With systems using material from
inter-feron-treated, infected cells there wasthe usual
inhibition of "C-amino acid or 3H-leucine in-corporation in response to EMC
RNA,
but no such inhibitionintheincorporationof35S-Fmetwas observed(Fig.3). Onoccasion,with
unfrac-tionated systems showing a profound effect
some inhibition (<50%) of Fmet incorporation
did occur, but this was always much less than
thatobservedwiththefree amino acids (> 80%).
Accepting that EMC-specific polypeptide
chains initiated with Fmet-tRNAF do escape the major inhibition one might, a priori, have expected to see a decrease in the inhibition of
3H-leucine incorporation in those systems
re-ceiving Fmet-tRNA as in the experiments
shown in Fig. 3B. However, the added Fmet
tRNAF is in competition with endogenous met
tRNA,
andonly 0.01 to 0.05 pmol of Fmet areincorporated per 0.34 pmol of added EMC
RNA. No accuratefigureforthe total number of
EMC-specific polypeptide chains initiated in
response totheadded RNA is yetavailable(not all EMC RNAmoleculesneed be active inthis
respect), but it is clearly possible that Fmet
from the added Fmet-tRNAF is only used in a small fraction of the initiation events (our
calculations suggest a figure of no more than
15%). Acceptingthis, no significant decrease in the inhibition of3H-leucineincorporation would be expected in the presence ofFmet-tRNAF.
There are a number of possible explanations
why incorporation of Fmet from
Fmet-tRNAF
might escape inhibition. It could be, for
exam-ple, that the major inhibition operates at the
level of charging or breakdown of the initiator
tRNAF
et. Thus, by adding 35S-Fmet-tRNAFone mightalready be beyond the site of inhibi-tion in the sequence of events involved in initiation. Indeed, we have some preliminary
evidence for a reduced ability to charge the methionine tRNAs in these systems, but on fractionation this does not appear to be respon-sible for the major inhibition observed (L. A. Ball, unpublished data). Nevertheless a re-duced charging activity could result in a re-duced pool of endogenous met-tRNAF in the cell-free system. This, in turn, would result in an increase in the effective specific activity of the added 35S-Fmet-tRNAF in the inhibited system and a spuriously increased level of incorporation. The results obtained with
AUG(U)yg
as message, however, argue againstany such explanation.
Incorporation from
Fmet-tRNAF
andmet-tRNAF in response to AUG(U)n-. When
as-sayed with
AUG(U)r,
as mRNA, incorporationof label from initiator tRNAF "let was inhibited under conditions where incorporation of Fmet in response to EMC RNA escaped inhibition
(Table 1). This was true with
AUG(U)3n,
asmessage both for the
formylated
Fmet-tRNAFand thenonformylated
met-tRNAF
species,
al-though in someexperimentsthe inhibition with
14 J. VIROL.
on November 10, 2019 by guest
http://jvi.asm.org/
INTERFERON MEDIATED INHIBITIONOFTRANSLATION
A
14C
7-Cpu
35s
5-n. 10
4 S
3-
~~~~~~~~6-2-
4--2 4~~~~~~~~~
0 10 20 30 4050 6 0 1020 3040 S0 6
min. min.
B
105
5
C~~~~~~PM.
~
~
~
~
CM
3
-5e
51 3SP
20
40
60
120
TIME
(mins).
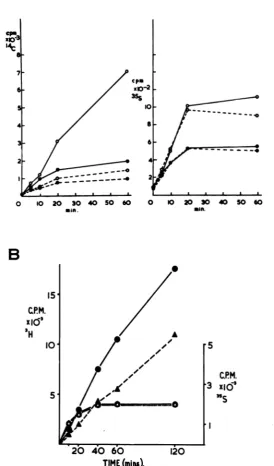
FIG. 3. Incorporation of "S-Fmet from 'S-Fmet-tRNAF in the cell-free system: absence of inhibition in extractsfrominterferon-treated, vacciniavirus-infectedcells. A,Unfractionated extracts. The graph on the left shows theincorporationof a mixture of14C-aminoacids, that on the right the incorporation in parallel assays of
35S-Fmet from 85S-Fmet tRNAF with time at 30 C in the cell-free system. Interferon-treated, vaccinia
virus-infected cellextractsplus O--Oandminus 0--* EMC RNA; vaccinia virus-infected cell extracts plusO-Oand minus0-* EMC RNA. Thesymbols are the same for both graphs. B, Assays of L-cell sap withpreincubatedmicrosomesfrom normal Krebs cells. Here the incorporation in response to EMC RNA of free 3H-leucine andof3'S-Fmetfrom3"S-Fmet-tRNAFincluded in the sameassay mix was followed with time at 30 C. 3H-leucine incorporation with vaccinia virus-infected cell extracts (0) and interferon-treated, vaccinia virus-infected cell extracts (A). 3"S-Fmet incorporation with vaccinia virus-infected cell extracts (0) and with interferon-treated, vaccinia virus-infected cell extracts (A). The assays were carried out as described in Materials and Methods with 0.25
,gCi
of"4C-amino
acid mix (50 mCi/mAtom of carbon)or55gCi
of3H-leucine (52 Ci/mmol) plus 35S-FmettRNAF equivalent to 3 x 101 to 4 x10&counts per min (4 pmol at 45 to 50 Ci/mmol). VOL.13, 1974on November 10, 2019 by guest
http://jvi.asm.org/
[image:7.495.123.398.82.549.2]TABLE 1. Incorporation of 35S-methionine from the initiator tRNAFMe' species in response to AUG(LT)n in cell-free systems frominterferon-treated, infected cells"
Incorporation in response toAUG(U)a-with Incorporation in response to EMC RNA with Cell-free systems from 35S-met-tRNAF
35S-Fmet-tRNAF
35S-Fmet-tRNAF
3H-phenylalanine(counts/min) (counts/min) (counts/min) (counts/min) Vacciniavirus-infected
cells ... 15,600 6,500 2,500 11,500
Interferon-treated,vaccinia
virus-infectedcells ... 4,400 (28%)b 2,200 (30%)b 2,000(80%)b 2,050(18%)b
aUnfractionated postmitochondrial supernatant (S10) fractions were used throughout. Met-tRNA and Fmet-tRNA were from the same preparation and had essentially the same specific activity. Two
Atliters
of one or other of the35S-met-labeled initiatortRNAFMet species (4 x 105counts/min,2.2 pmol) wereadded to each assay (25 ,liters). All assays were carried out in duplicate under the conditions already described (Materials and Methods). Incorporation from the abnormal Fmet-tRNA was apparently less efficient than from the normal met-tRNAF in accord with the results of Brown and Smith (3).b%ofincorporation in the vaccinia-infected system.
the formylated species was marginally less.
(With
AUG[U]n
as message, nonformylatedmet tRNAF can be used since the N-terminal
methionine is not cleaved from the relatively
shortnascent
met(phe)n oligopeptides
[3].)Initi-ation with saturating amounts of
AUG(U)n-
isover morerapidly (5to 10 min) inthese systems
than with EMC RNA (20 to 40 min, Fig. 3).
However, control experiments involving the
additionofthesemessengers progressivelywith
timehave shownnodifferenceinthestabilityof
functional Fmet and met-tRNAF species in
thesesystems. Itseemsunlikely,therefore, that
the apparent side-stepping by Fmet of the
inhibitory effect with EMCRNA (Fig. 3) simply
reflectsareductioninthe sizeofthe endogenous
met-tRNAF pool (Table 1), or anydifferencein
the relative stabilitiesofthenormal endogenous
(or added) initiator met-tRNAF as compared
with the formylated
Fmet-tRNAF
species.Characterization of the Fmet-labeled
poly-peptide products synthesized in response to
EMC RNAincell-free systems from
interfer-on-treated,
infected cells. Anobviouspossiblealternative explanation for the escape of the
Fmet incorporation from the major inhibition
would be provided if, in these particular sys-tems, initiation with Fmet occurred at an
ab-normal site on the EMC RNA genome. The
Fmet-labeled
polypeptide products
synthesized
in response to EMC RNA were, therefore,
analyzed
by electrophoresis
on SDSpolyacryl-amide gels (Fig. 4). In
addition,
themajor
EMC-specific Fmet-labeled initiation
peptide
released ontryptic digestionofthe
polypeptide
products was
compared
(Fig.
5)
with thatformed in responsetoEMC RNA in theKrebs
cell-free system. This latter has been shown
by
Smithtobeidentical
(plus
theformyl
group)
tothatnormally involvedinthe initiationofEMC
polypeptide
synthesis inthe intactinfected cell(27). Fromacomparisonofthe
autoradiographs
shown in Fig. 4 with those in Fig. 1 and 2and
previously (18), it is clear that
35S-Fmet
fromFmet-tRNAF is
incorporated
predominantly
into the same series of
high-molecular-weight
EMC-specific
polypeptides
as is freemethio-nine. Double labeling experiments with
35S-Fmet-tRNAF
and 3H-leucine have confirmedthis and similar results have been
reported by
Oberg and Shatkin for the Krebs cell-free
sys-tem (23). The
relatively
greater amounts ofradioactivity in the lower molecular
weight
material with 35S-Fmetaslabel(compareFig.4
with Fig. 1 and2) isconsistent with
incorpora-tion ofFmet
exclusively
at theN-terminus,
forover the molecular
weight
range of25,000
to130,000 observed here, the ratio ofN-terminal
Fmet per molecule to
randomly
distributedmethionine or leucine would be expectedto
de-crease
by
afactor of5.Accuratequantitation
inthese systems, whichseemto show less
prema-ture termination of translation of EMC RNA
thanisobservedinthe Krebssystem, is
compli-catedby thefact thatsome
cleavage
ofnascenthigh-molecular-weight
EMCprecursorpolypep-tide may beoccurring
(R.
M. Esteban andI. M.Kerr, manuscript in
preparation).
Thereis,
inaddition, a significant (e.g.,
Fig.
3A)
back-ground of
endogenous
incorporation
in thesenonpreincubated
systems.Nevertheless,
oncomplete digestion ofthese Fmet-labeled
prod-uctswithPronase, >95% of thelabel
co-electro-phoresed with Fmetand<5% withmethionine,
confirming thatat least95%ofthelabel
incor-porated isN-terminal. On
tryptic
digestion
thesameEMC-specificFmet initiation
peptide
(A,
Fig. 5A) wasobtained with all of thesesystems
16 KERR ET AL. J. VIROL.
on November 10, 2019 by guest
http://jvi.asm.org/
A
G-
1"
l 40E t.
a_h*
i2
-721
2
3
4
5
6
7
8
REO
B . v
i
55
G_
_t
140
t
S3V** -34
*:..Sf..1
A-1
2
3
4
5
6
REO
FIG. 4. Characterizationofthe35S-Fmet-labeledpolypeptide productssynthesizedin responsetoEMC RNA with time in the cell-free system. A, Unfractionated extracts from vaccinia virus-infected (gels 1 to4) and interferon-treated,vacciniavirus-infected (gels5to8)cellswereincubated in the presenceof8x10J countsper minof35S-Fmet-tRNAF (equivalentto100pmol ofI'S-metat45Ci/mmol)and
20,ug
ofEMC RNA(7.4pmol) per 0.5-ml assay.At 30min(gels1and5),60min(gels2and6),120min(gels3and7),and240min(gels4and 8)0.1-mIsampleswere treated with SDS andsubjectedtoelectrophoresisonSDS-polyacrylamidegelsin the presenceof reoviruspolypeptidesasmolecularweightmarkers. Thefigurestotherightrepresentthe molecular weightsinthousandsofthemajorreoviruspolypeptidesdetailed in thelegendtoFig.1.Autoradiographs ofthe driedgels are shown.Incorporation ofFmet in the interferon-treated, infected cell system (gels5to8) was inhibited to some extent (25 to 40%) in this experiment, but much less so than was the incorporation of 3H-leucine andfree 5S-methionine(70to80%)assayedinparallel. B,L-cell sapfrom vaccinia-infected(gels1 to3) andinterferon-treated, vaccinia-infected (gels4 to6)cellswereincubatedwithpreincubatedmicrosomes fromnormalKrebscells in the presenceof4 x10J countsperminof3S-FmettRNAF (equivalentto50pmol of 3"S-metat 45Ci/mmol)and20Mgg
ofEMC RNA (7.4pmoles) per 0.5-mI assay.At 30min(gels 1 and 4),60min (gels 2 and 5), and 120 min (gels 3 and 6) 0.125-mI samples were treated with SDS and subjected to electrophoresisasinA. Therewas nosignificantinhibition(<10%)of incorporation of Fmet in this experiment whereas therewas a50%o
inhibitionof incorporation of 3H-leucine and of free3S-methionine assayed inparallel. InbothA (240min) and B (120 min) samples were taken and completely digested with trypsin then with Pronasefor further electrophoretic analysis which showed that essentially all (> 95%) of the radioactivity incorporated into polypeptideswasrecoverableasFmet.on November 10, 2019 by guest
http://jvi.asm.org/
[image:9.495.104.393.76.460.2]!~~~~*
A
B~~~~~~~~~~~~~~~~B
I~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
l
2
3
4
5 6
7
8
9
10 11
[image:10.495.117.395.62.461.2]PR
0
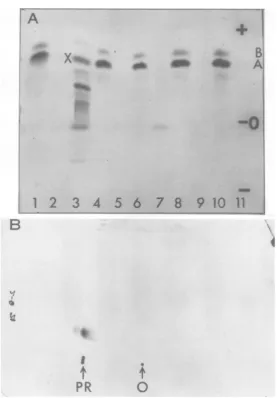
FIG. 5. Fmet-tryptic peptides formed in response to EMC RNA in Krebs and L-cell-free systems. The polypeptideproducts from the variouscell-freesystemslabeled with35S-Fmet-tRNAF or, in the case of band 3 in A,free3"S-methionine,weredigestedwithtrypsin.A,Electrophoresisonthin-layerplatesatpH 6.5.Samples (2to5uliters)wereplacedat0with theanodetothetopofthefigure.Krebscell-free system plus (1) andminus (2)EMC RNA. (3)Krebscell-freesystemplusEMC RNA withfree83S-methionine. Vaccinia-infected L-cell systemplus(4) andminus(5)EMC RNA. Interferon-treated, vaccinia-infected system plus (6) and minus(7) EMC RNA. Theremainingassayswerewithpreincubated microsomesfrom normal Krebs cells andvaccinia
virus-infected cellsapplus (8)and minus (9) EMC RNA and interferon-treated, vaccinia virus-infected cell
sapplus(10) and minus (11) EMC RNA. (B) Two-dimensional chromatography andelectrophoresis onthin layer plates (6). The digests were placed at the bottom center of the sheets (arrow) and subjected to
electrophoresisatpH6.5with the anodetotheleft.P.R. indicates thepositiontowhichaphenol red marker migrated during theelectrophoresis. Chromatography was towards thetopofthe sheet. The results withthe Fmet-trypticpeptides fromall ofthesystems inFig.5A wereessentially identical tothe examplefrom the Krebs cell system shown here. ThesamemajorFmetpeptidewasdetectedonanalysisoftheproduct after 5, 10, and120minofincubation in the Krebscell-freesystem.Experimentsin which thedigests labeled withFmet were runseparatelyand mixed withdigestslabeled withfreemethionine showed that themajor Fmetpeptide
migratedslightlyfasterin bothdimensions than the major peptideX(sample3 inFig. 5A) labeled withfree
methionine. The assays carriedoutasinMaterials andMethodswerescaled upto0.3mlforthe Krebscell systemand0.5mlforthe L-andKrebs microsome L-cellsapsystems.3BS-Fmet-tRNAFequivalentto4 x 10.
countsper min (22.6pmol ofmethionine at a specific activity of100 Ci/mmol) was added toeach assay. Incorporation in responsetoEMC RNA variedfrom 20,000 counts/min (0.11 pmol) forthe Krebssystemto
100,000counts/min (0.55pmol)forthe mixed system.
18 KERR ET AL. J. VIROL.
on November 10, 2019 by guest
http://jvi.asm.org/
(inhibited and control, unfractionatedand frac-tionated cell sap with Krebs microsomes) as
with the Krebs cell system(Fig. 5A). The minor Fmet-peptide (B, Fig. 5A) (usually<10%ofthe total radioactivity) is thought to be a further
breakdown product of the major (23, 27). It
seems reasonable to conclude that initiation with Fmet occurs at the normal site on EMC
RNAand that the Fmet is incorporatedintothe normalspectrumofEMC-specificpolypeptides in these systems. Thus, the ability of Fmet incorporation toescape the major inhibition is
not the result of its incorporation at an
abnor-mal site.
Comparing the resultsfor theanalysesofthe
Fmet N-terminally labeled polypeptide
prod-ucts synthesized at different times inthe cell-freesystemusing unfractionated (S10) material
frominterferon-treated and untreated, infected
cells (Fig. 4A), the same bias away from the
higher molecular weight products isseeninthe
interferon case as wasnoted for the
incorpora-tion of free amino acids (Fig. 1). Again, in agreementwith the results in Fig. 2, there isno
such bias in thesystemsinwhichcorresponding cell sap fractions were assayed with Krebs
microsomes (Fig. 4B). Thus, even though
poly-peptide chains initiated with Fmet escape the
major inhibition early in translation, they are
stillsubjecttothe minor inhibitionaffecting the accumulation ofhigh molecular weight material in the unfractionatedsystem.
DISCUSSION
It is impossible asyet to tell howthe inhibi-tion characterized here for systems from inter-feron-treated, vaccinia virus-infected cells, which apparently affects the translation of
EMC RNA, AUG(U)11 and mouse globin
mRNA (I. M. Kerr et al., in press; Friedman
unpublished data) but notofpoly U (11),
cor-relates withthe effectwe have seen in systems
from interferon-treated, EMC-infected L-cells,
which appears toshow somespecificity for
dif-ferent mRNAs (I. M. Kerr et al., in press; R.
M.Esteban, D. R. Tovel, and I. M. Kerr,
manu-script in preparation). Nor is it clear how this inhibition correlates with the much smaller effect of interferon treatment in the absence of infection that we have reported for these (11) and other systems (17), orwith the largeeffect
reported in the absence ofinfection by Falcoff et al. (9). The possibility exists that the inhibi-tion ofproteinsynthesis seenin the
interferon-treated, vaccinia-virus-infected cells, is a
sec-ondary effect of the abortive replicationof virus in these systems. Alternatively, it could
repre-sent the sum of a virus-mediated
inhibition-ofhost protein synthesis andan
interferon-me-diated inhibition ofvirus, rather than ablanket
interferon-mediated inhibition of both. On the
other hand, our preliminary analyses of these
different systemshave indicatedpoints in
com-mon as well as apparentdifferences, suggesting that a common underlying mechanism may be
involved. Moreover, the idea that
infection,
orother insultto the cell, mayberequiredto
trig-gerthe fulldevelopment of the
interferon-medi-ated inhibition has its attractions.
According
toonevariation onthis
hypothesis,
there could bea switch-offofproteinsynthesison infection of
the interferon-treated cell
during
which timeeither the events leadingtocell death are
initi-ated, i.e., the cell
dies,
thuspreventing
thespread of infection, or the
input
viral nucleicacid is inactivated so that, after a
period,
nor-mal host metabolismcan resume.Inmany cases
interferon pretreatment does not prevent cell
death afterinfection, it merely reduces the virus
yield and this hypothesis could
provide
thebasis for further work should it turn out that
there is no significant selection between host
andviral mRNAs in the effectupontranslation.
This,however, isonlyone
possibility;
theresultsin the interferon-treated
SV40-infected
cellsystems
(24)
strongly
suggest a specificin-hibition of viral functions, and the absence of
selectivity
forhost and viralmRNA inthe cell-free system hasby
no means beenestablished.Globin
mRNA,
particularly
intheformin whichit is presented tothe cell-free system may, for
example,
beatypical ofhost message inthein-tactcell. Amuchmoredetailedanalysis of these
cell-free systems and of the
possible
impor-tance of the different methods used in their
preparation and assay will be required before
the relationships between them and their
sig-nificancetointerferon actioncanberesolved.
Itis,
however,
withthe furthercharacteriza-tion of the enhanced inhibition of
translation,
triggered
by
infection intheinterferon-treated,
vaccinia-virus-infected cellsystemthatwehave
been concerned here. Even if this inhibition is
secondary
to the primary event of interferonaction (an event perhaps capable of affecting
transcription and translation in different
cell-virussystems in differentways), the absence of
aneffect ofthetranslation ofpoly U (11) and on
the incorporation of Fmet from Fmet-tRNAF
(Fig. 3) clearly indicates that a highly specific
mechanismisinvolved which must beof
inter-est in the control of protein synthesis. The
resultsofthe analysis of thepolypeptide
prod-ucts synthesized in response to EMC RNA in
cell-freesystems of this type are consistent with
VOL. 13, 1974
on November 10, 2019 by guest
http://jvi.asm.org/
KERR ET AL.
amajorinterferon-mediatedinhibition of
trans-lation at, or shortly after initiation, with a
minorsecondary inhibition ofchain elongation
(Fig. 1and2).On the basis of this data alone it
is possible that the major inhibition occurs
shortly after initiation whenthe productis too
smalltobeseen onacrylamidegels. It would be
premature to exclude such a mechanism, but
for it to be unique to an early event in chain
elongation it seems likely that it would be
closely linked to initiation, as indeed is
sug-gested by the results obtainedwith theinitiator
Fmet-tRNAF.
Fmet from Fmet-tRNAF is incorporated
N-terminally into the normal spectrum of high
molecular weight polypeptides synthesized in
response toEMC RNAinthese systems (Fig. 4),
but ispresent as a commonN-terminalpeptide
(Fig. 5). This, therefore, provides further
evi-dence in favour ofthe model thatwe (18) and
others (2) have previously proposed for the
translation of EMC RNA in the cell-free
sys-tem. According tothis model, initiation in the
cell-free system occurs at a unique site on the
EMC RNA as in the intact infected cell, the
majority of the highmolecular weight
polypep-tidesbeing formed
by
prematureterminationoftranslationatdifferent preferred sites (possibly
of
nucleolytic cleavage)
onthelarge EMC
RNAmessage.
Theinterestingaspect oftheresultsobtained
with Fmet-tRNAF here, however, is that the
incorporation of Fmetfrom it escapesthe
inhi-bition of translationseeninthe
interferon-treat-ed, infected-cell system. The results with
AUG(U)"
as message(Table
1) and thoseconcerning the
stability
ofthechargedinitiatortRNAsargueagainstany
simple
explanation
forthis escape, such as would be involved ifthe
inhibitory eventaffected the chargingor
break-down of thenormal initiatormet-tRNAFsothat
the inhibition
preceded
the reactions aftertheaddition of the
precharged Fmet-tRNAF.
Nordoes the
side-stepping
ofthe inhibitionby
theFmet-tRNAF reflect initiation at an abnormal
site(Fig. 5). The most likely explanationseems
to be that the abnormal formyl group on the
Fmet-tRNA is not
recognized
by
the inhibitor.But if this were thewholeanswer onewouldnot
have expected the incorporation of Fmet from
Fmet-tRNAF
tobeinhibitedwithAUG(U)n
as message(Table
1). Thereisevidence fromworkwith thereticulocytesystem, for
example,
thatboth with Fmet-tRNAF (5) and with
synthetic
messages(4),the factorrequirementsfor
initia-tion aredifferent from those observedwith the
nonformylated initiator met-tRNAF or natural
messages. This emphasizes,
incidentally,
thefact that the inhibition of incorporation in
response to
AUG(U)51
in these systems shouldnot be taken as proofthat all messages will be
affected: ifthere areadditional factors
specifi-cally required for the translation of host
mes-sages it is possible that they could provide the
specificity not apparent here. Returning to the
effect of Fmet-tRNAF, however, there are, in
fact, a numberofpossibleexplanations for this
intriguing phenomenon which are currently
under investigation. Meanwhile, taking these
results together, they suggest that the presence
of the abnormal formyl group on the initiator
Fmet-tRNAF allows a factor(s) involved in the
initiation oftranslationofEMCRNA, at least, to escape recognition at the site ofthe major
interferon-mediated inhibition oftranslation.
ACKNOWLEDGMENTS
Wethank D.Risby, A.Douglas,andSusan O'Connor for technical assistance and J. A. Sonnabend for much useful discussion.R.M.F., NationalCancerInstitute staffmember atthe National Institutes ofHealth,was avisiting scientist. The interferon was obtained from The Microbiological Re-search Establishment, Porton, Wiltshire, England (C. J.
Bradish)and from K.Paucker,andpurifiedreoviruswasthe generousgiftofJ. J.Skehel.
LITERATURE CITED
1. Boime,I.,H.Aviv,and P.Leder.1971.Proteinsynthesis
directedbyencephalomyocarditis virus RNA.II. The in vitro synthesis of high-molecular-weight proteins
and elements of the viral capsid. Biochem. Biophys.
Res. Commun.45:788-795.
2. Boime,I.,and P. Leder.1972.Proteinsynthesis directed
by encephalomyocarditis virus mRNA. III. Discrete
polypeptidestranslated fromamonocistronic messen-gerinvitro. Arch. Biochem.Biophys. 153:706-713. 3. Brown, J.C.,andA.E.Smith. 1970.Initiatorcodonsin
eukaryotes. Nature(London)226:610-612.
4. Crystal, R. G., and W. F. Anderson. 1972. Initiation of
haemoglobinsynthesis.Comparisonofmodel reactions that useartificialtemplateswith thoseusingnatural messenger RNA. Proc. Nat. Acad. Sci. U.S.A. 69:706-711.
5. Crystal, R. G., A. W. Nienhuis, P. M. Prichard, D. Picciano, N.A.Elson, W.C. Merrick,H.Graf,D. A.
Shafritz, D. G. Laycock, J. A. Last, and W. F. Anderson. 1972.Initiation ofglobinsynthesis. FEBS Lett. 24:310-314.
6. Dobos, P., I. M. Kerr, and E. M. Martin. 1971. The synthesis ofcapsid and non-capsid viral proteins in response to encephalomyocarditis virus ribonucleic acidinanimal cell-free systems. J.Virol. 8:491-499. 7. Drews,J., G. Hogenauer, F.Unger,and R. Weil. 1971.
Incorporation ofmethionine from met-tRNAFMe'into internalpositionsofpolypeptidesbymouseliver
poly-somes.Biochem. Biophys.Res.Commun.43:905-912. 8. Eggen,K.L.,andA.J.Shatkin.1972.Invitro translation ofcardiovirus ribonucleic acidbymammalian cell-free extracts.J. Virol. 9:636-645.
9. Falcoff, E., R. Falcoff, B. Lebleu, andM. Revel. 1972. InterferontreatmentinhibitsmengoRNAand haemo-globin mRNA translationincell-freeextractsof Lcells. Nature. N.Biol.(London)240:145-147.
20 J. VIROL.
on November 10, 2019 by guest
http://jvi.asm.org/
10 Friedman, R. M., R. M. Esteban, D. H. Metz, D. R. Tovell, I. M. Kerr, and R. Williamson. 1972. Transla-tion of RNAby L cell extracts: effect of interferon.
FEBSLett. 24:273-277.
11. Friedman, R. M., D. H. Metz, R. M. Esteban, D. R. Tovell, L. A. Ball, and I. M. Kerr. 1972. Mechanism of interferon action:inhibition of viral messenger ribonu-cleic acid translation in L-cell extracts. J. Virol. 10:1184-1198.
12. Gupta, N. K., N. K.Chatterjee, K. K. Bose,S.Bhaduri, and A. Chung. 1970. Roles of methionine transfer RNAs in protein synthesis in rabbit reticulocytes. J. Mol. Biol. 54:145-154.
13. Gupta, N. K., N. K. Chatterjee, C. L. Woodley, and K. K.Bose. 1971. Protein synthesis inrabbit reticulocytes: factors controlling internal and terminal methionine
codon recognitionby the methionyl tRNA species. J. Biol. Chem.246:7460-7469.
14. Housman, D., M.Jacobs-Lorena,U. L.Rajbhandary,and H. F.Lodish. 1970. Initiation of haemoglobin synthesis
by methionyl-tRNA. Nature(London) 227:913-918. 15. Hunter, A. R., and R. J. Jackson. 1971. The origin and
nature ofthe methionine residue initiating the synthe-sis of haemoglobin in vivo and in vitro. Eur. J. Biochem. 19:316-322.
16. Joklik, W. K., and T.C. Merigan. 1966. Concerning the mechanism of action of interferon. Proc. Nat. Acad. Sci. U.S.A. 56:558-565.
17. Kerr, I. M. 1971. Protein synthesis in cell-free systems; an effect of interferon.J. Virol. 7:448-459.
18. Kerr, I. M., R. E. Brown, and D. R. Tovell. 1972. Characterization of the polypeptides formed in re-sponse toencephalomyocarditisvirusribonucleic acid and in a cell-free system from mouse ascites tumor cells. J. Virol.10:73-81.
19. Kerr, I. M., N. Cohen, and T. S. Work. 1966. Factors controlling amino acid incorporation by ribosomes from
Krebs 2 mouse ascites tumour cells. Biochem. J. 98:826-835.
20. Kerr, I. M., and E. M. Martin. 1972. Simple method for the isolation ofencephalomyocarditisvirusribonucleic acid. J. Virol. 9:559-561.
21. Metz, D. H., and M. Esteban. 1972. Interferon inhibits
viral protein synthesis in L-cells infected with vaccinia virus. Nature (London). 238:385-388.
22. Muench, K., and P.Berg. 1966.Preparationofaminoacyl ribonucleic acidsynthetasesfrom Escherichia coli,p. 375-383. In G. L. Cantoni and D. R. Davies (ed.),
Procedures in nucleic acid research. Harper andRow, London.
23. Oberg, B. O., and A. J. Shatkin. 1972. Initiation of picornavirus protein synthesis in ascites cell extracts. Proc. Nat. Acad. Sci. U.S.A. 69:3589-3593.
24. Oxman, M. N. 1973. Interferon, tumors and tumor viruses, p. 391-479. In N. B. Finter (ed.), Interferon and interferoninducers, 2nd ed. North Holland Pub-lishingCompany, Amsterdam.
25. Paucker, K., B. J. Berman, R. R. Golgher, and D. Stancek. 1970. Purification, characterization and at-tempts at isotopic labeling of mouse interferon. J. Virol. 5:145-152.
26. Rabinowitz,J. C. 1963. Folic acid p. 185-252. In P. D. Boyer, K. Lardy, and K.Myrback (ed.),Theenzymes 2nd ed. 2. Academic PressInc., London.
27. Smith, A. E. 1973. The initiation of protein synthesis
directedby the RNAfromencephalomyocarditis virus. Eur. J. Biochem. 33:301-313.
28. Smith, A. E., and K. A. Marker. 1970. Cytoplasmic methionine transfer RNAs from eukaryotes. Nature (London) 226:607-610.
29. Smith, R.E., H. J. Zweerink, and W. K. Joklik. 1969. Polypeptide components of virions, top component and cores ofreovirus-type 3.Virology39:791-810. 30. Sundararajan, T. A., and R. E. Thach, 1966. Role of the
formylmethionine codonAUGinphasing translation of synthetic messenger RNA. J. Mol. Biol. 19:74-90. 31. VonEhrenstein, G. 1967. Isolation of sRNA from intact
E. coli cells. p. 588-592. In L. Grossman and K. Moldave(ed.).Methods inenzymology, vol. 12, part A. Academic Press Inc., London.
32. Waters, L. C., and G. D. Novelli. 1971. Analytical reversed-phase chromatography of Escherichia coli aminoacyl-tRNAs, p. 39-41. In K. Moldave and L. Grossman (ed.). Methods inenzymology,vol. 20, part C. Academic Press Inc., London.