DOI: 10.1534/genetics.105.044644
Note
Mutational Analysis of the pH Signal Transduction Component PalC of
Aspergillus nidulans
Supports Distant Similarity to BRO1 Domain
Family Members
Joan Tilburn,*
,1Juan C. Sa´nchez-Ferrero,
†Elena Reoyo,
†Herbert N. Arst, Jr.* and
Miguel A. Pen
˜alva
†*Department of Infectious Diseases, Faculty of Medicine, Imperial College London, London W12 0NN, United Kingdom and
†Departamento de Microbiologı´a Molecular, Centro de Investigaciones Biolo´gicas CSIC, Madrid 28040, Spain
Manuscript received April 21, 2005 Accepted for publication May 6, 2005
ABSTRACT
The alkaline ambient pH signal transduction pathway component PalC has no assigned molecular role. Therefore we attempted a gene-specific mutational analysis and obtained 55 newpalC loss-of-function alleles including 24 single residue substitutions. Refined similarity searches reveal conserved PalC regions including one with convincing similarity to the BRO1 domain, denoted PCBROH, where clustering of mutational changes, including PCBROH key residue substitutions, supports its structural and/or functional importance. Since the BRO1 domain occurs in the multivesicular body (MVB) pathway protein Bro1/Vps31 and also the pH signal transduction protein PalA (Rim20), both of which interact with MVB component (ESCRT-III protein) Vps32/Snf7, this might reflect a further link between the pH response and endocytosis.
R
EGULATION by ambient pH has been extensively studied inAspergillus nidulanswhere it is mediated by the PacC/Pal regulatory circuit and major contribu-tions have also been made by studies on the equivalent Rim systems in the yeastsSaccharomyces cerevisiae,Candida albicans, andYarrowia lipolytica.Response to ambient pH inA. nidulans(reviewed by Pen˜ alvaand Arst2002, 2004; Arstand Pen˜ alva2003)
is mediated by PacC (Caddicket al. 1986; Tilburnet al.
1995), which is activated by a two-step proteolysis of the full-length form, PacC72(Orejaset al. 1995; Mingotet al.
1999; Dı´ezet al. 2002), in response to the alkaline
ambi-ent pH signal transduced by the six-membered Pal sig-naling pathway (Arstet al. 1994). The functional PacC
250-residue form, PacC27, is an activator of
alkaline-expressed genes (Espeso and Pen˜ alva 1996) and
re-pressor of acid-expressed genes (Espesoand Arst2000).
Signal transduction components PalH and PalI (S. cerevisiaehomologs Rim21p and Rim9p, respectively) are predicted seven- and four-pass membrane proteins (Liand Mitchell1997; Denisonet al. 1998; Negrete
-Urtasunet al. 1999) and strong candidates as ambient
pH sensors. PalB (S. cerevisiae Rim13p), a calpain-like cysteine protease (Denisonet al. 1995; Lambet al. 2001;
Sorimachiand Suzuki2001), is probably responsible
for the first, pH-sensitive, signaling proteolysis. PalA (S. cerevisiaeRim20p) (Negrete-Urtasunet al. 1997; Xu
and Mitchell2001) contains the160-residue BRO1
domain (PFAM domain PF03097; http://www.sanger. ac.uk/Software/Pfam/index.shtml), first identified in yeast Bro1p (Nickasand Yaffe1996). PalA apparently
enables the signaling proteolysis by interacting both with PacC, through two YPXL/I motifs flanking the signaling proteolysis site, and with Vsp32/Snf7, the
endosomal sorting complex required for transport-III (ESCRT-III) protein, as demonstrated by Vincentet al.
(2003). This agrees with the model described for
S. cerevisiae (Xu and Mitchell 2001; Xu et al. 2004)
where Rim20p (PalA) interacts with both Rim101p (PacC) (Xu and Mitchell 2001) and Vsp32p/Snf7p,
which also interacts with Rim13p (PalB) (Ito et al.
2001), to form a scaffold-promoting interaction be-tween the Rim101p cleavage site and the Rim13p pro-tease. A functional link between pH signal transduction and multivesicular body (MVB) pathway sorting com-plexes has been firmly established in S. cerevisiae (Xu
et al. 2004) and shown to be conserved in C. albicans
(Kullaset al. 2004; Xuet al. 2004).
1Corresponding author: Department of Infectious Diseases, Imperial
College London, Hammersmith Hospital Campus, Ducane Rd., London W12 0NN, United Kingdom. E-mail: j.tilburn@imperial.ac.uk
Possible molecular roles remain elusive for PalF (S. cerevisiaeRim8p) and PalC, which has noS. cerevisiae
homolog (Li and Mitchell1997; Maccheroni et al.
1997; Negrete-Urtasun et al. 1999). As the PalC
primary amino acid sequence revealed no evident
sequence signature and PalC appeared absent from the hemiascomycete lineage, we carried out mutational analysis of this protein along with sequence profile similarity searching, exploiting the recent publication of a number of fungal genomes.
Mutational analysis: The GABA (g-aminobutyrate) technique is a powerful tool for the selection ofpacC
(Mingotet al. 1999; Ferna´ ndez-Martı´nezet al. 2003)
andpal(Arstet al. 1994; Denisonet al. 1998; Negrete
-Urtasunet al. 1999) loss-of-function mutations. It relies
on the ability of these acidity-mimicking mutations to suppressareAr(¼areA
, nitrogen metabolite repressed) mutations for utilization of g-aminobutyrate (GABA) as the nitrogen source, through derepression of acid-expressedgabAspecifying the GABA permease (Caddick
et al. 1986; Hutchings et al. 1999; Espeso and Arst
2000). In haploid strains, the GABA technique yields mutations in any of the seven pH regulatory genes. To target mutations topalC, we employed an areArpalC1
/
areArpalCdiploid. This diploid cannot use GABA as the
nitrogen source but suppression can be achieved by acidity mimicry, resulting from mutation of the palC1
allele or through mitotic recombination yielding palC
homozygosity. To avoid the latter, we constructed diploid R using inoB2 (inositol auxotrophy) distal to palC40 (Negrete-Urtasun et al. 1999) and in repulsion to
areAr18 (Arst et al. 1989), a reciprocal translocation
of chromosomes III and IV, including palC and inoB
(Figure 1). Consequently,palC40 homozygosity without inositol auxotrophy can occur only through alignment of the translocation-containing chromosome III and the untranslocated chromosome IV with recombination both between theareAr18 breakpoint andpalC40 (4 cM)
and betweenpalC40 andinoB2 (22 cM) (Figure 1). The first experiment (Table 1) yielded 48 acidity-mimicking—as determined by impaired growth on pH 8
Figure1.—Diploid R (used forpalCmutant selection). (A)
Chromosomes III and IV of diploid RareAr18/areAr3palC40
inoB2. The diploid was constructed using standard classical genetic techniques (Clutterbuck 1993) between parents
of the relevant partial genotypesareAr18 (nitrogen metabolite
repressed) (Arstet al.1989) andinoB2 (inositol requiring),
areAr3 (nitrogen metabolite repressed), andpalC40 (acidity
mimicking) (Negrete-Urtasun 1997; Negrete-Urtasun
et al. 1999). (B) Chromosomes III and IV after replication. (Ci) If alignment and recombination occur between dupli-cated homologous (with respect to the centromeres) non-sister chromatids of chromosome IV, homozygosity forpalC40 would always result in homozygosity forinoB2 and thus in ino-sitol auxotrophy, due to the areAr18 translocation that
pre-cludes recombination between palC40 and inoB2. Recovery ofpalC40inoB2 homozygotes is prevented by excluding inosi-tol from the selection medium. (Cii) If alignment occurs between the homologous regions of chromosomes III and IV (with respect to the centromeres) and if two recombina-tion events occur, one between the areAr18 breakpoint and
palC40 and a second betweenpalC40 andinoB2,inoB1 palC40 homozygotes can result.
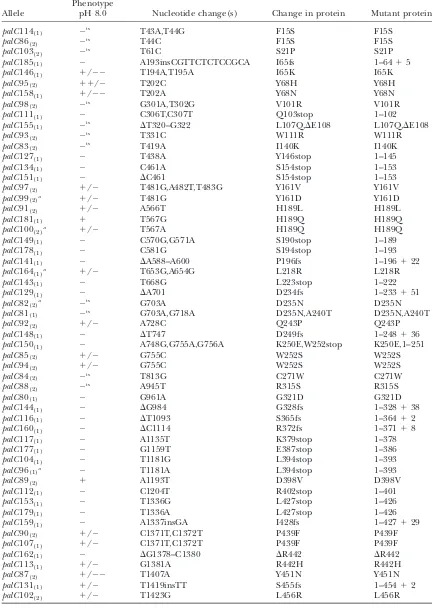
TABLE 1
palCmutations isolated in this work
Allele
Phenotype
pH 8.0 Nucleotide change(s) Change in protein Mutant protein
palC114(1) ts T43A,T44G F15S F15S
palC86(2) ts T44C F15S F15S
palC103(2) ts T61C S21P S21P
palC185(1) A193insCGTTCTCTCCGCA I65fs 1–6415
palC146(1) 1/ T194A,T195A I65K I65K
palC95(2) 11/ T202C Y68H Y68H
palC158(1) 1/ T202A Y68N Y68N
palC98(2) ts G301A,T302G V101R V101R
palC111(1) C306T,C307T Q103stop 1–102
palC155(1) ts DT320–G322 L107Q,DE108 L107Q,DE108
palC93(2) ts T331C W111R W111R
palC83(2) ts T419A I140K I140K
palC127(1) T438A Y146stop 1–145
palC134(1) C461A S154stop 1–153
palC151(1) DC461 S154stop 1–153
palC97(2) 1/ T481G,A482T,T483G Y161V Y161V
palC99(2)a 1/ T481G Y161D Y161D
palC91(2) 1/ A566T H189L H189L
palC181(1) 1 T567G H189Q H189Q
palC100(2)a 1/ T567A H189Q H189Q
palC149(1) C570G,G571A S190stop 1–189
palC178(1) C581G S194stop 1–193
palC141(1) DA588–A600 P196fs 1–196122
palC164(1)a 1/ T653G,A654G L218R L218R
palC143(1) T668G L223stop 1–222
palC129(1) DA701 D234fs 1–233151
palC82(2)a ts G703A D235N D235N
palC81(1) ts G703A,G718A D235N,A240T D235N,A240T
palC92(2) 1/ A728C Q243P Q243P
palC148(1) DT747 D249fs 1–248136
palC150(1) A748G,G755A,G756A K250E,W252stop K250E,1–251
palC85(2) 1/ G755C W252S W252S
palC94(2) 1/ G755C W252S W252S
palC84(2) ts T813G C271W C271W
palC88(2) ts A945T R315S R315S
palC80(1) G961A G321D G321D
palC144(1) DG984 G328fs 1–328138
palC116(1) DT1093 S365fs 1–36412
palC160(1) DC1114 R372fs 1–37118
palC117(1) A1135T K379stop 1–378
palC177(1) G1159T E387stop 1–386
palC104(1) T1181G L394stop 1–393
palC96(1)a T1181A L394stop 1–393
palC89(2) 1 A1193T D398V D398V
palC112(1) C1204T R402stop 1–401
palC153(1) T1336G L427stop 1–426
palC179(1) T1336A L427stop 1–426
palC159(1) A1337insGA I428fs 1–427129
palC90(2) 1/ C1371T,C1372T P439F P439F
palC107(1) 1/ C1371T,C1372T P439F P439F
palC162(1) DG1378–C1380 DR442 DR442
palC113(1) 1/ G1381A R442H R442H
palC87(2) 1/ T1407A Y451N Y451N
palC131(1) 1/ T1419insTT S455fs 1–45412
palC102(2) 1/ T1423G L456R L456R
NovelpalCmutations were selected after UV mutagenesis of diploid RpabaA1yA2areAr18/biA1areAr3palC40
inoB2fwA1 (see Clutterbuck1993 for gene symbols) on minimal medium (Cove1966) pH 6.5 with 1%
medium (Cove1976)—palCmutants. These included 24
new truncations, 8 single and one double missense, two with three-base deletions, and one rearrangement muta-tion (data not shown) plus 12palC40 mitotic recombi-nants, despite precautions. In a second attempt and to avoid null phenotypepalC truncations, we screened for leaky or temperature-sensitive growth on pH 8.0 medium and obtained another 20palCmissense mutants, thus to-taling 55 newpalCalleles (Table 1). Alignment (Figure 2) illustrates that most of the missense mutations affect amino acids conserved in the majority of the ascomycete PalCs shown. Of the single residue change mutations, only palC80 (Gly321Asp) and palC162 (DArg442) are complete loss-of-function mutations. All truncating mu-tations result in complete loss of function except the leakypalC131 (1–45412), which removes 53 C-terminal residues including a completely conserved C-terminal di-aromatic motif. This motif (di-tyrosine inA. nidulans
PalC) resembles the di-phenylalanine motif, a C-terminal transport motif facilitating ER export (Nufer et al.
2002). As palC131 is partially functional, this motif and other residues C-terminal to residue 454 cannot be completely essential for PalC structure and function. The complete loss-of-function truncating mutations
palC159 (1–427129) andpalC153 andpalC179, trun-cating the protein cleanly after residue 426, indicate that at least some of residues 427–454 are essential, which agrees with the clustering of mutations in this window. Mutations substituting conserved residues in regions ‘‘BRO1 similar,’’ ‘‘LALA,’’ and ‘‘ERRE’’ (Figure 4B) further support the structural and/or functional importance of these regions. We caution, however, that the phenotypes of any of thepalCmutations character-ized here might be due to reduced PalC protein levels resulting from protein misfolding with consequent in-stability or even from messenger inin-stability rather than from PalC dysfunction.
PalC orthologs: PalC orthologs are present in mem-bers of three major fungal phyla, ascomycota, basidio-mycota, and zygomycota (Figures 2 and 3). Among ascomycete PalCs theY. lipolyticaortholog (Figure 2) is the first to be detectable in hemiascomycetes (yeasts). This dispels the notion, based on the failure to detect PalC homologs in the previously available hemiascomy-cete genomes (supplementary Table S1 at http://www. genetics.org/supplemental/) ofAshbya gosypii,C. albi-cans, Candida glabrata, Debaryomyces hansenii, Kluyver-omyces lactis,S. cerevisiae, andSchizosaccharomyces pombe, that PalCs are exclusive to euascomycetes (filamentous fungi). As strong comparative genomic evidence indi-cates that Y. lipolytica separated early from the main hemiascomycete line and has not been subjected to the significant constraints in genome size characterizing other yeasts (Dujonet al. 2004), we suggest that palC
was present in a universal fungal ancestor and that
palCorthologs might have been lost from most hemi-ascomycetes.
Y. lipolytica palCis the only homolog of anA. nidulans
pH regulatory gene not identified by mutations affect-ing pH regulation of extracellular proteases (Tre´ ton
et al. 2000; Gonzalez-Lopezet al. 2002). Thus its
func-tional involvement in pH regulation remains to be established.
Refined similarity searching reveals the PalC-related protein family: As sequence similarity searches gave no indication of a possible molecular role for PalC, we also used the relatively sensitive procedures of HMMer (Eddy1998) and PSI-BLAST (Altschulet al. 1997) to
seek PalC-related protein families. Searching the nrdbembl ( January 2005) protein database with the 262 N-terminal residues of PalC detected, as well as PalC orthologs, we detected two members of the PFAM BRO1 domain family after the third round of iteration with
E-values markedly below the cutoff (E ¼ 0.005) and
acidity-mimicking mutants. Acidity-mimicking mutant diploids were haploidized on benlate-containing Aspergillus complete medium (Cove1966; Hastie1970) lacking inositol and with NaH
2PO4added to 1m. Haploid isolates were phenotype tested
and categorized by their growth on pH 8 medium (Cove1976), which is the most sensitive test to distinguish the relative leakiness
of mutations., complete loss-of-function, virtually no growth at 25°or 37°;ts, some growth at 25°but not at 37°;1/,1/,
and11/, increasing amounts of growth at 37°;1, similar to wild type on pH 8 medium yet acidity mimicking by more stringent criteria such as molybdate hypersensitivity (Caddick et al. 1986). All missense mutations except palC80 permitted significant
growth at pH 8 at 25°. The subscript after the allele number refers to the experiment in which the mutation was isolated (see text). The difference in phenotype between the haploidpalC181 (His189Gln) mutant and the aneuploidpalC100 (His189Gln), ‘‘1’’vs.‘‘1/,’’ respectively, suggests that the mutant phenotype of a new mutation in haploidy might be less pronounced than that in aneuploidy withpalC40. fs, frameshift at the indicated codon;1N, whereNis a number, indicates the number of frame-shifted amino acid residues. Due to the large number of mutations selected, a relatively small number of strains were analyzed in some cases and the possibility cannot be ruled out that the phenotype might be affected by a modifying mutation, although an unexpected phenotype led to analysis in a cross. Naked DNA or DNA from conidiospores from haploid isolates was used as tem-plate for PCR amplification usingpalC-specific primers. Mutations were detected by the direct sequencing of PCR products. The entire coding region was sequenced for all new mutant alleles. Nucleotide numbers refer to the published sequence (Negrete
-Urtasunet al. 1999).
a
The sequenced strain is probably aneuploid and carriedpalC40 in addition to the newpalCmutation. TABLE 1
Figure 2.—Alignment of
some ascomycete and basid-iomycete PalCs and a zygomy-cete PalC. The alignment was carried out using the T-Coffee multiple sequence alignment (Notredame et al. 2000).
Shading was according to the Blosum62 matrix:.90% similarity, solid background; 50–90% similarity, dark shad-ing; 30–50% similarity, light shading.Residue-substituting mutations are indicated by[; different substitutions of the same residue are indicated by /; double mutations in the same allele are indicated by parentheses;Zmarks the ultimate wild-type residue in PalC159, the most C-terminal total loss-of-function trun-cated mutant protein;Y indi-cates the ultimate wild-type residue in the leaky truncated mutant PalC131 protein. Res-idues 38–235 composing the PCBROH domain (see Fig-ure 4 and text) are italicized. A total of 73 Ustilago maydis
PalC residues with no similar-ity to any other protein shown have been removed from the fifth block of the alignment. They are: 315-HAGTQIGL SANHEHELASRLSASRDR ADHEHDDDMVETNRGA GAQATSKRNKLLGRFKLG SSKSSPPRSASVH-388. Ini-tials refer to species names used in A. With the sole exception of theA. nidulans
PalC, for which complete cDNA sequence is available, proteins from all other fungi were conceptually translated from predicted genes derived from genomic sequences. In
all 63 PFAM PF03097 BRO1-like domain-containing proteins after the eighth iteration and retrieved no new unrelated sequences subsequently. This relation-ship to BRO1 domain proteins was highly suggestive in view of the PalA BRO1 domain and was substantiated using a Hidden Markov model derived from the region of similarity corresponding toA. nidulansPalC 38–235 in the 14 PalCs available to query the nrdbembl, which detected three BRO1 domain proteins with reasonable scores (0.0018–0.046). In a third approach, a PalC Hidden Markov model in an HHpred (So¨ ding 2005)
search of the PFAM database gave only one significant hit (E-value 3e-05), the BRO1-like domain. These data provide statistical evidence that a region of PalC com-prising residues 28–235 is significantly, albeit remotely, related to the PFAM PF03097 BRO1-like domain and a region of similarity extending downstream from it (Figure 4A). We have denoted this region PCBROH (PalC-BRO1 homology). The frequency of mutational changes here and their occurrence in key PCBROH residues such as Tyr68 and Trp111 indicate a major functional and/or structural role in PalC (Figure 4B).
BRO1 domain proteins include, as well as PalA (Negrete-Urtasunet al.1997), PalA orthologs Rim20p
(S. cerevisiaeandC. albicans) (Xuand Mitchell2001),
the human PalA homolog AIP1/Alix (Missottenet al.
1999; Vitoet al. 1999), and the yeast Bro1p (Nickasand
Yaffe 1996), an MVB pathway protein recruiting the
deubiquitinase Doa4p to endosomes (Odorrizi et al.
2003; Luhtalaand Odorrizi2004). All these proteins
interact with Vps32p/Snf7p (a key component of ESCRT-III) or its homologs (Xu and Mitchell2001;
Katohet al. 2003; Stracket al. 2003; Vincentet al.2003; vonSchwedleret al. 2003; Katohet al. 2004; Pecket al.
2004; Xuet al. 2004).
It has been suggested that the BRO1 domain is the region of interaction between Rim20p-Bro1p family members and Snf7p family members (Xuet al. 2004).
However, the finding that the BRO1 domain-containing protein human rhophilin-2 did not interact with either of two human hSnf7 proteins tested (Pecket al. 2004)
argues against this generalization. Thus a precise role for the BRO1-like domain in PalC cannot be assigned. However, it is tempting to speculate that it reflects a further link between the pH response system and the MVB pathway multiprotein complexes at the endosome and/or cell membrane.
The PacC-mediated pH regulatory system is an important virulence determinant in plant (reviewed by Pen˜ alva and Arst 2004) and animal pathogens,
in-cluding C. albicans (reviewed by Fonzi 2002; Pen˜ alva
and Arst2002; Davis2003) andA. nidulans(Bignell
et al. 2005). In view of the growing frequency of invasive mold, and in particular of Aspergillus infections (re-viewed by Clarkand Hajjeh2002), the uniquely fungal
PalC might possibly be a suitable target for therapeutic intervention.
We are grateful to Lily Stanton for technical assistance; Elaine Bignell, Eduardo Espeso, and Olivier Vincent for comments on the manuscript; and Joanna Rudnicka for interesting discussion. We thank the Wellcome Trust, the Direccio´n General de Investigacio´n Cientı´fica y Te´cnica, and the Consejo Superior de Investigaciones Cientı´ficas (CSIC) for their support through grants 067878, BIO2003-0077, and for a CSIC Bioinformatics I3P studentship for J.C.S.-F., respectively.
Note added in proof: The structure of the Bro1 domain in yeast Bro1p has been determined ( J. Kim, S. Sitaraman, A. Hierro, B. M. Beach,
G. Odorriziand J. H. Hurley, 2005, Structural basis for endosomal
targeting by the Bro1 domain. Dev. Cell8:937–947) and establishes that the Bro1 domain contains 367 residues, thereby extending considerably beyond the 160-residue Bro1p PFAM domain. Sequence alignment suggests that the region of A. nidulans PalC corresponding to the
Figure 3.—Phylogenetic
tree relating PalC orthologs used in this work. This was constructed using MEGA version 3.0 neighbor-joining method (Kumaret al. 2004).
D is a measure of sequence divergence.
structurally based Bro1 domain of Bro1p includes the PalC N-terminal 446 residues, within which the majority of our single residue substitutions fall. Conserved PalC residues Arg47, Tyr68, Trp111, and Glu136 (Figures 2 and 4A) are likely to correspond, respectively, to Bro1p residues Arg51,
Tyr70, Trp94, and Glu116, which are central in one of two structure-stabilizing, buried, charged, and polar clusters. The loss-of-function phenotypes of palCTyr68His or -Asn and Trp111Arg mutations are consistent with the predicted structural importance of these residues.
Figure 4.—Conserved regions in PalC. (A) The PalC BRO1 homology. Multiple sequence alignment of five ascomycete PalC
proteins and representative members of each of five PF03097/BRO1-containing protein subfamilies illustrates sequence similarity between PalCs (indicated with a black bar on the left) and Bro1-like proteins. The five PF03097 subfamilies have been recognized by phylogenic analysis ( J. C. Sa´ nchez-Ferrero, O. Vincentand M. A. Pen˜ alva, unpublished results) and comprise PalA ascomycete
LITERATURE CITED
Altschul, S. F., T. L. Madden, A. A. Scha¨ ffer, J. Zhang, Z. Zhang
et al., 1997 Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res.25:
3389–3402.
Arst, H. N., Jr., and M. A. Pen˜ alva, 2003 pH regulation inAspergillus
and parallels with higher eukaryotic regulatory systems. Trends Genet.19:224–231.
Arst, H. N., Jr., D. Tollerveyand M. X. Caddick, 1989 A
translo-cation associated, loss-of-function mutation in the nitrogen metabolite repression regulatory gene of Aspergillus nidulans
can revert intracistronically. Mol. Gen. Genet.215:364–367. Arst, H. N., Jr., E. Bignelland J. Tilburn, 1994 Two new genes
involved in signalling ambient pH inAspergillus nidulans.Mol. Gen. Genet.245:787–790.
Bignell, E., S. Negrete-Urtasun, A. M. Calcagno, K. Haynes,
H. N. Arst, Jr.et al., 2005 TheAspergilluspH-responsive
tran-scription factor PacC regulates virulence. Mol. Microbiol.55:
1072–1084.
Caddick, M. X., A. G. Brownleeand H. N. Arst, Jr., 1986 Regulation
of gene expression by pH of the growth medium inAspergillus nidulans.Mol. Gen. Genet.203:346–353.
Clark, T. A., and R. A. Hajjeh, 2002 Recent trends in the
epidemi-ology of invasive mycoses. Curr. Opin. Infect. Dis.15:569–574. Clutterbuck, A. J., 1993 Aspergillus nidulans, pp. 3.71–3.84 in
Genetic Maps of Complex Genomes, edited by S. J. O’Brien. Cold
Spring Harbor Laboratory Press, Cold Spring Harbor, NY. Cove, D. J., 1966 The induction and repression of nitrate reductase
in the fungusAspergillus nidulans.Biochim. Biophys. Acta113:
51–56.
Cove, D. J., 1976 Chlorate toxicity inAspergillus nidulans.Studies of
mutants altered in nitrate assimilation. Mol. Gen. Genet.146:
147–159.
Davis, D., 2003 Adaptation to environmental pH inCandida albicans
and its relation to pathogenesis. Curr. Genet.44:1–7. Denison, S. H., M. Orejasand H. N. Arst, Jr., 1995 Signaling of
ambient pH inAspergillusinvolves a cysteine protease. J. Biol. Chem.270:28519–28522.
Denison, S. H., S. Negrete-Urtasun, J. M. Mingot, J. Tilburn, W. A.
Mayeret al., 1998 Putative membrane components of signal
transduction pathways for ambient pH regulation inAspergillus
and meiosis inSaccharomycesare homologous. Mol. Microbiol.
30:259–264.
Dı´ez, E., J. A´lvaro, E. A. Espeso, L. Rainbow, T. Sua´ rezet al.,
2002 Activation of theAspergilluszinc-finger transcription fac-tor requires two proteolytic steps. EMBO J.21:1350–1359. Dujon, B., D. Sherman, G. Fischer, P. Durrans, S. Casaregaloet al.,
2004 Genome evolution in yeasts. Nature430:35–44. Eddy, S. R., 1998 Profile hidden Markov models. Bioinformatics14:
755–763.
Espeso, E. A., and H. N. Arst, Jr., 2000 On the mechanism by which
alkaline pH prevents expression of an acid-expressed gene. Mol. Cell. Biol.20:3355–3363.
Espeso, E. A., and M. A. Pen˜ alva, 1996 Three binding sites for the
Aspergillus PacC zinc-finger transcription factor are necessary and sufficient for regulation by ambient pH of the isopenicillin N synthase gene promoter. J. Biol. Chem.271:28825–28830. Ferna´ ndez-Martı´nez, J., C. V. Brown, E. Dı´ez, J. Tilburn,
H. N. Arst, Jr.et al., 2003 Overlap of nuclear localisation signal
and specific DNA-binding residues within the zinc finger domain of PacC. J. Mol. Biol.334:667–684.
Fonzi, W. A., 2002 Role of pH response inCandida albicans
viru-lence. Mycoses45(Suppl. 1): 16–21.
Gonzalez-Lopez, C. I., R. Szabo, S. Blanchin-Roland and
C. Gaillardin, 2002 Genetic control of extracellular protease
synthesis in the yeastYarrowia lypolitica.Genetics160:417–427. Hastie, A. C., 1970 Benlate-induced instability of Aspergillus
dip-loids. Nature226:771.
Hutchings, H., K.-P. Stahmann, S. Roels, E. A. Espeso, W. E.
Timberlake et al., 1999 The multiply-regulated gabA gene
encoding the GABA permease ofAspergillus nidulans: a score of exons. Mol. Microbiol.32:557–568.
Ito, T., T. Chiba, R. Ozawa, M. Yoshida, M. Hattoriet al., 2001 A
comprehensive two-hybrid analysis to explore the yeast proteome interactome. Proc. Natl. Acad. Sci. USA98:4569–4574.
Katoh, K., H. Shibata, H. Suzuki, A. Nara, K. Ishidoh et al.,
2003 The ALG-2-interacting protein Alix associates with CHMP4b, a human homologue of yeast Snf7 that is involved in multivesicular body sorting. J. Biol. Chem.278:39104–39113. Katoh, K., H. Shibata, K. Hattaand M. Maki, 2004 CHMP4b is a
major binding partner of the ALG-2-interacting protein Alix among the three CHMP4 isoforms. Arch. Biochem. Biophys.421:159–165. Kullas, A. L., M. Liand D. A. Davis, 2004 Snf7p, a component of
the ESCRT-III protein complex, is an upstream member of the
RIM101pathway inCandida albicans.Eukaryot. Cell3:1609–1618. Kumar, S., K. Tamuraand M. Nei, 2004 MEGA3: integrated
soft-ware for Molecular Evolutionary Genetics Analysis and sequence alignment. Brief. Bioinform.5:150–163.
Lamb, T. M., W. Xu, A. Diamondand A. P. Mitchell, 2001 Alkaline
response genes ofSaccharomyces cerevisiaeand their relationship to the RIM101 pathway. J. Biol. Chem.276:1850–1856.
Li, W., and A. P. Mitchell, 1997 Proteolytic activation of Rim1p,
a positive regulator of yeast sporulation and invasive growth. Genetics145:63–73.
Luhtala, N., and G. Odorrizi, 2004 Bro1 coordinates
deubiquiti-nation in multivesicular body pathway by recruiting Doa4 to endosomes. J. Cell Biol.166:717–729.
Maccheroni, W., Jr., G. S. May, N. M. Martinez-Rossiand A. Rossi,
1997 The sequence of palF, an environmental pH response gene inAspergillus nidulans.Gene194:163–167.
Mingot, J. M., J. Tilburn, E. Dı´ez, E. Bignell, M. Orejaset al.,
1999 Specificity determinants of proteolytic processing of Asper-gillusPacC transcription factor are remote from the processing site, and processing occurs in yeast if pH signalling is bypassed. Mol. Cell. Biol.19:1390–1400.
Missotten, M., A. Nichols, K. Riegerand R. Sadoul, 1999 Alix, a
novel mouse protein undergoing calcium-dependent interaction with the apoptosis-linked-gene 2 (ALG-2) protein. Cell Death Differ.6:124–129.
Negrete-Urtasun, S., 1997 Aspects of the pH signal transduction
pathway in the filamentous fungus Aspergillus nidulans. Ph.D. Thesis, University of London, London.
Negrete-Urtasun, S., S. H. Denison and H. N. Arst, Jr.,
1997 Characterization of the pH signal transduction pathway genepalA ofAspergillus nidulansand identification of possible homologs. J. Bacteriol.179:1832–1835.
Negrete-Urtasun, S., W. Reiter, E. D´ezı , S. H. Denison, J. Tilburn
et al., 1999 Ambient pH signal transduction inAspergillus: com-pletion of gene characterization. Mol. Microbiol.33:994–1003. Nickas, M. E., and M. P. Yaffe, 1996 BRO1, a novel gene that
inter-acts with components of the Pkc1p-mitogen-activated protein kinase pathway in Saccharomyces cerevisiae. Mol. Cell. Biol. 16:
2585–2593.
Notredame, C., D. G. Higginsand J. Heringa, 2000 T-Coffee: a
novel method for fast and accurate multiple sequence alignment. J. Mol. Biol.302:205–217.
Nufer, O., S. Guldbransen, M. Degen, F. Kappeler, J. P. Paccaud
et al., 2002 Role of cytoplasmic C-terminal amino acids of mem-brane proteins in ER export. J. Cell Sci.115:619–628. Odorrizi, G., D. J. Katzmann, M. Babst, A. Audhyaand S. D. Emr,
2003 Bro1 is an endosome-assiociated protein that functions in the MVB pathway inSaccharomyces cerevisiae.J. Cell Sci.116:1893– 1903.
Orejas, M., E. A. Espeso, J. Tilburn, S. Sarkar, H. N. Arst, Jr.,et al.,
1995 Activation of the Aspergillus PacC transcription factor in response to alkaline ambient pH requires proteolysis of the carboxy-terminal moiety. Genes Dev.9:1622–1632.
Peck, J. W., E. T. Bowdenand P. D. Burbelo, 2004 Structure and
function of human Vps20 and Snf7 proteins. Biochem. J.377:
693–700.
Pen˜ alva, M. A., and H. N. Arst, Jr., 2002 Regulation of gene
expression by ambient pH in filamentous fungi and yeasts. Microbiol. Mol. Biol. Rev.66:426–446.
Pen˜ alva, M. A., and H. N. Arst, Jr., 2004 Recent advances in the
characterization of ambient pH regulation of gene expression in filamentous fungi and yeasts. Annu. Rev. Microbiol.58:425–451. So¨ ding, J., 2005 Protein homology detection by HMM-HMM
com-parison. Bioinformatics21:951–960.
Sorimachi, H., and K. Suzuki, 2001 The structure of calpain.
J. Biochem.129:653–664.
Strack, B., A. Calistri, S. Craig, E. Popova and H. G.
Go¨ ttlinger, 2003 AIP1/ALIX is a binding partner for
HIV-1 p6 and EIAV p9 functioning in virus budding. Cell114:
689–699.
Tilburn, J., S. Sarkar, D. A. Widdick, E. A. Espeso, M. Orejaset al.,
1995 TheAspergillusPacC transcription factor mediates regula-tion of both acid- and alkaline expressed genes by ambient pH. EMBO J.14:779–790.
Tre´ ton, B., S. Blanchin-Roland, M. Lambert, A. Le´ pingle and
C. Gaillardin, 2000 Ambient pH signalling in ascomycetous
yeasts involves homologues of theAspergillus nidulansgenespalF
andpalH.Mol. Gen. Genet.263:505–513.
Vincent, O., L. Rainbow, J. Tilburn, H. N. Arst, Jr. and
M. A. Pen˜ alva, 2003 YPXL/I is a protein interaction motif
recognized by Aspergillus PalA and its human homologue AIP1/Alix. Mol. Cell. Biol.23:1647–1655.
Vito, P., L. Pellegrini, C. Guietand L. D’Adamio, 1999 Cloning of
AIP1, a novel protein that associates with the apoptosis-linked gene ALG-2 in a Ca21-dependent reaction. J. Biol. Chem.274:
1533–1540.
VonSchwedler, U. K., M. Stuchell, B. Mu¨ ller, D. M. Ward, H. Y.
Chunget al., 2003 The protein network of HIV budding. Cell
114:701–713.
Xu, W., and A. P. Mitchell, 2001 Yeast PalA/AIP1/Alix homolog
Rim20p associates with a PEST-like region and is required for its proteolytic cleavage. J. Bacteriol.183:6917–6923.
Xu, W., F. J. Smith, Jr., R. Subaran and A. P. Mitchell,
2004 Multivesicular body-ESCRT components function in pH response regulation inSaccharomyces cerevisiaeandCandida albi-cans.Mol. Biol. Cell15:5528–5537.