Lineage and Serotype Identification of Listeria monocytogenes by Matrix-assisted Laser Desorption Ionization-time of Flight Mass Spectrometry
Full text
Figure
Related documents
The results of this study show that identification of diverse species of Mycobacterium , and strains within those species, is possible using whole-cell MALDI.. All of the 37
Whole-cell fingerprinting by matrix-assisted laser desorption ionization–time-of-flight mass spectrometry (MALDI-TOF MS) in combination with a dedicated bioinformatic software
Detection of carbapenemase activity by MALDI- TOF analysis is similar to the reference method, the detection of imipenem hydrolysis using UV spectrometry, but the assay described
The blind test of the reference database, in which two TABLE 2 Identification of ticks removed from patients using morphological, MALDI-TOF MS, and molecular methods and the results
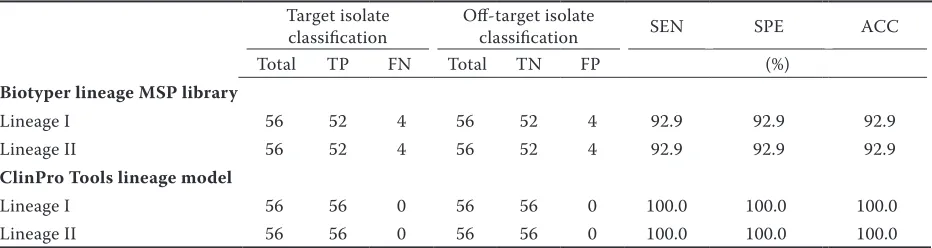
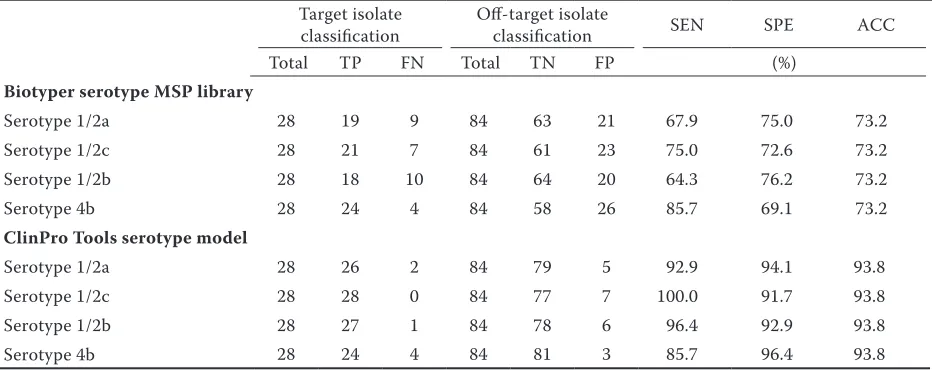
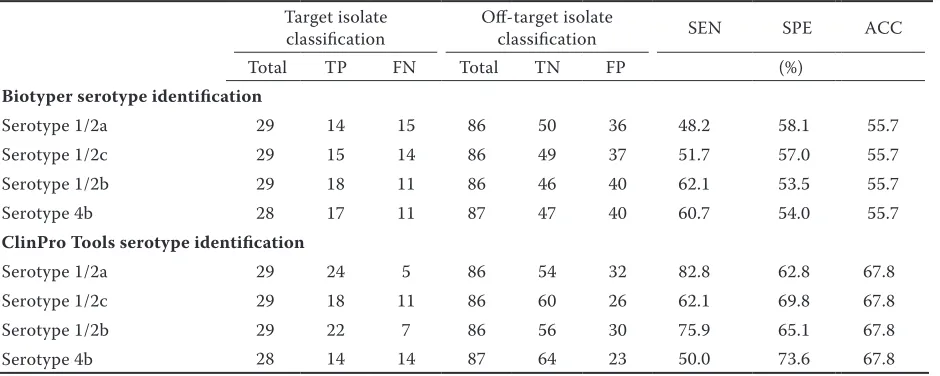
Here, matrix-assisted laser desorption ionization–time of flight mass spectrometry (MALDI-TOF MS) coupled to ClinProTools was used to discover MALDI-TOF MS biomarker peaks and
A rapid and sensitive (100%) matrix-assisted laser desorption ionization ⴚ time of flight mass spectrometry (MALDI-TOF MS) assay was developed to detect OXA-48-type producers, using
The results acquired using the online application also surpassed those obtained with the same set of strains identified against the Bruker database using the MALDI Biotyper
Identification by matrix- assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) and conventional identification of 318 microorgan- isms isolated from