MOST
UV-INDUCED RECIPROCAL TRANSLOCATIONS IN
SORDARIA MACROSPORA OCCUR I N OR NEAR
CENTROMERE REGIONS
G . LEBLON, D. ZICKLER AND S. LEBILCOT
Laboratoire de Ginktique, Bitiment 400, Universitk Paris-Sud, Centre d’Orsay, 91405 Orsay Cedex, France
Manuscript received March 18, 1985 Revised copy accepted October 4, 1985
ABSTRACT
In fungi, translocations can be identified and classified by the patterns of ascospore abortion in asci from crosses of rearrangement X normal sequence. Previous studies of UV-induced rearrangements in Sordaria macrospora revealed that a major class (called type 111) appeared to be reciprocal translocations that were anomalous in producing an unexpected class of asci with four aborted ascospores in bbbbaaaa linear sequence (b = black; a = abortive). T h e present study shows that the anomalous type 111 rearrangements are, in fact, reciprocal translocations having both breakpoints within or adjacent to centromeres and that bbbbaaaa asci result from 3: 1 disjunction from the translocation quadriva- lent.-Electron microscopic observations of synaptonemal complexes enable cen- tromeres to be visualized. Lengths of synaptonemal complexes lateral elements in translocation quadrivalents accurately reflect chromosome arm lengths, ena- bling breakpoints to be located reliably in centromere regions. All genetic data are consistent with the behavior expected of translocations with breakpoints at centromeres.-Two-thirds of the UV-induced reciprocal translocations are of this type. Certain centromere regions are involved preferentially. Among 73
type-I11 translocations, there were but 13 of the 21 possible chromosome com- binations and 20 of the 42 possible combinations of chromosome arms.
N fungi that produce tetrads of spores following meiosis, chromosome rear-
I
rangements of various types can be identified by visual observation of asci from crosses of translocation X wild type. They contain different numbers of defective (deficient) ascospores and occur with characteristic frequencies thatare often diagnostic (PERKINS 1974; PERKINS and BARRY 1977). Among the
UV-induced aberrations obtained in Sordaria macrospora, three types were dis- tinguishable by their patterns of aborted spores in ordered asci from crosses
of rearrangement X normal sequence (ARNAISE, LEBLON and LARES 1984).
Using the theoretical basis outlined by PERKINS (1974), in which the frequen- cies of tetrad types in heterozygous crosses provide information on the posi- tions of breakpoints relative to the centromeres, two of the three types could
be characterized (ARNAISE, LEBLON and LARES 1984). Type I were defined as
reciprocal translocations; they were distinguished by 8b:Oa (b = black, a =
184 G. LEBLON, D. ZICKLER AND S. LEBILCOT
abortive) and Ob:8a asci, derived from alternate and adjacent disjunction, and 2b:2a:2b:2a, 2b:4a:2b and 2a:4b:2a following exchange in one of the two
centromere-breakpoint intervals. Type I1 were distinguished by 8b:Oa and
6b:2a, the latter resulting from segregation of the elements of tip-to-nontip interchanges of arms or transposition of chromosome segments either within or between chromosomes, in which duplication- but not deficiency-bearing spores survive. Type I11 (the major class) did appear to be reciprocal translo- cations when their pairing structures were reconstructed from serial sections (ZICKLER et al. 1984), but were anomalous in producing an unexpected class of asci with four aborted ascospores in bbbbaaaa linear sequence.
In this paper, 73 rearrangements of the anomalous type I11 are analyzed genetically. In addition, electron microscopy is used to localize their centro- meres and breakpoints. T h e observations confirm and extend the conclusion that type I11 consists of reciprocal translocations having both breakpoints in centromeric regions and that bbbbaaaa asci result from 3:l disjunction from the quadrivalent. T h e chromosomes involved in type-I11 translocations are identified. Recovery of such translocations is nonrandom with respect to the chromosomes involved.
MATERIALS AND METHODS
Organism, media and crossing techniques: Sordaria macrospora is an homothallic ascomycete similar in many respects to Neurospora. T h e main characteristics have been described by ESSER and KUENEN (1967). T h e strains used in the present study were derived from a wild stock, St Ismier, FGSC4818. T h e media and crossing techniques were previously described (ZICKLER et al. 1984).
Genetic markers: For the phenotypes and descriptions of the markers used here, see ZICKLER et al. (1 984). Since each linkage group has now been assigned to its respective chromosome, the previous letters designating the linkage groups are changed as follows: LGa is designated as chromosome 1 , LGb as chromosome 5 , LGc as 6 , LGd as 7, LGe as 4 , LGf as 3 and LGg as 2. T h e genotypes of the multiply marked strains are sgrl
(chromosome I ) , arol (chromosome 5), Pam2 (chromosome 4 ) , sgn2 (chromosome 3)
for the strain 1342GL; sgrl (chromosome I ) , arol (chromosome 5 ) , Pam2 (chromosome 4 ) , sgn2 (chromosome 3), asyl (chromosome 2 ) for the strain 1348GL; and Pam1 (chro- mosome 6 ) , tsgl (chromosome 7), sgr2 (chromosome 4 ) , asyl (chromosome 2) for the strains 1488GL and 1487GL. T h e genetic marker per1 is located on chromosome 4 ,
typl on chromosome 1 and thrl on chromosome 5 . T h e spo76 phenotype is described in MOREAU, ZICKLER and LEBLON (1985).
Chromosomal rearrangements: Translocations will be symbolized in order to show the chromosomes involved once identified; each translocation of independent origin is identified by an arabic number following the symbol, e.g., T(1,5)116. T h e 7 3 chromo- somal rearrangements studied originated from t w o U V mutageneses of the mycelium. Translocations 14 through 49 were collected from different irradiated plates at UV doses of 200 J/m' in a preliminary experiment. T h e other 62 rearrangements, begin- ning with translocation 102 and ending with 218, were isolated by ARNAISE, LEBLON and LARES (1 984) from different irradiated plates, with the exception of translocation 195, which was collected from a nonirradiated control plate. Therefore, all type-111 rearrangements were independently collected. Moreover, in each of the two experi- ments, all irradiated and control mycelium came from a newly isolated haploid wild-
type nucleus.
was postfixed in 2% phosphate-buffered Os04 for 1 hr. The asci were dehydrated in a graded alcohol series, followed by propylene oxide. Each ascus was then embedded in
a drop of Epon and was observed in the light microscope in order to select the meiotic stage. Serial thin sections were picked up with a single hole grid without supporting film, transferred to a formvar-coated single hole grid and poststained ( 5 % aqueous uranyl acetate for 30 min, followed by lead citrate for 20 min). Three-dimensional reconstructions were performed, as described previously (ZICKLER 1977).
RESULTS
Patterns of ascospore abortion: Electron microscopic studies of the synap- tonemal complex of six type-I11 rearrangements (T14, T 4 1 , T123, T157, T165 and T21?) demonstrated that they correspond to reciprocal translocations
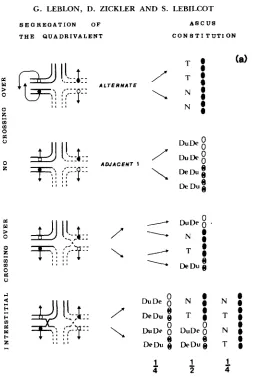
(ZICKLER et al. 1984; this work). Patterns of ascospore abortion, therefore, were examined carefully to see if they conformed with those expected of reciprocal translocations (Figure la). Ascus patterns in crosses of these six translocations by wild type are shown in Tables 1 and 2. Asci were observed both after the perithecia were crushed in water (Table 1) and after the asco- spores were discharged on Petri-dish lids (Table 2). T h e data from Tables 1 and 2 are compiled in Table 3. Three major kinds of asci are observed both in crushed perithecia and in discharged octads: 8 black, 4 black:4 nonblack
and 8 nonblack. These will be symbolized as 8b:Oa, 4b:4a and Ob:8a, where b
is used to signify black, and a (abortive) is used, for convenience, to signify all defective ascospores, including those that are underpigmented as well as those that are actually abortive. In the first ascus type, 8b:Oa, which is the product of alternate disjunction (Figure 1 b), most of the spores are able to germinate and give rise to viable progeny. In the second type, 4b:4a, spores are almost always arranged in bbbbaaaa sequence. Abortive spores never germinate, and only a fraction of black spores germinate (from 10 to 90% according to the type-I11 translocation studied). Moreover, for each viable ascus, the four black spores are either of wild genotype or of translocation genotype. As shown in Figure l b , a 3:l disjunction from the quadrivalent explains the occurrence of such type of ascus. T h e third type of ascus, Ob:8a, can be clearly distinguished from young asci of type I, which lack pigment because they are immature.
Immature asci of type I show homogeneous underpigmented or poorly devel-
oped spores, whereas in asci of the third type, Ob:8a, two spore phenotypes can be frequently distinguished, with arrangement alalalala2a2a2a2. More- over, Ob:8a asci are observed as a major class when ascospores are examined as discharged octads. Ascospores from immature type-I asci are not projected from the perithecia in crosses that are not heterozygous for a chromosome rearrangement. They are not expected to be projected when a translocation is present. This third type of ascus is interpreted as the product of adjacent 1 (or 2) disjunctions, and a1 and a2 are interpreted as reciprocally duplicate- deficient spores (Figure 1 b).
186 G . LEBLON, D. ZICKLER AND S. LEBILCOT
S E G R E G A T I O N O F T H E Q U A D R I V A L E N T
A L T E R N A T E
A D J A C E N T 1
/”
\
/”
A S C U S C O N 8 T I T U T I O N
T O (a)
0
/
7 - 0 0\
N :N :
S E O R E O A T I O N O F T H E Q U A D R I V A L E N T
A S C U S
C O N 8 T I T U T I O N
A D J A C E N T 1
/
\
DuDe
8
Du De
8
De Du 8
De Du 8
e
e
0
N + l 0
0
1
N + l 00
\
T-1 0 0T-1 0
:,I
:ef
D I S J U N C T I O N I ,I , I , I , 0 ,
FIGURE lb.
of asci flattened between slide and cover slip under the compound microscope than in discharged octads under a dissecting microscope. Third, preferential discharge of particular ascus types could have occurred.
Despite these discrepancies, an excess of adjacent vs. alternate disjunction is regularly observed (Table 3). A second observation, clearly shown in Table 1, is that asci diagnostic of interstitial crossing over (Figure la) were never ob- served. Similar results were found with the 67 other type-111 translocations.
T h e observations concerning the type-€11 translocations indicate that both breakpoints always segregate at the first division. These observations were complemented by genetic analysis of 522 asci originating from alternate dis- junction from crosses between a multiply marked normal strain and each of
23 type-111 translocations (Table 4). First-division segregation was systemati- cally observed for the breakpoints treated as a mutant allele with a phenotype of abortive spores in crosses to the normal strain, whereas only 95 first-division
segregation for the breakpoint was observed among 221 such asci from crosses
188 G . LEBLON, D. ZICKLER AND S. LEBILCOT
C O - 0 0 - 0
- 0 0 - 0 0
h se
-
c 0 0 m e - - vh se
* c v
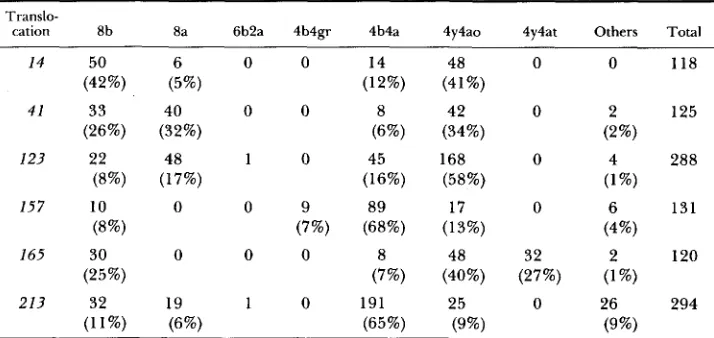
TABLE 2
Ascus patterns from discharged octads from crosses between type-111 translocation and normal sequence strains
Translo-
cation 8b 8a 6b2a 4b4gr 4b4a 4y4ao 4y4at Others Total
14 50 6 0 0 14 48 0 0 118
(42%) (5%) (12%) (41%)
4 1 33 40 0 0 8 42 0 2 125
(26%) (32%) (6%) (34%) (2%)
(8%) (17%) (16%) (58%) (1 %)
(8%) (7%) (68%) (13%) (4%)
123 22 48 1 0 45 168 0 4 288
157 10 0 0 9 89 17 0 6 131
165 30 0 0 0 8 48 32 2 120
(25%) (7%) (40%) (27%) (1%)
213 32 19 1 0 191 25 0 26 294
(11%) (6%) (65%) (9%) (9%)
Abbreviations: b = black wild-type; a = abortive; gr = gray; y = yellow; a0 = abortive opaque; at = abortive translucent.
TABLE 3
Ascus patterns from crosses between type-111 translocation and normal sequence strains
Percentage of 8bOa:Obaa:4b4a
Translocation Asci within perithecia Discharged octads
14 9:77:11 42:46: 12
4 1 10:79:7 26:66:6
I23 14:68:15 8:75: 16
157 14:52:31 8:13:75
I65 12:72: 12 25:67:7
213 11:62:24 11:15:65
In crosses between type-111 translocations and multiply marked normal strains, the frequency of second-division segregation of genetic markers present in the multiply marked strains and linked to the breakpoints (Table 5 ) suggests that the absence of second-division segregation for both breakpoints in type- I11 translocations is the consequence of their location very near to or at their respective centromeres and not the consequence of an inhibition of recombi- nation in the interstitial centromere-breakpoint intervals. This has been con- firmed by the fact that, for several type-I11 translocations, genetic markers located on either side of a centromere linked to the breakpoints showed a quasinormal level of second-division segregation (Table 5).
190
Second: first Tr;tnslocation division
Second: first Translocation division
30 0:2 1
40 0:24
41 0:33
44 0:30
46 0:35
4Y 0:24
104 0:19
107 0:36
116 0:18
I 1 9 0:13
I20 0:26
121 0:42
127 0:23
139 0:23
157 0:22
I 6 5 0:lO
181 0:25
191 0:14
206 0:18
210 0:12
211 0:12
213 0:14
215 0:28
between type-I11 translocations and multiply marked strains. Thus, in a first
experiment, chromosomes were identified for three type-I11 translocations (T41
and T46, Table 6; TI4, Table
7).
Next, a self-sterile cross-fertile mutation( s p l l or $55) was crossed into each of the three translocations in order to avoid selfing. Then, the three resulting double mutant strains (TI4 s p l l , T41 $55 and T46 $155) were crossed to the 70 other type-111 translocations and the hybrid perithecia were dissected. Among the 70 translocations, 20 were distinguished in which all asci from one of the three crosses contained only wild-type spores, indicating identity of both breakpoints or, at least, that they do not differ by so much as a single gene locus required for ascospore mat- uration (Table 8). T h e other 50 translocations produced hybrid asci containing mainly aborted spores in the three crosses. The chromosomes interchanged within three among the 50 translocations (T123, TI57 and T213) were iden- tified by crosses to strains containing centromeric markers (Tables 6 and
7).
A self-sterile cross-fertile mutation ($055) was crossed into each of the three
translocations, then the three resulting double mutant strains (TI23 sp55,
TI57
sp55 and T213 sp55) were crossed to the 47 other type-I11 translocations. Nineteen of them showed identical breakpoints as either T123, T157 or T213
(Table 8). Thus, the chromosomes interchanged within the 73 translocations
were identified by six successive steps involving both linkage mapping of the breakpoints by crosses to normal sequence strains and research of identity of the breakpoints in pairwise crosses between translocations. Results are sum- marized in Table 9.
T h e apparent identity of both breakpoints in several translocations has been confirmed, either by pairwise crosses between translocations each of which is associated with a different complementing self-sterile mutation (e.g., TI04 sp55
TABLE 5
Percentage of seconddivision segregation of the centromere markers linked to the
breakpoints in crosses of type-I11 translocation X multiply marked normal sequence strain
Second/(first plus second) division of the marker in cross
Segregating Chromosome Translocation X
Segregating translocation marker involved normal Normal X normal
T104, T107, T116, T119, T120, T121, T139, TI81 or T215 T30, T46, T157, T165, or T215 T40, T119, TIZI, T127, T139,
TI57 or T213 T4 1
T40, T41, T181, T191, T206, T211
or T213
T41, T49, TI16 or T I 9 1
T14, T44, T192, T195, T207 or
T210
T14, T30, T44, T46, T49, T104, T107, T120, T123, T127, T165, T192, T195, T206, T207, T210
or T211 T(1,7)120
T(1,7)120
T(4,5}19 I
T(4,5)19 1
sgr I
asyl
sgn2
per1 Pam2
arol pam I
tsgr
tYP 1
adel
thrl
arol
I 5/95 (5%) 5/103 (5%)
2 16/97 (16%) 11/52 (21%)
3 27/96 (28%) 7/22 (32%)
4 1/22 (5%) 2/54 (4%)
4 31/68 (46%) 77/111 (69%)
5 2/84 (2%) 8/119 (7%)
6 1/118 (1%) 8/848 (1%)
7 3/273 (1%) 2/149 (1%)
I 7/20 (35%) 20/67 (30%)
1 1/9 ( 1 1%) 21/33 (64%)
5 10/15 (67%) 10/41 (24%)
5 2/12 (17%) 8/119 (7%) (right arm)
(left arm)
(right arm)
(left arm)
quence strains (e.g.,
TI23
andT211;
T14,
T192,
T195,
T207
andT2IO;
T30
and
T165;
T119
andT139;
TI04
andT120;
Tables 6 and7).
We are confident that type-111 translocations within a group all originated separately from independent repetitions of similar genetic events. Evidence for independent origins comes, first, from the fact that each type-111 translocation originated from a different irradiated plate and, second, from the fact that type-111 translocations within the same group frequently include representatives from two distinct mutageneses, in each of which all irradiated material origi- nated from a different, newly isolated haploid wild-type nucleus (for example
T20
andT117;
TI4
andT102;
T44
andT112,
Table 9).192 G. LEBLON, D . ZICKLER A N D S. LEBILCOT TABLE 6
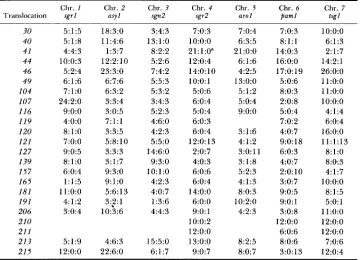
Linkage relationships between the breakpoints and the centromeric markers
Chr. 1 Chr. 2 Chr. 3 Chr. 4 Chr. 5 Chr. 6 Chr. 7 Translocation sgrl asyl sgn* sgr2 arol paml tsgl
30 40 41 44 46 49 104 107 116 119 120 121 127 139 157 165 181 191 206 210 21 1 213 215 5:1:5 5:1:8 4:4:3 10:0:3 5:2:4 6:1:6 7:l:O 24:2:0 9:o:o 4:O:O 8:l:O 7:O:O 9:0:5 8:l:O 6:0:4 1:1:5 11:o:o
4: 1 :2 3:0:4 5:1:9 12:o:o 18:3:0 11:4:6 1:3:7 12:2:10 23:3:0 6:7:6 6:3:2 3:3:4 3:0:5 7:l:l 3:3:5 5:8:10 3:3:3 3:1:7 9:3:0 9:l:O 5:6: 13
3:?:1 10:3:6 4:6:3 22:6:0 3:4:3 13:l:O 8:2:2 5:2:6 7:4:2
5 : 5 : 3
5 : 3 : 2 3:4:3
5 : 2 : 3
4:6:0 4:2:3 5:5:0 14:6:0 9:3:0 1 O : l : O 4:2:3 4:0:7 1:3:6 4:4:3
15:5:0 6: 1 :7
7:0:3 1o:o:o 21: 1 :O" 12:0:4 14:O:lO 1o:o: 1
5:0:6 6:0:4 5:0:4 6:0:3 6:0:4
120: 13
2:0:7 4:0:3 6:0:6 6:0:4 14:O:O 6:O:O 9:o: 1 10:0:2 12:o:o 13:O:O 9:0:7 7:0:4 6:3:5 21:o:o 6: 1 :6 4:2:5 13:O:O 5:1:2 5:0:4 9:o:o 3:1:6 4:1:2 3:O:ll 3: 1 :8 5:2:3 4:1:3 8:0:3 10:2:0 4:2:3 8:2:5 8:0:7
7:0:3 1o:o:o 8 : l : l 6:1:3 14:0:3 2:1:7 16:O:O 14:2:1 17:O: 19 26:O:O
5:0:6 11:o:o
8:0:3 11:o:o
2:0:8 1o:o:o 5:0:4 4: 1 :4 7:0:2 6:0:4 4:0:7 16:O:O 9:0:18 11:1:13
6:0:3 8:1:0
4:0:7 8:0:3 2:O:lO 4:1:7 3:0:7 1O:O:O 9:0:5 8:1:5 9:o: 1 5:O: 1 3:0:8 11:o:o
12:o:o 12:o:o
6:0:6 12:O:O 8:0:6 7:0:6 3:0:13 12:0:4
~~~~
Asci with eight viable spores were analyzed from crosses of the type-I11 rearrangement X
multiply marked Normal sequence strain. For each entry, the three values represent parental ditype, tetratype and nonparental ditype between the breakpoint and the corresponding centro- meric marker. Chr. = Chromosome.
T h e chromosome 4 centromere-linked marker used was per1 and not sgr2.
TABLE 7
New linkage relationships between centromeric markers
~~ Markers showing Interchanged Translocation new linkage PD T NPD chromosomes
14 paml tsgl 38:O:O 6, 7 44 paml tsgl 18:l:O 6, 7
121 sgrl sgn2 15:5:0 1, 3 123 tsgl sgr2 12:o:o 4, 7 191 sgr2 arol 9:6:0 4, 5 191 yasl arol 10:4:0 4, 5 192 paml tsgl 12:o:o 6, 7 I95 paml tsgl 9:o:o 6, 7 206 sgr2 tsgl 32:l:O 4, 7 207 paml tsgl 12:o:o 6, 7
Asci with eight viable spores were analyzed from crosses of the type-111 translocation X multiply marked normal sequence strain.
% % %
+ 2 z
n m + z
+ % 2
% + 2
+ % 2
+ % %
+ % %
% % +
+ E 2
% + %
+ % z
+ % % % + % + % % % + % + % % % + % % + % + % %
2 + 2
I * *
- * *
- + - U
2 - 3 2
2 z z
6
3
z
% 1 1 % I\ 9 2 2 3 2 4 a “i x 5 1 a h 1 1 1 I\ 1 1 a Pz
1 P 1 a 2 3 1 h 1 1 -5 I I194 G. LEBLON, D. ZICKLER AND S. LEBILCOT
T A B L E 9
Type-I11 translocations listed by interchanged chromosomes
Chronio-
sonies Translocations
1;5 116 1;7 107
1;4 181 1;3 I19 (= 139)
121 (= 194) 1;2 215
5;7
5;4 22 (= 191, 204)
193, 208, 212) 104 (= 120)
49 (= 132, 147, 200)
4 1 (= 45, 113, 136, 163, 179,
Chrorno- somes
6; 7
7;4
7; 3 7;2
4 9
Translocations
4 4 (= 112, 152, 155)
14 (= 102, 146, 168, 189, I92 195, 201, 202, 207, 210, 218) 206
20 (= 117, 123, 131, 138, 1 4 1 , 1 4 4 , 161, 171, 176, 178, 180, 182, 185, 188, 197, 211, 214) 103 (= 127)
46 (= 199)
30 (= 162, 165) 40
25 (= 213) 3;2 1 1 1 (= 157)
Translocations in parentheses appear to be identical to the translocation after which they are listed.
frequently involved (49 of the 73), whereas chromosome 6 is only interchanged
with chromosome 7.
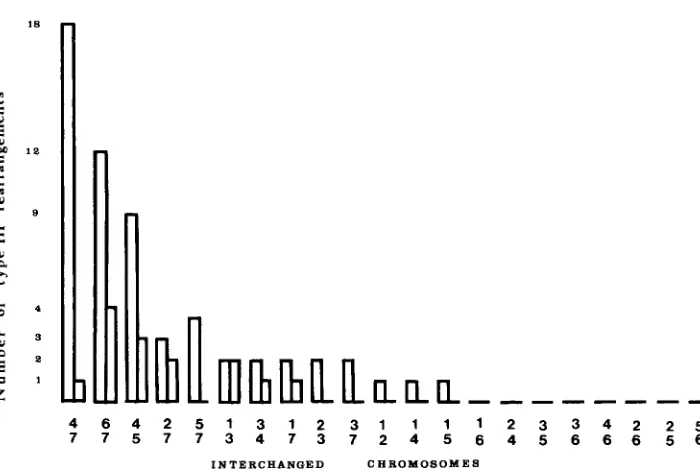
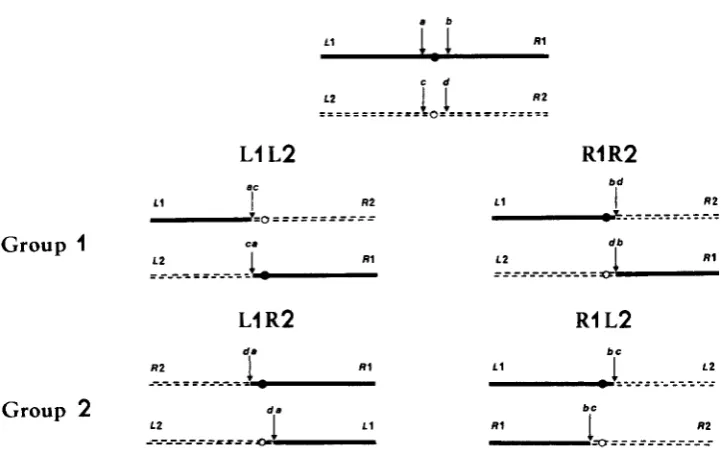
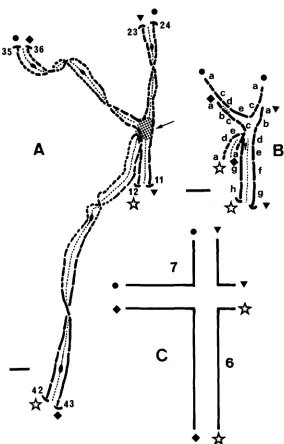
Translocations interchanging the same chromosome pair fall into only two groups of apparently isosequential rearrangements (Figure
2).
All crosses be- tween groups lead to high spore abortion, whereas all crosses between trans- locations in the same group lead to wild-type progeny. This result is expected if breaks leading to type-I11 translocations are adjacent to centromeres (Figure3). In this case, the two groups of apparently isosequential translocations cor- respond to the different possibilities for interchanging arms between the same two chromosomes, i.e., for one group, left arm of the first chromosome with left arm of the second ( L l L 2 , Figure 3 ) or right arm of the first with right arm of the second ( R l R 2 , Figure 3 ) , and for the second group, left arm of
the first with right arm of the second ( L l R 2 , Figure 3 ) or right arm of the first with left arm of the second ( R l L 2 , Figure 3 ) .
In order to know if the two groups of isosequential type-111 translocations correspond to the different possibilities of exchanging arms between the same two chromosomes, several crosses were made between two strains each carrying the same translocation but marked with two centromere linked genes located
on either side of the breakpoint on the same chromosome, or on different chromosomes involved in the translocation. T h e new linkage relationships were determined in the case of T(I,7)107 and T(1,7)120, each belonging to a distinct
group of isosequential rearrangements and involving chromosomes 1 and 7
4 6 4 2
7 7 5 7 57 13 34 71 23 37 12 41 51 16 42 53 63 46 62 52 56
I N T E R C H A N G E D C H R O Y O B O Y E B
FIGURE 2.-Distribution of the breakpoints on the seven chromosomes. T w o distinct groups of
apparently isosequential translocations were found, and each is represented by a distinct bar. If
the letters L and R represent left and right arms of the first and the second interchanged chro- mosomes, then one bar represents translocations L l L 2 and R l R 2 and the other R l L 2 and L l R 2 (see Figure 3 ) .
of isosequential rearrangements and involving chromosomes 4 and 5 . Results are shown in Table 10. If the letters L and R represent left and right arms of the linkage groups, then the constitution of T(1,7)107 is R l . R 7 and L l . L 7 ,
whereas T(1,7)120 is Ll.R7 and Rl.L7. Similarly, the constitution of T(4,5)41
is R4.R5 and L4.L5, whereas T(4,5)191 is R4.L5 and L4.R5.
Cytological mapping of the translocation: In Sordaria, the seven bivalents can be identified by their length. Central and lateral elements of the synap- tonemal complex (SC) in six type-I11 translocation/wild-type heterozygotes were reconstructed from serially sectioned zygotene and pachytene nuclei. Bi- valent SC lengths compared to the lengths of SC involved in the quadrivalent
arms permitted the identification of the interchanged chromosomes (ZICKLER
et al. 1984). T h e interchange regions were defined by the relative position of exchange of the lateral element pairing partners. In all type-I11 translocations
sharing a common chromosome (e.g., T(2,3)157 and T(2,7)165 for chromosome
2 ) , the interchange on this common chromosome was located in the same region in both translocations. These similarities in breakpoint localization sug-
gest a small interchange region on each chromosome as the origin of the type-
111 translocations.
196 G. LEBLON, D. ZICKLER AND S. LEBILCOT
a b
L1
1.1
R lL1 L2
Group
1
1 2
ca
I R1
R1 R 2
L1
R 2
RI L2
L2
Group
2
L1 R1
9"
R2+XI == = =====----= --
- - - -
-----,,,,-
a--
,-
FIGURE 3.-Consequence of breaks adjacent to centromeres on the constitution of two appar- ently isosequential groups of translocations. Segments originally in one of the wild-type chromo- somes are shown as solid lines with black centromeres, those in the other wild-type chromosome as dotted lines with white centromeres. In the progeny of crosses between translocations in the same group, segregants with two black or two white centromeres are possible and inferentially viable. In the progeny of crosses between groups, a spore abortion pattern similar to that observed from crosses of a type-I11 translocation X wild type is expected. LI = left arm of the first chromosome; R I = right arm of the first chromosome.
visualized as regions of differentiated SC structure were systematically inves- tigated. In all cases, a single dense structure associated with the synaptonemal complex was detected either in all or in some of the bivalents (Figures 4 and 5). T h e position of this structure corresponds to the position of the centromere when metaphases (especially clear for the four longest chromosomes during postmeiotic mitoses) are examined by light microscopy (ZICKLER et al. 1984).
These dense structures and the relative position of interchange in the trans- location heterozygotes are at corresponding positions along the chromosomes (Table 11). In three nuclei from 7'(6,7)14 heterozygotes, the region of ex- change seen at the lateral element level appeared to be in the centromere structures of chromosomes 6 and 7 (Figures 6 and
7).
DISCUSSION
Cytological evidence: The three-dimensional reconstructions of 28 nuclei heterozygous for reciprocal translocations confirm and extend the earlier ob- servations that, within the quadrivalents, measurement of arm lengths using synaptonemal complex lateral elements is sufficiently precise to allow localiza- tion of breakpoints in species, like fungi, with small chromosomes (GILLIES
TABLE 10
197
New linkage relationships between different arm markers in crosses homozygous for a type-
I11 translocation; translocations tested are members of the two different isosequential groups
for that pair of translocated chromosomes
~~
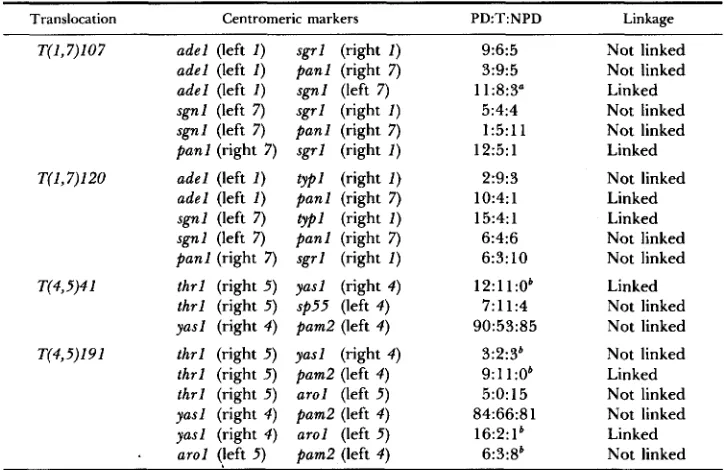
Translocation Centromeric markers PD:T:NPD Linkage
~~ ~
T(1,7)107 adel (left I )
adel (left 1 ) adel (left I ) sgnl (left 7 ) sgnl (left 7)
panl (right 7 ) T(I,7)120 adel (left I )
adel (left I )
sgnl (left 7 ) sgnl (left 7 ) panl (right 7 ) T(4,5)4 1 thrl (right 5 )
thrl (right 5 ) yasl (right 4 ) T(4,5)19 1 thrl (right 5 ) thrl (right 5 ) thrl (right 5 ) yasl (right 4 ) yasl (right 4 )
.
a r o l (left 5 )~
sgrl (right 1 )
panl (right 7 ) sgnl (left 7 )
sgrl (right I )
panl (right 7 )
sgrl (right I )
typl (right I )
panl (right 7 ) typl (right I ) panl (right 7)
sgrl (right I)
yasl (right 4 )
$55 (left 4 ) pam2 (left 4 ) yasl (right 4 ) Pam2 (left 4 ) arol (left 5 ) pamZ (left 4 ) arol (left 5 ) bum2 (left 4 )
9:6:5 3:9:5 11:8:3" 5:4:4 1:5:11 12:5: 1 2:9:3 10:4: 1 15:4: 1 6:4:6 6:3:10 12:l 1:Ob 7:11:4 90:53:85 3:2:3' 9:ll:Ob 5:0:15 84:66:81 16:2:lb 6:3:8' ~~
Not linked Not linked Linked Not linked Not linked Linked
Not linked Linked Linked Not linked Not linked
Linked Not linked Not linked
Not linked Linked Not linked Not linked Linked Not linked
Asci were analyzed from crosses between two parents containing the same type-I11 translocation but t w o different centromeric markers. Left I = the corresponding centromeric marker is located on the left arm of chromosome 1 . PD = parental ditype; T = tetratype; NPD = nonparental ditype.
For the three NPD asci, a second-division segregation of the two markers was observed. Asci with first-division segregation of the color marker were selected before analysis.
Somewhat differentiated structure in each pachytene bivalent consisting of condensed fibers within (and sometimes surrounding) the central region of the
synaptonemal complex has been described in several organisms (RASMUSSEN
and HOLM 1980). This structure is believed to mark the centromere. In fungi, these putative centromere regions have, so far, been found mainly in Basidi- omycetes (CARMI et al. 1 9 7 8 ; HOLM et al. 1981; SLEZEC 1981), and they are seen only during zygotene and early pachytene. In S. macrospora, such struc- tures are found during mid-zygotene, and they are preserved up to late zy- gotene only if their corresponding chromosomes are entrapped within another bivalent as pairing progresses (interlocking). As in the other cases, the likeli- hood that these dense structures correspond to centromeres is supported by the facts that only one is found per bivalent and that 'the location of the structure on its chromosome remains constant in different nuclei. Moreover, the structures identified by electron microscopy are in the same region of their respective chromosomes as are the centromeres seen at mitotic metaphase by light cytology (ZICKLER et al. 1984).
198 G. LEBLON, D. ZICKLER AND S. LEBILCOT
s-
1 -
-
a4
- s -
-
7
-4-
-
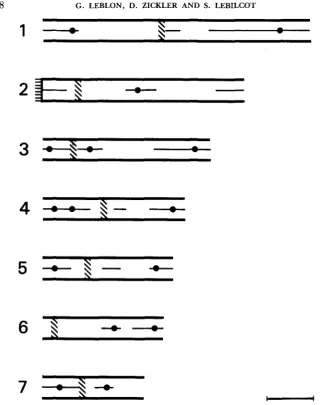
FIGURE 4.-Location of centromeric regions (hatched) in the seven bivalents of a mid-zygotene
nucleus. Horizontal lines at one end of bivalent 2 indicate its termination in the nucleolus. Central element initiation sites and recombination nodules (black circles on the central element) are also
shown. Bar = 2 pm.
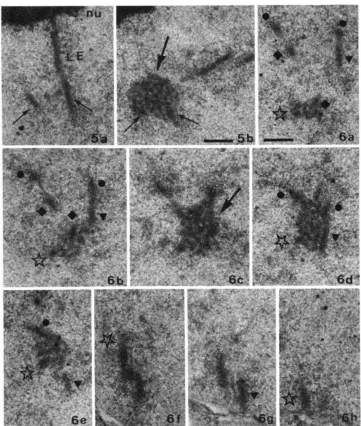
I ' I ( . ~ . R E . .-).-;I imd I). .r\vo (on\cc I I I I \ V V Y iioll\ i n ;I ~ v g o i c ~ I I U C I C U ~ showirlg tllc d t ~ - ; ~ t i r ~ ~ ( I I I I ' C
of the ccniromeric region near the nucleolar organizer of' chromosome 2. In a . one of the almost paired lateral elements (LE) is associated with the nucleolus (nu). In b, dense material (large arrow) fills and surrounds the central region formed between the two homologous lateral elements (small
arrows). Bar = 0.3 pm.
FIGURE 6.-a-h, Eight serial sections of a heterozygous T(6,7)14 pachytene nucleus. Sections 6a-e cross the centromeric region (arrow in c) that corresponds to the exchange region of bivalents 6 and 7. T h e four chromosomes involved in the translocation quadrivalent are marked by four different symbols. Rar = 0.3 pni.
200 G . LEBLON, D. ZICKLER AND S. LEBILCOT
2
2
Qi
I
Qi
2
Go
M
.E I
k
.I2
09r-
*
0 '?*
E.* I .0 I
B
= B
w p "
U:
0
4
0
0.I 4
9
.I
U
B 5 2 2 - 2 2 x x x x x
2 2 2 2 2
x x x x x
X
Ih h
A
20
0
0
B
*
+ *
FIGURE 7.-A, Complete reconstruction of the quadrivalent partially shown in Figure 6. The arrow points to the centromere region. Numbers indicate the section and terminal bars indicate the telomere attachment to the nuclear envelope. B, Interpretative drawing through the region of pairing exchange corresponding to the eight sections of Figure 6 . The sections (6a-h) are denoted by their corresponding letters, and the four chromosomes involved in the exchange have the same symbols as in Figure 6. C, Translocation quadrivalent corresponding to the reconstruction in A. The wild-type chromosomes 6 and 7 are located in opposite configuration. Bar = 0.3 pm.
this is a characteristic of breakpoints close to the centromeres (BURNHAM 1956,
202 G . LEBLON, D. ZICKLER AND S. LEBILCOT
is expected either for translocations having both breakpoints near centromere or for translocations having only one breakpoint near centromere.
High incidence of type-I11 translocations: In Sordaria, two-thirds of UV- induced reciprocal translocations are of this type, with breakpoints at centro-
meres (ARNAISE, LEBLON and LARES 1984). T h e occurrence of chromosomal
interchanges in centromeric regions is also a characteristic of Robertsonian translocations, which represent a widespread chromosomal polymorphism in several species and have been especially studied in mice (GROPP et al. 1972;
ADOLPH and KLEIN 1983). In man, the frequent occurrence of mitomycin-C-
induced damage in centromeric regions has been interpreted as due to inter-
changes promoted in regions of repetitive DNA (BOURGEOIS 1974). In asco-
mycetes where no such regions of repetitive DNA have been reported, some other explanation must be proposed.
Molecular studies of centromeres are now in progress in fungi. Four yeast
centromeres have been cloned, and their sequences are known (BLACKBURN
and SZOSTAK 1984). Comparisons of their sequences reveal a core region of 82-89 base pairs (region 11) composed of extremely AT-rich DNA (93-94%) with limited or no homology among the four centromeres. Moreover, this core region is flanked by two short conserved sequences (regions I and
HI),
one of which (region 111) is composed of 11 highly conserved base pairs. Both regions I1 and 111 are indispensable for centromere function. This organization and the fact that mitotic crossing over has repeatedly been shown to be clusterednear the centromere (MINET, GROSSENBACHER-GRUNDER and THURIAUX 1980;
MALONE, GOLIN and ESPOSITO 1980) suggest an attractive hypothesis for ex-
plaining the high incidence of UV-induced reciprocal translocations in centro- meric regions in S . macrospora. Photoreactivation showed that reciprocal trans-
locations are due mainly to formation of pyrimidine dimers (ARNAISE, LEBLON
and LARES 1984). If a core region exists in Sordaria similar to that in yeast, a high concentration of pyrimidine dimers could be formed in it (e.g., 47 possi-
bilities in yeast CEN 3). Thymine-thymine dimers are formed more readily
than cytosine-thymine or cytosine-cytosine dimers (see WITKIN 1969). If repair is associated with mitotic crossing over, as suggested by studies of FABRE (1972) in Schizosaccharomyces pombe, crossing over could occur in the small homologous sequences flanking the core region. When flanking regions belonging to two nonhomologous chromosomes recombine, reciprocal translocations are gener- ated. Recombination between sequences in nonhomologous positions is able to generate reciprocal translocations in yeast (SUGAWARA and SZOSTAK 1983).
203
half only interchanges that were outside the strict centromere region, left or right of region I1 and 111, could lead to viable reciprocal translocations.
We are much indebted to DAVID PERKINS for helpful discussions and for critical reading of the manuscript. We should like to thank JACQUES LAVAL and ALAIN NICOLAS for their suggestions, ALMUTH COLLARD for her technical assistance, MICHELLE DAHURON for typing the manuscript and the students of the DiplGme d’Etudes Approfondies d e Ginitique for the interest they took in this work. This investigation was supported by the “Centre National d e la Recherche Scientifique (L.A. no. 040086)” and the University of Paris X1.
LITERATURE CITED
AWLPH, S. and J. KLEIN, 1983
ARNAISE, S., G. LEBLON and L. LARES, 1984
Genetic variation of wild mouse populations in southern Ger-
A system for the detection of chromosomal rear-
T h e molecular structure of centromeres and telo- many. I. Cytogenetic study. Genet. Res. 41: 117-134.
rangements using Sordaria macrosflora. Mutat. Res. 125: 33-42.
meres. Annu. Rev. Biochem. 53: 163-194.
BLACKBURN, E. H. and J. W. SZOSTAK, 1984
BOURGEOIS, C. A., 1974 Distribution of Mitomycin C-induced damage in human chromosomes with special reference to regions of repetitive DNA. Chromosoma 48: 36 1-366.
BURNHAM, C. R., 1956
CARMI, P., P. B. HOLM, Y. KOLTIN, S. W. RASMUSSEN, J. SAGE and D. ZICKLER, 1978
Chromosomal interchanges in plants. Bot. Rev. 22: 419-552.
T h e pachytene karyotype of Schizophyllum commune analyzed by three-dimensional reconstructions of synaptonemal complexes. Carlsberg Res. Commun. 43: 1 1 7-132.
lntrachromosomal intergenic recombination. pp. 148-153. In:
Genetics of Fungi, Springer-Verlag, Berlin.
Relation between repair mechanisms and induced mitotic recombination after UV irradiation in the yeast Schizosaccharomyces pombe. Mol. Gen. Genet. 117: 153-166.
ESSER, K. and R. KUENEN, 1967
FABRE, F., 1972
GILLIES, C. B., 1979 T h e relationship between synaptonemal complexes, recombination nodules and crossing-over in Neurospora crassa bivalents and translocation quadrivalents. Genetics 91: 1-17.
GOSALVEZ, J., C. LOPEZ-FERNANDEZ and R. GARCIA-LAFUENTE, 1 982 A spontaneous translocation heterozygote involving centromere regions in Gomphocerus sibiricus (L.). Chromosoma 86: 49- 57.
Robertsonian chromosomal variation GROPP, A., H. WINKING, L. ZECH and H. MULLER, 1972
and identification of metacentric chromosomes in feral mice. Chromosoma 30: 265-288.
HOLM, P. B., S. W. RASMUSSEN, D. ZICKLER, B. C. Lu and J. SAGE, 1981 Chromosome pairing, recombination nodules and chiasma formation in the basidiomycete Coprinus cinereus. Carls- berg Res. Commun. 4 6 305-346.
Meiotic and mitotic recombination in Aspergillus and its chromosomal aberra- tions. Adv. Genet. 1 9 33-131.
Mitotic versus meiotic recombination in KAFER, E., 1977
MALONE, R. E., J. E. COLIN and M. S. ESPOSITO, 1980
Saccharomyces cerevisiae. Curr. Genet. 1: 241-248.
MINET, M., A.-M. GROSSENBACHER-GRUNDER and P. THURIAUX, 1980 T h e origin of a centromere effect on mitotic recombination. Curr. Genet. 2: 53-60.
204 G. LEBLON, D. ZICKLER AND S. LEBILCOT
PERKINS, D. D., 1974 T h e manifestation of chromosome rearrangements in unordered asci of Neurospora. Genetics 77: 459-489.
PERKINS, D. D. and E. G. BARRY, 1977 T h e cytogenetics of Neurospora. Adv. Genet. 1 9 133- 285.
RASMUSSEN, S. W . and P. B. HOLM, 1980
SLEZEC, A. M., 1981
Mechanics of meiosis. Hereditas 93: 187-216.
Mkiose d e I’agaricale Pleurotus eryngi (Dcex Fr.) Quel: description et dPter- mination du nombre chromosomique par reconstitution tridimensionnelle des complexes syn- aptonkmatiques.
c.
R. Seances Acad. Sci. [III] 292: 523-528.Recombination between sequences in nonhomologous positions. Proc. Natl. Acad. Sci. USA 80: 5675-5679.
Ultraviolet-induced mutation and DNA repair. Annu. Rev. Genet. 3: 525- 552.
Development of the synaptonemal complex and the “recombination nodules” during meiotic prophase in the seven bivalents of the fungus Sordaria macrospora Auersw. Chromosoma 61: 289-316.
Linkage group- chromosome correlations in Sordaria macrosflora: chromosome identification by three-dimen- sional reconstruction of their synaptonemal complex. Curr. Genet. 8: 57-67.
SUGAWARA, N. and J. W . SZOSTAK, 1983
WITKIN, E. M., 1969
ZICKLER, D., 1977
ZICKLER, D., G. LEBLON, V. HAEDENS, A. COLLARD and P. THURIAUX, 1984