Copyright © 1998, American Society for Microbiology. All Rights Reserved.
Extracellular and Cytosolic Iron Superoxide Dismutase from
Mycobacterium bovis BCG
SUNG-KOO KANG,
1YONG-JAE JUNG,
1CHEORL-HO KIM,
2ANDCHUL-YONG SONG
1*
Department of Biology, Chung-Ang University, Dongjak-ku, Seoul 156-756,
1and College of Oriental Medicine,
Dongguk University, Kyoungju City, Kyungpook 780-714,
2Korea
Received 6 March 1998/Returned for modification 24 June 1998/Accepted 1 September 1998
Two forms of iron superoxide dismutase (SOD) were purified from cell extract (CE) and culture filtrate (CF)
of Mycobacterium bovis BCG, respectively. The molecular weight of both enzymes was estimated to be
approx-imately 84,000 by gel filtration, whereas that of their subunits was 21,500, as determined by sodium dodecyl
sulfate-polyacrylamide gel electrophoresis, indicating that each of purified enzymes is composed of four
identical subunits. The specific activities of CE SOD and CF SOD were 3,850 and 4,040, respectively. The
purified enzymes were not joined by disulfide bonds and were, to some extent, resistant to sodium dodecyl
sulfate. Their activities were lost by H
2O
2, but not by azide and cyanide, indicating iron SODs. Enzyme
activities were detectable over a broad range of pHs, from 5.0 to 9.0, and were stable for 6 months at
2
20°C.
Each value of pI was 4.5. In Western blots, both enzymes reacted with sera of tuberculosis patients, but not with
normal sera. The N-terminal amino acid sequences of CE SOD and CF SOD were the same, suggesting that
there is no N-terminal signal sequence.
Superoxide dismutases (SODs), which catalytically scavenge
the superoxide radical (O2
2) to hydrogen peroxide and
mo-lecular oxygen, serve a protective role against oxygen toxicity in
all aerobic organisms (13, 37). Because the superoxide radical
is a normal product of the univalent reduction of molecular
oxygen, SODs are thought to be the primary defense against its
potential cytotoxicity (13). There are three common forms of
SODs, which differ in the metal ion cofactor at the active site.
Manganese-containing SODs (MnSODs) are found in bacteria
and mitochondria, while iron-containing SODs (FeSODs) are
mainly found in the cytosol of prokaryotes, in primitive
eu-karyotes, and in some green plants (4, 5). In contrast,
copper-zinc SODs (Cu-ZnSODs) are mostly found in the cytosol of
eukaryotes, surprisingly in an increasing although limited
num-ber of bacteria (10, 18).
Until now, numerous papers have reported the distribution,
characterization, and biological significance of SODs in
myco-bacteria (17, 19, 23). However, only a few myco-bacteria have been
shown to secrete SOD during growth. It was shown previously
that Mycobacterium tuberculosis, a human pathogen, secreted
iron SOD, while the nonpathogenic species, Mycobacterium
smegmatis and Mycobacterium phlei, did not (19). Similarly, it
was shown previously that the virulent strain Nocardia
aster-oides GUH-2 secretes the enzyme into the growth medium (7).
SODs could also be involved in the pathogenicity of
Mycobac-terium leprae (33), although the secretion of the enzyme has
not been demonstrated and remains controversial.
Members of the mycobacteria have emerged as major
op-portunistic pathogens in humans, with the advent of the AIDS
epidemic (16). Recently, such a case was also reported in a
human immunodeficiency virus-infected individual who
dis-played reactivation of Mycobacterium bovis BCG 30 years after
vaccination (3), although M. bovis BCG has been used in many
countries for vaccination to prevent tuberculosis. BCG
vacci-nation may cause disseminated mycobacterial infection, an
ill-ness caused by the vaccine itself in patients suffering from
severe immune deficiency.
In this study, we explored the production and secretion of
SOD by M. bovis BCG by investigating the purification and
some properties of M. bovis BCG SOD along with evidence of
the association of this enzyme with the cytosol and the growth
medium of the organism.
MATERIALS AND METHODS
Bacterial strain and culture condition.The M. bovis BCG strain used in this study was Pasteur strain 1173P2 and was obtained from the Korea Institute of Tuberculosis. The bacterium was cultured at 37°C in Sauton’s medium (30) without Triton.
Preparation and fractionation of CEs and CFs.Cells were collected on What-man filter paper, washed three times with phosphate-buffered saline, and sus-pended with sterilized distilled water. After homogenization, cells were disrupted with a French press (Aminco, Rochester, N.Y.) at a pressure of 18,000 lb/in2and centrifuged at 15,000 rpm for 30 min at 4°C. Culture filtrates (CFs) were filtered through a 0.2-mm-pore-diameter membrane (Gelman, Ann Arbor, Mich.) and concentrated by an Amicon concentrator with a YM-10 membrane (Amicon, Beverly, Mass.). Each of the cell extracts (CEs) and CFs was precipitated by 70 to 90% ammonium sulfate, suspended with D.W., lyophilized, and stored at
220°C.
Enzyme assay and protein determination.The activity of the enzyme was assayed at 37°C for 30 min. The assay mixtures (0.5 ml) contained 50ml of 0.5 M potassium phosphate (pH 7.5), 25ml of 16% Triton X-100, 2.5ml of 10 mM EDTA, 75ml of 1.2 mM NTC, 2.5ml of xanthine oxidase (1.0 U), the sample, 25
ml of 4 mM hypoxanthine, and distilled water. The A540was monitored (Shi-madzu UV-240) after addition of 0.5 ml of a solution containing 1 M formate buffer (pH 3.5), 10% Triton X-100, and 40% formaldehyde. An enzymatic unit was defined as the amount of the enzyme required to cause a 50% inhibition in the rate of reduction of NTC under the assay conditions (27). The standard protein used in unit determination was SOD from bovine kidney (Sigma, St. Louis, Mo.).
The protein concentration was determined by the Lowry method (21) with bovine serum albumin as a standard.
Purification of CE SOD and CF SOD.Crude CE and CF extracts were dialyzed against 50 mM Tris-HCl (pH 8.0) containing 0.5 M ammonium sulfate, briefly centrifuged, and chromatographed on phenyl Sepharose 4B (1.6 by 15 cm [Sigma]) at room temperature in the same buffer. The gradient procedure was performed with D.W. at a flow rate of 30 ml/h. SOD fractions were dialyzed against D.W. (pH 7.0) and lyophilized (Labconco, Kansas City, Mo.). The ly-ophilized material was dissolved with 50 mM Tris-HCl (pH 7.5) containing 0.15 M NaCl prior to chromatography at a flow rate of 20 ml/h on a Sephacryl S-200 (Pharmacia, Uppsala, Sweden) gel filtration (1.6 by 85 cm) system calibrated with standard proteins (Pharmacia) at 4°C.
* Corresponding author. Mailing address: Department of Biology,
Chung-Ang University, 221, hueksuk-dong, Dongjak-ku, Seoul
156-756, Korea. Phone: 82 02 820 5208. Fax: 82 02 816 6710. E-mail: cysong
@cau.ac.kr.
784
on August 17, 2020 by guest
http://cvi.asm.org/
SDS-PAGE and native electrophoresis.Sodium dodecyl sulfate-polyacrylam-ide gel electrophoresis (SDS-PAGE) was performed as described previously (20) on a minigel comprising a 12.5% acrylamide continuous resolving gel, and native electrophoresis was carried out by the method of Bollag and Edelstein (11).
Enzyme activity staining following native electrophoresis.SOD activity was visualized on a polyacrylamide gel as described previously (9). The SOD-active area appeared as a clear zone on a blue-violet background.
Inhibition test.To test inhibition, each SOD (1mg) was preincubated with 3 mM cyanide, 20 mM azide, and 1 mM H2O2for 30 min and electrophoresed on 12.5% polyacrylamide gel prior to enzyme activity staining.
Enzyme stability and isoelectric focusing.After 12.5% polyacrylamide native gel electrophoresis, each lane was soaked for 12 h at pH 4.5 to 5.0 (sodium acetate buffer), 6.0 to 7.5 (sodium phosphate), 8.0 to 8.5 (Tris-HCl), and 9.0
(glycine-NaOH), and enzyme activity staining was carried out before the results were read at 550 nm in a densitometer (LKB, Bromma, Sweden). Isoelectric focusing was carried out automatically with Phastsystem (Pharmacia, Sweden).
N-terminal sequencing of the purified SOD. Approximately 20mg of the purified enzyme was applied to an SDS-PAGE gel. After electrophoresis, the protein was electrophoretically transferred to a polyvinylidene difluoride mem-brane. The area containing the band was cut out and subjected to N-terminal amino acid sequence analysis with a protein sequencer (Milligen 6600B).
Western blot analysis.Western blotting was carried out as described previ-ously (34). The sera used were from tuberculosis patients and were obtained from the Korea Institute of Tuberculosis.
RESULTS
Purification and properties of CE and CF SODs.
The SODs
were effectively purified by 70 to 90% ammonium sulfate
pre-cipitation and underwent hydrophobic interaction
chromatog-raphy with phenyl Sepharose CL-6B and Sephacryl S-200 gel
filtration. The results showed the SDS-PAGE pattern of
puri-fied CE SOD (20
m
g) and CF SOD (20
m
g) (Fig. 1a). A single
band with a molecular weight of about 21,500 was stained by
Coomassie brilliant blue. The specific activity of CE-SOD was
3,850 U/mg of protein, representing a purification of 30.1-fold
with 53.5% recovery. CF SOD was purified approximately
28.57-fold with a specific activity of 4,040 U/mg of protein and
29.1% recovery (Tables 1 and 2). The activities of purified
enzymes were stable at
2
20°C for 6 months (data not shown).
The molecular weights of the native enzymes, as determined by
gel filtration on Sephacryl S-200 (Fig. 2b) and native
electro-phoresis (Fig. 1b), were about 84,000. These results indicated
that each of the purified SODs is a tetramer composed of
identical polypeptides with a molecular weight of about 21,500.
The stability of both enzymes was detectable over a broad
range of pHs, 5.0 to 9.0, but CF SOD was a little more stable
than CE SOD at pH 4.0 (Fig. 3). For unknown reasons, the
specific activity and stability of CF SOD are higher than those
of CE SOD.
The pI value of both enzymes was 4.5, and those areas of
activity were visualized in the same bands by zymography (Fig.
4).
FIG. 1. SDS-PAGE and native electrophoresis analysis of purified enzymes. (a) SDS-PAGE (12.5% polyacrylamide) pattern. Lanes: M, standard marker proteins; 1, purified CE SOD (20mg); 2, purified CFSOD (20mg). Molecular weight (shown in parentheses) standard markers included the following proteins (Pharmacia): phosphorylase b (94,000), bovine serum albumin (67,000), ovalbu-min (43,000), carbonic anhydrase (30,000), trypsin inhibitor (20,100), anda -lact-albumin (14,400). (b) Native (12.5% polyacrylamide) electrophoresis pattern. Lanes: M, standard marker proteins; 2, purified CE SOD; 4, purified CF SOD; 1 and 3, no protein. Standard marker proteins (Pharmacia, Sweden) were aldo-rase (158,000) and bovine serum albumin (66,000).
TABLE 2. Purification of SOD from CFs of M. bovis BCG
Purification step proteinTotal (mg)
Total activity
(U)a
Sp act
(U/mg) purificationFold Recovery%
Crude extract
b10.08
1,527.1
141.4
1.00
100.0
HIC
c0.21
933.3
3,794.0
26.83
60.1
Gel filtration
d0.11
444.4
4,040.0
28.57
29.1
aA unit is defined as the amount of SOD needed to cause a 50% inhibition in
the rate of oxidation of neotetrazolium chloride at 37°C.
bSeventy to 90% ammonium sulfate precipitation. cSepharose CL-6B hydrophobic interaction chromatography. dSephacryl S-200 superfine molecular sieve chromatography.
on August 17, 2020 by guest
http://cvi.asm.org/
Subunit molecular weight.
The enzymes preincubated with
sample buffer for 2 h at 37°C in the presence and absence of
2-mercaptoethanol in sample buffer were subjected to
SDS-PAGE (4, 37). Comparison of their mobility to that of
molec-ular weight of the standards yielded a subunit with a molecmolec-ular
mass of 21,500 Da in both the absence and presence (Fig. 5) of
mercaptoethanol, indicating that the enzymes are not affected
by mercaptoethanol. Even though a 66,000-Da protein band
was also detected on the electrophoresis gel, it was suggested
that the native enzyme structures are resistant to SDS because
FIG. 2. Molecular weight determination of purified SODs. (a) Molecular weight determination by SDS-PAGE (12.5% polyacrylamide). The marker pro-teins were the same as in Fig. 1a. (b) Molecular weight determination by Sephacryl S-200 molecular sieve chromatography. The molecular weight (shown in parentheses) marker proteins (Pharmacia) were as follows: aldorase (158,000), bovine serum albumin (67,000), ovalbumin (43,000), chymotrypsinogen A (25,000), and RNase A (13,700). Arrows indicated the purified SODs.
FIG. 3. Effect of stability of pH on the activity of purified enzymes. After native gel (12.5% polyacrylamide) electrophoresis, sliced gels were soaked in various pH buffers (pH 4.0 to 9.0) at 37°C for 12 h, incubated in staining solution containing 0.25 mM nitro blue tetrazolium for 30 min at 37°C, and read at 550 nm in a densitometer (LKB). Maximal activity was determined as 100%. The buffers used were 0.1 M sodium acetate (pH 4.0 to 5.0), 0.1 M sodium phosphate (pH 6.0 to 7.5), 0.1 M Tris-HCl (pH 8.0 to 8.5), and 0.1 M glycine-NaOH (pH 9.0).■, CE;E, CF.
FIG. 4. Isoelectric focusing analysis of purified enzymes. (a) Isoelectric fo-cusing in 3-9 IEF premade gel (Pharmacia). (b) Activity staining. Lanes: M, standard markers; 1, purified CE SOD; 2, purified CF SOD. Isoelectric point (shown in parentheses) markers included the following proteins (Pharmacia): trypsinogen (9.30), lentil lectin basic band (8.65), lentil lectin middle band (8.45), lentil lectin acidic band (8.15), horse myoglobin basic band (7.35), horse myo-globin acidic band (6.85), human carbonic anhydrase B (6.55), bovine carbonic anhydrase B (5.85),b-lactoglobin (5.20), soybean trypsin inhibitor (4.55), and amyloglucosidase (3.50). Note that not all marker positions are labeled on the figure.
on August 17, 2020 by guest
http://cvi.asm.org/
of their partial disruption by SDS treatment. From this result,
it is clear that the enzymes are composed of four subunits of
equal size and that these subunits are not joined by interchain
disulfide bonds.
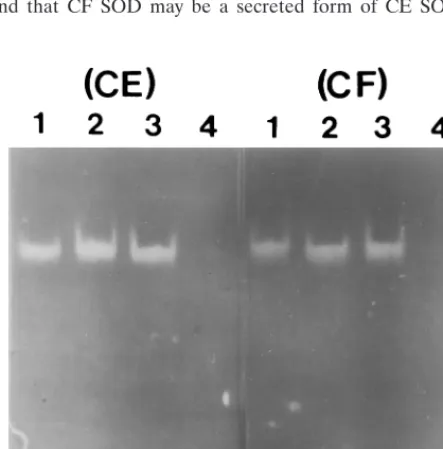
Effect of inhibition.
In Fig. 6, both of the purified CE and CF
SODs were inhibited completely by 1 mM H2O2
(CE and CF
in lane 4), but not by 10 mM azide (CE and CF in lane 3) and
3 mM cyanide (CE and CF in lane 2), as shown by zymography.
This indicated that the purified enzymes are iron-SOD (4, 35)
and that CF SOD may be a secreted form of CE SOD. To
confirm this hypothesis, when a 3-day culture filtrate was
pre-cipitated by 85% ammonium sulfate and dialyzed with D.W., a
protein band with the same molecular weight of the purified
CF SOD appeared by activity-staining electrophoresis (data
not shown).
N-terminal amino acid sequence comparison.
When
puri-fied enzymes were subjected to Edman degradation and
deter-mination of N-terminal amino acid sequences, the N-terminal
amino acid sequence of CE SOD was AXYTLPDLDX and
that of CF SOD was AEYTLPDLDXDYGAL. Ten of the
N-terminal amino acid sequences in CE SOD and 15 of those
in CF SOD in this study showed a very high degree of similarity
to those of other mycobacterial species and N. asteroides and
low homology with those of other bacteria (Table 3). It is
interesting that the N-terminal sequence of CE SOD was
iden-tical to that of CF SOD, implying that there are no N-terminal
signal peptides responsible for secretion. Although an
uniden-tified sequence existed, it was predicted that the first X of CE
SOD is glutamic acid (E), as judged from Edman degradation
data.
Western blot analysis.
To examine whether the purified
en-zymes are working as antigenic molecules in tuberculosis
pa-tients, Western analysis was carried out with sera from the
patients. Both CE SOD (Fig. 7, CE, lanes 1 to 8) and CF SOD
(Fig. 7, CF, lanes 1 to 4) were reacted with sera of tuberculosis
patients, but were not reacted with normal sera (Fig. 7, CE,
lanes 9 to 12, and CF, lanes 5 to 8). Even though only one
result is presented, the Western blotting data strongly
indi-cated that both of the purified SODs have the same antigenic
determinant, as shown by antigenicity against the sera of
tu-berculosis patients.
DISCUSSION
Iron-SOD has been purified from CE and CF of M. bovis
BCG grown in Sauton’s medium. Purified enzymes showed
similarities with respect to molecular weight, pH profile,
sub-unit structure, substrate specificity, sensitivity to inhibitors, and
antigenicity to tuberculosis patients. The molecular weight and
tetrameric form of the enzymes (Fig. 1) were similar to those
of SODs of M. tuberculosis (19), Mycobacterium avium (12),
and N. asteroides (7). However, there is a slight difference from
those of M. leprae, M. smegmatis, Mycobacterium lepraemurium,
Mycobacterium intracellulare, and Mycobacterium duvalii, even
with BCG (22). Since adoption of a tetrameric form is thought
to confer the stability of the enzyme in cellular and
extracel-FIG. 5. Determination of subunit molecular weight of purified enzymes by 12.5% acrylamide gel electrophoresis. Lanes: M, marker proteins; 1 and 2, CE SOD; 3 and 4, CF SOD. Lanes 1 and 3 show results in the absence of mercap-toethanol, and lanes 2 and 4 show results in the presence of it. Each of the samples was incubated at 37°C for 2 h. The marker proteins used were the same as in Fig. 1a.
FIG. 6. Effects of inhibitors on purified enzymes. One microgram of the enzyme was preincubated with 3 mM cyanide (lane 2), 20 mM azide (lane 3), and 1 mM H2O2(lane 4), respectively, for 30 min and electrophoresed on a 12.5% acrylamide gel prior to enzyme activity staining. Lane 1 contained the control.
P. aeruginosa A---FELPPLPYEKNAL Hassett et al. (14) B. fragilis (GenBank
accession no. M96560)
TYEMPKLPYANNAL Unpublished
observations (1992)
aX, undetected amino acid sequence. Boldface indicates identical amino
ac-ids.
on August 17, 2020 by guest
http://cvi.asm.org/
lular fluids (15), the purified SODs showed a broad pH stability
(Fig. 3). These results were similar to those with many other
SODs (24, 28, 31).
CE SOD and CF SOD shared strong N-terminal amino acid
sequence homologies with SODs from mycobacteria, such as
M. tuberculosis (93%), Mycobacterium fortuitum (93%), M.
avium (87%), M. leprae (87%), and N. asteroides (87%), but
low homologies with those from other bacteria, such as
Esch-erichia coli (60%), Pseudomonas aeruginosa (40%), and
Bacte-roides fragilis (33%) (Table 3). The correspondence in
N-ter-minal amino acid sequence between CE SOD and CF SOD
indicated that the SOD of M. bovis BCG is not preceded by
signal peptides. Thus, it was suggested that the structural gene
of the enzyme from M. tuberculosis is not preceded by a signal
peptide sequence (38). Although M. bovis BCG iron SOD
appeared to be cytosolic as well as a secreted protein, it was not
the product of autolysis, because it was found in CF within 3
days of growth under identical culture conditions. SODs from
CE and CF showed identical molecular weights, pIs, metal
cofactors, N-terminal amino acid sequences, and Western
blots, suggesting that CF SOD should be due to direct
secre-tion from CE SOD. How SOD is exported in mycobacteria
remains unknown, since there are no possible signal peptides
(12, 38). Probably, some specific systems for protein
exporta-tion exist in mycobacteria. Further investigaexporta-tion is in progress
to explain the secretion mechanism of the enzyme in the outer
cell surface of the bacterium and at the gene level.
It has been reported that superoxide enhances formation of
hydroxyl radical (OH
z
), a highly reactive molecule that will
react with various biomolecules, including lipids, proteins, and
DNA, both by reducing Fe
31to Fe
21and by serving as a
source of H2O2
(26). Most bacteria contain SODs and catalase
as means of eliminating superoxide and H2O2, respectively.
Pathogenic microorganisms are exposed to exogenous
super-oxide and H2O2
generated by host neutrophils and other
phagocytes (26). The secreted SOD has been documented to
occur in a few virulent mycobacteria (2, 7, 12), showing that it
could be involved in pathogenicity (6, 12, 19). The secretion of
SOD in virulent forms of N. asteroides, as well as its association
with the outer cell envelope, could provide protection against
killing by superoxide radicals (6), which are produced during
active phagocytosis. More direct evidence for such a protective
role had been obtained previously via administration of a
monoclonal antibody specific for the SOD of N. asteroides (8).
The fact that exogenously added SOD has protected bacteria
against phagocytic attack (36) also illustrates the importance of
SOD. Therefore, the SOD secreted by mycobacteria is more
important, because it could function in the bacteria as a first
line of defense against oxygen-mediated killing.
It has been also reported that the SOD in the purified
protein derivative of M. tuberculosis used for the skin test is a
general antigen (29). In this study (Fig. 7), both M. bovis BCG
CE SOD and CF SOD obviously showed antigenicity against
sera from patients with tuberculosis and had the same
anti-genic determinant. This result showed that this SOD could be
used as a marker protein for diagnosis of tuberculosis.
How-ever, the specificity and sensitivity need to be determined with
a larger number of sera from patients with different stages of
tuberculosis and other mycobacterial infections and from
BCG-vaccinated or nonvaccinated individuals.
ACKNOWLEDGMENTS
We thank Yong-Gil Park of the Korea Institute of Tuberculosis for
providing the bacterial strain and patients’ sera, Samuel Len of the
Korea Herald for proofreading the manuscript, and Chul-Sun Choi of
Chung-Ang University for helpful review. This study was supported in
part by a Special Research grant from Chung-Ang University.
REFERENCES
1. Alcendor, D. J., G. D. Chapman, and B. L. Beaman. 1995. Isolation, se-quencing and expression of the superoxide dismutase-encoding gene (sod) of
Norcardia asteroides strain GUH-2. Gene 164:143–147.
2. Anderson, P., D. Askgaard, L. Ljungqvist, J. Bennedsen, and I. Heron. 1991. Proteins released from Mycobacterium tuberculosis during growth. Infect. Immun. 59:1905–1910.
3. Armbruster, C., W. Junker, N. Vetter, and G. Jaksch. 1990. Disseminated Bacille Calmette-Guerin infection in an AIDS patient 30 years after BCG vaccination. J. Infect. Dis. 162:1216.
4. Asada, K., K. Yoshikawa, M. Takahashi, Y. Maeda, and K. Enmanji. 1975. Superoxide dismutase from a blue-green algae, Plectonema boryanum. J. Biol. Chem. 242:4087–4096.
5. Bannister, J. V., and G. Rotillo. 1984. A decade of superoxide dismutase activity. Dev. Biochem. 26:146–189.
6. Beaman, B. L., C. M. Black, F. Doughty, and L. Beaman. 1985. Role of superoxide dismutase and catalase as determinants of pathogenicity of
No-cardia asteroides: importance in resistance to microbicidal activities of human
polymorphonuclear neutrophils. Infect. Immun. 47:135–141.
7. Beaman, B. L., M. S. Steve, E. M. Stephen, D. Richard, and H. P. Misra. 1983. Purification and properties of a unique superoxide dismutase from
Nocardia asteroides. J. Biol. Chem. 258:91–96.
8. Beaman, L., and B. L. Beaman. 1990. Monoclonal antibodies demonstrate that superoxide dismutase contributes to protection of Nocardia asteroides within the intact host. Infect. Immun. 58:3122–3128.
9. Beauchamp, C., and I. Fridovich. 1971. Superoxide dismutase: improved assays and an assay applicable to acrylamide gels. Anal. Biochem. 44:276– 287.
10. Beck, B. L., L. B. Tabatabai, and J. E. Mayfield. 1990. A protein isolated from Brucella abortus is a Cu-Zn superoxide dismutase. Biochemistry 29: 372–376.
11. Bollag, D. M., and S. T. Edelstein. 1991. Protein methods, p. 143–160. Wiley Interscience, New York, N.Y.
FIG. 7. Western blot analysis of purified enzymes. SDS-PAGE (12.5% polyacrylamide) and electrophoretic transfer of purified SOD onto nitrocellulose membranes were performed as described in Materials and Methods. Arrows indicate the purified SOD bands of CE (a) and CF (b). Lanes 1 to 8 in panel a (CE) and 1 to 4 in panel b (CF) show the reaction with sera from patients infected with M. tuberculosis; lanes 9 to 12 in panel a (CE) and 5 to 8 in panel b (CF) show results with normal sera from healthy individuals.
on August 17, 2020 by guest
http://cvi.asm.org/
17. Ichihara, K. E., M. Kusunose, E. Kusunose, and T. Mori. 1977. Superoxide dismutase from Mycobacterium lepraemurium. J. Biochem. 81:1427–1433. 18. Kroll, J. S., P. R. Langford, and B. M. Loynds. 1991. Copper-zinc superoxide
dismutase of Haemophilus influenzae and H. parainfluenzae. J. Bacteriol.
173:7449–7457.
19. Kusunose, E., K. Ichihara, Y. Nada, and M. Kusunose. 1976. Superoxide dismutase from Mycobacterium tuberculosis. J. Bacteriol. 80:1342–1352. 20. Laemmli, U. K. 1970. Cleavage of structural proteins during the assembly of
the head of bacteriophage T4. Nature (London) 227:680–685.
21. Lowry, O. H., N. J. Rosebrough, A. L. Farr, and R. J. Randall. 1951. Protein measurement with the Folin phenol reagent. J. Biol. Chem. 193:265–275. 22. Lygren, S. T., O. Closs, H. Bercouvier, and L. G. Wayne. 1986. Catalases,
peroxidases, and superoxide dismutases in Mycobacterium leprae and other mycobacteria studied by crossed immunoelectrophoresis and polyacrylamide gel electrophoresis. Infect. Immun. 54:666–672.
23. Mayer, B. K., and J. O. Falkinham III. 1986. Superoxide dismutase activity of Mycobacterium avium, M. intracellulare, and M. scrofulaceum. Infect. Im-mun. 53:631–635.
24. McCord, J. M., and I. Fridovich. 1969. An enzymatic function for erythro-cuprein (hematoprein). J. Biol. Chem. 244:6049–6055.
25. Menendez, M. C., P. Domenech, J. Prieto, and M. J. Garcia. 1995. Cloning
monads. J. Bacteriol. 162:1255–1260.
32. Takeda, Y., and H. Avila. 1986. Structure and gene expression of the E. coli Mn-superoxide dismutase gene. Nucleic Acids Res. 14:4577–4589. 33. Thangaraj, H. S., F. I. Lamb, E. O. Davis, P. J. Jenner, L. H. Jeyakumar, and
M. J. Colston.1990. Identification, sequencing, and expression of
Mycobac-terium leprae superoxide dismutase, a major antigen. Infect. Immun. 58:
1937–1942.
34. Tsang, V. C. W., J. M. Peralta, and A. Bartlett. 1983. Enzyme-linked immu-noelectrotransfer blot technique (EITB) for studying the specificities of antigens and antibodies separated by gel electrophoresis. Methods Enzymol.
92:377–391.
35. Yost, F. J., and I. Fridovich. 1973. An iron containing superoxide dismutase from Escherichia coli. J. Biol. Chem. 248:4905–4908.
36. Yost, F. J., and I. Fridovich. 1974. Superoxide radicals and phagocytosis. Arch. Biochem. Biophys. 161:395–401.
37. Yost, F. J., Jr., and I. Fridovich. 1973. An iron-superoxide dismutase from
Escherichia coli. J. Biol. Chem. 14:4905–4908.
38. Zhang, Y., R. Lathigra, T. Garbe, D. Catty, and D. Young. 1991. Genetic analysis of superoxide dismutase, the 23 kilodalton antigen of Mycobacterium
tuberculosis. Mol. Microbiol. 5:381–391.