Note
Incorporation of Large Heterologies Into Heteroduplex DNA During
Double-Strand-Break Repair in Mouse Cells
Steven J. Raynard* and Mark D. Baker*
,†,1*Department of Molecular Biology and Genetics, College of Biological Science and†Department of Pathobiology, ppOntario Veterinary College, University of Guelph, Guelph, Ontario N1G 2W1, Canada
Manuscript received February 5, 2002 Accepted for publication July 1, 2002
ABSTRACT
In this study, the formation and repair of large (⬎1 kb) insertion/deletion (I/D) heterologies during double-strand-break repair (DSBR) was investigated using a gene-targeting assay that permits efficient recovery of sequence insertion events at the haploid chromosomal immunoglobulin (Ig)-locus in mouse hybridoma cells. The results revealed that (i) large I/D heterologies were generated on one or both sides of the DSB and, in some cases, formed symmetrically in both homology regions; (ii) large I/D heterologies did not negatively affect the gene targeting frequency; and (iii) prior to DNA replication, the large I/D heterologies were rectified.
H
OMOLOGOUS recombination represents a ma- potential to accommodate sequence heterogeneities in-jor pathway for the repair of chromosomal dou- cluding single-base-pair mismatches or unpaired multi-ble-strand breaks (DSBs; Resnick and Martin 1976; base insertion/deletion (I/D) loops. Currentinforma-RudinandHaber1988;Nickoloffet al.1989;Sargent tion indicates that most, if not all, meiotic and mitotic
et al.1997;Lianget al.1998;Takataet al.1998;Linet gene conversion of single-base-pair mismatches and
al.1999). In yeast, current evidence suggests that the small I/D loops results from mismatch repair (MMR) ends of the DSB are resected by a 5⬘-to-3⬘exonuclease of hDNA (Peteset al.1991;Bollaget al.1992;Elliott
to produce long, 3⬘-ended, single-strand tails (Figure et al.1998;NgandBaker1999;Nickoloffet al.1999; 1A;Caoet al.1990;WhiteandHaber1990;Sunet al. Elliott andJasin 2001). Meiotic gene conversion of 1991; Sugawara and Haber 1992). In the modified large (⬎1 kb) insertions and deletions occurs with an version of the double-strand-break-repair (DSBR) model efficiency similar to that of small I/D heterologies and (Szostaket al.1983;Sunet al.1991), one 3⬘end invades point mutations in yeast (Petes et al.1991). Recently, a homologous duplex, anneals with its complementary a high rate of segregation of very large heterozygous sequence to generate a region of asymmetric hetero- insertions (2.6 and 5.6 kb) was observed during mitotic duplex DNA (hDNA), and primes DNA repair synthesis, and meiotic recombination in yeast MMR mutants, sug-displacing a D loop. The D loop is enlarged and anneals gesting that large I/D heterologies can be incorporated to the second 3⬘end forming another region of asym- into hDNA and are available to cellular MMR activities metric hDNA from which repair synthesis also initiates. (Clikemanet al.2001;Kearneyet al.2001).
This culminates in the formation of two Holliday junc- Less is known about the behavior of large I/D heterol-tions (Figure 1B). If the amount of homologous flank- ogies in mammalian cells. In mouse cells, spontaneous ing DNA is sufficient, Holliday junction formation and conversion of a 1.5-kb insertion by intrachromosomal branch migration may extend the heteroduplex in a recombination between tandem repeats has been re-symmetric fashion on one or both sides of the DSB ported (LetsouandLiskay1987;GodwinandLiskay (Figure 1C). Resolution of the two Holliday junctions 1994). The rate of conversion of the large insertion was in opposite planes results in crossover, integrating the up to two orders of magnitude lower than that of single-vector into the chromosome and duplicating the region base-pair and small insertions at the same site. Wald-of shared homology (Figure 1D). manet al.(1999) reported a similar low rate of intrachro-The formation of hDNA is important since it has the mosomal gene conversion of a 2.4-kb insertion in mouse cells. Inserts of 114, 200, and 800 bp converted with roughly equal frequency during spontaneous
intrachro-1Corresponding author:Department of Pathobiology, Ontario
Veteri-mosomal recombination between tandem repeats in
nary College, University of Guelph, Guelph, ON N1G 2W1, Canada.
E-mail: mdbaker@uoguelph.ca Chinese hamster ovary (CHO) cells (Sargentet al.1996).
several features consistent with the DSBR model of re-combination (Figure 1;Orr-Weaveret al.1981; Szos-taket al.1983;Sunet al.1991), including the formation of hDNA on both sides of the initiating DSB (Li and
Baker 2000a; Baker and Birmingham 2001). In this system, it should be noted that only single crossover events result in vector integration. Previously, it was reported that repair of DSB-induced intrachromosomal homologous recombination occurs primarily through gene conversion unassociated with reciprocal exchange in mammalian cells (Johnson andJasin2000). There-fore, the recombinants recovered in this study may rep-resent a fraction of the total recombination events.
The 12.6-kb vector, pTC⌬2.8, bears a 2.8-kb deletion
(denoted M1* to distinguish it from the wild-type M1 chromosomal sequence) on the 5⬘side of the DSB (Fig-ure 2). Targeted integration of a single copy of the vector into the chromosomal -locus generates a tan-dem duplication of the shared C-region of homology. As shown in Figure 3A, four possible marker patterns are distinguishable in Southern analysis by diagnostic
EcoRI fragments. Four separate electroporations yielded 45 independent, single-copy targeted recombinants (Table 1). The marker pattern in each recombinant is presented in Figure 3B. As an example of the Southern analysis, Figure 3C presents some representative G418R
transformants.
The majority of the recombinants (33/45) contained a recombinant -locus that bears the chromosomal
Figure1.—DSBR model of recombination. The targeting (M1) marker in the 5⬘C-region and the vector-borne
vector and chromosomal sequences are indicated by the thick
(M1*) marker in the 3⬘ C-region [pattern in Figure
and thin lines, respectively. Regions of newly synthesized DNA
3A(i) and recombinants 9/1, 41/1, 49/1, and 52/1 in
primed by invading 3⬘ vector ends are indicated by broken
Figure 3C]. As can be deduced from Figure 2, this class
lines. Arrowheads denote positions where strand nicks in the
joint molecule (C) would permit vector integration. Regions is explained most simply by a single reciprocal crossover
of asymmetric hDNA and symmetric hDNA are indicated with event on the DSB proximal side of the heterology. Five A and S, respectively. For further details, see text.
recombinants (7/1, 10/1, 12/1, 27/1, and 33/1) had chromosomal (M1) markers in both C-regions of ho-mology [pattern in Figure 3A(ii) and recombinant 27/1 However, since the nature of the initiating lesion was
in Figure 3C]. These recombinants are consistent with not known in these studies, a distinction between a
a gene conversion event toward the chromosomal se-double-strand gapped and an hDNA intermediate could
quence, which might arise as a consequence of enlarge-not be made. Formation and efficient repair of 26-base
ment of the initial DSB to form a double-strand gap loop mismatches during extrachromosomal
recombina-(DSG) that, if large enough to remove the vector-borne tion in CHO cells has been reported (Billet al.2001).
M1* marker, could be repaired by DNA synthesis as We report here evidence that I/D heterologies of 1.3
shown in Figure 1. Alternatively, the M1* marker may and 2.8 kb are incorporated into hDNA during DSB
remain and, following strand invasion or Holliday junc-repair in mammalian cells. We utilized a gene-targeting
tion branch migration, be incorporated into hDNA of assay in which the haploid chromosomal
immunoglobu-the general structure depicted in Figure 1D. In this case, lin (Ig)-gene in the murine hybridoma cell line, Sp6/
the conversion tracts in the 5⬘and 3⬘C-regions in the HL (Ko¨ hlerandShulman1980;Ko¨ hleret al.1982),
recombinants would be consistent with MMR of hDNA, serves as the recipient for transformation with
enhancer-such as observed previously for simple mismatches (Ng
trap insertion vectors (pTC⌬2.8and pTC⌬2.8/1.3)
bear-andBaker1999;Nickoloffet al.1999). ing large deletions within the region of shared
homol-The marker patterns in the remaining seven recombi-ogy (Figure 2). As described previously (Valanciusand
nants provide strong support for incorporation of the
Smithies1991;Hastyet al.1992;Denget al.1993;Li
large heterology into an hDNA intermediate. Recombi-and Baker 2000a,b; Baker and Birmingham 2001),
nants 24/1 and 42/1 contained the vector-borne (M1*) mammalian gene targeting with sequence insertion
Figure 2.—Gene target-ing at the chromosomal im-munoglobulin-locus. The structure of the recipient haploid chromosomal-gene in the Sp6/HL hybridoma cell line and the enhancer-trap insertion vectors, pTC ⌬2.8 and pTC⌬2.8/1.3, is
shown. The 12.6-kb vec-tor, pTC⌬2.8, contains the
cloned SpeI/XbaI fragment encompassing the -gene switch (S) and constant (C) regions from the Sp6/ HL hybridoma (Ko¨ hler
and Shulman 1980; Ko¨
h-ler et al. 1982; Ochi et al. 1983) inserted into a pSV2neo vector backbone
(SouthernandBerg1982).
During cloning inE. coli, a 2.8-kb segment was deleted from the S-region, leaving a residual, ⵑ0.4-kb S
-seg-ment (Ochiet al. 1983; in
this study, the deletion is de-noted M1* to distinguish it from the wild-type chromo-somal sequence M1). The S-deletion does not affect the ability of this frag-ment to encode a func-tional -chain (Ochiet al. 1983). The 11.3-kb vector, pTC⌬2.8/1.3, was derived from pTC⌬2.8by deleting the 1.3-kbDraIII/ EcoRV segment (denoted M2* to distinguish it from the wild-type chromosomal sequence, M2) from the C-region. In each vector backbone, the 372-bp NsiI/NdeI fragment encompassing the SV40 early region enhancer was deleted. As described previously (Bautistaand Shulman1993;Ngand Baker1998; NgandBaker1999), the enhancer-trap feature enriches for gene-targeting events at the chromosomal Ig-locus. In both vectors, cutting at the uniqueXbaI site creates the DSB. Transfection of vector DNA (8.7 pmol) into 2 ⫻ 107 recipient Sp6/HL hybridoma cells was performed by electroporation according to
conditions described previously (Bakeret al.1988). According to trypan blue exclusion, hybridoma cell survival averagesⵑ50%. Following electroporation, the surviving hybridoma cells were distributed at a low cell density to the individual wells of 96-well tissue culture plates and placed under G418 selection as described earlier (NgandBaker1999;LiandBaker2000a,b). These procedures ensure, with high likelihood, that individual transformants represent the progeny of single G418Rcells deposited in
the culture wells (NgandBaker1999;LiandBaker2000a,b). Genomic DNA was prepared from each G418Rtransformant by
the method ofGross-Bellardet al.(1973) and subjected to Southern analysis to identify correctly targeted clones and to assign chromosomal I/D markers in the recombinant-locus. For the recipient Sp6/HL-gene, the fragment sizes generated following digestion withEcoRI andPacI/PaeR7I are presented. Probe F is an 870-bpXbaI/BamHI C-region fragment. VHTNP, TNP-specific -heavy-chain variable region; S,-gene switch region; C,-gene constant region;neo, neomycin phosphotransferase gene; E,EcoRI; Pa,PaeR7I; Pc,PacI; Sp,SpeI; Xb,XbaI. The figure is not drawn to scale.
3A(iv) and recombinant 42/1 in Figure 3C], signifying this small region. Although we cannot exclude this mechanism, we believe the results are more consistent events in which genetic information was transferred
from the linearized vector to the unbroken chromo- with initiation of homologous recombination at the site of the initial DSB atXbaI, with the final products being some. This situation is most consistent with MMR of the
hDNA intermediate shown in Figure 1D in the direction generated by MMR of the hDNA intermediate pre-sented in Figure 1D.
of the M1* marker. In the five remaining recombinants
(2/1, 16/1, 20/1, 36/1, and 38/1), the M1* and M1 Of the remaining 714 G418Rtransformants, three bore EcoRI C-fragments of 16.2, 12.6, and 8.9 kb. As indi-markers reside in the 5⬘and the 3⬘C-regions,
respec-tively [pattern in Figure 3A(iii) and recombinants 2/1 cated above, the unit length pTC⌬2.8vector is 12.6 kb.
Therefore, these fragment sizes are diagnostic of recom-and 16/1 in Figure 3C]. These recombinants could have
been generated by a crossover event initiating in the binants bearing a C-region triplication resulting from the targeted integration of two copies of the transfer 1.3 kb of homology on the 5⬘side of marker M1*.
Figure 3.—Analysis of-gene struc-tures in recombinants generated by tar-geted integration of the vector, pTC ⌬2.8. (A) Targeted integration of a
sin-gle copy of the vector, pTC⌬2.8, into
the Sp6/HL chromosomal-gene gen-erates a duplication of the C-region of homology separated by the integrated vector sequences. As explained in the text, combinations of the M1 chromo-somal and M1* vector markers in the 5⬘ and 3⬘ C-regions yields four possible targeted vector integration patterns in the recombinants (designated i–iv). For each recombinant-gene structure, the diagnostic fragment sizes generated fol-lowing digestion withEcoRI and hybrid-ization with C-probe F (Figure 2) are presented. (B) Analysis of recombinant -gene marker patterns in recombinants generated by targeted integration of pTC⌬2.8. For clarity, only the genetic
markers are shown. (C) Southern analy-sis of the-gene structure in representa-tive G418R transformants. Hybridoma
genomic DNA was digested withEcoRI, electrophoresed through a 0.7% agarose gel, blotted to nitrocellulose, and hy-bridized with 32P-labeled C-specific
probe F (Figure 2). The hybridoma cell line, igm10, is an Sp6-derived mutant that has lost the chromosomal Ig-gene
(Ko¨ hlerand Shulman1980) and was
TABLE 1
Gene-targeting frequencies
Transfer Surviving recipient Total G418R Targeted Gene-targeting frequency
vector Sp6/HL cells cells recombinants (recombinants/cell)a
pTC⌬2.8 4⫻107 759 45 1.1⫻10⫺6
pTC⌬2.8/1.3 2⫻107 691 73b 3.7⫻10⫺6
aGene-targeting frequencies were calculated as a function of the number of hybridoma cells surviving
electroporation which, according to trypan blue exclusion, averagedⵑ50%.
bSouthern analysis was performed on the first 300 G418Rtransformants generated following transfection
with pTC⌬2.8/1.3. In our experience, most targeted recombinants form colonies early in the G418 selection
process. Likely, this results from a growth advantage over random transformants due to the placement of the enhancerlessneogene in the expressed, chromosomal-locus. Therefore, the 73 recombinants analyzed here are assumed to represent the vast majority of those that were generated in these experiments.
vector integration (for example, in Figure 3C, trans- vector integration via a single crossover at or near the DSB (class I). Class II is represented by five recombi-formants 34/1, 40/1, and 48/1). The majority of these
(703/711) retained the endogenous, 12.5-kbEcoRI-gene nants (49/2, 52/2, 74/2, 88/2, and 95/3). In cell lines 52/2, 74/2, 88/2, and 95/3, vector-borne markers (M1* fragment with evidence in many of one or more
variable-sized fragments, likely representing cases of random and/or M2*) are present at equivalent positions in both regions of homology. Cell line 49/2 contains the M1* integration of the targeting vector into the hybridoma
genome. The 8 remaining transformants contained C- and M2 markers in the 5⬘ C-region, while in the 3⬘ C-region, the M1 and M2* markers are present. The hybridizing fragments of unexpected size. Hybridoma
cell lines with unexpected C-region fragments have marker patterns in these recombinants are most consis-tent with MMR of an hDNA tract that encompassed the been observed in previous gene-targeting studies (Baker
and Read 1993; Ng and Baker 1999; Li and Baker 2.8-kb and/or the 1.3-kb heterologies in both participat-ing regions of homology. Seven class III recombinants 2000b;BakerandBirmingham 2001). Although they
have not been fully characterized, these may represent (2/2, 10/2, 24/2, 32/2, 47/2, 55/3, and 68/3) display a pattern in which, on one side of the DSB, equivalent cases where one arm of the vector has been degraded,
forcing the cells to undergo one-sided recombination positions in both regions of homology contain either the chromosomal M1 or M2 marker, while, in equivalent with the target locus or perhaps illegitimate
recombina-tion such as has been reported previously (Kang and positions on the opposite side of the DSB, both the vector-borne and chromosomal markers are present.
Shulman1991;Berinsteinet al. 1992;BakerandRead
1993). While consistent with the possibility of MMR of the hDNA intermediate shown in Figure 1D, chromosomal The 11.3-kb vector, pTC⌬2.8/1.3, differs from the
en-dogenous-locus by a large deletion on each side of sequences residing on only one side of the DSB might also arise by asymmetric DSG formation and repair. the DSB. The vector-borne markers are denoted M1*
and M2* to distinguish them from the corresponding The final class III recombinant (109/3) contains all chromosomal markers in both the 5⬘and 3⬘regions of wild-type -locus sequences, M1 and M2, respectively
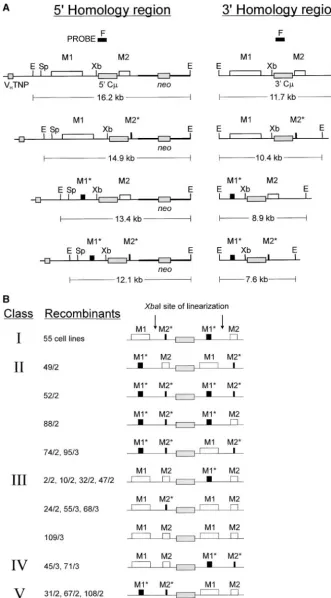
(Figure 2). As depicted in Figure 4A, four possible homology. This marker pattern could have arisen by MMR of hDNA or by repair of a DSG that encompassed marker combinations can be generated in each of the
5⬘and 3⬘C-regions. For simplicity, they are illustrated the vector-borne M1* and M2* markers. Class IV is rep-resented by the two recombinants, 45/3 and 71/3. In separately; however, each pattern in the left column can
potentially be found with one in the right, yielding a these cell lines, the M1 and M2 markers reside in the 5⬘ C-region, while the M1* and M2* markers reside total of 16 possible C-region marker patterns.
South-ern blot analysis identified 73 transformants from two in the 3⬘C-region. Similarly, in class V recombinants, 31/2, 67/2, and 108/2, the M1* and M2* markers reside separate electroporations of pTC⌬2.8/1.3that bore
diag-nosticEcoRI C-fragments expected for integration of in the 5⬘ C-region, while the M1 and M2 markers reside in the 3⬘C-region. As with the five recombinants a single copy of the vector into the target locus (Table
1). The marker pattern in each targeted recombinant generated by transfer of pTC⌬2.8described above (2/1,
16/1, 20/1, 36/1, and 38/1), we cannot exclude the is presented in Figure 4B. The criteria used in
interpre-ting the marker patterns in each recombinant were the possibility that the class IV and V recombinants might have arisen as a consequence of crossover within the same as those utilized in the preceding section. On
the basis of the mechanism(s), the 16 possible marker 1.0 kb of homology on the 3⬘side of the vector-borne M2* marker (class IV) or within the 1.3 kb of homology patterns are divided into five classes. As was seen with
pTC⌬2.8, the vast majority of recombinants (55/73) on the 5⬘side of the vector-borne M1* marker (class V).
Of the remaining 227 G418Rtransformants analyzed,
Figure 4.—Analysis of-gene struc-tures in recombinants generated by the targeted integration of the vector, pTC ⌬2.8/1.3. (A) Targeted integration of a
single copy of the vector pTC⌬2.8/1.3into
the Sp6/HL chromosomal-gene gen-erates a duplication of the C-region of homology separated by the integrated vector sequences. As described in the text, the various combinations of chro-mosomal (M1 and M2) and vector-borne (M1* and M2*) markers in the 5⬘ and 3⬘C-regions yield 16 possible targeted vector integration patterns. To unambig-uously distinguish the diagnostic 12.1-kb EcoRI fragment in transformants bearing the M1* and M2* markers in the 5⬘ C-region from the endogenous 12.5-kb EcoRI fragment that would be expected in transformants bearing a randomly in-tegrated vector, genomic DNA from ap-plicable cell lines was digested with the combination PacI/PaeR7I, which cuts outside the transferred vector, and sub-jected to Southern analysis with C-probe F (Figure 2). Random trans-formants contain the endogenous 14.8-kb PacI/PaeR7I C-region frag-ment (Figure 2) together with a second, C-hybridizing fragment of a variable size from the ectopic vector integration site, whereas correctly targeted cell lines contain both C-regions linked on a PacI/PaeR7I fragment whose size de-pends on the genetic marker combina-tion present. (B) Analysis of recombi-nant-gene marker patterns in targeted recombinants generated by targeted in-tegration of pTC⌬2.8/1.3. For clarity, only
the genetic markers are shown. The marker patterns are grouped into classes I–V on the basis of their likely mecha-nism(s) of formation, as described in the text. Abbreviations are the same as in Figure 2. The figures are not drawn to scale.
6 containedEcoRI fragments of 14.9, 11.3, and 8.9 kb fragments is consistent with a C-region triplication re-sulting from targeted integration of two tandem vector and 1 contained EcoRI fragments of 14.9, 11.3, and
7.6 kb. The 11.3-kb band is the size of the unit length copies. The balance of the 220 G418Rtransformants did
not contain theEcoRI fragments indicative of targeted pTC⌬2.8/1.3vector, and its presence with the 14.9- and
Recombinants bearing vector-borne markers at equiv- side of the heterologies. The length of uninterrupted homology adjacent to the DSB in our study is expected alent positions in both participating regions of
homol-ogy (recombinants 24/1, 42/1, 52/2, 74/2, 88/2, and to be sufficient for efficient initiation of recombina-tion (Waldman and Liskay 1988; Priebe et al. 1994; 95/3) are consistent with the possibility of the I/D
heter-ology being incorporated in symmetric hDNA, with sub- Elliottet al.1998). Thereafter, strand transfer might be able to propagate through the heterologies, such as sequent repair of the mismatches occurring toward the
vector-borne sequences. This would occur if 5⬘-to-3⬘re- has been suggested previously (Waldman andLiskay
1988). This is also suggested byin vitrostudies that have section of the DSB did not proceed past the heterology
and, following strand invasion by the 3⬘ end (Figure shown that the ability of Escherichia coli RecA protein and its eukaryotic homolog Rad51 to bypass heterolo-1B), the Holliday junction was translated through the
heterology by branch migration (Figure 1C). In support gies during strand transfer is limited to insertions (or deletions) of⬍250 and⬍9 bp, respectively (Iypeet al.
of this, we have previously demonstrated the occurrence
of symmetric hDNA during targeted vector integration 1994;Morelet al.1994;Holmeset al.2001). However, in the presence of single-strand binding protein, the in mammalian cells, as evidenced by sectoring of small
palindromic markers at equivalent positions in both E. coliRuvAB complex can act on RecA strand exchange intermediates to mediate bypass of large (⬎1 kb) heter-participating regions of homology at the recombinant
-locus (BakerandBirmingham2001). Alternatively, ologous insertions (Iypeet al.1994;Parsonset al.1995;
AdamsandWest1996). Recent studies have identified extensive resection of the broken DNA ends might
oc-cur past the heterology. Strand invasion by a 3⬘ end protein complexes in mammalian cells that exhibit branch migration and Holliday junction resolution ac-would then incorporate the heterology in a region of
asymmetric hDNA (Figure 1B) that is repaired in favor tivities similar to RuvABC (Chen et al. 2001; Con-stantinouet al.2001).
of the vector-borne marker. DNA repair synthesis
di-rected from the second 3⬘ end fills in the gap using Previous reports showed that spontaneous intrachro-mosomal gene conversion of a 1.5-kb insertion was up the chromosomal strand in the displaced D loop as a
template (Figure 1B). Reverse branch migration through to two orders of magnitude lower than that of single-base-pair and small insertions at the same site in mouse the heterology would generate a region of symmetric
hDNA, as in the first mechanism. Other mechanisms cells (Letsou and Liskay 1987; Godwin and Liskay
1994). The authors postulated that the large heterolo-could also produce this marker pattern (Allers and
Lichten2001). gies interfered with the formation of the conversion intermediate. In this study, the apparent efficiency with The inhibitory influence of the MMR system on
ho-mologous recombination between diverged substrates which large heterologies are encompassed within hDNA during recombination between a transferred plasmid has been well documented in both yeast and
mamma-lian cells (reviewed inEvans andAlani2000; Harfe and chromosome might suggest that topological con-straints interfere with this process during intrachromo-and Jinks-Robertson 2000). Even a single-base-pair
mismatch within a region of otherwise perfect identity somal recombination between closely linked substrates. Alternatively, the disparity in the results might reflect is sufficient to reduce spontaneous mitotic
intrachromo-somal recombination rates up to fivefold (Dattaet al. differences in the pathways of spontaneous intrachro-mosomal gene conversion and DSB-induced homolo-1997;ChenandJinks-Robertson1999;Lukacsovich
andWaldman1999). The absolute frequencies of gene gous recombination between a transfected plasmid and a chromosome.
targeting obtained in this study (ⵑ1 ⫻ 10⫺6
recombi-nants/cell and ⵑ3 ⫻ 10⫺6 recombinants/cell for the Inclusion of large I/D heterologies in a heteroduplex
region is expected to generate large, single-strand loop vectors, pTC⌬2.8 and pTC⌬2.8/1.3, respectively; Table
1) are similar to those reported previously (1.7⫻10⫺6 mismatches. As described previously in studies utilizing
small palindrome genetic markers (LiandBaker2000a,b; recombinants/cell) for enhancer-trap vectors in which
the C-region was either wild type (NgandBaker1998) BakerandBirmingham2001), the recombinant isola-tion procedures utilized here would have permitted the or marked with simple restriction enzyme site
polymor-phisms (NgandBaker1999). Thus, the results suggest detection of any unrepaired mismatch as sectored (mixed) colonies. Thus, the absence of sectoring dem-that the presence of a large heterology on one or both
sides of the DSB did not negatively impact the efficiency onstrates that the mismatches were efficiently repaired
in vivo prior to DNA replication and division of the of the mammalian gene-targeting reaction. The reason
for the discrepant results might be due to the large single cell undergoing recombination. Previously, re-pair of large loop mismatches in mammalian cells has amount of perfect homology (⬎1.6 kb) between the
DSB and the beginning of the I/D heterologies. This been suggested, but from the results of injecting pre-formed heteroduplexes (Ayareset al.1987;Weissand might make it less likely for hDNA to have the
opportu-nity of forming and explain why the marker pattern in Wilson1987, 1989) as well as from studies examining recombination between extrachromosomal plasmids the majority of recombinants (88/118) is consistent with
Bollag, R. J., D. R. Elwood, E. D. Tobin, A. R. GodwinandR. M.
Current evidence suggests that multiple, overlapping
Liskay, 1992 Formation of heteroduplex DNA during
mamma-loop repair pathways are active in yeast cells. Small mamma-loops lian intrachromosomal gene conversion. Mol. Cell. Biol. 12:
1546–1552.
[⬍15 nucleotides (nt)] are efficiently rectified by the
Cao, L., E. AlaniandN. Kleckner, 1990 A pathway for generation
general MMR pathway (Krameret al.1989;Tranet al.
and processing of double-strand breaks during meiotic
recombi-1996; Sia et al. 1997; Luhr et al. 1998). Larger (16- nation inS. cerevisiae.Cell61:1089–1101.
Chen, W., andS. Jinks-Robertson, 1999 The role of the mismatch
to 283-nt) loops are repaired primarily via an
MMR-repair machinery in regulating mitotic and meiotic
recombina-independent mechanism (Tran et al. 1996; Sia et al.
tion between diverged sequences in yeast. Genetics151:1299–
1997; Corrette-Bennettet al.1999;HarfeandJinks- 1313.
Chen, X. B., R. Melchionna, C. M. Denis, P. H. Gaillard, A. Blasina Robertson1999;Corrette-Bennettet al.2001). In
con-et al., 2001 Human Mus81-associated endonuclease cleaves
Hol-trast, others have reported a requirement for MMR
pro-liday junctionsin vitro.Mol. Cell8:1117–1127.
teins in the repair of 26- and 94-nt loops (Kirkpatrick Clikeman, J. A., S. L. WheelerandJ. A. Nickoloff, 2001 Efficient
incorporation of large (⬎2 kb) heterologies into heteroduplex
and Petes 1997; Harfe and Jinks-Robertson 1999).
DNA:Pms1/Msh2-dependent and -independent large loop
mis-Both MMR-dependent and -independent repair of very
match repair inSaccharomyces cerevisiae.Genetics157:1481–1491.
large loops (⬎2 kb) have been reported in yeast (Clike- Constantinou, A., A. A. Davies and S. C. West, 2001 Branch
migration and Holliday junction resolution catalyzed by activities
man et al. 2001;Kearney et al.2001). Repair of both
from mammalian cells. Cell104:259–268.
small and large loops also requires the nucleotide
exci-Corrette-Bennett, S. E., B. O. Parker, N. L. MohlmanandR. S.
sion repair Rad1/Rad10 junction-specific endonuclease Lahue, 1999 Correction of large mispaired DNA loops by
ex-tracts ofSaccharomyces cerevisiae.J. Biol. Chem.274:17605–17611.
complex (Kirkpatrick andPetes 1997;Kirkpatrick
Corrette-Bennett, S. E., N. L. Mohlman, Z. Rosado, J. J. Miret,
1999;Nicholsonet al.2000;Kearneyet al. 2001).
Evi-P. M. Hesset al., 2001 Efficient repair of large DNA loops in
dence for multiple loop repair pathways has also been Saccharomyces cerevisiae.Nucleic Acids Res.29:4134–4143.
Datta, A., M. Hendrix, M. LipsitchandS. Jinks-Robertson, 1997
suggested from studies performed in mammalian cells
Dual roles for DNA sequence identity and the mismatch repair
(Umaret al.1994;Littmanet al.1999;Billet al.2001).
system in the regulation of mitotic crossing-over in yeast. Proc.
The Ercc1/Xpf complex, considered to be the mamma- Natl. Acad. Sci. USA94:9757–9762.
Deng, C., K. R. ThomasandM. R. Capecchi, 1993 Location of
lian equivalent to the yeast Rad1/Rad10 complex (
Sij-crossovers during gene targeting with insertion and replacement
berset al. 1996), may also play a role in the repair of
vectors. Mol. Cell. Biol.13:2134–2140.
loop mismatches, although, in one study, loop repair Elliott, B., andM. Jasin, 2001 Repair of double-strand breaks by
homologous recombination in mismatch repair-defective
mam-was independent of this complex (Littmanet al. 1999).
malian cells. Mol. Cell. Biol.21:2671–2682.
We thank Leah Read for providing excellent technical assistance Elliott, B., C. Richardson, J. Winderbaum, J. A. Nickoloffand and the members of our laboratory for helpful comments during the M. Jasin, 1998 Gene conversion tracts from double-strand course of this work. This work was supported by an operating grant break repair in mammalian cells. Mol. Cell. Biol.18:93–101.
Evans, E., andE. Alani, 2000 Roles for mismatch repair factors in from the Canadian Institutes of Health Research (CIHR) to M.D.B.
regulating genetic recombination. Mol. Cell. Biol.20:7839–7844. and a CIHR studentship to S.J.R.
Godwin, A. R., andR. M. Liskay, 1994 The effects of insertions on mammalian intrachromosomal recombination. Genetics136: 607–617.
Gross-Bellard, M., P. Oudet and P. Chambon, 1973 Isolation
LITERATURE CITED
of high-molecular-weight DNA from mammalian cells. Eur. J. Biochem.36:32–38.
Adams, D. E., andS. C. West, 1996 Bypass of DNA heterologies
during RuvAB-mediated three- and four-strand branch migration. Harfe, B. D., andS. Jinks-Robertson, 1999 Removal of frameshift intermediates by mismatch repair proteins inSaccharomyces
cerevis-J. Mol. Biol.263:582–596.
Allers, T., andM. Lichten, 2001 Intermediates of yeast meiotic iae.Mol. Cell. Biol.19:4766–4773.
Harfe, B. D., andS. Jinks-Robertson, 2000 DNA mismatch repair recombination contain heteroduplex DNA. Mol. Cell8:225–231.
Ayares, D., D. Ganea, L. Chekuri, C. R. CampbellandR. Kucherla- and genetic instability. Annu. Rev. Genet.34:359–399.
Hasty, P., J. Rivera-PerezandA. Bradley, 1992 The role and fate
pati, 1987 Repair of single-stranded DNA nicks, gaps, and loops
in mammalian cells. Mol. Cell. Biol.7:1656–1662. of DNA ends for homologous recombination in embryonic stem cells. Mol. Cell. Biol.12:2464–2474.
Baker, M. D., andE. C. Birmingham, 2001 Evidence for biased
Holliday junction cleavage and mismatch repair directed by junc- Holmes, V. F., K. R. Benjamin, N. J. CrisonaandN. R. Cozzarelli, 2001 Bypass of heterology during strand transfer by
Saccharo-tion cuts during double-strand-break repair in mammalian cells.
Mol. Cell. Biol.21:3425–3435. myces cerevisiaeRad51 protein. Nucleic Acids Res.29:5052–5057.
Iype, L. E., E. A. Wood, R. B. InmanandM. M. Cox, 1994 RuvA
Baker, M. D., andL. R. Read, 1993 Analysis of mutations introduced
into the chromosomal immunoglobulingene. Somat. Cell Mol. and RuvB proteins facilitate the bypass of heterologous DNA insertions during RecA protein-mediated DNA strand exchange. Genet.19:299–311.
Baker, M. D., N. Pennell, L. BosnoyanandM. J. Shulman, 1988 J. Biol. Chem.269:24967–24978.
Johnson, R. D., andM. Jasin, 2000 Sister chromatid gene conversion Homologous recombination can restore normal
immunoglobu-lin production in a mutant hybridoma cell immunoglobu-line. Proc. Natl. Acad. is a prominent double-strand break repair pathway in mammalian cells. EMBO J.19:3398–3407.
Sci. USA85:6432–6436.
Bautista, D., andM. J. Shulman, 1993 A hit-and-run system for Kang, Y., andM. J. Shulman, 1991 Effect of vector cutting on its recombination with the chromosomal immunoglobulin gene in introducing mutations into the Ig H chain locus of hybridoma
cells by homologous recombination. J. Immunol.151:1950–1958. hybridoma cells. Somat. Cell Mol. Genet.17:525–536.
Kearney, H. M., D. T. Kirkpatrick, J. L. GertonandT. D. Petes,
Berinstein, N., N. Pennell, C. A. Ottaway andM. J. Shulman,
1992 Gene replacement with one-sided homologous recombi- 2001 Meiotic recombination involving heterozygous large inser-tions inSaccharomyces cerevisiae: formation and repair of large, nation. Mol. Cell. Biol.12:360–367.
Bill, C. A., D. G. Taghian, W. A. DuranandJ. A. Nickoloff, 2001 unpaired DNA loops. Genetics158:1457–1476.
Kirkpatrick, D. T., 1999 Roles of the DNA mismatch repair and Repair bias of large loop mismatches during recombination in
mammalian cells depends on loop length and structure. Mutat. nucleotide excision repair proteins during meiosis. Cell. Mol. Life Sci.55:437–449.
Kirkpatrick, D. T., andT. D. Petes, 1997 Repair of DNA loops Priebe, S. D., J. Westmoreland, T. Nilsson-TillgrenandM. A. Resnick, 1994 Induction of recombination between homolo-involves DNA-mismatch and nucleotide-excision repair proteins.
Nature387:929–931. gous and diverged DNAs by double-strand gaps and breaks and
role of mismatch repair. Mol. Cell. Biol.14:4802–4814.
Ko¨ hler, G., andM. J. Shulman, 1980 Immunoglobulin M mutants.
Eur. J. Immunol.10:467–476. Resnick, M. A., andP. Martin, 1976 The repair of double-strand
breaks in the nuclear DNA ofSaccharomyces cerevisiaeand its
ge-Ko¨ hler, G., M. J. Potash, H. LehrachandM. J. Shulman, 1982
Deletions in immunoglobulin mu chains. EMBO J.1:555–563. netic control. Mol. Gen. Genet.143:119–129.
Rudin, N., andJ. E. Haber, 1988 Efficient repair of HO-induced
Kramer, B., W. Kramer, M. S. WilliamsonandS. Fogel, 1989
Het-eroduplex DNA correction inSaccharomyces cerevisiaeis mismatch chromosomal breaks inSaccharomyces cerevisiaeby recombination between flanking homologous sequences. Mol. Cell. Biol. 8: specific and requires functionalPMSgenes. Mol. Cell. Biol.9:
4432–4440. 3918–3928.
Sargent, R. G., R. V. Merrihew, R. Nairn, G. Adair, M. Meuthet Letsou, A., andR. M. Liskay, 1987 Effect of the molecular nature
of mutation on the efficiency of intrachromosomal gene conver- al., 1996 The influence of a (GT)29microsatellite sequence on
homologous recombination in the hamster adenine phosphori-sion in mouse cells. Genetics117:759–769.
Li, J., andM. D. Baker, 2000a Formation and repair of heteroduplex bosyltransferase gene. Nucleic Acids Res.24:746–753.
Sargent, R. G., M. A. BrennemanandJ. H. Wilson, 1997 Repair DNA on both sides of the double-strand break during mammalian
gene targeting. J. Mol. Biol.295:505–516. of site-specific double-strand breaks in a mammalian
chromo-some by homologous and illegitimate recombination. Mol. Cell.
Li, J., andM. D. Baker, 2000b Use of a small palindrome genetic
marker to investigate mechanisms of double-strand-break repair Biol.17:267–277.
Sia, E. A., R. J. Kokoska, M. Dominska, P. GreenwellandT. D.
in mammalian cells. Genetics154:1281–1289.
Liang, F., M. Han, P. J. RomanienkoandM. Jasin, 1998 Homology- Petes, 1997 Microsatellite instability in yeast: dependence on repeat unit size and DNA mismatch repair genes. Mol. Cell. Biol. directed repair is a major double-strand break repair pathway in
mammalian cells. Proc. Natl. Acad. Sci. USA95:5172–5177. 17:2851–2858.
Sijbers, A. M., W. L. de Laat, R. R. Ariza, M. Biggerstaff, Y. F. Lin, Y., T. LukacsovichandA. S. Waldman, 1999 Multiple
path-ways for repair of DNA double-strand breaks in mammalian chro- Weiet al., 1996 Xeroderma pigmentosum group F caused by a defect in a structure-specific DNA repair endonuclease. Cell86: mosomes. Mol. Cell. Biol.19:8353–8360.
Littman, S. J., W. H. FangandP. Modrich, 1999 Repair of large 811–822.
Southern, P. J., andP. Berg, 1982 Transformation of mammalian insertion/deletion heterologies in human nuclear extracts is
di-rected by a 5⬘ single-strand break and is independent of the cells to antibiotic resistance with a bacterial gene under control of the SV40 early region promoter. J. Mol. Appl. Genet.1:327–341. mismatch repair system. J. Biol. Chem.274:7474–7481.
Luhr, B., J. Scheller, P. MeyerandW. Kramer, 1998 Analysis of Sugawara, N., andJ. E. Haber, 1992 Characterization of double-strand break-induced recombination: homology requirements
in vivocorrection of defined mismatches in the DNA mismatch
repair mutants msh2,msh3andmsh6ofSaccharomyces cerevisiae. and single-stranded DNA formation. Mol. Cell. Biol.12:563–575.
Sun, H., D. TrecoandJ. W. Szostak, 1991 Extensive 3⬘ -overhang-Mol. Gen. Genet.257:362–367.
Lukacsovich, T., andA. S. Waldman, 1999 Suppression of intra- ing, single-stranded DNA associated with the meiosis-specific dou-ble-strand breaks at theARG4recombination initiation site. Cell chromosomal gene conversion in mammalian cells by small
de-grees of sequence divergence. Genetics151:1559–1568. 64:1155–1161.
Szostak, J. W., T. L. Orr-Weaver, R. J. RothsteinandF. W. Stahl,
Morel, P., A. Stasiak, S. D. EhrlichandE. Cassuto, 1994 Effect of
length and location of heterologous sequences on RecA-mediated 1983 The double-strand-break repair model for recombination. Cell33:25–35.
strand exchange. J. Biol. Chem.269:19830–19835.
Ng, P., andM. D. Baker, 1998 High efficiency site-specific modifica- Takata, M., M. S. Sasaki, E. Sonoda, C. Morrison, M. Hashimoto et al., 1998 Homologous recombination and non-homologous tion of the chromosomal immunoglobulin locus by gene
tar-geting. J. Immunol. Methods214:81–96. end-joining pathways of DNA double-strand break repair have
overlapping roles in the maintenance of chromosomal integrity
Ng, P., andM. D. Baker, 1999 Mechanisms of double-strand-break
repair during gene targeting in mammalian cells. Genetics151: in vertebrate cells. EMBO J.17:5497–5508.
Thaler, D. S., andF. W. Stahl, 1988 DNA double-chain breaks in 1127–1141.
Nicholson, A., M. Hendrix, S. Jinks-RobertsonandG. F. Crouse, recombination of phage lambda and of yeast. Annu. Rev. Genet. 22:169–197.
2000 Regulation of mitotic homeologous recombination in
yeast. Functions of mismatch repair and nucleotide excision re- Tran, H. T., D. A. GordeninandM. A. Resnick, 1996 The preven-tion of repeat-associated delepreven-tions inSaccharomyces cerevisiaeby pair genes. Genetics154:133–146.
Nickoloff, J. A., J. D. Singer, M. F. HoekstraandF. Heffron, mismatch repair depends on size and origin of deletions. Genetics 143:1579–1587.
1989 Double-strand breaks stimulate alternative mechanisms of
recombination repair. J. Mol. Biol.207:527–541. Umar, A., J. C. BoyerandT. A. Kunkel, 1994 DNA loop repair by
Nickoloff, J. A., D. B. Sweetser, J. A. Clikeman, G. J. Khalsa human cell extracts. Science266:814–816.
andS. L. Wheeler, 1999 Multiple heterologies increase mitotic Valancius, V., and O. Smithies, 1991 Double-strand gap repair double-strand break-induced allelic gene conversion tract lengths in a mammalian gene targeting reaction. Mol. Cell. Biol. 11:
in yeast. Genetics153:665–679. 4389–4397.
Ochi, A., R. G. Hawley, T. Hawley, M. J. Shulman, A. Traunecker Waldman, A. S., andR. M. Liskay, 1988 Dependence of
intrachro-et al., 1983 Functional immunoglobulin M production after mosomal recombination in mammalian cells on uninterrupted transfection of cloned immunoglobulin heavy and light chain homology. Mol. Cell. Biol.8:5350–5357.
genes into lymphoid cells. Proc. Natl. Acad. Sci. USA80:6351– Waldman, A. S., H. Tran, E. C. GoldsmithandM. A. Resnick, 1999
6355. Long inverted repeats are an at-risk motif for recombination in
Orr-Weaver, T. L., J. W. SzostakandR. J. Rothstein, 1981 Yeast mammalian cells. Genetics153:1873–1883.
transformation: a model system for the study of recombination. Weiss, U., andJ. H. Wilson, 1987 Repair of single-stranded loops
Proc. Natl. Acad. Sci. USA78:6354–6358. in heteroduplex DNA transfected into mammalian cells. Proc.
Parsons, C. A., A. StasiakandS. C. West, 1995 TheE. coliRuvAB Natl. Acad. Sci. USA84:1619–1623.
proteins branch migrate Holliday junctions through heterolo- Weiss, U., andJ. H. Wilson, 1989 Effects of nicks on repair of gous DNA sequences in a reaction facilitated by SSB. EMBO J. single-stranded loops in heteroduplex DNA in mammalian cells.
14:5736–5744. Somat. Cell Mol. Genet.15:13–18.
Petes, T. D., R. E. MaloneandL. S. Symington, 1991 Recombina- White, C. I., andJ. E. Haber, 1990 Intermediates of recombination tion in yeast, pp. 407–521 in The Molecular and Cellular Biology during mating type switching inSaccharomyces cerevisiae.EMBO J.
of the Yeast Saccharomyces: Genome Dynamics, Protein Synthesis and 9:663–673. Energetics, edited by J. R.Broach, J. R.Pringleand E. W.Jones.
![Figure 3C]. As can be deduced from Figure 2, this class](https://thumb-us.123doks.com/thumbv2/123dok_us/1645111.1205750/2.612.47.280.44.387/figure-c-deduced-figure-class.webp)