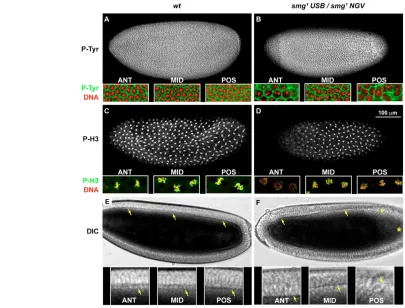
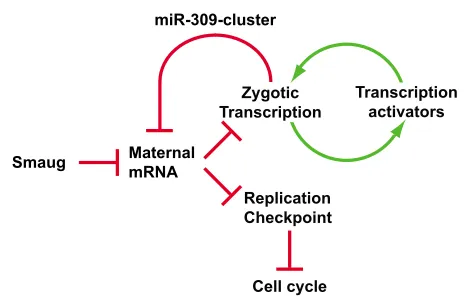
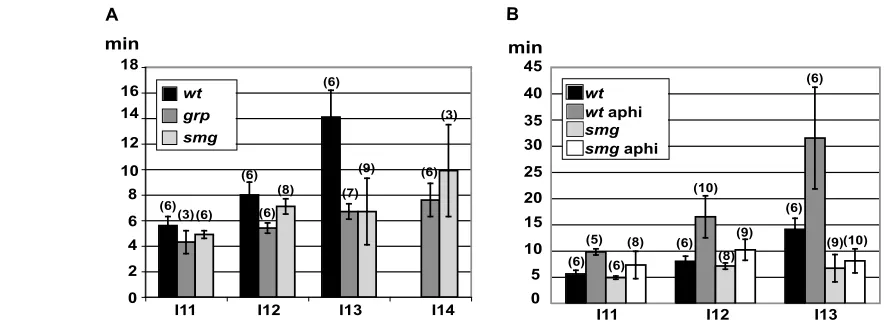
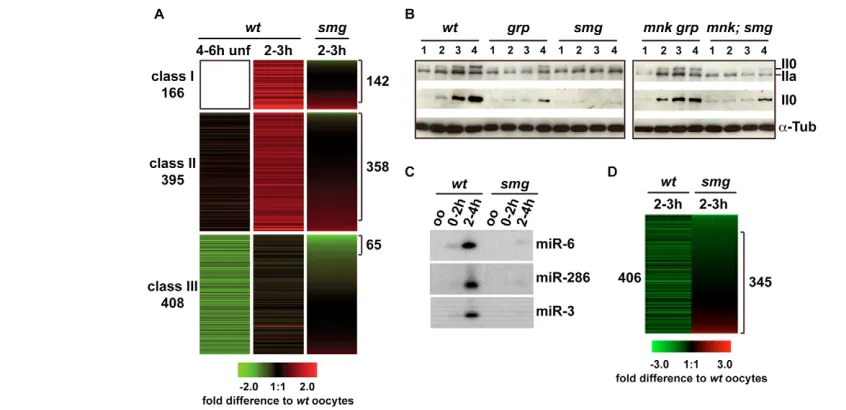
An essential role for the RNA binding protein Smaug during the Drosophila maternal to zygotic transition
Full text
Figure
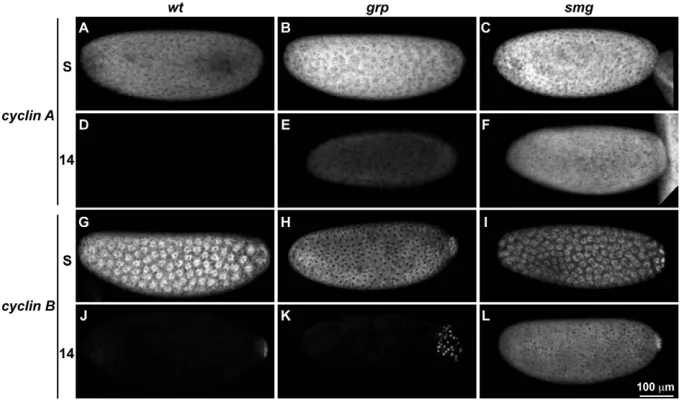
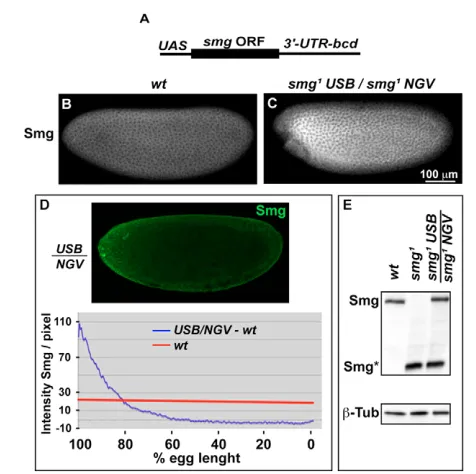
Related documents
This study showed that initially from Weeks 1-3, there was no significant increase in ALP values of rats fed with aqueous and methanolic extracts of Xylopia aethiopica until in
Here Resistive Temperature detector Pt 100 (RTD PT 100) is used to measure the temperature, RT pressure switch is used to measure the pressure inside the boiler and
Social networking texts among college students : identity and imagination online..
Consider a 3-variate multivariate variance components estimation problem with an among and within classification for the random components and with E[Yi]=lPi for i=l, 2 and 3.
The present study, which looked at undergraduate level students, found that there was a statistically significant relationship, where Spearman’s r = .476, between online
Socially Aware D2D Cooperative Communications for Enhancing Internet of Thing Application Yan et al EURASIP Journal on Wireless Communications and Networking (2018) 2018 132 https
Specifically, the risk is considered as the area of the distribution of the arrival time fell after the latest time of the time window, and the mean of the arrival time is
Actuarial approach in a mixed fractional Brownian motion with jumps environment for pricing currency option Shokrollahi and K?l??man Advances in Difference Equations (2015) 2015 257 DOI