CHARACTERIZATION OF
SALMONELLA SPP
– PRODUCING
EXTENDED-SPECTRUM Β-LACTAMASE (ESBL) ISOLATED FROM
CHICKEN SLAUGHTERHOUSES PROCESSING LINE IN
NAMAKKAL
*1R. Sowmiyadevi, 1M. Anitha, 2D. Jegadeeshkumar and 2B. Madhumathi
1
Department of Biotechnology, Selvamm Arts and Science College, Namakkal, Tamilnadu,
India.
2
Chromopark, Namakkal, Tamilnadu, India.
ABSTRACT
The aim of the current work was to evaluate the prevalence,
antimicrobial susceptibility biofilm formation and ESBL producing
Salmonella species isolated from different poultry meat and poultry
processing line. The sampling at slaughterhouses was performed for
Knife, blender, Chopping board, poultry feed, feces, cage and meat.
Totally 32 isolates were identified as Salmonella typhi and Salmonella
typhimurium and were tested for susceptibility to 10 antimicrobial
agents by the disk diffusion method. The highest probability of
antimicrobial resistance occurrence of Salmonella spp was noticed for
Gentamyicin and second most was kanamycin and Ciprofloxacin. A
more frequent profile of multidrug resistance was noticed for isolates
from cutting board, fecal and meat samples. In addition, 54% and 61.5% of biofilm and
betalactamase producers were observed respectively. These results reinforce the need of
efficient strategy implementation to control and reduce Salmonella spp in chickens at
slaughter levels, and the necessity to reduce the use of antimicrobials in the poultry sector.
KEYWORDS: Salmonella spp, Biofilm, Betalactamase, ESBL, poultry processing line.
INTRODUCTION
Salmonella is one of the most important food borne pathogens in developing countries.[1] A
wide range of food products is noticed as possible sources of human Salmonella infection,
Volume 7, Issue 7, 1229-1237. Research Article ISSN 2277– 7105
Article Received on 10 Feb. 2018,
Revised on 02 Mar. 2018, Accepted on 23 Mar. 2018,
DOI: 10.20959/wjpr20187-11677
*Corresponding Author
R. Sowmiyadevi
Department of
Biotechnology, Selvamm
Arts and Science College,
Namakkal, Tamilnadu,
but poultry products have been identified as the most important transmission route.
Processing in poultry slaughterhouses is an important source of Salmonella contamination in
chicken meat[2] especially in small scale poultry slaughterhouses where traditional
slaughtering processes are commonly followed[3] Mostly salmonella present in the
gastrointestinal tract of chicken, during the slaughtering process it may be damaged, resulting
in contaminated carcasses. Furthermore, cross-contamination can occur from a
Salmonella-positive flock or the slaughter equipment to the carcasses of a Salmonella-free flock.
The prevalence of antimicrobial resistance among food-borne pathogens has increased
including in Salmonella[4] This increase is attributed to the selection pressure created by using
antimicrobials in food-producing animals, in addition to the unregulated use of antibiotics by
humans in developing countries.[5]
One of the most important mechanism was responsible for antimicrobial resistance in
organisms producing extended spectrum of betalactamase enzymes. The ESBLs are plasmid
mediated enzymes and capable of hydrolyzing and inactivating a wide variety of β-lactams,
including third generation cephalosporins and aztreonam. The plasmid mediated antibiotic
resistance facilitate the easy spread between species and most likely to occur in the
gastrointestinal tract.[6,7]
The most important mechanisms were organisms producing biofilm, it may be stuck
penetration of the antimicrobial agents through the biofilm matrix, altered the growth rate of
biofilm organisms. Thus, the ability to form biofilm could be an effective strategy to enhance
the survival and persistence under stressed conditions like host invasion or following
antibiotic treatment.[8] Recently, the emergence of antimicrobial resistance in Salmonella, has
also led to ineffective treatment of salmonellosis. Thus, the objectives of this study were to
isolate and estimate the prevalence of Salmonella in poultry slaughterhouses in Namakkal, to
identify their species and to assess their antimicrobial resistance.
MATERIALS AND METHODS Sample collection
The samples were collected from slaughter house (meat and poultry related products) in
afternoons, in order to minimize the microbial changes due to environmental temperatures
and post-slaughter timings.
Sample preparation
Ten grams of collecting meat and feed samples were weighed and transferred to sterile flasks
containing 10 ml of phosphate buffer saline (PBS).[9] Samples were homogenized using a
meat grinder under aseptic conditions. In case of fecal samples, one gram was inoculated into
peptone water and incubated at 370C for overnight. The swab samples were taken from
cutting board, blender, knife and cage, which were inoculated into peptone water and
incubated for overnight. After, loop full of culture was inoculated into different selective
media such as SS agar, MacConkey and Chromogenic media. The plates were incubated for
48 hours at 370C. Colony morphology on the plate was observed and Gram staining was
conducted. Biochemical tests were performed to identify pathogenic bacteria related to food
contamination. These tests included Oxidase, TSI, Urease, Motility, Catalase, Indole,
Simmons citrate and Methyl red and Veges proskauer.
Antibiotic resistance test
The disc diffusion method was followed[10] to determine the antibacterial activity. The Petri
plate containing 20 ml of Mueller Hinton agar was seeded with 24 hours old fresh culture of
bacterial isolates. By making use of template drawn discs were dispensed on the solidified
Mueller Hinton agar with test organisms. This was incubated at 370C for 24 hours in an
incubator. The zone of inhibition was measured by making use of Antibiotic zone scale (Hi -
media). The resistance patterns were interpreted as per CDC recommendations.
Isolation of biofilm producing isolates
The Congo red agar medium was prepared by adding 37g of the BHI powder, 50 g of sucrose
and 10 g of agar in 1 L of distilled water. The mixture was then autoclaved for 15 min at
121°C. Once the agar solution has cooled down to about 50°C, a solution of Congo red (8
g/L) was added and mixed again and then the media were poured into the Petri plates and
allowed to solidify. Once the media had settled, the plates were inoculated with the
microorganisms and incubated at 37°C for 24 h. The black colour colonies were indicated as
Double disc synergy test
By this method, a synergy between a disc of augmenting (ceftazidime and clavulanic acid)
and third generation cephalosporins was detected. The clavulanate in augmenting disc
diffuses through the agar and inhibits the β-lactamases surrounding third generation
cephalosporin disc. Discs containing 30µg of Ceftazidime + Clavulanic acid (CAC),
cefoxitin, Cefpodoxime, Ceftazidime and Cefixime and were placed over inoculated
Mueller-Hinton agar plates 20 mm apart from centrally placed ceftazidime -clavulanic acid disc
(20/10µg). Following overnight incubation at 37°C, diameter of zone of inhibition was
measured. Extension of the edge of the inhibition zone of ceftazidime, cefixime, cefoxitin,
cefpodoxime disc on the side exposed to the disc containing ceftazidime -clavulanic acid was
positive for ESBL.
PCR amplification for detection of beta lactamase genes from food samples
All isolates were screened for the resistance genes SHV, TEM, CTX-M, and OXA by a
multiplex PCR assay using Hong Fang procedure.[12] The plasmid DNA was separated
according to procedure of Sathasivam and Manickam.[13] PCR amplification reactions were
performed in a volume of 25 µl containing 12.5 µl of 2x Promega PCR Master Mix (USA),
0.2 µM concentrations of each primer (1µl), 2µl of plasmid DNA template and make up 25 µl
with molecular grade water. The cycling parameters were as follows: an initial denaturation
at 95°C for 15 min; followed by 30 cycles of 94°C for 30s, 62°C for 90s, and 72°C for 60s;
and with a final extension at 72°C for 10 min. The amplified PCR products were subjected to
electrophoresis at 1.5% agarose gel in 1XTBE buffer. A 100 bp ladder molecular weight
[image:4.595.131.465.539.740.2]Fig. 2: Amplification of ESBL genes from Salmonella species.
RESULT AND DISCUSSION
In our studies, 41% of Salmonella spp were observed from total samples. It was found that all
the sample sources (except the cage sample) showed the positive response to the presence of
life threatening pathogenic bacteria Salmonella isolates. Among the 41% of isolates, 38% (5)
of were Salmonella typhi and 61.5% (8) of being Salmonella typhimurium, which was
confirmed with SS agar and salmonella Chromogenic agar media and biochemical test.
Contamination of chicken meat by Salmonella spp. may indicate hygienic and sanitary issues
in breeding sites, during slaughter or during handling of animals thereafter, as reported by
several authors. In fact, Prakash et al.,[14] in a study that analyzed 82 samples from 8 different
processing plants, identified Salmonella spp. in 15% of tested samples. Contamination with
Salmonella spp. was also detected by Hafiz et al.,[15] in a study that analyzed 182 poultry
slaughter house samples, the pathogen was identified in 88.46% of samples. They were
observed higher percentage of occurrence from cage, blender and cutting board than present
study.
Presence of bacteria on surfaces of cutting boards can lead to transmission of bacterium to
uncontaminated meat products. This chopping board is an inanimate object which doesn’t
have basic nutrients for survival of microbes. However, microbe’s survival was due to the
handling of equipment, improper cleaning and failing to disinfect the equipment which is
used during the slaughtering process.
The prevalence of Salmonella in live birds arriving in wet markets might be very low, but
during processing under unhygienic conditions led to the amplification of contamination of
the carcass from various stages along the processing continuum. Salmonellae usually infect
their hosts via the gastrointestinal tract, the organisms are apparently able to adhere, multiply,
and colonize at any point along the GI tract of chicks[16] which may be shed in the feces, and
hence a source of contamination of other animals, humans and the environment.[17]
Food contamination with antibiotic-resistant bacteria can be a major threat to public health
and this phenomenon was increased during recent decades.[4] This increase could be
attributed to the selection pressure created by using antimicrobials in food-producing animals,
in addition to the unregulated use of antibiotics by humans in developing countries.[5]
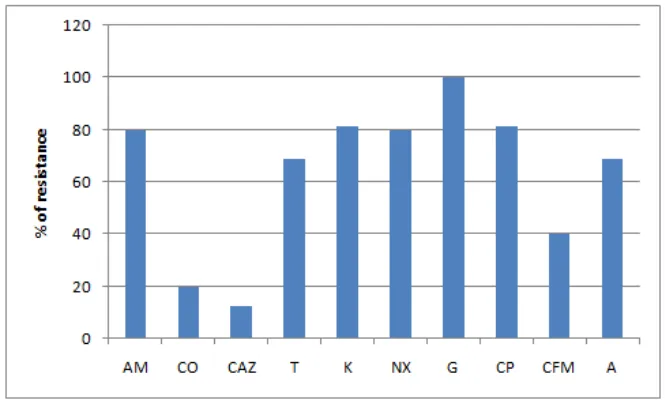
The isolates were 100% resistant to Gentamyicin and second most was kanamycin and
Ciprofloxacin (81.2%). The increasing resistance was observed for Tetracycline and
Ampicillin (69%). The lowest resistance was observed in cefotaxime (12.5%). Totally 62.3%
of isolates was resistances to such antibiotics (Fig.1). This percentage was higher than the
previous studies of Ali and Anil[18] they were observed 36.36% of gentamycin and
Tetracycline resistance salmonella spp from poultry related samples.
Pathogenicity of Salmonella species is mainly due to the acquisition and expression of
various virulence factors, especially due to biofilm formation. During the production of
biofilm producing isolates, they express several virulence factors and an increased resistance
against phagocytosis and other host defense mechanisms[19], In the present study, out of 13
isolates, 7 (54%) of were biofilm producers, which were highest predominant on meat,
cutting board and blender samples. Some studies have demonstrated that Salmonella spp.
from different poultry sources may form biofilm[20], poultry farm and poultry feed materials,
fecal samples[21], feed and water[22]; inadequate disinfection of abattoir trucks.[23] In this
present study most of the biofilm producers were highly resistant to antibiotic than biofilm
negative isolates.
to be the most prescribed worldwide in hospitals. Over 50 years of inappropriate or abusive
use of β-lactamase has created a tremendous selective pressure on the bacterial ecosystem. This high selective pressure selected resistant strains which produced new β-lactamases or
variants of classical plasmid-mediated β-lactamase which are frequently found in
Enterobacteriaceae.[24]
In the present study, isolates of Salmonella species were subjected to double disc diffusion
synergy method for determination of ESBL isolates. Totally 61.5% of isolates were shown
positive for ESBL producers. Most of the ESBLs producing organisms under this study were
also found to be co-resistant to other groups of antibiotics. This phenomenon was accordance
with the study done by Randall et al.,[25] Lalzampuia, et al.,[26] also reported similar results,
where the ESBLs producing enteric bacteria were also resistant to another group of
antibiotics including aminoglycosides, tetracycline. Furthermore, in the present study, ESBLs
genes were amplified by multiplex PCR, among the 4 genes, highest prevalent gene was
TEM and second most was SHV and followed by OXA and CTXM. This study was similar
to previous reports of Prakash et al.,[14] they were amplified ESBL genes from poultry
slaughter house samples (Fig.2).
This study describes the prevalence of Salmonella contamination in the chicken meat and
poultry slaughter house line from the small-scale processing plant in Namakkal, Tamilnadu,
India. The relatively high prevalence in chickens and its small scale processing plant indicate
that poultry is undoubtedly a major potential source of human salmonellosis. These results
call for urgent attention as such prevalence is imminent risk to public health.
REFERENCE
1. Sudershan RV, Naveen Kumar R, Kashinath L, Bhaskar V, Polasa K. Foodborne
Infections and Intoxications in Hyderabad India. Epidemiology Research International,
2014; 1-4.
2. Rasschaert G, Houf K, De Zutter L. Impact of the slaughter line contamination on the
presence of Salmonella on broiler carcasses. J Appl Microbiol, 2007; 103: 333–341.
3. Padungtod P, Kaneene JB. Salmonella in food animals and humans in northern Thailand.
Int. J. Food Microbiol, 2006; 108: 346–354.
4. Angulo FJ, Johnson KR, Tauxe RV, Cohen ML. Origins and consequences of
antimicrobial-resistant nontyphoidal Salmonella: implications for the use of
5. Aarestrup FM, Seyfarth AM, Emborg HD, Pedersen K, Hendriksen RS, Bager F. Effect
of abolishment of the use of antimicrobial agents for growth promotion on occurrence of
antimicrobial resistance in fecal enterococci from food animals in
Denmark. Antimicrobial Agents and Chemotherapy, 2001; 45: 2054–2059.
6. Kruse H, Sorum H. Transfer of multiple drug resistance plasmids between bacteria of
diverse origins in natural microenvironments. Appl. Environ Microbiol, 1994; 60:
4015–4021.
7. Silva J, Castillo G, Callejas L, López H, Olmos J. Frequency of transferable multiple
antibiotic resistance amongst coliform bacteria isolated from a treated sewage effluent in
Antofagasta, Chile. Electronic J Biotechnol, 2006; 9: 533–540.
8. Jeetendra Gurung, Annie Bakorlin Khyriem, Amit Banik, Wihiwot Valarie
Lyngdoh, Basabdatta Choudhury, Prithwis Bhattacharyya. Association of biofilm
production with multidrug resistance among clinical isolates
of Acinetobacter baumannii and Pseudomonas aeruginosa from intensive care unit.
Ind.jor.cra, 2013; 17(4): 214-218.
9. Shafini AB, Son R, Mahyudin NA, Rukayadi Y, Tuan Zainazor. Prevalence of
Salmonella spp. in chicken and beef from retail outlets in Malaysia. IFRJ, 2017; 24(1):
437-449.
10.Bauer AW, Kirby WM, Sherries JC, Turck M. Antibiotic susceptibility testing by a
standardized single disk method. American Journal of Clinical Pathnology, 1966; 45:
493–496.
11.Freeman DJ, Falkiner FR, Keane CT. New method for detecting slime production by
coagulase negative staphylococci. J Clin Pathol., 1989; 42: 872–874.
12.Hong Fang, Ferda Ataker, Göran Hedin, and Kathrine Dornbusch. 2008. Molecular
Epidemiology of Extended-Spectrum β-Lactamases among Escherichia coli Isolates
collected in a Swedish Hospital and Its Associated Health Care Facilities from 2001 to
2006. J Clin Microbiol, 2008; 46(2): 707–712.
13.Sadasivam S, Manickam A. Biochemical methods. 2nd edition, New Age International
(p) Ltd. Publisher, New Delhi, 1996; 169–169.
14.Prakash P, Perumal KD, Jegadeeshkumar D, Tamilmani P. Molecular characterization of
plasmid mediated esbl resistant Salmonella isolated from poultry environment in
Salmonella in poultry processing environments in wet markets in Penang and Perlis,
Malaysia. Veterinary World, 2017; 10(3): 286-292.
16.Desmidt, R. Ducatelle, and F. Haesebrouck. Immunohistochemical Observations in the
Ceca of Chickens Infected with Salmonella enteritidis Phage Type Four. Poultry Science,
1998; 77: 73–74.
17.Poppe C. Salmonella Infections in the Domestic Fowl. In: Salmonella in Domestic
Animals, Wray, C. and A. Wray (Eds.). CAB International, UK, 2000; pp: 107-132.
18.Ali Akbar and Anil Kumar Anal. Prevalence and antibiogram study
of Salmonella and Staphylococcus aureus in poultry meat. Asian Pac J Trop Biomed,
2013; 3(2): 163–168.
19.Costerton JW, Geesey GG, Cheng KJ. How bacteria stick. Sci. Am, 1978; 238: 86–95.
20.Yan Lu, Hongyan Dong, Sujuan Chen, Yiping Chen, Daxin Peng and Xiufan Liu.
Characterization of biofilm formation by Salmonella enterica Serovar Pullorum strains.
African Journal of Microbiology Research., 2011; 5(17): 2428-2437.
21.Marin C, Balasch S, Vega, Lainez M. Sources of Salmonella contamination during broiler
production in Eastern Spain, Prev. Vet. Med., 2011; 98: 39–45.
22.Heyndrickx M, Vandekerchove D, Herman L, Rollier I, Grijspeerdt K, Zutter L. Routes
for Salmonella contamination of poultry meat: Epidemiological study from hatchery to
slaughterhouse. Epidemiol. Infect, 2002; 129: 253–265.
23.Ramesh NS, Joseph W, Carr LE, Douglass LW, Wheaton FW. Evaluation of chemical
disinfectants for the elimination of Salmonella Biofilms from poultry transport containers.
Poult. Sci, 2002; 81: 904 - 910.
24.Nicole A Christian, Karen Roye-Green, and Monica Smikle. Molecular epidemiology of
multidrug resistant extended spectrum beta-lactamase producing Klebsiella
pneumoniae at a Jamaican hospital, 2000 – 2004. l. BMC Microbiology, 2010; 10(27):
1-8.
25.Randall LP, Clouting C, Horton RA, Coldham NG, Clifton-Hadley G, Davies FA, Teale
CJ. Prevalence of Escherichia coli carrying extended-spectrum beta-lactamases (CTX-M
and TEM-52) from broiler chickens and turkeys in Great Britain between 2006 and 2009.
J. Antimicrob. Chemother, 2010; 66(1): 86-95.
26.Lalzampuia H, Dutta TK, Warjri I, Chandra R. Detection of extended-spectrum
β-lactamases (blaCTX-M-1 and blaTEM) in Escherichia coli, Salmonella spp., and
Klebsiella pneumoniae isolated from poultry in North Eastern India, Veterinary World,