metal-organic papers
Acta Cryst.(2005). E61, m2093–m2094 doi:10.1107/S1600536805025626 Niet al. [Cu(C
11H14NO4)Cl]
m2093
Acta Crystallographica Section E Structure Reports
Online
ISSN 1600-5368
Chloro{2-[tris(hydroxymethyl)methyliminomethyl]-phenolato}copper(II)
Jia Ni, Yao-Wen Chen* and Haidan Zhang
Central Laboratory of Shantou University, Shantou, Guangdong 515063, People’s Republic of China
Correspondence e-mail: jni@stu.edu.cn
Key indicators
Single-crystal X-ray study T= 293 K
Mean(C–C) = 0.005 A˚ Rfactor = 0.035 wRfactor = 0.077
Data-to-parameter ratio = 17.7
For details of how these key indicators were automatically derived from the article, see http://journals.iucr.org/e.
#2005 International Union of Crystallography Printed in Great Britain – all rights reserved
In the title compound, [Cu(C11H14NO4)Cl], the tridentate
Schiff base ligand coordinates to the metal atom through the N and O atoms, forming a square-planar coordination geometry.
Comment
The chemistry of transition metal ion complexes of hydroxy (aryl and alkyl OH) rich molecules containing imine/amine groups is important in biomimetic chemistry (Cornmanet al., 1992). Many complexes of this kind have been reported (Asgedom & Rao, 1996; Dey, Rao, Saarenketo & Rissanen, 2002; Dey, Rao, Saarenketo, Rissanen & Kolehmainen, 2002). We report here a new copper(II) complex, (I), with a triden-tate Schiff base ligand.
In compound (I), the CuII center is four-coordinated in a square-planar configuration by one N and two O atoms of the Schiff base ligand and one Cl atom. The Cu—O bond lengths
[image:1.610.273.392.346.437.2] [image:1.610.206.457.505.713.2]Received 13 July 2005 Accepted 10 August 2005 Online 28 September 2005
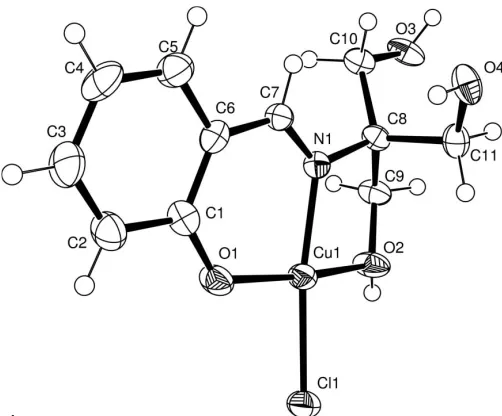
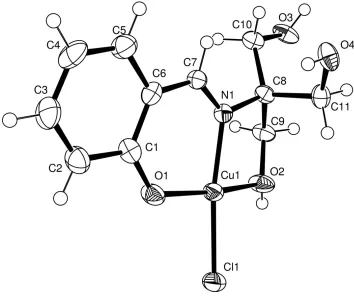
Figure 1
are 1.885 (2) and 1.979 (2) A˚ ; the shorter distance between copper and the phenoxy O atom indicates that the electro-negativity of atom O1 is stronger than that of the other O atoms of the ligand.
Experimental
The ligand 2-[tris(hydroxymethyl)methyliminomethyl]phenol was prepared according to the literature procedure of Asgedom et al. (1996). Cuprous chloride (0.105 g, 0.5 mmol) was added to a solution of the ligand (0.111 g, 0.5 mmol) in water (10 ml). After stirring for a short time, the solution turned dark green. The filtrate was left for 2 d at room temperature and green needle-shaped crystals were obtained in about 62% yield.
Crystal data
[Cu(C11H14NO4)Cl] Mr= 323.22 Tetragonal,P421c a= 16.7345 (6) A˚ c= 8.7634 (6) A˚ V= 2454.1 (2) A˚3 Z= 8
Dx= 1.750 Mg m 3
MoKradiation Cell parameters from 1879
reflections
= 2.4–23.3
= 2.00 mm1 T= 293 (2) K Needle, green 0.210.090.07 mm
Data collection
Bruker APEX area-dectector diffractometer
’and!scans
Absorption correction: multi-scan (SADABS; Bruker, 2002) Tmin= 0.679,Tmax= 0.873
20984 measured reflections
2918 independent reflections 2661 reflections withI> 2(I) Rint= 0.049
max= 27.9
h=21!21 k=21!21 l=11!11
Refinement
Refinement onF2 R[F2> 2(F2)] = 0.035 wR(F2) = 0.077 S= 1.01 2918 reflections 165 parameters
H-atom parameters constrained
w= 1/[2(F
o2) + (0.0354P)2
+ 1.812P]
whereP= (Fo2+ 2Fc2)/3
(/)max= 0.001 max= 0.36 e A˚
3 min=0.21 e A˚
3
Absolute structure: Flack (1983), 1265 Friedel pairs
[image:2.610.314.565.93.155.2]Flack parameter: 0.007 (15)
Table 1
Selected geometric parameters (A˚ ,).
Cu1—O1 1.885 (2)
Cu1—N1 1.951 (2)
Cu1—O2 1.979 (2)
Cu1—Cl1 2.2366 (8)
O1—Cu1—N1 94.92 (10) O1—Cu1—O2 169.50 (10) N1—Cu1—O2 81.81 (10)
O1—Cu1—Cl1 93.56 (7) N1—Cu1—Cl1 171.25 (8) O2—Cu1—Cl1 90.22 (7)
H atoms were placed in idealized positions [N—H = 0.82 A˚ , C— H = 0.93–0.97 A˚ andUiso(H) = 1.2Ueq(C,N)] and were included in the refinement in the riding-model approximation.
Data collection:SMART(Bruker, 2002); cell refinement:SAINT (Bruker, 2002); data reduction:SAINT; program(s) used to solve structure:SHELXS97(Sheldrick, 1997); program(s) used to refine structure: SHELXL97 (Sheldrick, 1997); molecular graphics: ORTEPII (Johnson, 1976); software used to prepare material for publication:SHELXL97.
The authors thank the Central Laboratory of Shantou University for supporting this study.
References
Asgedom, G. & Rao, C. P. (1996).Inorg. Chem.35, 5674–5683.
Bruker (2002).SADABS,SAINTandSMART. Bruker AXS Inc., Madison, Wisconsin, USA.
Cornman, C. R., Colpas, G. J., Hoeschele, J. D., Kampf, J. & Pecoraro, V. L. (1992).J. Am. Chem. Soc.114, 9925–9933.
Dey, M., Rao, C. P., Saarenketo, P. K. & Rissanen, K. (2002).Inorg. Chem. Comm.pp. 380–383.
Dey, M., Rao, C. P., Saarenketo, P., Rissanen, K. & Kolehmainen, E. (2002). Eur. J. Inorg. Chem.pp. 2207–2215.
Flack, H. D. (1983).Acta Cryst.A39, 876–881.
Johnson, C. K. (1976).ORTEPII. Report ORNL-5138. Oak Ridge National Laboratory, Tennessee, USA.
supporting information
sup-1
Acta Cryst. (2005). E61, m2093–m2094supporting information
Acta Cryst. (2005). E61, m2093–m2094 [doi:10.1107/S1600536805025626]
Chloro{2-[tris(hydroxymethyl)methyliminomethyl]phenolato}copper(II)
Jia Ni, Yao-Wen Chen and Haidan Zhang
S1. Comment
The chemistry of transition metal ion complexes of hydroxy (aryl and alkyl OH) rich molecules containing imine/amine
groups is important in biomimetic chemistry (Cornman et al., 1992). Many complexes of this kind have been reported
(Asgedom & Rao, 1996; Dey, Rao, Saarenketo & Rissanen, 2002; Dey, Rao, Saarenketo, Rissanen & Kolehmainen,
2002). We report here a new copper(II) complex, (I), with a tridentate Schiff base ligand.
In compound (I), the CuII center is four-coordinated in a square-planar configuration by one N and two O atoms of the
Schiff base ligand and one Cl atom. The Cu—O bond lengths are 1.885 (2) and 1.979 (2) Å; the shorter distance between
copper and the phenoxy O atom indicates that the electronegativity of atom O1 is stronger than that of the other O atoms
of the ligand.
S2. Experimental
The ligand 2-[tris(hydroxymethyl)methyliminomethyl]phenol was prepared according to the literature procedure of
Asgedom et al. (1996). Cuprous chloride (0.105 g, 0.5 mmol) was added to a solution of the ligand (0.111 g, 0.5 mmol) in
water (10 ml). After stirring for a short time, the solution turned dark green. The filtrate was left for 2 d at room
temperature and green needle-shaped crystals were obtained in about 62% yield.
S3. Refinement
H atoms were placed in idealized positions [N—H = 0.82 Å, C—H = 0.93–0.97 Å and Uiso(H) = 1.2Ueq(C)] and were
Figure 1
An ORTEPII plot (Johnson, 1976) of (I), showing the atom-numbering scheme. Displacement ellipsoids are drawn at the
50% probability level. H atoms are drawn as spheres of arbitrary radii.
Chloro{2-[tris(hydroxymethyl)methyliminomethyl]phenolato}copper(II)
Crystal data
[Cu(C11H14NO4)Cl]
Mr = 323.22
Tetragonal, P421c
Hall symbol: P -4 2n
a = 16.7345 (6) Å
c = 8.7634 (6) Å
V = 2454.1 (2) Å3
Z = 8
F(000) = 1320
Dx = 1.750 Mg m−3
Mo Kα radiation, λ = 0.71073 Å Cell parameters from 1879 reflections
θ = 2.4–23.3°
µ = 2.00 mm−1
T = 293 K Needle, green
0.21 × 0.09 × 0.07 mm
Data collection
Bruker APEX area-dectector diffractometer
Radiation source: fine-focus sealed tube Graphite monochromator
φ and ω scans
Absorption correction: multi-scan (SADABS; Bruker, 2002)
Tmin = 0.679, Tmax = 0.873
20984 measured reflections 2918 independent reflections 2661 reflections with I > 2σ(I)
Rint = 0.049
θmax = 27.9°, θmin = 1.7°
h = −21→21
k = −21→21
supporting information
sup-3
Acta Cryst. (2005). E61, m2093–m2094Refinement
Refinement on F2 Least-squares matrix: full
R[F2 > 2σ(F2)] = 0.035
wR(F2) = 0.077
S = 1.01 2918 reflections 165 parameters 0 restraints
Primary atom site location: structure-invariant direct methods
Secondary atom site location: difference Fourier map
Hydrogen site location: inferred from neighbouring sites
H-atom parameters constrained
w = 1/[σ2(F
o2) + (0.0354P)2 + 1.812P] where P = (Fo2 + 2Fc2)/3
(Δ/σ)max = 0.001 Δρmax = 0.36 e Å−3 Δρmin = −0.21 e Å−3
Absolute structure: Flack (1983), 1265 Friedel pairs
Absolute structure parameter: 0.007 (15)
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
x y z Uiso*/Ueq
Cu1 0.18286 (2) 0.741879 (19) 0.09321 (5) 0.02794 (11) Cl1 0.28495 (5) 0.76534 (5) −0.06532 (11) 0.0405 (2) N1 0.09148 (14) 0.73905 (15) 0.2314 (3) 0.0234 (5) O1 0.21555 (13) 0.63628 (13) 0.1344 (3) 0.0335 (5) O2 0.13204 (14) 0.84338 (13) 0.0301 (3) 0.0351 (5)
H2A 0.1380 0.8636 −0.0544 0.053*
O3 −0.08821 (14) 0.86992 (15) 0.2373 (2) 0.0337 (5)
H3 −0.1088 0.8869 0.3158 0.051*
O4 0.07400 (15) 0.84538 (14) 0.5015 (3) 0.0337 (5)
H4 0.1005 0.8046 0.5127 0.051*
C1 0.1834 (2) 0.58627 (18) 0.2323 (3) 0.0289 (6) C2 0.2141 (2) 0.5080 (2) 0.2414 (4) 0.0376 (8)
H2 0.2556 0.4931 0.1770 0.045*
C3 0.1843 (2) 0.4534 (2) 0.3426 (4) 0.0439 (9)
H3A 0.2054 0.4020 0.3446 0.053*
C4 0.1232 (2) 0.4736 (2) 0.4422 (4) 0.0421 (9)
H4A 0.1051 0.4370 0.5140 0.051*
C5 0.0901 (2) 0.54798 (19) 0.4334 (4) 0.0354 (7)
H5 0.0483 0.5611 0.4987 0.043*
C6 0.11758 (19) 0.60572 (18) 0.3277 (4) 0.0278 (6) C7 0.07552 (17) 0.68002 (18) 0.3202 (3) 0.0249 (6)
H7 0.0324 0.6862 0.3860 0.030*
C8 0.04322 (17) 0.81324 (18) 0.2348 (3) 0.0234 (6) C9 0.05021 (18) 0.84912 (19) 0.0741 (4) 0.0298 (7)
H9A 0.0167 0.8198 0.0031 0.036*
H9B 0.0333 0.9046 0.0747 0.036*
C10 −0.04503 (17) 0.79829 (18) 0.2701 (3) 0.0273 (6)
H10A −0.0651 0.7547 0.2080 0.033*
H10B −0.0515 0.7840 0.3766 0.033*
C11 0.0808 (2) 0.87221 (18) 0.3487 (4) 0.0295 (7)
H11A 0.1368 0.8793 0.3239 0.035*
Atomic displacement parameters (Å2)
U11 U22 U33 U12 U13 U23
Cu1 0.02737 (19) 0.02232 (17) 0.03414 (19) 0.00267 (14) 0.00942 (16) 0.00233 (16) Cl1 0.0386 (4) 0.0303 (4) 0.0527 (5) 0.0000 (3) 0.0219 (4) 0.0010 (4) N1 0.0232 (12) 0.0231 (12) 0.0239 (12) −0.0004 (10) 0.0004 (9) −0.0011 (10) O1 0.0352 (11) 0.0274 (10) 0.0378 (13) 0.0095 (10) 0.0115 (10) 0.0053 (9) O2 0.0352 (12) 0.0351 (12) 0.0350 (12) 0.0087 (10) 0.0143 (10) 0.0136 (10) O3 0.0281 (11) 0.0502 (14) 0.0228 (11) 0.0163 (11) 0.0025 (9) 0.0021 (10) O4 0.0450 (14) 0.0304 (12) 0.0258 (12) 0.0083 (10) −0.0084 (10) −0.0055 (9) C1 0.0315 (16) 0.0292 (15) 0.0261 (15) 0.0050 (13) −0.0033 (13) −0.0009 (12) C2 0.0416 (19) 0.0308 (16) 0.0404 (19) 0.0082 (15) 0.0033 (16) 0.0012 (15) C3 0.050 (2) 0.0251 (16) 0.056 (2) 0.0072 (16) −0.0002 (19) 0.0070 (15) C4 0.044 (2) 0.0311 (17) 0.051 (2) −0.0064 (15) 0.0007 (17) 0.0187 (16) C5 0.0349 (16) 0.0338 (16) 0.0376 (19) −0.0021 (14) 0.0031 (15) 0.0046 (14) C6 0.0310 (17) 0.0234 (15) 0.0291 (16) −0.0039 (12) −0.0021 (13) 0.0024 (13) C7 0.0228 (14) 0.0268 (15) 0.0251 (14) −0.0017 (12) 0.0024 (11) −0.0033 (13) C8 0.0231 (13) 0.0272 (14) 0.0197 (13) 0.0033 (13) 0.0002 (11) −0.0003 (12) C9 0.0282 (14) 0.0331 (15) 0.0282 (16) 0.0081 (12) 0.0063 (13) 0.0062 (14) C10 0.0235 (14) 0.0352 (17) 0.0232 (14) 0.0031 (13) −0.0003 (11) 0.0002 (13) C11 0.0331 (16) 0.0210 (14) 0.0344 (17) 0.0042 (13) −0.0034 (13) −0.0021 (13)
Geometric parameters (Å, º)
Cu1—O1 1.885 (2) C3—C4 1.386 (5)
Cu1—N1 1.951 (2) C3—H3A 0.93
Cu1—O2 1.979 (2) C4—C5 1.365 (5)
Cu1—Cl1 2.2366 (8) C4—H4A 0.93
N1—C7 1.286 (4) C5—C6 1.415 (4)
N1—C8 1.481 (4) C5—H5 0.93
O1—C1 1.313 (4) C6—C7 1.430 (4)
O2—C9 1.426 (3) C7—H7 0.93
O2—H2A 0.82 C8—C11 1.538 (4)
O3—C10 1.429 (4) C8—C10 1.529 (4)
O3—H3 0.82 C8—C9 1.535 (4)
O4—C11 1.417 (4) C9—H9A 0.97
O4—H4 0.82 C9—H9B 0.97
C1—C2 1.408 (4) C10—H10A 0.97
C1—C6 1.421 (4) C10—H10B 0.97
C2—C3 1.367 (5) C11—H11A 0.97
C2—H2 0.93 C11—H11B 0.97
O1—Cu1—N1 94.92 (10) C5—C6—C1 118.7 (3)
O1—Cu1—O2 169.50 (10) C5—C6—C7 117.6 (3)
N1—Cu1—O2 81.81 (10) C1—C6—C7 123.6 (3)
O1—Cu1—Cl1 93.56 (7) N1—C7—C6 126.4 (3)
N1—Cu1—Cl1 171.25 (8) N1—C7—H7 116.8
supporting information
sup-5
Acta Cryst. (2005). E61, m2093–m2094C7—N1—C8 121.2 (2) N1—C8—C11 109.1 (2)
C7—N1—Cu1 123.9 (2) N1—C8—C10 113.2 (2)
C8—N1—Cu1 114.80 (18) C11—C8—C10 111.6 (2)
C1—O1—Cu1 127.2 (2) N1—C8—C9 105.5 (2)
C9—O2—Cu1 113.26 (17) C11—C8—C9 108.3 (3)
C9—O2—H2A 109.5 C10—C8—C9 108.9 (2)
Cu1—O2—H2A 123.7 O2—C9—C8 107.2 (2)
C10—O3—H3 109.5 O2—C9—H9A 110.3
C11—O4—H4 109.5 C8—C9—H9A 110.3
O1—C1—C2 118.7 (3) O2—C9—H9B 110.3
O1—C1—C6 123.7 (3) C8—C9—H9B 110.3
C2—C1—C6 117.5 (3) H9A—C9—H9B 108.5
C3—C2—C1 121.7 (3) O3—C10—C8 108.1 (2)
C3—C2—H2 119.1 O3—C10—H10A 110.1
C1—C2—H2 119.1 C8—C10—H10A 110.1
C2—C3—C4 120.9 (3) O3—C10—H10B 110.1
C2—C3—H3A 119.5 C8—C10—H10B 110.1
C4—C3—H3A 119.5 H10A—C10—H10B 108.4
C5—C4—C3 119.1 (3) O4—C11—C8 112.2 (2)
C5—C4—H4A 120.5 O4—C11—H11A 109.2
C3—C4—H4A 120.5 C8—C11—H11A 109.2
C4—C5—C6 121.9 (3) O4—C11—H11B 109.2
C4—C5—H5 119.1 C8—C11—H11B 109.2
C6—C5—H5 119.1 H11A—C11—H11B 107.9