INDUCTION OF INTRACHROMOSOMAL RECOMBINATION
IN YEAST BY INHIBITION
OF THYMIDYLATE
BIOSYNTHESIS
B.
A. KUNZ,' G . R. TAYLOR AND R. H. HAYNES Department of Biology, York University, Toronto, Ontario, Canada M3J l P 3Manuscript received October 28, 1985 Revised copy accepted July 1 , 1986
ABSTRACT
The biosynthesis of thymidylate in the yeast Saccharomyces cerevisiae can be inhibited by antifolate drugs. We have found that antifolate treatment enhances the formation of leucine prototrophs in a haploid strain of yeast carrying, on the same chromosome, two different mutant leu2 alleles separated by Escherichia coli plasmid sequences. That this effect is a consequence of thymine nucleotide depletion was verified by the finding that provision of exogenous thymidylate eliminates the increased production of Leu+ colonies. DNA hybridization analysis revealed that recombination, including reciprocal exchange, gene conversion and unequal sister-chromatid crossing over, between the duplicated genes gave rise to the induced Leu+ segregants. Although gene conversion unaccompanied by crossing over was responsible for the major fraction of leucine prototrophs, events involving reciprocal exchange exhibited the largest increase in frequency. These data show that recombination is induced between directly repeated DNA sequences under conditions of thymine nucleotide depletion. In addition, the results of this and previous studies are consistent with the possibility that inhi- bition of thymidylate biosynthesis in yeast may create a metabolic condition that provokes all forms of mitotic recombination.
NHIBITION of thymidylate (dTMP) biosynthesis in actively growing cells
I
results in thymine nucleotide depletion that can induce a variety of genetic
alterations
in
vitro
(for reviews, see KUNZ 1982; MEUTH
1984; HAYNES
1985;
HAYNES
and KUNZ 1986). Evidence that reduction of thymine nucleotide levels
may occur
in vivo
is based on the association between folate metabolism and
the synthesis of thymidylate. Mammalian cells cannot manufacture folate,
which is an essential vitamin and the precursor of a tetrahydrofolate cofactor
required for dTMP biosynthesis (BLAKLEY
1969; ERBE 1979; SHANE and
STOKSTAD
1985). Diminution of tetrahydrofolate pools leads to thymine nu-
cleotide depletion and produces various genetic, biochemical and clinical symp-
toms (COLMAN
1977; ERBE 1979; KUNZ 1982; KRUMDIECK 1983; SHANE
and
STOKSTAD
1985; HAYNES
and KUNZ 1986). Furthermore, folate and dTMP
'
Present address: Department of Microbiology, The University of Manitoba, Winnipeg, Manitoba, Canada R3T 2N2.376
B. A. KUNZ, G. R . TAYLOR AND R . H. HAYNES TABLE 1Genotypes of strains
Strain Genotype"
BKSo-1A M A T a leu2-3,112 canl-52 his4-519 met8-1 SUP+ BKSo-1AT M A T a leu2-3,112 canl-52 his4-519 met8-1 SUP-o tup-
KT-1AD M A T a leu2-1 pBR322 leu2-3,112 c a n l - 5 2 his4-519 met8-1
KT-IATD M A T a leu2-1 pBR322 leu2-3,112 canl-52 has4-519 met8-I tup-
KT-IAS M A T a leu2-1 c a n l - 5 2 his4-519 met8-1
KT-IATS M A T a leu2-1 canl-52 his4-519 mst8-1 tup-
.,
T h e order of LEU2 alleles on chromosome 111 (Figure 1 ) is included in the genotype of the duplication strains and is as follows: centromere distal LEV2 allele; plasmid sequence; centromereproximal LEU2 allele. As the tup- strains of the K T series are derived from BKSo-lAT, all tup- alleles are identical.
deprivation produce very similar, if not identical, types of
DNA
and chromo-
some damage
(KUNZ 1982; HAYNES
and
KUNZ
1986).
We have been using the yeast
Saccharomyces cerevisiae
as a model eukaryotic
system to study the genetic effects of antifolate drug-induced thymine nucleo-
tide
depletion. Previously, we found that inhibition of dTMP biosynthesis in
yeast is lethal, weakly mutagenic, induces
DNA
strand breakage and enhances
mitotic interchromosomal reciprocal (crossing over) and nonreciprocal (gene
conversion) recombination between homologous
DNA
regions on homologous
chromosomes
(KUNZ
et al. 1980; BARCLAY
et al. 1982;
KUNZ
and
HAYNES
1982;
KUNZ
et al. 1982;
ECKARDT,
KUNZ
and
HAYNES
1983).
More recently, we
demonstrated that thymidylate depletion also provokes mitotic unequal sister-
chromatid crossing over between reiterated ribosomal
DNA
genes and mating-
type switching between the MAT
and
HML
or
HMR
loci
(KUNZ
et al. 1984;
KUNZ, TAYLOR
and
HAYNES
1985).
These latter events are considered forms
of intrachromosomal recombination, a term applied to the transfer of genetic
information between repeated genes on the same chromosome or chromatid,
or
between sister chromatids
(PETES 1980; SZOSTAK
and
Wu
1980; JACKSON
and
FINK 1981; KLEIN
and
PETES
1981;
KLEIN 1984; HABER
and
HEARN
1985;
JACKSON
and
FINK 1985).
T h e data suggest that thymine nucleotide depletion
induces various types of intrachromosomal recombination.
To
explore such
phenomena more extensively, we have now examined the effect of inhibiting
dTMP biosynthesis by antifolate treatment
on genetic exchange between du-
plicated
LEU2
regions separated by
Escherichia coli
plasmid sequences. Recip-
rocal recombination and gene conversion, as well as unequal sister-chromatid
crossing over,
wereinduced between the directly repeated
LEU2
sequences by
dTMP deprivation. We found that, although the majority of induced recom-
binants was d u e to gene conversion alone, the largest increase in recombination
frequency was observed for those events involving reciprocal exchange.
MATERIALS AND METHODS
INTRACHROMOSOMAL RECOMBINATION
377
d T M P , a nucleotide permeable (tup-) derivative (BKSo-1 A T ) of BKSo-1 A was selected on YPDP medium containing antifolates plus dTMP, as described by LITTLE and HAYNES (1979). BKSo-1A o r BKSo-1AT were used for all constructions in this study.Plasmids:
pYE16HI was obtained from T. D. PETES and carries the yeast leu2-1 allele. Other plasmids were constructed for this study. pGTlO consists of the pYE16HI BgZII fragment bearing leu2-1 inserted into the E. coli vector pBR322 (BOLIVAR et al. 1977) at the BamHI site. pGT12 has the YEp13 (BROACH, STRATHERN and HICKS 1979) XhoI-Sal1 fragment that carries the yeast LEU2 gene inserted at the Sal1 site of the E. coli vector pUC8 (MESSING and VIEIRA 1982).Media:
YPDP and minimal omission media have been described previously (BARCLAY et al. 1982).DNA
isolation:
Total yeast DNA was isolated by the mini-prep technique described by SHERMAN, FINK and HICKS (1983). Plasmid DNA was isolated by the alkaline ex- traction procedure of BIRNBOIM and DOLY (1 979).Yeast transformation:
Transformation of yeast strains was carried out using lithium acetate, as described by ITO et al. (1983).Hybridization analyses:
For colony hybridization, individual yeast colonies were sus- pended in 0.25 ml 0.9 M sorbitol, 0.1 M disodium ethylenediaminetetraacetate, pH 7.5, containing0.5
mg/ml zymolyase 100,000 (Miles Laboratories) and incubated at 37" for 30 min t o induce spheroplast formation. T e n microliters of each spheroplast suspension was spotted onto a nitrocellulose filter that was air dried briefly. T h e filter was then treated with 0.5 M sodium hydroxide t o denature the DNA and was neutralized and dried, as described by SHERMAN, FINK and HICKS (1983). For DNA hybridization, total yeast DNA was digested withBglII,
XhoI o r Hind111 endonuclease and was fractionated by electrophoresis on a horizontal0.7%
agarose gel run at 1 V/cm. Electrophoresis was carried out in 40 mM Tris-acetate, 20 mM sodium acetate, 1 mM disodium ethylenedi- aminetetraacetate, pH 8.0. T h e gel was then soaked for 1 h r in 0.5 M sodium hydroxide t o denature the DNA, neutralized in 1 M Tris-hydrochloride, pH 8.0, for 1 h r and dried a t 60" under vacuum for 45 min (TSAO, BRUNK and PEARLMAN 1983). T h e dried nitrocellulose filters (for colony hybridization) o r gels (for DNA hybridization) were hybridized with "P-labeled pBR322 or pGT12 DNA, respectively. Nick translation, t o label the plasmid DNA, and hybridizations were performed according t o the methods of MANIATIS, FRITSCH and SAMBROOK (1 982). Following hybridization, the filters and gels were exposed t o Curix RP1 X-ray film at -70" using Dupont Cronex lightning plus intensifying screens.378
B. A. KUNZ, G . R . TAYLOR AND R . H. HAYNESixxB
Bocteriol vector DNA0
leu 2 - 1 DNAT
891 11T
8gl I / / B a m H It s o 1 I
leu 2-3. i12 DNA
7
Hind Ill-
Yeost chromosomal DNA
- Yeost wecror DNA I X h o 1
yeast leu 2-3.11zlocus
-A
Replacement w t h X h o / - S o l I frogmen? Directed integrotton ot
Xhol restrtction sife
k I I x + u
FIGURE 1 .-Formation of yeast strains having either directly repeated copies of Eeu2-I and leu& ?,112 or leu2-3,212 replaced by l e d - 1 . Following digestion with XhoI and transformation, the plasmid pGTlO integrates by homologous recombination at the XhoI endonuclease site 5’ to the leu2-3,IZZ locus on chromosome III of strain BKSo-1A or BKSo-1AT. Digestion of pGTlO with
XhoI and Sal1 before transformation results in replacement of the chromosomal XhoI-Sal1 fragment carrying leu2-3,llZ with the pGTlO XhoI-Sal1 fragment carrying leuZ-I. The structures predicted for the duplication and replacement strains were confirmed by DNA hybridization analysis, as described in MATERIALS AND METHODS. T h e XhoI and Sal1 endonuclease sites are centromere distal and centromere proximal to the LEU2 locus, respectively (DOBSON, KINGSMAN and KINCSMAN
198 1).
INTRACHROMOSOMAL RECOMBINATION
1
2
3
4
5
0
23.7.
9.5.
6.7.
4.3.
2.3.
4 0 . 4
+
3.0
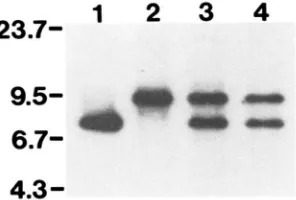
FIGURE 2.-Analysis of DNA insertions in duplication and lcu2-I replacement strains. Total yeast DNA was digested with BglIl and hybridized with a LEU2 probe, as described in MATERIALS
AND METHODS. DNA was isolated from the parental strain BKSo-1A (lane I), the duplication strains KT-IAD and KT-IATD. respectively (lanes 2 and 3), the leu2-I replacement strains KT-1AS and KT-IATS, respectively (lanes 4 and 5). T h e DNA size markers are in kilobases.
1
2
3
4
23.7-
9.5-
gl
6.7-
4.3
-
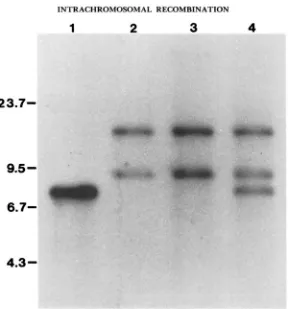
FIGURE 3.-Analysis of pGTIO integration in duplication strains. DNA was digested with Xhol and hybridized with a LEU2 probe, as described in MATERIALS AND METHODS. DNA analyzed was
pGTl0 (lane 1) or total yeast DNA isolated from the parental strain BKSo-IA (lane 2) and the duplication strains KT-1AD and KT-IATD, respectively (lanes 3 and 4). T h e DNA size markers are in kilobases.
380
B. A. KUNZ, G. R . TAYLOR AND R . H. HAYNES TABLE 2Segregation of spontaneous leucine prototrophs: frequencies and rates Frequency Rate" Strain Relevant genotype (X 106) (X 108)
KT-IAD l e d - 1 leu2-3,112 43.9 250 KT- 1 ATD leu2-I leu2-3,112 tup- 39.1 223
KT-I AS Eeu2-1 1.3 7.3
K T - 1 ATS leu2-I tup- 1.9 10.7
a Frequencies are expressed as events per surviving cell. Rates are expressed
as events per cell division.
had integrated at the XhoI endonuclease site adjacent to the LEU2 locus as depicted in Figure 1. However, these patterns also would have been observed if the positions of the leu2-I and leu2-3,112 alleles were switched, perhaps as a consequence of gene conversion during integration (ORR-WEAVER, SZOSTAK and ROTHSTEIN 198 1). T o es- tablish the orientation of the leu2 alleles relative to the centromere of chromosome ZZZ, total DNA from strains KT-1AD a n d KT-1ATD was digested with XhoI, ligated and transformed into E. coli strain JF1754 (TAYLOR et al. 1982) to retrieve plasmid pGTlO. Bacterial transformants were obtained using both yeast DNA samples, and plasmid DNA was isolated from these transformants, digested separately with XhoI and Hind111 and found to be 7.4 kb in size, as expected for pGT10. If the leu2 alleles were ordered in the duplication strains as shown in Figure 1, then the recovered plasmids should carry the leu2-1 allele and, consequently, be able to transform yeast strain BKSo-1 A to leucine prototrophy (see above). No Leu+ transformants would be expected if the plasmid carried the leu2-3,112 allele. T h e plasmid DNA retrieved from KT-1AD and KT-1ATD transformed BKSo-1 A to Leu+. Leucine auxotrophy in the transformants reappeared upon selection for loss of suppressor activity (see above), confirming that the plasmids carried the leu2-I allele. Given that the XhoI site is centromere distal to the LEU2 locus (DOBSON, KINGSMAN a n d KINGSMAN 1981), the order of the LEU2 alleles on chromo- some
ZZZ
in the duplication strains is leu2-I, pBR322, leu2-3,112, centromere.Antifolate treatment:
Yeast cells in the logarithmic phase of growth were suspended in YPDP broth (2 X l o 6 cells/ml) with or without d T M P (300 rg/ml) and incubated with shaking at 34" for 30 min (KUNZ et al. 1980). They were then transferred to an equal volume of YPDP broth, prewarmed to 3 4 " , containing methotrexate (100 pg/ ml) plus sulfanilamide (5 mg/ml) with or without d T M P (300 rg/ml) and incubated at 34" with shaking.Determination of rates
ofleucine prototroph formation:
For each strain, ten tubes of YPDP broth (3 ml) were inoculated with approximately ten cells per tube and were incubated with shaking at 30" for 48 hr. T h e cells were harvested, washed three times with sterile, distilled H 2 0 ; diluted where appropriate; plated on complete minimal me- dium a n d leucine omission medium; and incubated for 72 h r at 30". T h e frequency of prototroph formation for each independent culture was then determined. T h e rates of leucine prototroph formation per round of replication were calculated according to the following expression (DRAKE 1970): rate = (0.4343 mean frequency)/(log mean final cell number-log mean initial cell number).RESULTS
381
BKSo-1
A T , having only the
leu2-3,112
allele, failed to produce Leu+ colonies
at a detectable frequency (the
Zeu2-3,112
allele has been reported to have a
spontaneous reversion frequency of
<1
Ow’*
(BROACH,
STRATHERN
and HICKS
1979)). T h e strains used in this study are isogenic except for the
LEU2
locus
and the presence or absence of the
tup-
mutation. In addition, all the nucleo-
tide-permeable strains carry the same
tup-
mutation. Furthermore, there is no
significant difference
(P
>
0.99) in the rates of leucine prototroph formation
when corresponding
TUP+
and
tup-
strains are compared. Finally, while both
the
leu2-1
and
canl-52
alleles present in the duplication strains are ochre
suppressible, the majority (585/604
=
95%) of the spontaneous KT-1 ATD
Leu+ segregants tested had not concurrently become canavanine-sensitive.
These various facts indicate that the
tup-
mutation did not influence the seg-
regation of leucine prototrophs and argue that the enhanced formation of
Leu+ colonies in the duplication strains was not due to reversion or suppression
of
leu2-1.
Intrachromosomal recombination also could produce Leu+ segregants. It has
been demonstrated in a similar system that spontaneous intrachromosomal
gene conversion occurs more frequently than events involving reciprocal ex-
change
(JACKSONand FINK 1981), and hybridization analysis of DNA from
leucine prototrophs recovered here (see below) was consistent with this finding.
Thus, we conclude that the increased frequency of spontaneous Leu+ segre-
gants in the
LEU2
duplication strains was, in large part, a result of intrachro-
mosomal gene conversion.
Induction
of
leucine prototrophs by antifolate treatment:
The methylation
of dUMP to dTMP by thymidylate synthase in yeast (Figure 4) requires the
methyl donor
N5,N’o-methylene-tetrahydrofolate
(BISSON
and THORNER
1977;
TAYLOR
et
al.
1982). As
S .
cerevisiae
lacks the enzyme thymidine kinase (GRI-
VELL
and
JACKSON1968) and the thymidylate synthase reaction constitutes a
significant drain on intracellular tetrahydrofolate pools (BLAKLEY
1969; ERBE
1979; SHANE
and STOKSTAD
1985), dTMP biosynthesis can be blocked by
limiting the supply of reduced folates. Treatment of yeast cells with the anti-
folate drugs methotrexate plus sulfanilamide effectively causes dTMP depletion
(KUNZ
etal.
1984). When logarithmic phase cells of the
LEU2
duplication strain
382
B. A. KUNZ, G. R . TAYLOR AND R . H . HAYNES2-AMlN0-4-HYDRoX Y-
6-METHYLPERIN PARA-AMINOBENZOATE
GLUTAMATE
FIGURE 4.-A scheme for the inhibition of thymidylate biosynthesis by antifolates. Thymidylate synthase methylates dUMP to dTMP and NJ,N’o-methylene-tetrahydrofolate (FH4: tetrahydrofo- late) functions as the methyl donor in reaction 1 . In general, the dihydrofolate (FHz) analogue methotrexate inhibits dihydrofolate reductase (reaction 2), whereas sulfanilamide, a p-aminoben- zoate analogue, poisons de novo folate synthesis (reaction 3) (BLAKLEY 1969). In yeast, treatment with methotrexate plus sulfanilamide blocks dTMP biosynthesis presumably by inhibiting FH4 synthesis, so that N4,N”-methylene-FH4 consumed in reaction 1 is not replaced.
neously arising KT-1 ATD Leu+ colonies were treated with the folate antago-
nists. T h e survival after
9 hr exposure to the drugs was identical for both cell
types. Thus, leucine independence did not confer a selective advantage during
antifolate treatment.
Previously, we have determined that thymine nucleotide depletion in yeast
results in weak, allele-specific mutagenesis
(ECKARDT,KUNZ
and
HAYNES
1983).
However, treatment of the
leu2-I
replacement strains K T - I
AS
and KT-1
ATS
with methotrexate plus sulfanilamide did not increase the frequency of leucine
prototrophs (Table
3).
In
addition, when Leu+ segregants induced by antifolate
treatment of KT-1 ATD were examined, the majority (>95%) was found to
have remained canavanine-resistant (Table
4),although both
leu2-I
and
canI-
52
are ochre suppressible alleles. These data demonstrate that thymine nucleo-
tide depletion does not lead to reversion or suppression of
leu2-I.
This, plus
the fact that
leu2-3, I12
is not detectably reverted by physical or chemical
mutagens
(HINNEN,HICKS
and
FINK
1978), indicates that the antifolate-induced
leucine prototrophs did not emerge as a consequence of mutation.
INTRACHROMOSOMAL RECOMBINATION
383
I
c
/
_/o----*----oc----. t dTMP
0--0
-
dTMP /16 /
/O
/
I 3
L
/
/O
I
/lot
//
TIME ( h o u r s )
FIGURE 5.-Induction of leucine prototrophs by dTMP depletion. Strain KT-1 ATD was treated with methotrexate plus sulfanilamide, as described in MATERIALS AND METHODS. At 3-hr intervals,
samples were withdrawn and were washed three times with sterile H,O; cell titers (A) were determined by Coulter Counter, and washed cells were diluted if necessary and were plated onto appropriately supplemented minimal media to detect survivors and leucine prototrophs. Surviving fractions (B: viable cells per milliliter divided by Coulter counts per milliliter) and the frequencies of leucine prototrophs (C) were determined after incubating the plates at 30" for 4 days. The data are expressed as the means (normalized to time zero) of four independent experiments: C---., with dTMP; 0- -0, without dTMP.
locus. To our knowledge, the exact positions of all three mutations have not
been established. In principle, three orientations are possible (Figure 6). Within
these orientations, specific recombination events can
be
characterized
by
loss
(pBR322-) or retention (pBR322+) of the pBR322
DNA
sequences.
In orientation
I
(Figure
6A),
a simple intrastrand reciprocal recombination
event between the mutant
leu2
alleles could
yield
a Leu+ recombinant and
result in excision of the pBR322
DNA
sequences on a circular
DNA
molecule
containing the three
leu2
mutations in a single allele,
leu2-3,112,1.
Following
384
B. A. KUNZ, G.R.
TAYLOR ANDR.
H. HAYNES TABLE 3Antifolate treatment of TUP+ and tup- strains"
Treatment Surviving Leu+ frequency Strain Relevant genotype period (hr) fraction (X 106)
KT-1AD leu2-I leu2-3,112 0
6 12 KT-IATD leu2-I leu2-3,IIZ tup- 0 6 12
KT- 1 AS leu2-I 0
6 12
KT-IATS leu2-I tup- 0
6 12
1
.oo
200.59 93
0.13 377
1
.oo
160.64 98
0.12 336
1
.oo
1.80.64 2.1
0.08 1.3
1
.oo
2.90.65 2.2
0.06 2.5
a Strains were treated with antifolates. as described in MATERIALS AND METHODS.
TABLE 4
Canavanine-resistant colonies among Leu+ segregants
Treatment Total no. of period (hr) colonies tested
0 200
3 400
6 400
9 400
12 400
Canavanine- resistant colonies 195 388 386 395 39 1
7% canavanine- resistant colonies 97.5 97.0 96.5 98.7 97.7
Leu+ colonies that emerged following antifolate treatment of strain KT- 1 ATD for specific periods were replicated to appropriately supple- mented minimal medium containing canavanine (30 pg/ml), incubated at 30" for 48 hr and then scored for canavanine resistance.
centromere distal to the pBR322
DNA
sequences on the other chromatid also
could produce a Leu+ pBR322- recombinant having a single copy of the
LEU2
region. In orientations I1 and I11 (Figures 6B and C), no simple reciprocal
crossover could result in a Leu+ pBR322- recombinant. However, for all three
orientations, gene conversion of one
leu2
allele to
LElJ2,
in
G1
or
GP,
followed
by reciprocal recombination could generate a Leu' recombinant lacking the
pBR322
DNA
sequences. It might be suggested that, if intrastrand gene con-
version was initiated by double-strand gap repair (SZOSTAK,
ORR-WEAVER
and
ROTHSTEIN
1983), then the pBR322
DNA
sequences might be eliminated by
gap formation rather than by reciprocal exchange. If such elimination could
occur, generation of an intact chromosome, and
so
recovery of Leu+ recom-
binants, would still require reciprocal exchange. Thus, for these orientations,
we would classify the various events that could lead to loss of the pBR322
A. ORIENTATION
I
p B R 3 2 2
Q
B. ORIENTATION
GI
pWR322
/-c77
I3 1-
+
C.
ORIENTATION
IU
FIGURE 6.-Formation of Leu+ recombinants by reciprocal exchange. A, Orientation I: In G I , an intrastrand event yields a Leu' pBR322- recombinant and a circular molecule carrying the pBR322 sequences and the three leu2 mutations. In G s , an unequal sister-chromatid event yields a Leu+ pBR322- recombinant and a Leu- recombinant carrying the three leu2 mutations, three copies of the LEU2 region and two copies of the pBR322 sequences, B, Orientation 11: In G I , an intrastrand event yields a leu- pBR322- recombinant carrying the three leu2 mutations and a circular DNA molecule carrying a Leu' allele and the pBR322 sequences. In Gs, an unequal sister- chromatid event yields a Leu+ recombinant with three copies of the LEU2 region and two copies of the pBR322 sequences and a leu- pBR322- recombinant carrying the three leu2 mutations. C, Orientation 111. In G I , an intrastrand event in any of the intervals A-D will not yield a Leu+ recombinant. In G P , an unequal sister-chromatid event in any of the intervals A-D will not yield a Leu+ recombinant.
386
B. A. KUNZ, G. R. TAYLOR AND R . H. HAYNES TABLE 5Frequencies of gene conversion, reciprocal recombination and unequal sistershromatid crossing over among LEU2+ recombinants
Colony hybridization DNA hybridization
Treatment No. of % Leu+ % Leu+ No. of % single % pBR322
period (hr) colonies tested pBR322’ pBR322- colonies tested copy pBR322 duplication
0 100 95 5 18 83 17
12 100 73 27 20 85 15
Mean Frequency (X
Total recombination
Gene, conversion
Unequal Reciprocal sister-chromatid
exchange crossing over
0 1.15 0.91 (79)” 0.06 (5) 0.18 (16)
24.6 15.3 (62) 6.6 (27) 2.7 (11)
12
The mean frequency of total recombination is the arithmetic average of frequencies from four independent experiments. The mean frequencies of gene conversion, reciprocal exchange and unequal sister-chromatid crossing over were determined by multiplying the percentage of gene conversion, reciprocal exchange or unequal sister-chromatid crossing over by the mean frequency of recombination.
a The numbers in parentheses show the percentage of gene conversion, reciprocal exchange and
unequal sister-chromatid crossing over as determined by colony and DNA hybridization.
rocal exchange and could not be distinguished from each other in our system.
Colony hybridization using pBR322 DNA as the probe showed that antifolate
treatment caused a significant
(P
<
0.001) fivefold increase in the fraction of
KT-IATD Leu+ segregants that had lost the pBR322 DNA sequences (Table
387
1
2
3
4
23.7-
9.5
=6.7-
4.3
9FIGURE 7.-Quantitation of LEU2 regions in antifolate-induced Leu+ isolates. Total yeast DNA was digested with Hindlll and hybridized with a LEU2 probe, as described in MATERIALS AND METHODS. DNA was pGTlO (lane 1) or total yeast DNA isolated from untreated Leu- KT-IATD cells having a duplication of the LEU2 region (lane 2 ) and antifolate-induced Leu+ KT-1 ATD cells having a duplication and a triplication, respectively, of the LEU2 region (lanes 3 and 4). T h e DNA size markers are in kilobases.
hybridiiation bands, neither the size of pGT10, would
be expected if the Leu+
pBR322+ recombinant examined was the product of gene conversion alone.
T w o hybridization bands were detected for 15 of 18 spontaneous and 17 of
20 antifolate-induced KT-1 ATD Leu+ pBR322+ recombinants tested
by
such
an analysis; the remainder gave three hybridization bands (Figure 7, Table 5).
This indicates that the majority of recombinants examined resulted from gene
conversion. However, it should
be
noted that antifolate treatment caused sim-
ilar increases, 17- and 15-fold, respectively, in the mean frequencies of events
that could
be
characterized as intrachromosomal gene conversion
or
unequal
sister-chromatid crossing over, compared to the corresponding spontaneous
values (Table 5).
It is
well known that, for
diploid
yeast, different allelic combinations at a
particular locus exhibit different frequencies of induced mitotic interchromo-
somal recombination in response to the same treatment
(KUNZ
and
HAYNES
388
B. A. KUNZ, G.R.
TAYLOR ANDR.
H. HAYNESinduction of intrachromosomal recombination
by
dTMP depletion might pro-
duce results quantitatively different from those reported here, but qualitative
differences would not be expected.
DISCUSSION
Antifolate treatment
toinhibit thymidylate biosynthesis induced the forma-
tion of leucine prototrophs in a haploid strain of yeast carrying different mu-
tant
leu2 alleles directly repeated on the same chromosome. This induction of
Leu+ colonies was prevented by concurrent provision of dTMP. Appropriate
control experiments showed that induced mutation was not responsible for the
enhanced formation of leucine prototrophs. Hybridization analysis revealed
that the Leu+ segregants arose as a consequence of intrachromosomal recip-
rocal exchange and gene conversion, as well as unequal sister-chromatid cross-
ing over. Thus, the results presented here demonstrate that depletion of thy-
mine nucleotides in
S . cerevisiae
induces various forms of intrachromosomal
recombination between repeated alleles separated by plasmid
DNA sequences.
If, as it seems reasonable to assume (BARCLAY
et
al.
1982),
the genetic effects
of dTMP deprivation are dependent on DNA replication, then the reciprocal
exchange and gene conversion events may represent intrachromatid recombi-
nation. We have found previously that inhibition
of
dTMP biosynthesis en-
hances unequal sister-chromatid crossing over in the reiterated ribosomal
DNA
sequences of yeast (KUNZ et
al.
1984).
Here, it is shown that this type of
recombination also can be induced in other chromosomal regions.
Gene conversion was the major event detected; however, the fraction of
recombinants attributable to induced conversion alone was somewhat lower
than the fraction of spontaneous convertants. By contrast, the proportion of
events classified as reciprocal exchange was enhanced fivefold by antifolate
treatment. This resulted in a 110-fold increase in the frequency
of
reciprocal
recombination, compared to the
1
'I-fold increase observed for the frequency
of antifolate-induced gene conversion. Other evidence also suggests that en-
hanced intrachromosomal reciprocal exchange may be characteristic of thy-
mine nucleotide depletion. First, in an earlier study, we scored deletion of a
LEU2
insert from the ribosomal DNA gene cluster to assay the induction of
unequal sister-chromatid crossing over by inhibition of dTMP biosynthesis
(KUNZ
etal. 1984). While this event produced Leu+/Leu- sectored colonies,
isolates that were entirely Leu- also were induced. T h e data suggest that
reciprocal exchange between the tandemly arrayed ribosomal DNA genes to
delete
the LEU2 insert led to the emergence of the Leu- colonies. Second,
preliminary experiments indicate that antifolate treatment also can induce ex-
cision of a transposon located 5' to the yeast gene for glucose repressible
alcohol dehydrogenase
(B.
A.
KUNZ,
unpublished observations). This type of
event is believed to occur via reciprocal recombination between the homolo-
gous
delta sequences flanking the larger portion of the transposon (ROEDER
and FINK
1983).
Finally, thymidylate starvation of dTMP auxotrophs bearing
two directly repeated, partially deleted copies of the thymidylate synthase gene
INTRACHROMOSOMAL
389
panied by
loss
of the intervening plasmid sequences
(G.
R. TAYLOR,
unpub-
lished observations). The restoration of a functional
TMPl
gene appears to be
due to intrachromosomal reciprocal exchange between the duplicated defective
copies of
TMPl.
Not only are these various findings consistent with the induc-
tion of intrachromosomal reciprocal recombination by thymine nucleotide de-
pletion, but they also indicate that such induced exchange is not limited to
particular regions of the genome or to specific gene arrangements.
T h e orientation of the
leu2-1
and
leu2-3,112
mutational sites within the
LEU2
locus cannot be established from our data. However, certain of these
findings suggest that the
leu2
mutations are ordered as diagrammed
for
ori-
entation I (Figure 6A). First, following antifolate treatment, there was a large
increase in the mean frequency of Leu+ pBR322- recombinants (Table
5).
Only in orientation I can a simple reciprocal exchange in
G I
or
G2
produce
this type of Leu+ segregant. Second, although the frequency of events involv-
ing reciprocal recombination
is enhanced dramatically by dTMP depletion, the
frequency of identified unequal sister-chromatid crossovers is affected much
less markedly (Table
5).
Leucine prototrophs resulting from unequal sister-
chromatid crossing over in orientation
I
would lose the pBR322 DNA se-
quences, retain a single copy of the LEU2 region and so be characterized by
hybridization analysis merely as the product of a reciprocal exchange event,
rather than unequal sister-chromatid recombination. This could account for
the observed difference in the extent of induction of reciprocal recombination
and unequal sister-chromatid crossing over. In orientation I, identification of
a Leu+ unequal sister-chromatid recombinant as such would require that the
recombinant result from a gene conversion event associated with reciprocal
exchange. Thus, the relative paucity of characterized unequal sister-chromatid
recombinants also would be consistent with the finding that antifolate-induced
intrachromosomal gene conversion is largely unassociated with crossing over
(Table
5).
If the
leu2
mutations were in orientation 11, a much larger increase
in the frequency of identified unequal sister-chromatid recombinants would be
expected, as this orientation allows
a simple crossover to generate a leucine
prototroph with a triplication of the LEU2 region. Arrangement of the muta-
tions in orientation
I11
would necessitate formation of Leu+ pBR322- recip-
rocal recombinants and Leu+ pBR322+ unequal sister-chromatid recombinants
by crossing over following gene conversion. In this case, both types of recom-
binant would be expected to occur with similar frequencies. Although these
results argue in favor of orientation I, the order of the
leu2-1
and
leu2-3,112
mutations can be established unequivocally only by DNA sequencing. However,
knowledge of the precise orientation of the
leu2
mutations would not alter the
categories of intrachromosomal recombination as defined here, the fractions
of leucine prototrophs attributed to each category or the fact that dTMP
depletion preferentially induces intrachromosomal events that involve recip-
rocal exchange.
390
B. A . KUNZ, G. R. TAYLOR AND R. H. HAYNESresults of this investigation demonstrate that dTMP deprivation provoked by
folate antagonists also enhances various types of intrachromosomal exchange.
Taken collectively, these findings indicate that, in actively growing cells, inhi-
bition of dTMP biosynthesis may create a metabolic condition able to induce
all recognized forms of mitotic recombination. If intrachromosomal recombi-
nation plays a role in the evolution and stability of multigene families
(SMITH
1973;
TARTOFF
1973;
SZOSTAK
and
WU
1980;
BALTIMORE
1981;
NAGYLAKI
and
PETES
1982), our results also suggest that such phenomena may be influ-
enced by any factor(s) that causes thymine nucleotide deficiencies.
We thank D. C. HAWTHORNE for supplying yeast strains used to construct BKSo-IA, T. D. PETS for suggesting the use of leu2-I and providing plasmid pYE16H1, L. E. SPRACKLIN and L. T. JANDRISITS for technical assistance and I. FULTON for typing the manuscript. L. E. S. was supported in part by a grant from the E. I. Canada Career Access Program to B. A. K. This work was supported by the Natural Sciences and Engineering Research Council of Canada.
LITERATURE CITED
BALTIMORE, D., 1981 Gene conversion: some implications for immunoglobulin genes. Cell 24:
Genetic and biochemical
592-594.
BARCLAY, B. J., B. A. KUNZ, J. G. LITTLE and R. H. HAYNES, 1982
consequences of thymidylate stress. Can. J Biochem. 6 0 172-194.
BIRNBOIM, H. C. and J. DOLY, 1979
binant plasmid DNA. Nucleic Acids Res. 7: 1513-1523.
BISSON, L. and J. THORNER, 1977
myces cereuisiae are deficient in thymidylate synthetase. J. Bacteriol. 132: 44-50.
BLAKLEY, R. L., 1969
A rapid alkaline extraction procedure for screening recom-
Thymidine-5’-monophosphate requiring mutants of Saccharo-
The Biochemistry of Folic Acid and Related Pteridines. North Holland, Am- sterdam.
BOLIVAR, F., R. L. RODRIGUEZ, P. J. GREENE, M. C. BETLACH, H. L. HEYNEKER and H. W. BOYER, Construction and characterization of new cloning vehicles. 11. A multipurpose cloning
Transformation in yeast: development
1977
system. Gene 2: 95-1 13.
of a hybrid cloning vector and isolation of the CANZ gene. Gene 8: 121-133. BROACH, J. R., J. N. STRATHERN and J. B. HICKS, 1979
COLMAN, N., 1977
DOBSON, M. J., S. M. KINGSMAN and A. J. KINGSMAN, 1981
Folate deficiency in humans. Adv. Nutr. Res. 1: 77-124.
Sequence variation in the LEU2 region of the Saccharomyces cerwisiae genome. Gene 16: 133-1 39.
DRAKE, J. W., 1970 The Molecular Basis of Mutation. pp. 44-45. Holden Day, Oakland, California. ECKARDT, F., B. A. KUNZ and R. H. HAYNES, 1983 Variation of mutation and recombination frequencies over a range of thymidylate concentrations in a diploid thymidylate auxotroph of yeast. Curr. Genet. 7: 399-402.
ERBE, R. W., 1979
GRIVELL, A. R. and J. F. JACKSON, 1968
Genetic aspects of folate metabolism. Adv. Hum. Genet. 9 293-354.
Thymidine kinase: evidence for its absence from Neu- rospora crassa and some other microorganisms and the relevance of this to the specific labeling of deoxyribonucleic acid. J. Gen. Microbiol. 54: 307-31 7.
rad52-independent mitotic gene conversion in Saccharomyces cerevisiae frequently results in chromosomal loss. Genetics 11 1: 7-22.
Molecular mechanisms in genetic stability and change: the role of deoxy- HABER, J. E. and M. HEARN, 1985
INTRACHROMOSOMAL RECOMBINATION
39
1
ribonucleotide pool balance. pp. 1-23. In: Genetic Consequences of Nucleotide Pool Imbalance,
Edited by F. J. DE SERRES. Plenum, New York.
HAYNES, R. H. and B. A. KUNZ, 1986 The influence of thymine nucleotide depletion on genetic stability and change in eukaryotic cells. Curr. Sci. (Bangalore) 5 5 1 - 1 1 .
HINNEN, A., J. B. HICKS and G. R. FINK, 1978 Transformation of yeast. Proc. Natl. Acad. Sci. ITO, Y., Y. FUKADA, K. MURATA and A. KIMURA, 1983 Transformation of intact yeast cells
treated with alkali ions. J. Bacteriol. 153: 163-168.
Gene conversion between duplicated genetic elements in
Meiotic recombination between duplicated genetic elements
Lack of association between intrachromosomal gene conversion and recip-
Intrachromosomal gene conversion in yeast. Nature 2 8 9 USA 7 5 1929-1933.
JACKSON, J. A. and G. R. FINK, 1981 yeast. Nature 292: 306-31 1 . JACKSON, J. A. and G. R. FINK, 1985
in Saccharomyces cerevisiae. Genetics 109: 303-332. KLEIN, H. L., 1984
rocal exchange. Nature 310: 748-753. KLEIN, H. L. and T. D. PETES, 1981
144-1 48.
KRUMDIECK, C. L., 1983 Role of folate deficiency in carcinogenesis. pp. 225-245. In: Nutrition Factors in the Induction and Maintenance of Malignancy, Edited by C. E. BUTTERWORTH and M. L. HUTCHINSON, Academic Press, New York.
Genetic consequences of deoxyribonucleotide pool imbalances. Environ. Mu- tagen. 4: 695-725.
Induction of mitotic recombination in yeast by starvation for thymine nucleotides. Proc. Natl. Acad. Sci.
Thymineless recombination in
Saccharomyces cerevisiae is independent of the ability to undergo meiosis. Curr. Genet. 5: 29- 31.
Phenomenology and genetic control of mitotic recombi- nation in yeast. Annu. Rev. Genet. 15: 57-89.
KUNZ, B. A. and R. H. HAYNES, 1982 DNA repair and the genetic effects of thymidylate stress in yeast. Mutat. Res. 93: 353-375.
Mating-type switching in yeast is induced by thymine nucleotide depletion. Mol. Gen. Genet. 1 9 9 540-542.
Inhibition of thymidylate biosynthesis induces mitotic unequal sister chromatid recombination in Saccha- romyces cerevisiae. Curr. Genet. 8: 2 1 1-2 17.
LITTLE, J. G. and R. H. HAYNES, 1979 Isolation and characterization of yeast mutants auxo- trophic for Z’-deoxyuridine 5’-monophosphate. Mol. Gen. Genet. 1 6 8 141-1 5 1 .
MANIATIS, T., E. F. FRITSCH, and J. SAMBROOK, 1982 Molecular cloning-A Laboratory Manual.
Cold Spring Harbor Laboratory, Cold Spring Harbor, New York.
MESSING, J. and J. VIEIRA, 1982 A new pair of M13 vectors for selecting either DNA strand of double-digest restriction fragments. Gene 1 9 269-276.
KUNZ, B. A., 1982
KUNZ, B. A., B. J. BARCLAY, J. C. GAME, J. G. LITTLE and R. H. HAYNES, 1980 USA 7% 6057-6061.
KUNZ, B. A., F. ECKARDT, J. G. LITTLE and R. H. HAYNES, 1982
KUNZ, B. A. and R. H. HAYNES, 1981
KUNZ, B. A., G. R. TAYLOR and R. H. HAYNES, 1985
KUNZ, B. A., G. R. TAYLOR, B. KONFORTI, B. W. GLICKMAN and R. H. HAYNES, 1984
MEUTH, M., 1984 The genetic consequences of nucleotide precursor pool imbalance in mam- malian cells. Mutat. Res. 126: 107-112.
392
ORR-WEAVER, T. L., J. W. SZOSTAK and R. J. ROTHSTEIN, 1981 PETES, T. D., 1980
ROEDER, G. S. and G. R. FINK, 1983 ROTHSTEIN, R. J., 1983
SHANE, B. and E. L. R. STOKSTAD, 1985 Vitamin BIZ-folate interrelationships. Annu. Rev. Nutr. SHERMAN, F., G. R. FINK and J. B. HICKS, 1983 Methods in Yeast Genetics. Cold Spring Harbor
Laboratory, Cold Spring Harbor, New York.
SMITH, G. P., 1973 Unequal crossover and the evolution of multigene families. Cold Spring Harbor Symp. Quant. Biol. 38: 507-513.
SZOSTAK, J . W., T. L. ORR-WEAVER and R. J. ROTHSTEIN, 1983 The double-strand-break repair model for recombination. Cell 33: 25-35.
SZOSTAK, J . W. and R. Wu, 1980 Unequal crossing over in the ribosomal DNA of Saccharomyces
TARTOFF, K., 1973 Unequal mitotic sister chromatid exchange and disproportionate replication as mechanisms regulating ribosomal DNA gene redundancy. Cold Spring Harbor Symp. Quant. Biol. 37: 491-500.
Isolation of the thymidylate synthetase gene by complementation in Saccharomyces cereuisiae. Mol. Cell Biol. 2: 437-442.
Hybridization of nucleic acids directly
Communicating editor: E. JONES B. A. KUNZ, G. R. TAYLOR AND R. H. HAYNES
Yeast transformation: a model system for the study of recombination. Proc. Natl. Acad. Sci. USA 78: 6354-6358.
DNA genes. Cell 1 9 765-774.
Genetic Elements, Edited by J. A. SHAPIRO. Academic Press, New York.
Unequal meiotic recombination within tandem arrays of yeast ribosomal
Transposable elements in yeast. pp. 299-328. In: Mobile
One-step gene disruption in yeast. Methods Enzymol. 101: 202-211.
5: 115-141.
cereuksiae. Nature 284 426-430.
TAYLOR, G. R., B. J. BARCLAY, R. K. STORMS, J. D. FREISEN and R. H. HAYNES, 1982