Copyright © 2000, American Society for Microbiology. All Rights Reserved.
Evaluation of the LiPA MYCOBACTERIA Assay for Identification
of Mycobacterial Species from BACTEC 12B Bottles
NANCIMAE MILLER,* SUSANNA INFANTE,
ANDTIM CLEARY
Department of Pathology, Jackson Memorial Medical Center, University of Miami, Miami, Florida
Received 22 November 1999/Returned for modification 22 January 2000/Accepted 28 February 2000
The LiPA MYCOBACTERIA (Innogenetics NV, Ghent, Belgium) assay was used to identify mycobacterial
isolates using culture fluid from positive BACTEC 12B bottles. The LiPA method involves reverse hybridization
of a biotinylated mycobacterial PCR fragment, a 16 to 23S rRNA spacer region, to oligonucleotide probes
arranged in lines on a membrane strip, with detection via biotin-streptavidin coupling by a colorimetric system.
This system identifies
Mycobacterium
species and differentiates
M. tuberculosis
complex,
M. avium-M.
intracel-lulare
complex, and the following mycobacterial species:
M. avium
,
M. intracellulare
,
M. kansasii
,
M. chelonae
group,
M. gordonae
,
M. xenopi
, and
M. scrofulaceum
. The mycobacteria were identified in the laboratory by a
series of tests, including the Roche AMPLICOR
Mycobacterium tuberculosis
(MTB) test, the Gen-Probe
AC-CUPROBE, and a PCR-restriction fragment length polymorphism (PCR-RFLP) analysis of the 65-kDa heat
shock protein gene. The LiPA MYCOBACTERIA assay detected 60 mycobacterium isolates from 59 patients.
There was complete agreement between LiPA and the laboratory identification tests for 26
M. tuberculosis
complex, 9
M. avium
, 3
M. intracellulare
complex, 3
M. kansasii
, 4
M. gordonae
, and 5
M. chelonae
group (all were
M. abscessus
) isolates. Three patient samples were LiPA positive for
M. avium-M. intracellulare
complex, and all
were identified as
M. intracellulare
by the PCR-RFLP analysis. Seven additional mycobacterial species were
LiPA positive for
Mycobacterium
spp. (six were
M. fortuitum
, and one was
M. szulgai
). The LiPA
MYCOBAC-TERIA assay was easy to perform, and the interpretation of the positive bands was clear-cut. Following PCR
amplification and gel electrophoresis, the LiPA assay was completed within 3 h.
Although more than 70 mycobacterial species have been
described, relatively few of them are strictly pathogenic for
man or animals (19). While
Mycobacterium tuberculosis
com-plex strains are still responsible for the majority of
Mycobac-terium
infections worldwide, opportunistic infections due to
mycobacteria other than tuberculosis (MOTT) have been on
the increase, mainly as a consequence of the AIDS epidemic
(8, 21, 23). Among the mycobacterial species often implicated
in MOTT infections are
M. avium-M. intracellulare
complex,
M.
chelonae,
M. abscessus,
M. kansasii, and
M. xenopi
(8, 19, 33).
M. gordonae
does not usually cause human infection but is
often encountered as a contaminant in clinical samples, and
discrimination from pathogenic species is a relevant diagnostic
issue (4).
The use of liquid cultures in the clinical laboratory improves
the ability to detect the growth of mycobacteria (14, 17, 26).
The radiometric method (BACTEC; Becton Dickinson, Sparks,
Md.) is a fast and sensitive liquid culture system (20). When a
BACTEC bottle is detected as positive, confirmation of the
presence of acid-fast bacilli is done by acid-fast staining and
the broth is plated on solid media.
M. tuberculosis
complex can
be identified rapidly by a variety of nucleic acid amplification
procedures that are commercially available (9, 30, 32). Rapid
identification of MOTT isolates growing on solid media can be
done by techniques such as thin-layer chromatography (11),
gas-liquid chromatography (30), high-performance liquid
chro-matography (10), and analysis with DNA probes (18). Recently
developed molecular methods, such as DNA probe tests (25)
and PCR-restriction fragment length polymorphism (RFLP)
analysis (28, 29), offer identification of this complex group of
organisms from a positive liquid culture medium prior to
de-tection of growth on solid media (3, 7). DNA probes
(ACCU-PROBE; Gen-Probe, Inc., San Diego, Calif.) can be used for
the rapid identification of
M. tuberculosis,
M. avium
and
M.
intracellulare,
M. gordonae, and
M. kansasii
from solid culture
and directly from liquid culture systems (2, 3, 6, 7, 16, 26).
Unfortunately, the DNA probes are available for a limited
number of species, and without colonial morphology to guide
probe selection, testing with multiple probes may be necessary.
An algorithm based on growth rate in the BACTEC 12B bottle
and a fluorochrome smear quantitation to guide DNA probe
selection has been reported (16). PCR-RFLP analysis is a
reliable method for identification of MOTT, encompassing
identification of the entire range of organisms normally
iso-lated in a clinical laboratory (28, 29). The LiPA
MYCOBAC-TERIA test offers identification of a limited number of
com-mon mycobacterial species by PCR amplification of the 16 to
23S rRNA spacer region of
Mycobacterium
species followed by
hybridization of the biotinylated amplified DNA product with
14 specific oligonucleotide probes. The specific probes are
immobilized as parallel lines on membrane strips. The
objec-tive of this study was to evaluate the LiPA MYCOBACTERIA
(Innogenetics NV, Ghent, Belgium) assay for identification
and differentiation of specific mycobacterial species from
pos-itive BACTEC 12B liquid cultures. The assay was evaluated for
specificity, ease of use, and interpretation of results in a
rou-tine clinical laboratory.
MATERIALS AND METHODS
Culture and identification.Respiratory specimens submitted for culture were decontaminated with an equal volume of 5%N-acetyl cysteine-NaOH and con-centrated by centrifugation (24). The sediment was used to prepare two smears and to inoculate a selective 7H11 agar plate and a BACTEC 12B bottle (Becton Dickinson). BACTEC 12B bottles were incubated at 37°C in 5% CO2 and
monitored for growth for 6 weeks with a BACTEC 460 instrument according to the manufacturer’s instructions. When the growth index of a bottle reachedⱖ50,
* Corresponding author. Mailing address: Department of Pathology,
University of Miami, P.O. Box 016960, Miami, FL 33101. Phone: (305)
585-6258. Fax: (305) 585-0008. E-mail: nmiller@med.miami.edu.
1915
on May 15, 2020 by guest
http://jcm.asm.org/
a smear was prepared to confirm the presence of acid-fast organisms and the liquid medium was subcultured onto a blood agar and a 7H11 agar plate. Isolates of mycobacteria growing on solid media were identified by DNA probes (AC-CUPROBE; Gen-Probe, Inc.) forM. avium,M. intracellulare,M. gordonae, and
M. kansasiior by conventional biochemical tests performed according to
stan-dard protocols (13, 19).
AMPLICORM. tuberculosisPCR test.Respiratory specimens submitted for culture that were acid-fast organism smear positive were processed for PCR directly from the decontaminated concentrated specimen according to the in-structions of the package insert for the AMPLICORM. tuberculosistest (Roche Diagnostics, Indianapolis, Ind.), as previously described (5). In addition, PCR testing was also performed from positive BACTEC 12B bottles using 0.5 ml of the culture fluid concentrated by centrifugation at 16,000⫻gfor 15 min in a 1.5-ml screw-cap microcentrifuge tube. The pellet was resuspended in 100l of lysis buffer and processed in the same way as direct clinical specimens for the remainder of the procedure. All manipulations of positive smear specimens and BACTEC 12B bottles were performed in a biological safety hood. PCR ampli-fication and detection were performed according to the manufacturer’s guide-lines.
LiPA MYCOBACTERIA assay.Specimens were prepared for PCR amplifica-tion by removal of 0.2 ml from BACTEC 12B bottles. The pellet was resus-pended in 20l of TE buffer (10 mM Tris-HCl, 1 mM EDTA [pH 8.0]) and placed in a 95°C heat block for 30 min, followed by centrifugation at 16,000⫻g
for 10 s. The tubes were placed in a⫺20°C freezer for 30 min. Upon thawing, samples were vortexed and centrifuged at 16,000⫻gfor 10 s. A reaction mix of 40l was prepared from the supplied amplification mixture containing de-oxynucleoside triphosphates, biotinylated primers, and thermostable DNA poly-merase, to which 10l of the processed specimen was added. The PCR was done in a Perkin-Elmer 9600 thermocycler with the following amplification profile: 95°C for 30 s, 62°C for 30 s, and 72°C for 30 s for 40 cycles. The presence of the amplified product was verified by electrophoresis of 10l of the amplified product in a 2.0% agarose gel followed by staining with ethidium bromide. The expected size of the amplicon was a single band with a length of 400 to 550 bp. For hybridization, a 10-l sample of the amplified product was denatured in the hybridization trough, followed by addition of the hybridization solution provided in the assay kit and the membrane strip. The hybridization solution was pre-warmed to 62°C. The tray of strips was placed in a 62°C shaking water bath (80 rpm) with a lid and incubated for 30 min (model Gemini II incubator; Robbins Scientific, Sunnyvale, Calif.). After hybridization, two stringent washes were done at 62°C. The remainder of the procedure was done at room temperature using a rotary shaker at 80 rpm. Each strip was washed twice for 1 min using 2.0 ml of rinse solution, followed by addition of alkaline phosphatase-streptavidin conjugate solution for 30 min. Each strip was washed twice with rinse solution and once with 2.0 ml of the substrate buffer prior to incubation with the substrate
(5-bromo-4-chloro-3-indoylphosphate and nitroblue tetrazolium) solution for 30 min while being shaken. The color development was stopped by washing the strips twice in 2.0 ml of distilled water with shaking for 3 min. After hybridization and detection, each strip was aligned along the reading chart for interpretation using a green line at the top of the strip for reference.
PCR-RFLP analysis.The PCR-RFLP analysis to identify theMycobacterium
species was done by PCR amplification of a 439-bp segment of the mycobacterial 65-kDa heat shock protein gene (27, 28). Specimens from positive BACTEC 12B bottles were processed for PCR using the same method as that used for the LiPA assay. The PCR was done in a Perkin-Elmer 9600 thermocycler with the follow-ing amplification profile: 95°C for 30 s, 62°C for 30 s, and 72°C for 30 s for 35 cycles.BstEII andHaeII (Sigma, St. Louis, Mo.) enzyme digestions of the am-plification product were performed, and restriction fragments were separated by agarose gel electrophoresis of a 3.0% gel composed of 2.0% high-resolution agarose (Sigma) and 1.0% routine-use agarose (Sigma). The molecular size standard (MspI-digested pUC18; Sigma) was placed in the gel every four lanes to reduce migration-related errors in interpretation of fragment sizes. Photographs were taken of the gels after ethidium bromide staining and were analyzed visually to determine the number and the sizes of the fragments present. Isolates were identified using a published PCR-RFLP analysis algorithm (28, 29). A website (www.hospvd.ch/prasite) was also used for pattern analysis and species identifi-cation.
RESULTS
LiPA PCR amplification of the 16 to 23S rRNA.
Amplified
product was detected in 57 of 60 patient samples after the first
amplification. The three negative samples were positive when
they were diluted 1:2 or 1:10 prior to amplification. The
am-plified product yielded a clear band in the range of 400 to 500
bp.
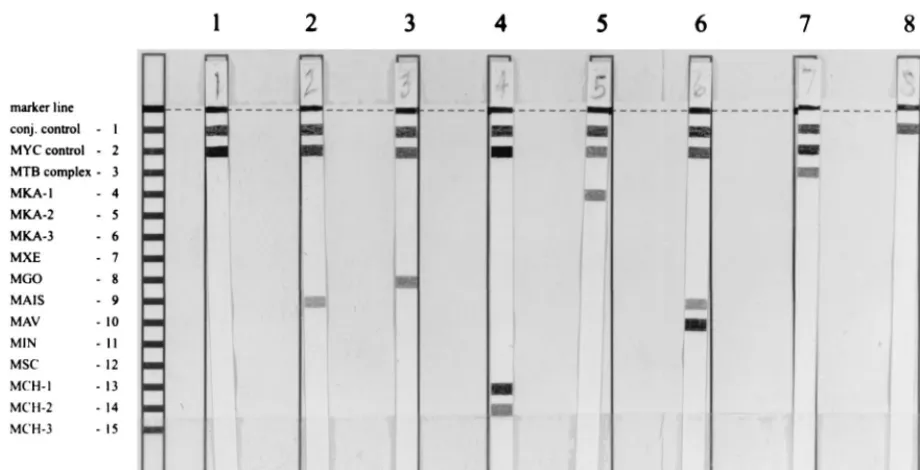
LiPA MYCOBACTERIA assay identification.
Figure 1
shows a representative sampling of results of the assay. Each
line number which was positive on the LiPA
MYCOBACTE-RIA strip was noted and used to determine the
Mycobacterium
species by using the probe alignment guide included in the kit
shown on the left side in Fig. 1. The conjugate control line and
Mycobacterium
species positive control line must be positive
for a valid result. The LiPA assay identified 60 mycobacterium
isolates from 59 patients (Table 1). One patient sample
pro-FIG. 1. Representative examples of results of the LiPA MYCOBACTERIA assay. The positions of the conjugate (conj.) control,Mycobacteriumgenus-positive control, and the 13 specific probes are shown on the left. The marker line at the top of the strip is used for orientation of the strip for analysis. Lanes: 1,M. fortuitum;2,M. avium-M. intracellularecomplex; 3,M. gordonae; 4,M. chelonae; 5,M. kansasii; 6,M. avium; 7,M. tuberculosis; 8, conjugate control withoutMycobacteriumDNA
present. MYC,Mycobacteriumcomplex; MTB,M. tuberculosiscomplex; MKA,M. kansasii; MXE,M. xenopi; MGO,M. gordonae; MAIS,M. avium-M. intracellulare
complex; MAV,M. avium; MIN,M. intracellulare; MSC,M. scrofulaceum; MCH,M. chelonae.
on May 15, 2020 by guest
http://jcm.asm.org/
[image:2.612.74.534.72.307.2]TABLE 1. Line probe assay results and identification of mycobacterial species by PCR and RFLP analysis of 60 patient samples
aSample Smearb GIc Gel resultd LIPA result LIPA ID Other testinge RFLP
1 Neg 999 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
2 1⫹ 999 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
3 Neg 999 Smear 480–550 MYC Mycobacteriumspecies MFO biochem ID MFO
4 2⫹ 428 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
5 3⫹ 193 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
6 Neg 211 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
7 4⫹ 533 Clear 400, weak 850 MYC, MTB M. tuberculosis MTB PCR MTB
8 4⫹ 999 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
9 Neg 783 Clear 400, weak 850 MYC Mycobacteriumspecies MSZ biochem ID MSZ
10 Neg 999 Clear 400 MYC, MGO M. gordonae MGO probe MGO
11 Neg 71 Clear 400, weak 850 MYC, MGO M. gordonae MGO probe MGO
12 Neg 303 Clear 400, weak
clear 500 MYC, MKA-1 M. kansasii MKA probe MKA
13 2⫹ 999 Clear 400 MYC, MKA-1 M. kansasii MKA probe MKA
14 Neg 999 Clear 450 MYC Mycobacteriumspecies MFO biochem ID MFO
15 Neg 205 Clear 400 MYC, MAIS MAIS complex MIN probe MIN
16 Neg 61 Clear 400 MYC, MAIS, MIN M. intracellulare MIN probe MIN
17 4⫹ 999 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
18 3⫹ 622 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
19 2⫹ 999 Smear 450–500 MYC Mycobacteriumspecies MFO biochem ID MFO
20 Neg 653 Clear 400 MYC, MGO M. gordonae MGO probe MGO
21 Neg 999 Clear 400, clear 280 MYC, MTB M. tuberculosis MTB PCR MTB
22 Neg 999 Smear 400–550 MYC, MTB M. tuberculosis MTB PCR MTB
23 Neg 999 Smear 400–550 MYC, MCH-1, MCH-2 M. chelonaegroup III MCH group and MFO
biochem ID MAB and MFO
24 Neg 999 Smear 400–550 MYC, MCH-1, MCH-2 M. chelonaegroup III MCH group biochem ID MAB
25 4⫹ 999 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
26 Neg 999 450, smear 480–520 MYC Mycobacteriumspecies MFO biochem ID MFO
27 Neg 999 Clear 400 MYC, MAIS, MAV M. avium MAV probe MAV
28 Neg 999 Smear 400–450 MYC, MAIS, MIN, MCH-1,
MCH-2 M. intracellulareM. chelonaeIIIand MIN probe and MCHgroup biochem ID MIN and MAB
29 Neg 711 Clear 400 MYC, MAIS, MAV M. avium MAV probe MAV
30 3⫹ 999 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
31 Neg 999 Clear 400 MYC, MCH-1, MCH-2 M. chelonaegroup III MCH group biochem ID MAB
32 Neg 999 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
33 Neg 999 Clear 400 MYC, MAIS, MAV M. avium MAV probe MAV
34 Neg 999 Clear 400 MYC, MAIS, MIN M. intracellulare MIN probe MIN
35 Neg 999 Clear 400 MYC, MCH-1, MCH-2 M. chelonaegroup III MCH group biochem ID MAB
36 Neg 999 Clear 400 MYC, MAIS, MAV M. avium MAV probe MAV
37 Neg 999 Clear 400 MYC, MAIS, MAV M. avium MAV probe MAV
38 Neg 999 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
39 3⫹ 999 Clear 400 MYC, MAIS, MAV M. avium MAV probe MAV
40 2⫹ 999 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
41 3⫹ 999 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
42 Neg 999 Clear 400 MYC, MGO M. gordonae MGO probe MGO
43 Neg 999 Clear 400, smear
450–520 MYC Mycobacteriumspecies MFO biochem ID MFO
44 Neg 999 Clear 460 MYC Mycobacteriumspecies MFO biochem ID MFO
45 Neg 999 Clear 400 MYC, MAIS, MAV M. avium MAV probe MAV
46 1⫹ 999 Clear 400 MYC, MAIS, MAV M. avium MAV probe MAV
47 4⫹ 999 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
48 Neg 999 Clear 400 MYC, MAIS, MAV M. avium MAV probe MAV
49 Neg Neg Clear 380, 550,
smear 400⫹ Conjugate control only Nonmycobacterial No mycobacteria isolated
50 4⫹ 999 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
51 1⫹ 999 Clear 400 MYC, MAIS MAIS complex MIN probe MIN
52 Neg 999 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
53 Neg 999 Clear 400 MYC, MKA-1 M. kansasii MKA probe MKA
54 4⫹ 999 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
55 4⫹ 999 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
56 4⫹ 999 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
57 2⫹ 548 Smear 450–500 MYC, MAIS MAIS complex MAIS probe MIN
58 3⫹ 34 Clear 400 MYC, MTB M. tuberculosis MTB PCR MTB
59 3⫹ 66 Weak 400 MYC, MTB M. tuberculosis MTB PCR MTB
60 3⫹ 23 Weak 450 MYC, MTB M. tuberculosis MTB PCR MTB
aMYC,Mycobacteriumspecies; MTB,M. tuberculosis; MGO,M. gordonae; MAIS,M. avium-M. intracellularecomplex; MIN,M. intracellulare; MAV,M. avium;
MKA-1,M. kansasiigroup I; MCH-1,M. chelonaegroups I, II, III, and IV; MCH-2,M. chelonaegroup III; MFO,M. fortuitum; MSZ,M. szulgai; MAB,M. abscessus; ID, identification; biochem, biochemical testing.
bSmear, concentrated acid-fast organism smear negative (Neg, negative; 1⫹to 4⫹, positive). cGI, growth index of BACTEC 12B bottle.
dAppearance and size(s) of band(s) (in base pairs). eOther testing, method of identification.
on May 15, 2020 by guest
http://jcm.asm.org/
duced amplified product of the correct size but was negative in
the LiPA assay.
There was complete agreement between the LiPA
MYCO-BACTERIA assay and the laboratory identification tests for 26
M. tuberculosis, 9
M. avium, 3
M. intracellulare, 3
M. kansasii, 4
M. gordonae, and 5
M. chelonae
group (all were
M. abscessus)
isolates from positive BACTEC 12B bottles (Table 2). In one
patient sample, bands were present for both
M. intracellulare
and
M. chelonae. This patient has a history of cultures positive
for both organisms, and the PCR-RFLP yielded the same
iden-tification. There were six isolates of
M. fortuitum
and one
isolate of
M. szulgai; all were positive for the
Mycobacterium
species probe, which identified them as mycobacterial species.
The assay is not capable of species identification for
M.
fortui-tum
and
M. szulgai.
The nine
M. avium
isolates were reactive with the
M.
avium-M. intracellulare
complex probe (M. avium,
M. intracellulare,
M. scrofulaceum,
M. malmoense, and
M. haemophilum) and
the
M. avium
probe (M. avium,
M. paratuberculosis, and
M.
sil-vaticum). The three
M. intracellulare
isolates were reactive with
the
M. avium-M. intracellulare
probe and the
M. intracellulare
probe. The four
M. gordonae
isolates were reactive with the
M. gordonae
probe. The three
M. kansasii
isolates were all
reactive with the
M. kansasii
group I probe (MKA-1) and
negative with the MKA-2 (group II) and MKA-3 (groups III,
IV, and V and
M. gastri) probes.
M. kansasii
isolates are
di-vided into five groups based on data derived from 16 to 23S
rRNA spacer nucleotide sequences (1). The MKA-1 probe
reacts with
M. kansasii
type I, the most frequent
M. kansasii
isolate from human sources worldwide (1).
M. kansasii
group
II, which is detected by the MKA-2 probe, is isolated from both
humans and the environment and is characterized by negative
hybridization in the ACCUPROBE assay.
M. kansasii
groups
III, IV, and V have rarely been isolated from humans but have
been found in environmental samples (1). The five
M. chelonae
solates were reactive with the MCH-1 probe (M. chelonae
groups
I, II, III, and IV) and the MCH-2 probe (M. chelonae
group
III).
M. chelonae
isolates are divided into four genotypical
clusters based on 16 to 23S rRNA nucleotide sequences (22).
The MCH-1 probe reacts with all four clusters, and the MCH-2
probe reacts with cluster III isolates, which encompass
M.
che-lonae
and
M. abscessus. The PCR-RFLP analysis identified the
five
M. chelonae
isolates as
M. abscessus. One patient isolate
that was identified as an
M. chelonae
group isolate by the LiPA
assay and as
M. abscessus
by PCR-RFLP analysis had been
identified as
M. fortuitum
by conventional methods. Closer
inspection found the culture to contain two organisms,
M.
for-tuitum
and
M. abscessus. We were unable to determine if both
species were detected in the LiPA assay since a specific probe
for
M. fortuitum
is not available and the reactivity with the
Mycobacterium
species probe may have been due to either or
both organisms. The presence of both organisms was
con-firmed by PCR-RFLP analysis of two different colonies from
the 7H11 agar plate.
DISCUSSION
The differentiation of species of
Mycobacterium
has
tradi-tionally been done by evaluation of growth characteristics and
biochemical testing. Rapid methodologies such as those using
DNA probes are limited by the number of available
commer-cial probes. In our laboratory, 50% of the acid-fast isolates
recovered from specimens are not MOTT; therefore, a
com-prehensive rapid detection method capable of identifying
mul-tiple species of mycobacteria in a single test would have a
significant impact. This paper describes the evaluation of a
practical method for the identification of mycobacterial DNA
amplified by PCR from acid-fast-bacillus-positive BACTEC
12B bottles. The LiPA line probe assay employs a reverse
hybridization reaction of biotin-labeled amplified DNA with
specific oligonucleotide probes fixed as parallel lines on a
membrane strip. This method was compared to a PCR-RFLP
procedure that is capable of differentiating 28 species of
clin-ically encountered mycobacteria (28, 29).
The LiPA MYCOBACTERIA test was easy to perform, and
the interpretation of the results was clear-cut and objective.
The LiPA assay identified 60 mycobacterium isolates from 59
patients. Six of seven of the isolates were
M. fortuitum
and one
was
M. szulgai, for which the assay does not have specific
probes; therefore, they were identified as
Mycobacterium
spe-cies in the LiPA assay. The assay correctly identified 50 of 53
isolates to the species level. The remaining three isolates were
identified as
M. avium-M. intracellulare
group isolates by LiPA
and were identified as
M. intracellulare
by PCR-RFLP analysis
and with the ACCUPROBE DNA probe. One culture was
found to contain two organisms by RFLP analysis,
M. fortuitum
and
M. abscessus. M. abscessus
was correctly identified in the
LiPA assay with the
M. chelonae
group probes. Since the
pres-ence of
M. fortuitum
would not be distinguishable from
M.
che-lonae, we cannot determine whether both organisms were
am-plified and detected in the assay.
In a smaller study of 27 specimens from liquid culture (S. A.
Watterson, B. A. Hussein, and F. A. Drobniewski, Abstr. 99th
Gen. Meet. Am. Soc. Microbiol. 1999, abstr. U-29, p. 639, 1999),
there was agreement with standard methods of identification
for 26 of 27 of the isolates. The discrepant sample was
identi-fied by standard biochemical methods as
M. fortuitum
but as
M.
chelonae
group III by the LiPA assay. However, a subsequent
sample from the same patient was identified as
M. chelonae
by
standard biochemical methods.
The main advantage of LiPA compared to testing with DNA
probes is that LiPA can identify a wide range of species in a
single assay instead of a technician performing a different test
for each species or waiting for growth on solid media to guide
the choice of DNA probe. The PCR-RFLP method has the
same advantage over the use of DNA probes, although the
PCR-RFLP results require more expertise to interpret than
the LiPA results. It was not surprising that the three
M.
kan-TABLE 2. Summary of mycobacterial species identification
by all methods tested
Organism isolatesNo. of
No. of isolates positive by:
LiPA M. tuber-culosis
PCR Probe RFLPanalysis Bio-chemical
testing
M. tuberculosis 26 26 26 26
M. kansasii 3 3 3 3
M. avium-M.
intracellu-larecomplex 15 15 15 15
M. avium 9 9 9 9
M. intracellulare 6 3b 5c 6
M. gordonae 4 4 4 4
M. fortuitum 7 7d,e 7 7
M. chelonaegroupa 5 5 5 5
M. szulgai 1 1d 1 1
aIncludesM. chelonaeandM. abscessus.
bAll six were positive by theM. avium-M. intracellularecomplex probe. cAll were positive by theM. avium-M. intracellularecomplex probe
(ACCU-PROBE).
dIdentified asMycobacteriumspecies only.
eOne sample was positive for two different species by culture, but we were
unable to determine if two species were present in the LiPA samples.
on May 15, 2020 by guest
http://jcm.asm.org/
[image:4.612.53.294.90.231.2]sasii
isolates were all reactive with the
M. kansasii
group I
probe (MKA-1) and negative with the probes for group II
(MKA-2) and group III, IV, and V and
M. gastri
(MKA-3)
isolates since
M. kansasii
group I represents the most common
clinical isolate from humans (1). Group I has a PCR-RFLP
pattern distinguishable from those of other
M. kansasii
groups,
and in our laboratory all clinical isolates have been group I.
In our laboratory, 50% of mycobacterial isolates in 1998
were
M. tuberculosis; the majority of isolates were identified by
an
M. tuberculosis
complex PCR assay. An additional 34.7% of
our isolates were identified using DNA probes for
M. avium-M.
intracellulare
complex (29.7%),
M. kansasii
(3.3%), and
M.
gordonae
(1.7%). The LiPA assay was able to identify these
isolates directly from the positive BACTEC 12B bottles with a
single assay and could also identify
M. chelonae, which
com-prised 5.7% of our isolates. In total, the LiPA assay could have
identified 90.6% of our isolates to the species level (1998 data).
The remaining 9.4% of isolates in our lab are composed mainly
of miscellaneous organisms, including
M. fortuitum, for which
the LiPA assay does not have a specific probe. In conclusion,
we found the LiPA MYCOBACTERIA assay to be an easy
test to perform in the clinical setting and one that provides
identification of a large variety of mycobacterial species in a
single test. As yet, the cost of the assay has not been set by the
manufacturer (Innogenetics NV).
The line probe assay technology has also been used for
detection of mutations in the
rpoB
gene of
M. tuberculosis
that
confer resistance to rifampin (12). The rate of concordance
with phenotypic rifampin susceptibility testing results was
92.2%. Another application of the line probe assay is to detect
and identify human papillomavirus (HPV) strains using a strip
with 28 specific probes for each of the 25 HPV genotypes (15).
Since HPV cannot be cultured efficiently, diagnosis of HPV
infection is based on cytology and molecular tools, which
makes this organism a perfect candidate for this type of assay.
REFERENCES
1.Alcaide, F., I. Richter, C. Bernasconi, B. Springer, C. Hagenau, R. Schulze-Ro¨bbecke, E. Tortoli, R. Martı´n, E. C. Bo¨ttger, and A. Telenti.1997. Het-erogeneity and clonality among isolates ofMycobacterium kansasii: implica-tions for epidemiological and pathogenicity studies. J. Clin. Microbiol.35: 1959–1964.
2.Badak, F. Z., S. Goskel, R. Sertoz, B. Nafile, S. Ermertcan, C. Cavusoglu, and A. Bilgic.1999. Use of nucleic acid probes for identification of
Myco-bacterium tuberculosisdirectly from MB/BacT bottles. J. Clin. Microbiol.37:
1602–1605.
3.Body, B. A., N. G. Warren, A. Spicer, D. Henderson, and M. Chery.1990. Use of Gen-Probe and Bactec for rapid isolation and identification of mycobac-teria. Correlation of probe results with growth index. Am. J. Clin. Pathol.93: 415–420.
4.Chan, J., J. C. McKitrick, and R. S. Klein.1984.Mycobacterium gordonaein the acquired immunodeficiency syndrome. Ann. Intern. Med.101:400. 5.D’Amato, R. F., A. A. Wallman, L. H. Hochstein, P. M. Colaninno, M.
Scardamaglia, E. Ardila, M. Ghouri, K. Kim, R. C. Patel, and A. Miller. 1995. Rapid diagnosis of pulmonary tuberculosis by using Roche
AMPLI-CORMycobacterium tuberculosisPCR test. J. Clin. Microbiol.33:1832–1834.
6.Ellner, P. D., T. E. Kiehn, R. Cammarata, and M. Hosmer.1988. Rapid detection and identification of pathogenic mycobacteria by combining radio-metric culture and nucleic acid probe methods. J. Clin. Microbiol.26:1349– 1352.
7.Evans, K. D., A. S. Nakasone, P. A. Sutherland, L. M. de la Maza, and E. M. Peterson.1992. Identification ofMycobacterium tuberculosisand
Mycobacte-rium avium-M. intracellularedirectly from primary BACTEC cultures by
us-ing acridinium-ester-labeled DNA probes. J. Clin. Microbiol.30:2427–2431. 8.Falkinham, J. O., III.1996. Epidemiology of infection by nontuberculosis
mycobacteria. Clin. Microbiol. Rev.9:177–215.
9.Gamboa, F., G. Fernandez, E. Padilla, J. M. Manterola, J. Lonca, P. J. Cardona, L. Matas, and V. Ausina.1998. Comparative evaluation of initial and new versions of the Gen-Probe amplifiedMycobacterium tuberculosis
direct test for direct detection ofMycobacterium tuberculosisin respiratory and nonrespiratory specimens. J. Clin. Microbiol.36:684–689.
10. Glickman, S. E., J. O. Kilburn, W. R. Butler, and L. S. Ramos.1994. Rapid identification of mycolic acid patterns of mycobacteria by high-performance
liquid chromatography using pattern recognition software and a Mycobacte-riumlibrary. J. Clin. Microbiol.32:740–745.
11. Hines, M. E., II, and K. S. Frazier.1993. Differentiation of mycobacteria on the basis of chemotype profiles using matrix solid-phase dispersion and thin-layer chromatography. J. Clin. Microbiol.31:610–614.
12. Hirano, K., C. Abe, and M. Takahashi.1999. Mutations in therpoBgene of rifampin-resistant Mycobacterium tuberculosis strains isolated mostly in Asian countries and their rapid detection by line probe assay. J. Clin. Mi-crobiol.37:2663–2666.
13. Kent, P. T., and G. P. Kubica.1985. Public health mycobacteriology: a guide for the level III laboratory. Centers for Disease Control, U.S. Department of Health and Human Services, Atlanta, Ga.
14. Kirihara, J. M., S. L. Hillier, and M. B. Coyle.1985. Improved detection times forMycobacterium aviumcomplex andMycobacterium tuberculosiswith the BACTEC radiometric systems. J. Clin. Microbiol.22:841–845. 15. Kleter, B., L. van Doorn, L. Schrauwen, A. Molijn, S. Sastrowijoto, J. ter
Schegget, J. Lindeman, B. ter Harmsel, M. Burger, and W. Quint.1999. Development and clinical evaluation of a highly sensitive PCR-reverse hy-bridization line probe assay for detection and identification of anogenital human papillomavirus. J. Clin. Microbiol.37:2508–2517.
16. Metchock, B., and L. Diem.1995. Algorithm for use of nucleic acid probes for identifying Mycobacterium tuberculosis from BACTEC 12B bottles. J. Clin. Microbiol.33:1934–1937.
17. Morgan, M. A., C. D. Horstmeier, D. R. DeYoung, and G. D. Roberts.1983. Comparison of radiometric method (BACTEC) and conventional culture media for recovery of mycobacteria from smear-negative specimens. J. Clin. Microbiol.18:384–388.
18. Musial, C. E., L. S. Tice, L. Stockman, and G. D. Roberts.1988. Identifica-tion of mycobacteria from culture by using the Gen-Probe Rapid Diagnostic System forMycobacterium aviumcomplex andMycobacterium tuberculosis
complex. J. Clin. Microbiol.26:2120–2123.
19. Nolte, F. S., and B. Metchock.1995. Mycobacterium, p. 400–437.InP. R. Murray, E. J. Baron, M. A. Pfaller, F. C. Tenover, and R. H. Yolken (ed.), Manual of clinical microbiology, 6th ed. ASM Press, Washington, D.C. 20. Pfyffer, G. E., H. M. Welscher, P. Kissling, C. Cieslak, M. J. Casal, J.
Gutierrez, and S. Rusch-Gerdes.1997. Comparison of the Mycobacteria Growth Indicator Tube (MGIT) with radiometric and solid culture for re-covery of acid-fast bacilli. J. Clin. Microbiol.35:364–368.
21. Portaels, F.1996. Epidemiology of mycobacterial diseases. Clin. Dermatol. 13:207–222.
22. Portaels, F., P. de Rijk, G. Jannes, R. Lemans, W. Mijs, L. Rigouts, and R. Rossau.1996. The 16S–23S rRNA spacer, a useful tool for taxonomical and epidemiological studies for theM. chelonaecomplex. Tubercle Lung Dis. 77(Suppl. 1):17.
23. Pozniak, A. L., A. H. C. Uttley, and R. J. Kent.1996.Mycobacterium avium
complex in AIDS: who, when, where, why and how? Soc. Appl. Bacteriol. Symp. Ser.25:40S–46S.
24. Ratnam, S., F. A. Stead, and M. Howes.1987. Simplified acetylcysteine-alkali digestion-decontamination procedure for isolation of mycobacteria from clinical specimens. J. Clin. Microbiol.25:1428–1432.
25. Reisner, B. S., A. M. Gatson, and G. L. Woods.1994. Use of Gen-Probe AccuProbes to identifyMycobacterium aviumcomplex,Mycobacterium
tuber-culosiscomplex,Mycobacterium kansasii, andMycobacterium gordonae
di-rectly from BACTEC TB broth cultures. J. Clin. Microbiol.32:2995–2998. 26. Roberts, G. D., N. L. Goodman, L. Heiferts, H. W. Larsh, T. H. Lindner, J. K.
McClatchy, M. R. McGinnis, S. H. Siddiqi, and P. Wright.1983. Evaluation of the BACTEC radiometric method for recovery of mycobacteria and drug susceptibility testing of Mycobacterium tuberculosisfrom acid-fast smear-positive specimens. J. Clin. Microbiol.18:689–696.
27. Schinnick, T.1987. The 65-kilodalton antigen ofMycobacterium tuberculosis. J. Bacteriol.169:1080–1086.
28. Taylor, T. B., C. Patterson, Y. Hale, and W. W. Safranek.1997. Routine use of PCR-restriction fragment length polymorphism analysis for identification of mycobacterium growing in liquid media. J. Clin. Microbiol.35:79–85. 29. Telenti, A., F. March, M. Bald, F. Badly, E. Bottger, and T. Bodmer.1993.
Rapid identification of mycobacteria to the species level by PCR and restric-tion enzyme analysis. J. Clin. Microbiol.31:175–178.
30. Tisdall, P. A., G. D. Roberts, and J. P. Anhalt.1979. Identification of clinical isolates of mycobacteria with gas-liquid chromatography alone. J. Clin. Mi-crobiol.10:506–514.
31. Tortoli, E., F. Lavinia, and M. T. Simonetti.1997. Evaluation of a commer-cial ligase chain reaction kit (Abbott Lcx) for direct detection of
Mycobac-terium tuberculosisin pulmonary and extrapulmonary specimens. J. Clin.
Microbiol.35:2424–2426.
32. Vuorinen, P., A. Miettinen, R. Vuento, and O. Ha¨llstrom.1995. Direct detection ofMycobacterium tuberculosiscomplex in respiratory specimens by Gen-Probe Amplified Mycobacterium Tuberculosis Direct Test and Roche Amplicor Mycobacterium Tuberculosis Test. J. Clin. Microbiol.33:1856– 1859.
33. Wayne, L. G., and C. Sramek II.1992. Agents of newly recognized or infre-quently encountered mycobacterial diseases. Clin. Microbiol. Rev.5:1–25.