organic papers
o724
Christian Peiferet al. C21H16N2O5C2H6OS doi:10.1107/S160053680500485X Acta Cryst.(2005). E61, o724–o725
Acta Crystallographica Section E
Structure Reports
Online
ISSN 1600-5368
5,6,7-Trimethoxy-2,3-dihydro-1
H
,8
H
-benzo[
a
]-pyrrolo[3,4-
c
]carbazole-1,3-dione dimethyl
sulfoxide solvate
Christian Peifer,a* Dieter Schollmeyerb and Gerd Dannhardtc
aPharmazeutisches Institut, Auf der Morgenstelle
8, Universita¨t Tu¨bingen, 72076 Tu¨bingen, Germany,b
Institut fu¨r Organische Chemie der Universita¨t Mainz, Duesbergweg 10-14, D-55099 Mainz, Germany, andc
Institut fu¨r Pharmazie, Staudingerweg 5, D-55099 Mainz, Germany
Correspondence e-mail: christian.peifer@uni-tuebingen.de
Key indicators
Single-crystal X-ray study
T= 295 K
Mean(C–C) = 0.005 A˚
Rfactor = 0.076
wRfactor = 0.187
Data-to-parameter ratio = 14.2
For details of how these key indicators were automatically derived from the article, see http://journals.iucr.org/e.
#2005 International Union of Crystallography
Printed in Great Britain – all rights reserved
The crystal structure of the title compound, C21H16N2O5
-C2H6OS, was determined to investigate the electrocyclic
reactivity of 3,4-diaryl-1H-pyrrole-2,5-diones (3,4-bisarylmal-eimides) to the yield corresponding carbazole derivatives.
Comment
The title compound, (III), bearing the carbazole moiety as a core structure, was accidentally isolated from an ethyl acetate solution of 3-(indol-3-yl)-4-(3,4,5-trimethoxyphenyl)-1H -pyrrole-2,5-dione, (I) (Peiferet al., 2005) at room temperature. The reaction scheme below shows the disrotatory cyclization of (I) and subsequent oxidation to yield (II).
The analytically pure 1H-pyrrole-2,5-dione derivative was found to undergo a reaction (monitored by thin-layer chro-matography) producing (II). A comparable mechanism of reactions of the class of 1H-pyrrole-2,5-diones had been reported by Sanchez-Martinez et al.(2003) and Harris et al.
(1993). However, after 24 h in an ethyl acetate solution, approximately 10% of (II) could be determined by high-performance liquid chromatographic analysis. Compound (II) was subsequently isolated by column chromatography and found to be chemically stable. Crystals of (I) precipitated at 278 K from DMSO. We now report the X-ray crystal structure analysis of carbazole (III), which is the DMSO solvate of carbazole (II) and which confirms the structure and strongly supports the mechanism of oxidative cyclization of the 1H
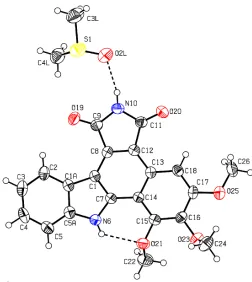
pyrrole-2,5-dione derivative to generate compound (II). The solvent DMSO molecule is linkedviaa hydrogen bond to the carbazole molecule (see Table 1 and Fig. 1).
Experimental
The title compound was obtained by crystallization of a DMSO solution of (II).
Crystal data
C21H16N2O5C2H6OS
Mr= 454.50
Monoclinic,P21=c
a= 7.994 (2) A˚
b= 20.040 (4) A˚
c= 13.644 (4) A˚
= 106.586 (12)
V= 2094.8 (9) A˚3
Z= 4
Dx= 1.441 Mg m
3
CuKradiation Cell parameters from 25
reflections
= 30–44
= 1.76 mm1
T= 295 (2) K Needle, yellow 0.240.060.04 mm
Data collection
Enraf–Nonius CAD-4 diffractometer
!/2scans
Absorption correction: scan (CORINC; Dra¨ger & Gattow, 1971)
Tmin= 0.783,Tmax= 0.932
4534 measured reflections 4228 independent reflections
2943 reflections withI> 2(I)
Rint= 0.062 max= 74.0
h=9!0
k= 0!24
l=16!17 3 standard reflections
frequency: 60 min intensity decay: 5%
Refinement
Refinement onF2
R[F2> 2(F2)] = 0.076
wR(F2) = 0.187
S= 1.11 4228 reflections 298 parameters
H-atom parameters constrained
w= 1/[2(F
o2) + (0.0644P)2
+ 1.9378P] whereP= (Fo
2
+ 2Fc 2
)/3 (/)max< 0.001
max= 0.32 e A˚ 3
min=0.35 e A˚ 3
Table 1
Hydrogen-bonding geometry (A˚ ,).
D—H A D—H H A D A D—H A
N6—H6 O21 0.79 2.20 2.741 (4) 126
N10—H10 O2L 0.87 2.00 2.849 (5) 164
H atoms attached to N atoms were located in a difference map. All H atoms were refined as riding with idealized geometry; C—H = 0.93 or 0.96 A˚ and the N—H distances (Table 1) were optimised to fit the electron density. Uiso(H) values were refined freely for H atoms bonded to N atoms, but was constrained to 1.5Ueq(C) for methyl groups and to 1.2Ueq(C) for H atoms bonded to other C atoms. The methyl groups were allowed to rotate but not to tip.
Data collection: CAD-4 Software (Enraf–Nonius, 1989); cell refinement: CAD-4 Software; data reduction: CORINC (Dra¨ger & Gattow, 1971); program(s) used to solve structure:SIR92(Altomare et al., 1994); program(s) used to refine structure:SHELXL97 (Shel-drick, 1997); molecular graphics:PLATON(Spek, 2003); software used to prepare material for publication:SHELXL97.
References
Altomare, A., Cascarano, G., Giacovazzo, C., Guagliardi, A., Burla, M. C., Polidori, G. & Camalli, M. (1994).J. Appl. Cryst.27, 435.
Dra¨ger, M. & Gattow, G. (1971).Acta Chem. Scand.25, 761–762.
Enraf–Nonius (1989).CAD-4 Software. Version 5. Enraf–Nonius, Delft, The Netherlands.
Harris, W., Hill, C. H., Keech, E. & Malsher, P. (1993).Tetrahedron Lett.34, 8361–8364.
Johnson, C. K. (1976).ORTEPII. ORNL-5138, revised. Oak Ridge National Laboratory, Tennessee, USA.
Peifer, C., Schollmeyer, D. & Dannhardt, G. (2005).Acta Cryst.E61, o720– o722.
Sanchez-Martinez, C., Faul, M. M., Shih, C., Sullivan, K. A., Grutsch, J. L., Cooper, J. T. & Kolis, S. P. (2003).J. Org. Chem.68, 8008–8014.
[image:2.610.312.564.66.348.2]Sheldrick, G. M. (1997).SHELXS97. University of Go¨ttingen, Germany. Spek, A. L. (2003).J. Appl. Cryst.36, 7–13.
Figure 1
supporting information
sup-1
Acta Cryst. (2005). E61, o724–o725
supporting information
Acta Cryst. (2005). E61, o724–o725 [https://doi.org/10.1107/S160053680500485X]
5,6,7-Trimethoxy-2,3-dihydro-1
H
,8
H
-benzo[
a
]pyrrolo[3,4-
c
]carbazole-1,3-dione dimethyl sulfoxide solvate
Christian Peifer, Dieter Schollmeyer and Gerd Dannhardt
5,6,7-Trimethoxy-2,3-dihydro-1H,8H-benzo[a]pyrrolo[3,4-c]carbazole-1,3-dione dimethyl sulfoxide solvate
Crystal data
C21H16N2O5·C2H6OS
Mr = 454.50
Monoclinic, P21/c
Hall symbol: -P 2ybc a = 7.994 (2) Å b = 20.040 (4) Å c = 13.644 (4) Å β = 106.586 (12)° V = 2094.8 (9) Å3
Z = 4
F(000) = 952 Dx = 1.441 Mg m−3
Melting point: 272 K
Cu Kα radiation, λ = 1.54178 Å Cell parameters from 25 reflections θ = 30–44°
µ = 1.76 mm−1
T = 295 K Needle, yellow 0.24 × 0.06 × 0.04 mm
Data collection
Enraf–Nonius CAD-4 diffractometer
Radiation source: rotating anode Graphite monochromator θ/2ω scans
Absorption correction: psi-scan (CORINC; Dräger & Gattow, 1971) Tmin = 0.783, Tmax = 0.932
4534 measured reflections
4228 independent reflections 2943 reflections with I > 2σ(I) Rint = 0.062
θmax = 74.0°, θmin = 4.0°
h = −9→0 k = 0→24 l = −16→17
3 standard reflections every 60 min intensity decay: 5%
Refinement
Refinement on F2
Least-squares matrix: full R[F2 > 2σ(F2)] = 0.076
wR(F2) = 0.187
S = 1.11 4228 reflections 298 parameters 0 restraints
Primary atom site location: structure-invariant direct methods
Secondary atom site location: difference Fourier map
Hydrogen site location: inferred from neighbouring sites
H-atom parameters constrained w = 1/[σ2(F
o2) + (0.0644P)2 + 1.9378P]
where P = (Fo2 + 2Fc2)/3
(Δ/σ)max < 0.001
Δρmax = 0.32 e Å−3
Special details
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes.
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2,
conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used
only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2
are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger.
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
x y z Uiso*/Ueq
C1 0.7962 (4) 0.51606 (18) 0.3833 (3) 0.0311 (8) C1A 0.8386 (4) 0.52185 (18) 0.2874 (3) 0.0330 (8) C2 0.8208 (5) 0.4797 (2) 0.2033 (3) 0.0417 (9)
H2 0.7697 0.4378 0.2011 0.050*
C3 0.8809 (6) 0.5018 (2) 0.1238 (3) 0.0498 (10)
H3 0.8715 0.4741 0.0678 0.060*
C4 0.9549 (6) 0.5645 (2) 0.1252 (3) 0.0500 (11)
H4 0.9930 0.5780 0.0699 0.060*
C5 0.9731 (5) 0.6068 (2) 0.2065 (3) 0.0458 (10)
H5 1.0241 0.6486 0.2078 0.055*
C5A 0.9127 (5) 0.58497 (19) 0.2869 (3) 0.0375 (8) N6 0.9150 (4) 0.61706 (17) 0.3766 (2) 0.0368 (7) H6 0.954 (2) 0.6524 (19) 0.3967 (11) 0.039 (12)* C7 0.8459 (4) 0.57645 (17) 0.4360 (3) 0.0318 (8) C8 0.7298 (4) 0.46665 (18) 0.4343 (3) 0.0311 (8) C9 0.6783 (5) 0.39624 (19) 0.4048 (3) 0.0361 (8) N10 0.6322 (4) 0.36876 (17) 0.4856 (3) 0.0419 (8) H10 0.6202 (8) 0.326 (3) 0.4940 (6) 0.072 (16)* C11 0.6463 (5) 0.41469 (19) 0.5643 (3) 0.0361 (8) C12 0.7110 (4) 0.47758 (18) 0.5301 (3) 0.0331 (8) C13 0.7554 (4) 0.53893 (18) 0.5834 (3) 0.0308 (8) C14 0.8275 (4) 0.58967 (17) 0.5343 (3) 0.0304 (7) C15 0.8802 (4) 0.65043 (18) 0.5883 (3) 0.0340 (8) C16 0.8666 (5) 0.65931 (18) 0.6853 (3) 0.0367 (8) C17 0.7865 (5) 0.6089 (2) 0.7305 (3) 0.0372 (8) C18 0.7331 (4) 0.55044 (19) 0.6801 (3) 0.0348 (8)
H18 0.6813 0.5179 0.7104 0.042*
O19 0.6745 (4) 0.36796 (14) 0.3257 (2) 0.0470 (7) O20 0.6124 (4) 0.40231 (15) 0.6433 (2) 0.0497 (7) O21 0.9541 (3) 0.69891 (13) 0.5426 (2) 0.0440 (7) C22 0.8477 (7) 0.7568 (2) 0.5112 (4) 0.0611 (13)
H22A 0.7408 0.7443 0.4616 0.092*
H22B 0.9088 0.7885 0.4815 0.092*
H22C 0.8221 0.7764 0.5695 0.092*
supporting information
sup-3
Acta Cryst. (2005). E61, o724–o725
C24 1.0928 (5) 0.7125 (2) 0.8090 (3) 0.0554 (12)
H24A 1.0922 0.6782 0.8580 0.083*
H24B 1.1240 0.7543 0.8437 0.083*
H24C 1.1763 0.7014 0.7730 0.083*
O25 0.7720 (4) 0.62545 (15) 0.8243 (2) 0.0495 (7) C26 0.6739 (6) 0.5813 (3) 0.8681 (3) 0.0555 (12)
H26A 0.5638 0.5719 0.8190 0.083*
H26B 0.6542 0.6016 0.9275 0.083*
H26C 0.7375 0.5404 0.8874 0.083*
S1 0.62426 (15) 0.19399 (6) 0.38603 (10) 0.0582 (4) O2L 0.5746 (5) 0.22835 (17) 0.4712 (3) 0.0696 (10) C3L 0.5485 (8) 0.1110 (3) 0.3870 (4) 0.0769 (16)
H3L1 0.6110 0.0897 0.4497 0.115*
H3L2 0.5672 0.0870 0.3302 0.115*
H3L3 0.4261 0.1115 0.3815 0.115*
C4L 0.4746 (7) 0.2194 (3) 0.2697 (4) 0.0652 (14)
H4L1 0.3576 0.2120 0.2728 0.098*
H4L2 0.4942 0.1940 0.2143 0.098*
H4L3 0.4908 0.2660 0.2587 0.098*
Atomic displacement parameters (Å2)
U11 U22 U33 U12 U13 U23
C24 0.043 (2) 0.056 (3) 0.060 (3) 0.000 (2) 0.004 (2) −0.018 (2) O25 0.0596 (18) 0.0523 (18) 0.0409 (16) −0.0060 (14) 0.0211 (14) −0.0066 (13) C26 0.059 (3) 0.075 (3) 0.036 (2) −0.010 (2) 0.020 (2) −0.003 (2) S1 0.0432 (6) 0.0558 (7) 0.0736 (8) −0.0053 (5) 0.0133 (5) −0.0108 (6) O2L 0.088 (3) 0.057 (2) 0.064 (2) −0.0114 (19) 0.0214 (19) −0.0147 (17) C3L 0.092 (4) 0.050 (3) 0.083 (4) 0.006 (3) 0.016 (3) −0.007 (3) C4L 0.075 (3) 0.057 (3) 0.063 (3) 0.008 (3) 0.018 (3) −0.006 (2)
Geometric parameters (Å, º)
C1—C8 1.399 (5) C15—C16 1.371 (5)
C1—C7 1.406 (5) C15—O21 1.375 (4)
C1—C1A 1.447 (5) C16—O23 1.378 (4)
C1A—C5A 1.398 (5) C16—C17 1.427 (5)
C1A—C2 1.400 (5) C17—O25 1.358 (4)
C2—C3 1.380 (6) C17—C18 1.364 (5)
C2—H2 0.9300 C18—H18 0.9300
C3—C4 1.385 (6) O21—C22 1.429 (5)
C3—H3 0.9300 C22—H22A 0.9600
C4—C5 1.372 (6) C22—H22B 0.9600
C4—H4 0.9300 C22—H22C 0.9600
C5—C5A 1.390 (5) O23—C24 1.434 (5)
C5—H5 0.9300 C24—H24A 0.9600
C5A—N6 1.378 (5) C24—H24B 0.9600
N6—C7 1.371 (4) C24—H24C 0.9600
N6—H6 0.7902 O25—C26 1.423 (5)
C7—C14 1.416 (5) C26—H26A 0.9600
C8—C12 1.375 (5) C26—H26B 0.9600
C8—C9 1.493 (5) C26—H26C 0.9600
C9—O19 1.211 (4) S1—O2L 1.499 (3)
C9—N10 1.374 (5) S1—C4L 1.768 (5)
N10—C11 1.394 (5) S1—C3L 1.770 (5)
N10—H10 0.8698 C3L—H3L1 0.9600
C11—O20 1.209 (4) C3L—H3L2 0.9600
C11—C12 1.487 (5) C3L—H3L3 0.9600
C12—C13 1.421 (5) C4L—H4L1 0.9600
C13—C18 1.400 (5) C4L—H4L2 0.9600
C13—C14 1.427 (5) C4L—H4L3 0.9600
C14—C15 1.423 (5)
C8—C1—C7 116.5 (3) C16—C15—O21 120.6 (3)
C8—C1—C1A 136.6 (3) C16—C15—C14 121.1 (3)
C7—C1—C1A 106.8 (3) O21—C15—C14 118.2 (3)
C5A—C1A—C2 119.1 (3) C15—C16—O23 120.4 (3) C5A—C1A—C1 106.6 (3) C15—C16—C17 119.6 (3)
C2—C1A—C1 134.3 (4) O23—C16—C17 119.8 (3)
C3—C2—C1A 118.3 (4) O25—C17—C18 125.9 (4)
supporting information
sup-5
Acta Cryst. (2005). E61, o724–o725
C1A—C2—H2 120.9 C18—C17—C16 120.3 (3)
C2—C3—C4 121.6 (4) C17—C18—C13 120.7 (3)
C2—C3—H3 119.2 C17—C18—H18 119.6
C4—C3—H3 119.2 C13—C18—H18 119.6
C5—C4—C3 121.2 (4) C15—O21—C22 114.4 (3)
C5—C4—H4 119.4 O21—C22—H22A 109.5
C3—C4—H4 119.4 O21—C22—H22B 109.5
C4—C5—C5A 117.6 (4) H22A—C22—H22B 109.5
C4—C5—H5 121.2 O21—C22—H22C 109.5
C5A—C5—H5 121.2 H22A—C22—H22C 109.5
N6—C5A—C5 129.4 (4) H22B—C22—H22C 109.5
N6—C5A—C1A 108.4 (3) C16—O23—C24 113.2 (3)
C5—C5A—C1A 122.1 (4) O23—C24—H24A 109.5
C7—N6—C5A 110.0 (3) O23—C24—H24B 109.5
C7—N6—H6 120.7 H24A—C24—H24B 109.5
C5A—N6—H6 129.1 O23—C24—H24C 109.5
N6—C7—C1 108.2 (3) H24A—C24—H24C 109.5
N6—C7—C14 128.1 (3) H24B—C24—H24C 109.5
C1—C7—C14 123.7 (3) C17—O25—C26 116.7 (3)
C12—C8—C1 121.3 (3) O25—C26—H26A 109.5
C12—C8—C9 108.1 (3) O25—C26—H26B 109.5
C1—C8—C9 130.5 (3) H26A—C26—H26B 109.5
O19—C9—N10 125.7 (4) O25—C26—H26C 109.5
O19—C9—C8 128.1 (4) H26A—C26—H26C 109.5
N10—C9—C8 106.2 (3) H26B—C26—H26C 109.5
C9—N10—C11 111.9 (3) O2L—S1—C4L 107.6 (2)
C9—N10—H10 124.3 O2L—S1—C3L 105.0 (2)
C11—N10—H10 122.3 C4L—S1—C3L 97.6 (3)
O20—C11—N10 124.1 (4) S1—C3L—H3L1 109.5
O20—C11—C12 129.8 (4) S1—C3L—H3L2 109.5
N10—C11—C12 106.1 (3) H3L1—C3L—H3L2 109.5
C8—C12—C13 123.0 (3) S1—C3L—H3L3 109.5
C8—C12—C11 107.8 (3) H3L1—C3L—H3L3 109.5
C13—C12—C11 129.2 (3) H3L2—C3L—H3L3 109.5
C18—C13—C12 123.0 (3) S1—C4L—H4L1 109.5
C18—C13—C14 120.1 (3) S1—C4L—H4L2 109.5
C12—C13—C14 116.9 (3) H4L1—C4L—H4L2 109.5
C7—C14—C15 123.7 (3) S1—C4L—H4L3 109.5
C7—C14—C13 118.5 (3) H4L1—C4L—H4L3 109.5
C15—C14—C13 117.9 (3) H4L2—C4L—H4L3 109.5
C2—C3—C4—C5 0.7 (7) C11—C12—C13—C14 175.2 (3) C3—C4—C5—C5A −0.8 (6) N6—C7—C14—C15 0.3 (6) C4—C5—C5A—N6 179.9 (4) C1—C7—C14—C15 179.0 (3) C4—C5—C5A—C1A 1.2 (6) N6—C7—C14—C13 −178.4 (3) C2—C1A—C5A—N6 179.5 (3) C1—C7—C14—C13 0.4 (5) C1—C1A—C5A—N6 −0.5 (4) C18—C13—C14—C7 −179.2 (3) C2—C1A—C5A—C5 −1.5 (5) C12—C13—C14—C7 1.5 (5) C1—C1A—C5A—C5 178.4 (3) C18—C13—C14—C15 2.0 (5) C5—C5A—N6—C7 −178.3 (4) C12—C13—C14—C15 −177.2 (3) C1A—C5A—N6—C7 0.5 (4) C7—C14—C15—C16 −176.9 (3) C5A—N6—C7—C1 −0.3 (4) C13—C14—C15—C16 1.7 (5) C5A—N6—C7—C14 178.6 (3) C7—C14—C15—O21 −0.4 (5) C8—C1—C7—N6 177.1 (3) C13—C14—C15—O21 178.3 (3) C1A—C1—C7—N6 −0.1 (4) O21—C15—C16—O23 1.7 (5) C8—C1—C7—C14 −1.9 (5) C14—C15—C16—O23 178.3 (3) C1A—C1—C7—C14 −179.0 (3) O21—C15—C16—C17 178.8 (3) C7—C1—C8—C12 1.4 (5) C14—C15—C16—C17 −4.7 (5) C1A—C1—C8—C12 177.5 (4) C15—C16—C17—O25 −176.3 (3) C7—C1—C8—C9 −174.8 (3) O23—C16—C17—O25 0.8 (5) C1A—C1—C8—C9 1.3 (7) C15—C16—C17—C18 3.9 (5) C12—C8—C9—O19 178.8 (4) O23—C16—C17—C18 −179.0 (3) C1—C8—C9—O19 −4.6 (6) O25—C17—C18—C13 −179.9 (3) C12—C8—C9—N10 −0.4 (4) C16—C17—C18—C13 −0.2 (5) C1—C8—C9—N10 176.2 (3) C12—C13—C18—C17 176.4 (3) O19—C9—N10—C11 −178.3 (4) C14—C13—C18—C17 −2.8 (5) C8—C9—N10—C11 0.9 (4) C16—C15—O21—C22 −72.9 (5) C9—N10—C11—O20 179.7 (4) C14—C15—O21—C22 110.5 (4) C9—N10—C11—C12 −1.0 (4) C15—C16—O23—C24 −99.1 (4) C1—C8—C12—C13 0.5 (5) C17—C16—O23—C24 83.8 (4) C9—C8—C12—C13 177.5 (3) C18—C17—O25—C26 −8.0 (6) C1—C8—C12—C11 −177.2 (3) C16—C17—O25—C26 172.3 (3) C9—C8—C12—C11 −0.2 (4)
Hydrogen-bond geometry (Å, º)
D—H···A D—H H···A D···A D—H···A
N6—H6···O21 0.79 2.20 2.741 (4) 126